Abstract
Repurposing existing medicines to target disease-associated genes represents a promising strategy for developing effective treatments for complex diseases. However, progress has been hindered by a lack of viable candidate drug targets identified through genome-wide association studies. Gene-based association tests provide a more powerful alternative to traditional SNP-based methods, yet current approaches often fail to leverage shared heritability across populations and to effectively integrate functional genomic data. To address these challenges, we develop GenT and its various extensions, comprising a framework of gene-based tests utilizing summary-level data from genome-wide association studies. Using GenT, we identify 16, 15, 35, and 83 candidate genes linked to Alzheimer’s disease, amyotrophic lateral sclerosis, major depression, and schizophrenia, respectively, not detected by Genome-Wide Association Studies (GWAS). Additionally, we use our multi-ancestry gene-based test (MuGenT) to identify 28 candidate genes associated with type 2 diabetes. By integrating brain expression and protein quantitative trait loci into our analysis, we identify 43 candidate genes associated with Alzheimer’s disease that have supporting xQTL evidence. We also perform experimental assays to demonstrate that the NTRK1 inhibitor GW441756 significantly reduces tau hyper-phosphorylation (including p-tau181 and p-tau217) in Alzheimer’s disease patient-derived iPSC neurons, providing mechanistic support for our predictions.
Similar content being viewed by others
Introduction
For decades, genetic epidemiologists have endeavored to identify disease-associated genes using findings derived from human genome sequencing1. The most common approach to discovering these genes has been through genome-wide association studies (GWAS), which estimate marginal associations between millions of single nucleotide polymorphisms (SNPs) and various phenotypes2. Typically, gene-level inferences are drawn by assigning the SNP with the smallest association p-value in a locus (known as the lead SNP) to its nearest physical gene3. However, functional genomic studies indicate that only one-third of lead SNPs accurately tag causal genes4,5, and only about 5% of lead SNPs are likely to be causal6. Consequently, this conventional methodology necessitates statistical correction for all genome-wide tested SNPs, rather than just the genes to which they are presumed related. This strict correction imposes limitations on statistical power, undermining the ability to detect disease-associated genes that interact with available drugs, referred to as ‘druggable genes’7,8,9.
A more powerful approach for generating clinically relevant insights into biology and drug discovery is gene-based association testing10. Gene-based association testing performs a joint test on a gene-specific SNP set. While existing gene-based association tests using summary GWAS data represent efficient screening tools, each standard method relies on a statistic with an unknown null distribution. Accurately estimating this distribution is critical for reducing false discoveries of disease-causing druggable genes. Past efforts to address this issue have involved numerical approximations (i.e., Pascal11; MAGMA12), transformation based on linkage disequilibrium (LD) structure that is seldom precisely known (DOT13), simulation (Pascal; VEGAS14), or permutations (MAGMA, LDAK15). However, these various methods often lack strong theoretical grounding and can inflate both false positive and negative rates16,17,18,19. Other gene-based inference methods attempt to assess causality using summary data by integrating GWAS results with transcriptomic information (e.g., S-PrediXcan, TWAS, SMR, EMAGMA, cis-MR4,5,20,21,22), but these approaches rely on additional assumptions that may be difficult to verify and are not universally applicable.
The principal strength of gene-based methods is their capacity to perform a single test per gene, typically requiring only GWAS summary statistics and an LD reference panel. Nevertheless, these methods are limited by their inability to integrate heterogeneous GWAS and LD information across different ancestral populations, to accommodate diverse transcriptomic data types such as gene expression and protein abundance, and to prioritize driver genes in a gene-dense locus. Given that many complex diseases emerge from multidimensional risk factors within diverse biological contexts, there is an urgent need to harness functional genomic data for efficiently screening disease-associated drug targets.
In this study, we present a statistical framework, GenT, designed to test the gene-based association null hypothesis with greater theoretical support. This framework states the exact null distribution of a gene-based test statistic when LD is known, and a highly precise null distribution when it is estimated. We leverage these derived properties to extend standard gene-based association testing to integrate multi-ancestry data (MuGenT), test for the presence of a shared disease-xQTL (molecular quantitative trait locus) signal, and finemap multiple genes in a locus using their test statistics to identify putative driver genes. We additionally propose three adaptations of the multi-ancestry test to better understand gene-disease association heterogeneity across ancestral populations. We applied these methods to identify candidate druggable genes associated with several complex diseases, including Alzheimer’s disease (AD), major depression (MDD), schizophrenia (SCZ), amyotrophic lateral sclerosis (ALS), and type 2 diabetes (T2D). Our analyses uncovered multiple potential druggable genes for each disease, supported by empirical evidence from experimental and clinical studies, providing valuable insights to guide future mechanistic investigations and drug discovery efforts.
Results
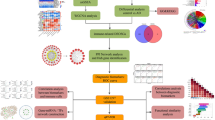
Figure 1 summarizes our set of marginal gene-based association tests and illustrates their application to real data through our single-trait association (GenT; Fig. 1b), multi-ancestry/trait association (MuGenT; Fig. 1c), and xQTL-weighted GenT (xGenT; Fig. 1d) tests. All methods test for an association between a phenotype and any SNP in a gene-specific set of SNPs, and require only GWAS summary statistics and an LD reference panel. GenT tests the null hypothesis that no SNP in the gene set is associated with the trait. MuGenT applied to a single trait measured in multiple populations tests the null hypothesis that no SNP in the gene set are associated with the trait in any population. xGenT tests the null hypothesis that no SNP is simultaneously associated with the trait and any xQTL trait which is modeled (e.g., gene expression in cortex tissue). Gene-based test statistics from all methods can be fine-mapped to identify putative driver genes in a locus after applying a simple asymptotic transformation (cf. Methods). The GenT test employs the standard gene-based association method using summary-level GWAS data, which calculates the sum of correlated SNP chi-square statistics for SNPs in/around a given gene body (see Supplementary Notes 1–2 for functional SNP-gene assignment results). Here we introduce a formulation of its null distribution, which we demonstrate in the Supplementary Note 2 to be exact under transformation when the LD structure is known and extremely precise when estimated across a range of conditions. Supplementary Figs. 1–3 provide comprehensive visual descriptions of the computational pipeline that each method follows, Supplementary Figs. 4–33 evaluate the performance of our analytic approach, and Supplementary Figs. 34–88 provide additional computational evaluations of our real data analyses.
a visual descriptions of the gene-based association tests that we introduce and their corresponding null hypotheses. We introduce GenT, MuGenT, xGenT, and three follow-up multi-ancestry tests: MuGenT-PH, MuGenT-Pleio, MuGenT-Sel. Parameters of association in each DAG are displayed as bold symbols. There are p populations/traits in the MuGenT DAG, which are indexed by k and s. There are k tissues in the xGenT DAG. The full null and alternative hypotheses of each method are stated in the Methods sections. b Motivating example of GenT which detects a potential druggable gene associated with Alzheimer’s disease (AD) that has supporting gene- and SNP-level fine-mapping evidence. c Motivating example of MuGenT which detects a potential druggable gene associated with T2D by leveraging genetic similarity across populations. d Motivating example of xGenT which detects an association between EZH1 and AD risk with colocalized evidence from cis expression quantitative loci (eQTL) from five brain tissues. In the far-right panel, annotated SNPs are jointly significant in the AD GWAS and spinal cord eQTL studies and their 95% confidence ellipses are displayed around their Z-statistics of marginal association. Source data are provided as a Source Data file.
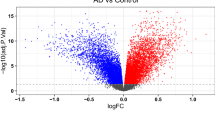
GenT controls its false positive rate at nominal levels better than existing gene-based association test methods (e.g., VEGAS, GATES, ACAT, MAGMA; Figs. 2a, 3a, 4a and Supplementary Figs. 4–12, 21, 25 to 27). For example, in simulations using data generated from the 1000 Genomes LD matrices23, the genome-wide inflation factor of MAGMA was 0.69 and of GenT was 1.00. This suggests that MAGMA has reduced power compared to GenT, which is supported by power comparisons both in simulation (Fig. 2a, Supplementary Figs. 28–30) and real data analyses with AD (Supplementary Figs. 23 and 24). Supplementary Fig. 21 shows that GenT can better control its Type I error rate at nominal levels as the magnitude of LD and sample size of reference panel varies than ACAT, GATES, and VEGAS. When the Type I error rate of these methods is controlled, they have the similar statistical power to GenT (Supplementary Fig. 22). The Supplementary Notes 3–7 provides complete details of all additional comparisons we made between GenT and other methods.
a results of simulations performed to evaluate the statistical properties of MuGenT. \({\lambda }_{{{{\rm{GC}}}}}\) in the left panel represents the genomic control inflation factor and has value 1 when no inflation is present. The blue value corresponds to MAGMA and the green to GenT. b Manhattan plots of two-sided gene-based test (GenT) p-values (bottom) and GWAS SNP-based tests (top) for four neuropsychiatric traits. Annotated genes are druggable and undetected in the respective GWAS. Dotted horizontal lines represent the Bonferroni-corrected significance thresholds of each test. c Examples of druggable genes detected in novel loci for AD and SCZ using GenT and posterior inclusion probabilities from gene- and SNP-level fine-mapping. Displayed under SNP-level locus plots are cell-specific gene expression proportions in human cortex tissue from Zhang et al. ‘Astro.’ fetal and mature astrocytes; ‘Endo.’ endothelial cells; ‘Oligo.’ Oligodendrocytes. Larger circles and darker red colors indicate greater FPKM values.
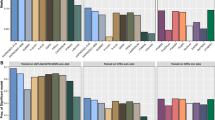
a results of simulations performed to evaluate the statistical properties of MuGenT. b Estimated genetic correlations of T2D between populations (top). Bar heights in the lower figure represent estimated SNP heritability values, and error bars indicate the 95% confidence interval. Estimated SNP heritability of T2D in each population (bottom). Source data are provided as a Source Data file. c Gene-level fine-mapping in two loci detecting potential druggable genes with GenT. These results provide supporting evidence for the CDC42BPA association with T2D but not FKBP5. d Results of two-sided MuGenT testing (top) and MuGenT-PH heterogeneity testing (bottom). Horizontal solid lines represent the Bonferroni-corrected significance threshold. e Volcano plots of ancestry-specific T2D GWAS p values, MuGenT and MuGenT-PH p-values, and SuSiE fine-mapping posterior inclusion probabilities for two example druggable genes in novel T2D loci. Horizontal dashed lines represent the two-side Bonferroni-corrected gene-based testing significance threshold. Solid lines represent the two-sided Bonferroni-corrected SNP-based testing significance threshold.
a results of simulations performed to evaluate the statistical properties of xGenT. b Gene-level fine-mapping results for three candidate druggable genes detected using xGenT with AD. In the right-most plot, shaded areas represent 95% confidence intervals. c Locus-specific association plots for three examples of candidate druggable genes and the gene expression tissues with corresponding evidence. P-values are from SNP-based testing in the AD GWAS (bottom) and eQTL data sets. d SNP-level locus plots for RIPK2 illustrating potential xQTL tissue and functional specificity. Warmer colors indicate stronger LD with the lead SNP; cooler colors indicate lower LD. The yellow region represents the hg19 start and end positions of RIPK2 and the horizontal dashed line indicates the level of genome-wide significance. P values are from two-sided tests of association with the trait labeled in each panel. e Scatterplots of SNP-specific marginal Z-statistics measuring association with AD and cerebellum eQTLs (left) and cortex pQTLs (right) in the RIPK2 locus. Error bars represent 95% confidence intervals. The blue line is the Mendelian Randomization causal estimate using MRBEE, and the reported P value is from a two-sided test that the causal estimate is equal to 0. Source data are provided as a Source Data file.
MuGenT expands upon GenT to accommodate multiple ancestral populations, harnessing their shared heritability and genetic correlation to enhance the power of the joint testing. This approach can also be applied to multiple phenotypes measured within the same population. xGenT extends GenT by incorporating xQTL effect sizes from diverse sources as weights, enabling an inference of shared disease and xQTL signals in a gene-specific SNP set. Each of these methods (GenT, MuGenT, and xGenT) leverages polygenicity and requires a less stringent corrected significance threshold than conventional SNP-based gene inference from GWAS. This relaxation increases the power of gene-based association testing to identify druggable genes associated with disease.
The results presented in Fig. 1b–d highlight examples of candidate genes detected exclusively through gene-based association testing, and not through traditional GWAS, for AD and T2D. ‘Druggable genes’ are defined as genes whose product (protein) has available binding ligands from currently FDA-approved or clinically investigative drugs identified by cross-referencing the ChEMBL24, BindingDB25, and GtoPdb26 databases with DrugBank27 (cf. Methods). This list included 3,369 genes interacting with at least one drug/compound. These genes represent candidate drug targets previously not detected in lead SNP analyses. For example, PDE3A exhibited a GenT false discovery rate (FDR) q value of 7.6 × 10−5, while the smallest FDR q-value for any SNP in this locus (±1 Mb) from AD GWAS was 0.148. Gene-level fine-mapping via SuSiE28 of genes in this locus prioritized only PDE4A (PIP = 1.00), and SNP-level fine-mapping of SNPs ±100Kb of PDE3A suggest the presence of a 95% credible set containing a causal SNP (PIP = 1.00). Similarly, the PPP3CA gene did not achieve genome-wide significance in any ancestry-specific GWAS for T2D, whether from European (EUR), African (AFR), East or South Asian (EAS, SAS), or Hispanic (HIS) populations, but its MuGenT FDR q-value was 4.9 × 10-45. Gene-level fine-mapping in the PPP3CA locus prioritized PPP3CA and BANK1 (PIPs=1.00), and SNP-level fine-mapping supports the likelihood of causal T2D SNPs in the PPP3CA locus ±100Kb across all populations except for SAS.
Figure 1d illustrates that xGenT, integrating eQTLs from five brain tissues, can detect an association between EZH1 expression and AD that GWAS alone fails to identify. Additionally, it demonstrates that MuGenT can simultaneously test multiple xQTL types within the same population. Simulation results showcased in Figs. 2a, 3a, and 4a suggest that our gene-based association tests maintain controlled false positive rates despite variations in GWAS sample sizes, SNP heritability, LD structure or estimation accuracy, and the number of tested SNPs. These tests generally exhibit greater power than gene-based inference from lead SNPs, benefitting not only from reduced multiple testing but also from leveraged polygenicity, while lead SNP-based methods lose power in such scenarios (Fig. 2a). The Supplementary Note 6 provides additional evaluations of Type I error for GenT, MuGenT and its post-hoc tests, and xGenT, demonstrating that each test has well-controlled false positive rates. Results presented in Supplementary Figs. 22, 24, 26, 29 and 30 suggest that GenT has comparable power to VEGAS and superior power to MAGMA. Supplementary Figs. 21, 25 and 26 show that GenT is the controls false positive rates better than MAGMA, VEGAS, ACAT, and GATES.
A compelling case study highlighting the advantages of GenT is the association between SYK and AD (Fig. 2c). The smallest AD GWAS P-value within 50kb of the SYK gene body was 2.1 × 10−5 (rs10512201), which falls short of genome-wide significance. However, gene-level fine-mapping in the SYK locus (1 Mb including DIRAS2, SYK, AUH, NFIL3, and ROR2) prioritized only SYK (PIP = 1.00), and SNP-level fine-mapping identified seven SNPs within 100Kb of SYK that had PIPs greater than 0.95. As the number of causal SNPs at the SYK locus increases, traditional gene-based inference using the lead SNP loses power. By contrast, gene-based association testing using GenT with 1492 proximal SNPs substantially enhanced the power to detect the SYK association with AD (FDR q = 8.0 × 10−5; Fig. 2a). Remarkably, it is approximated that the AD GWAS would have needed to recruit approximately 670 K additional participants to achieve genome-wide significance at 90% power to detect the SYK association using the lead SNP rs10512201.
GenT discovers potential druggable targets
We employed GenT to analyze 18,257 genes across four complex diseases: Alzheimer’s disease (AD), amyotrophic lateral sclerosis (ALS), major depressive disorder (MDD), and schizophrenia (SCZ), with results presented in Fig. 2b, c. All analyses utilized GWAS summary statistics from predominantly European (EUR) populations and EUR LD reference panels from the 1000 Genomes Phase 323. We identified 89 druggable genes associated with AD, 28 druggable genes associated with ALS, 65 druggable genes associated with MDD, and 258 druggable genes associated with SCZ, using a Bonferroni-corrected 5% significance threshold. Of these, 49 were finemapped (PIP > 0.9) in their loci for AD, 23 for ALS, 58 for MDD, and 177 for SCZ.
Notably, within the set of associated genes, we identified 75 druggable genes for AD, 26 druggable genes for ALS, 58 druggable genes for MDD, and 218 druggable for SCZ that were undetectable by the conventional lead SNP method (Fig. 2b). We also found 16, 15, 35, and 83 genes linked to AD, ALS, MDD, and SCZ, respectively, that reside beyond 1 Mb from any previously identified GWAS genes for each disease, spanning 10, 12, 24, and 57 independent loci (> 1 Mb separation). These genes are located within loci not captured by standard GWAS and encode druggable proteins. Examples of genes with functional studies supporting their disease associations, yet not meeting the threshold for genome-wide significance in GWAS, include KCNN4 in AD29, PISD in ALS30, EPHB2 in MDD31, and KCNQ2 in SCZ32.
Two examples of druggable genes with supporting gene- and SNP-level fine-mapping evidence are displayed in Fig. 2c and demonstrate that disease-associated druggable genes can still be prioritized as putative local signal drivers even when GWAS evidence falls short of genome-wide significance. We found at least one overlapping druggable gene for each pair of diseases, for example: EPHX2 for AD and SCZ, PPP3CA and CD40 for MDD and SCZ, and ATP6V1G2, PRKD1, and PSEN2 for ALS and SCZ. Comprehensive GenT fine-mapping results for all tested genes are available in Supporting Data 1–23 for AD, ALS, MDD, and SCZ, along with their SNP-level SuSiE fine-mapping results and gene-level fine-mapping results. In total, we applied gene-based association testing using GenT to 38 complex traits, and these results are publicly accessible at the repositories listed in the Data Availability section.
Multi-ancestry/trait gene-based association tests (MuGenT) identifies druggable targets across diverse populations
We applied our multi-ancestry/trait gene-based association test (MuGenT) to analyze five ancestral populations to identify druggable genes associated with T2D. The results are displayed in Fig. 3b–e. We utilized GWAS summary statistics from European, African, East Asian, South Asian, and Hispanic cohorts, along with population-specific LD reference panels from 1000 Genomes Phase 323. Our analysis revealed 1,211 independent genome-wide significant SNPs (defined as r2 < 0.01 in a 1 Mb window) in the trans-ancestry T2D GWAS, suggesting that T2D is highly polygenic.
MuGenT has optimal power when there is shared heritability and positive genetic correlation between populations (Fig. 3a). We estimated the genetic correlations of T2D between the five populations to range from 0.69 to 0.98 (Fig. 3b) using Popcorn33 (cf. Methods). In our analysis, MuGenT identified 269 genes significantly associated with T2D at the Bonferroni significance level despite lacking any genome-wide significant SNPs. Notably, 45 of these (7.6%; Fig. 3c, d) were druggable and not located within 1 Mb of any lead SNP from the trans-ancestry T2D GWAS (GWAS p < 5 × 10-8, LD r2 < 0.01 in a 1Mb window). Two examples of these potential druggable genes are FKBP5 (MuGenT p = 1.1 × 10-9) and CDC42BPA (MuGenT p = 1.3 × 10-7 (Fig. 3e). While FKBP5 methylation has been shown to associate with cardiometabolic risk34, follow-up gene-based fine-mapping in the FKBP5 locus (>1.5 Mb) identified TAF11, ANKS1A, TCP11 and SCUBE3 as the putative drivers (PIP = 1.00) and not FKBP5 (PIP = 0). Alternatively, gene-based fine-mapping in the CDC42BPA locus (>1.5 Mb) identified it as a putative driver, and it is a hypothesized regulator of insulin secretion35.
To further explore population heterogeneity in gene associations with T2D, we employed our MuGenT-PH post-hoc test across the African, European, East and South Asian, and Hispanic cohorts (Fig. 1a). The Spearman rank correlation between gene association p-values from MuGenT and population heterogeneity P values from MuGenT-PH was 0.37 (p < 2.2×10-16). MuGenT-PH aggregates SNP-level effect size heterogeneity at the gene level and conducts statistical testing.
Notable examples of between-population heterogeneity in the strength of druggable gene associations with T2D include the EIF4B and KIF11 genes. In the African cohort, the rs374136 SNP in the EIF4B gene showed a T2D GWAS association p-value of 2.6×10-18, while the next lowest p-value in other populations was 0.09 (observed in Europeans, the largest sample group). The MuGenT-PH p-value for heterogeneity was 1.2×10-8. Follow-up tests using our MuGenT-Sel post-hoc test (Fig. 1a) indicated an African-specific association of EIF4B with T2D, achieving a posterior probability greater than 0.95.
KIF11 is another druggable gene associated with T2D (MuGenT p = 4.5×10-263) that exhibited significant population heterogeneity in effect size (MuGenT-PH p = 1.2×10-68). In the E. Asian cohort (GWAS n = 247 K), the lead variant rs4933731 had a T2D GWAS p-value of 1.1×10-26, but all other populations reported GWAS p-values exceeding 5.5×10-3, including the EUR cohort, which was three times larger. Fine-mapping of the KIF11 locus suggests that the rs7911264 variant is likely causal across all populations except for SAS, with stronger LD patterns observed in EAS and SAS cohorts compared to others (Supplementary Fig. 85). All results from MuGenT and MuGenT-PH analyses for T2D are available in Supporting Data 5 and 6.
xGenT identifies druggable genes in AD
In addition to identifying T2D-associated genes, we evaluated AD-associated risk genes for colocalization and genetic correlation with expression and protein quantitative trait loci (e/pQTL) in brain tissues using our xQTL gene-based association test (xGenT). The results are displayed in Fig. 4. The eQTL data were sourced from the cerebellum, cortex, frontal cortex, spinal cord, and hippocampus, while the pQTL data were obtained from cortical tissue.
We show in Methods that in the absence of horizontal pleiotropy the xGenT test statistic is proportional to the total/marginal Mendelian Randomization causal effect sizes of expression in the tested brain tissue on AD risk. Of the 82 genes defining independent loci reported in the largest publicly available AD GWAS to date from Bellenguez et al.36, xGenT provides additional evidence of shared eQTL signals for 46 genes (56%) and shared pQTL signals for 32 genes (39%) at the Bonferroni significance level in our tested xQTLs. xGenT results from all gene-based tests using brain eQTLs and pQTLs are available in Supporting Data 7 and 8.
We identified 26 lead druggable genes (e.g., CHRM5, EGLN1, EZH1, RIOK1, RIPK2, SYK, NTRK1) with significant Bonferroni-adjusted xGenT p-values in eQTL and pQTL analyses that were located outside 1 Mb of any of the 82 AD-associated loci (Fig. 4b-c). SNP-level AD and eQTL associations for three of these genes (RIPK2, NTRK1, RIOK1) are displayed in Fig. 4c and Fig. 4b, which demonstrate that gene-level fine-mapping identified NTRK1 and RIPK2 as putative drivers with high confidence (PIP = 1.00) and RIOK1 with moderate confidence (PIP = 0.58). An example demonstrating the statistical power gained by xGenT through shared disease GWAS and xQTL signals is presented for RIPK2 in Fig. 4d. SNP-level fine-mapping using revealed that each of these three genes had at least one corresponding SNP with a causal PIP greater than 0.90, indicating their status as likely causal AD SNPs.
Using single cell RNA sequencing data from the Religious Orders Study/Memory and Aging Project (ROSMAP37), we found that all three genes were differentially expressed in glial cells within the cerebral cortex (Supplementary Fig. 87). This is consistent with existing literature implicating glial cells in AD pathology38,39. Furthermore, Mendelian Randomization (MR) analyses (cf. Methods) of RIPK2 gene expression in the cerebellum and protein abundance in the cortex suggest a potentially causal role for RIPK2 in protecting against AD (Fig. 4f).
Multi-omics validated xGenT-predicts druggable genes in AD
We detected 320 independent genes which had an xGenT Bonferroni p-value less than 0.05 in eQTL or pQTL testing and an AD GenT nominal p-value less than 0.05. These genes had correlations between their GenT test statistics less than 0.01 in a 1 Mb window around their base pair midpoints (cf. Methods). To validate these prioritized genes, we evaluated multi-omic evidence and displayed the top 150 genes with the strongest evidence in Fig. 5a. Our analysis highlighted several potent drug targets for AD, including ABAT, ATP2B1, HEXB, CHRM5, KIF1A, GLUD1, KCNS1, RCVRN, and RIPK2. For example, GLUD1 is upregulated in AD-associated microglia, and its protein (UniProt ID: P0036740) is elevated in the frontal cortex and hippocampus of the 5xFAD mouse model of AD41. Similarly, RCVRN is overexpressed in AD patients compared to controls in bulk RNA-seq data from hippocampal tissue42,43.
a Multi-omic evidence of association of top 150 of 616 xGenT-prioritized AD risk genes using the sources introduced in Methods. Genes included in this plot had the top 150 smallest two-sided xGenT P-values when tested with brain eQTLs and pQTLs. DEG/P: Differentially expressed gene/protein. b Local genetic correlations between gene expression and AD effects that are marginal, sparse, and calculated using the procedure described in Methods. Mendelian Randomization causal estimates are from univariable analyses and use the procedure described in Methods. Only genes with MR P-values less than 0.05/190 gene-tissue pairs have colored squares. c AD patient iPSC-derived neurons were treated with NTRK1 inhibitor GW441756 for 24 h. Western blot analysis was performed for indicated proteins. Upper: Representative blot. Lower: Quantification of p-tau/total tau for three independent blots. P-values are two-sided and test null hypotheses stating equality between mean p-tau/total tau levels in the DMSO vs each of the other non-DMSO groups. Error bars represent 95% confidence intervals. Source data are provided as a Source Data file. d Variants (triangles) and gene locations (gray areas) are annotated and using Ensembl (www.ensembl.org). Points are colored according to the LD with the lead SNP with warmer colors (red, orange) indicating stronger LD and cooler colors (yellow, green) weaker LD. P-values are directly from the AD GWAS data and are from two-sided tests of association of each SNP with AD. Experimental structures of HDGF (left, Protein Data Bank [PDB] ID: 1RI0), NTRK1 (right, PDB ID: 8J5X), and a predicted complex structure (center) generated using ColabFold (https://github.com/sokrypton/ColabFold). Unstructured and non-interacting regions pruned for clarity. Experimental structures downloaded from the RCSB Protein Data Bank. p values for ‘eQTL’ are from MuGenT applied to eQTLs for the five brain tissues; P values for ‘pQTL’ are from GenT applied to pQTLs from the ROSMAP case-control sample (see “Methods”); p values for ‘GenT’ are from GenT applied to the AD GWAS summary statistics.
Figure 5b presents marginal genetic correlation and Mendelian Randomization (MR) causal effect estimates for each gene expression-brain tissue pair with AD. These estimates suggest that the positive expression of genes such as EARS2, EZH1, NARS2, NTRK1, RCVRN, and RIOK1 are associated with increased AD risk across multiple brain regions. NTRK1 consistently demonstrated a strong association with AD risk, as evidence by analyses from GenT, gene-level GenT fine-mapping, xGenT, genetic correlation, and MR across various brain tissue types with a relatively strong positive effect on AD risk in the hippocampus (Fig. 5b). NTRK1 has been shown to associate with other conditions related to aging and cognitive decline such as Parkinson’s disease44, early AD subtype45, arthritis46, and hippocampal volume and cell survival47,48, but literature support for the association between NTRK1 and AD directly remains limited. MR sensitivity analyses using 11 other MR methods support our findings in Fig. 5b for NTRK1 and other genes (see Supplementary Fig. 33).
Figure 5d illustrates the distribution of AD GWAS SNP associations within the NTRK1 locus, highlighting two missense mutations that are marginally associated with AD: rs926103 on the third exon of SH2D2A and rs4399146 on the fifth exon of HDGF49. These SNPs serve as pQTLs for HDGF (FDR < 0.05), but not for NTRK1 (FDR > 0.05) in cortical tissue37. Gene-based testing of all cis-pQTL SNPs in the cortex with GenT, and cis-eQTL SNPs from multiple brain regions with MuGenT, indicated significant pQTLs for HDGF but no eQTLs, while NTRK1 exhibited significant eQTLs without significant pQTLs. Additionally, the NTRK1 protein physically interacts with CDK5 (Supplementary Fig. 88), a known AD risk gene associated with tau phosphorylation50,51.
Experimental validation of NTRK1 association with AD in iPSC-derived neurons
Given the strong evidence linking NTRK1 to AD and the limited supporting literature, we pharmacologically targeted NTRK1 to evaluate its functional connectivity to AD. NTRK1 plays a crucial role in the development and survival of nerve cells and is also specifically involved with subtyping of sensory neurons52. It is primarily expressed in the brain and adrenal glands53, and mutations in NTRK1 are associated with various cancers54 and pain insensitivity55.
Our western blot results demonstrated that targeting NTRK1 reduced the levels of Tau phosphorylation at two key sites (pTau181, p-value = 0.008; pTau217, p-value = 0.003) in a dose-dependent manner in female patient iPSC-derived neurons (IUGB416, cf. Methods) (Fig. 5c). Additionally, a NTRK1 inhibitor (GW441756, cf. Methods) significantly decreased both pTau181 and pTau217 (Fig. 5c) in another patient-derived iPSC neuronal cell line (IUGB269.1). Since hyperphosphorylation of Tau leads to the formation of neurofibrillary tangles, a hallmark AD pathology, our in vitro findings suggest that targeting NTRK1 may help alleviate Tauopathy in AD neurons. Further experimental observations on NTRK1 functional roles in AD are warranted in the future.
Discussion
Gene-based association testing offers a powerful alternative to standard SNP-based association methods most commonly used for discovering druggable genes in genome-wide association studies (GWASs). While SNP-based gene discovery faces challenges related to gene assignment and uncontrolled false positive and negative rates, gene-based methods can provide greater robustness against these issues. However, existing gene-based tests lack strong theoretical support, leading to inflated false positive and negative rates.
To address these challenges, we introduced a set of gene-based tests that employ a inferential approach, effectively controlling false positive and negative rates. This allows for the identification of numerous druggable genes associated with complex diseases. We demonstrated the effectiveness of our approach for AD, ALS, MDD, SCZ, and T2D (Supplementary Figs. 34–37) and highlighted select examples of druggable genes with supporting functional evidence.
Through gene-based association testing, we identified 415 potential druggable genes associated with at least one of these diseases. Notable examples include SYK for AD and THRB for schizophrenia. SYK is highly expressed in brain tissue56 and shows strong genetic correlations with 17 neuropsychiatric and metabolic phenotypes (Supplementary Fig. 35). Although the lead SNP in the SYK locus (±50Kb) did not reach genome-wide significance (p = 2.1 × 10−5; Bellenguez et al.36), we found a significant association with AD using both GenT and xGenT, with brain eQTLs (p = 2 × 10−9) and pQTLs (p = 2.1 × 10−9). SYK works synergistically with TREM2 in microglia to clear amyloid beta57, the accumulation of which is a key hallmark of AD pathology58. This study provides genome-wide association evidence linking SYK to AD, which has previously only been documented in experimental settings.
Additionally, we identified an association between the thyroid hormone receptor gene (THRB) and SCZ in a locus that was not detected in previous GWAS. Variants in THRB are linked to various phenotypes, including cognitive function59, asthma60, telomere length61, myopia62, and atrial fibrillation63. THRB is primarily expressed in cerebral cortical neurons56 and functions as a thyroid hormone receptor. Functional studies suggest that thyroid hormone is vital for normal brain development64. Given that many SCZ risk factors relate to disruptions in normal brain development65,66,67,68, we hypothesize that THRB may confer SCZ risk by disrupting thyroid signaling pathways during development. Both THRB and SYK warrant further functional investigation as potential therapeutic targets for SCZ and AD, respectively.
We also applied a multi-ancestry gene-based association test (MuGenT) to T2D across African, European, East Asian, South Asian, and Hispanic populations, identifying 45 druggable genes in loci that were not detected using SNP-level testing in the largest T2D GWAS, which involved over 1.2 million individuals (Supplementary Figs. 34–37). Our gene-based test demonstrated greater power than trans-ancestry SNP-level GWAS that relies on individual level data, as it reduced the multiple testing burden and did not assume homogenous effect sizes across populations. Potential druggable genes include FKBP5 and CDC42BPA. Functional studies have suggested that FKBP5 expression in subcutaneous adipose tissue may induce insulin resistance69,70, and its methylation could increase the risk of T2D and hyperlipidemia34. Additionally, increased activation of the protein encoded by FKBP5 has been linked to tau aggregation in AD71. CDC42BPA has been associated with metabolic traits such as aberrant low-density lipoprotein72 and diastolic blood pressure73, as well as smoking initiation74. Furthermore, it has been implicated in inflammatory responses using CRISPRi screening in human-derived astrocytes75.
We also detected four potential T2D-associated genes using our gene-based test of population heterogeneity: CA6, CCT3, KRT8, and ASIC2. Previous studies indicate that CA6 protein abundance in plasma differs between T2D patients with and without β-cell dysfunction76. CCT3 may be a marker of diabetic nephropathy77 and KRT8 has been shown to be involved in the regulation of blood glucose78. ASIC2, primarily expressed in neurons, is related to cardiovascular function79 and has been linked to metabolic syndrome in mice80. Collectively, these findings provide epidemiological and biological support for the association between these genes and T2D risk.
Additionally, we highlighted three druggable genes (RIPK2, NTRK1 and RIOK1) associated with AD using our gene-based test that integrated brain xQTLs. Although the lead GWAS SNPs for these genes did not achieve genome-wide significance, they exhibited strong xQTL signals, resulting in high significance when tested with xGenT. Each of these genes has previously been validated for association with AD endophenotypes using CRISPRi. For example, CRISPRi testing suggested that NTRK1 over-expression can cause AD-like neuronal loss81,82. Additionally, knocking down RIPK2 in astrocytes is associated with increased inflammatory reactivity, while RIOK1 knockdown is linked to reduced reactivity75. These associations align with observed positive genetic correlations between AD risk and gene expression in the cerebellum (RIPK2; Supplementary Figs. 16 and 33) and hippocampus (RIOK1; Supplementary Figs. 15 and 32), indicating a potential role for brain over-expression of RIPK2 and RIOK1 in influencing neuroinflammation associated with AD83,84,85.
Our work demonstrates several strengths, including robust statistical properties of our gene-based tests, the discovery of empirically supported genes across multiple disease phenotypes, the integration of transcriptomic information, and the availability to apply fine-mapping to gene-based test statistics in large genomic regions. First, our gene-based tests are theoretically grounded and extensively validated through statistical theory and simulations. They require only GWAS summary statistics and an LD reference panel, making them applicable for inter-population gene association inference. Our standard test, GenT, utilizes a standard gene-based test statistic with a now clearly defined null distribution, overcoming challenges faced by previous methods that led to inflated false positive and negative rates. The extensions developed from GenT leverage its theoretical foundations, enabling a broader range of gene-based inferences than previously available to researchers.
We identified potential druggable genes using real-world data across multiple complex diseases, revealing numerous candidate targets that warrant additional experimental investigation. Many of these genes also have historical support from candidate gene or functional studies, strengthening the evidence that our tests can reveal true disease associations that escape detection by traditional SNP-based GWAS methods, a common standard for detecting disease-associated genes. Furthermore, by integrating transcriptomic data in our xGenT tests, we can provide additional insights into the contexts in which specific disease risks arise. Our framework can include any xQTL data, such as those for protein abundance, DNA methylation, gene expression, alternative splicing, and metabolomics.
We acknowledge several potential limitations in our current gene-based framework. Fully leveraging GWAS data from multiple ancestral populations can enhance candidate druggable gene discovery only if corresponding population-specific data exists, which is often not the case for many complex diseases. The list of such diseases is extensive, and future research should focus on disseminating all population-specific GWAS and xQTL data. Even when sample sizes are small, these data can still be utilized by meta-analytic techniques such as MuGenT to recover some lost power. Additionally, optimal assignment of SNPs to genes remains ambiguous in the literature. Consequently, we conservatively assigned SNPs to all genes within 50Kb of the gene body. As shown in Supplementary Note 12, 10Kb and 100Kb windows explain approximately the same trait heritability as that which is explained by SNPs in 50Kb windows. Increasing window sizes on average reduce the power of gene-based association testing if they do not capture additional causal SNPs, but increasing SNP set sizes makes transformed gene-based test statistics for fine-mapping converge to their asymptotic distribution faster.
An alternative approach assigns SNPs to gene-specific sets based on their functional (e.g., transcriptomic, proteomic) consequences. Our development of xGenT is an attempt at prioritizing SNPs with functional consequences in gene-specific sets formed using positional information, though gene-specific sets formed only from functional SNPs were not formed in our analyses. One challenge of this approach is that it potentially introduces a sequential testing procedure which requires stricter control of the Type I error rate. Supplementary Note 15 discusses and compares positional SNP-gene assignment to functional SNP-gene assignment using hippocampal eQTL data with AD and shows that functional SNP-gene assignment is less powerful but may prioritize genes better than distance-based SNP-gene assignment (Supplementary Figs. 31 and 32). This subject is a focus of our future work. Finally, while our results provide preliminary evidence of associations between druggable genes and disease phenotypes, further functional analyses are necessary before these genes can be considered safe and effective therapeutic targets. Gene-based fine-mapping can provide supporting genetic evidence for a trait since it evaluates gene association using a different inferential framework. However, high-throughput functional assays are still the best next step towards identifying candidate drug targets with real-world potential for a therapeutic effect on disease since in silico methods are subject to theoretical assumptions.
Within the theoretical frameworks of in silico methods, we still recommend the incorporation of gene prioritization approaches such as gene-level finemapping and/or xQTL-integrated analyses. These approaches may help to identify the most disease relevant genes from a large set of disease association genes. It should also be noted that standard assumptions of independence between genes based on the observed absence of LD between their SNP sets is violated if non-neighboring genes are functionally related. Functional relationships between multiple genes may be established in distinct biological pathways via protein-protein interactions (PPIs), promoter/enhancer relationships, and/or transcription factor binding. Within these pathways, many genes which are not in LD may be present. As a result, these genes may not be functionally independent with each other, even if gene prioritization methods such as finemapping or xQTL-integrated analyses detect each of them. Current standard applications of GWAS and subsequent gene-based testing can identify many prioritized candidate genes because of the assumed independence via absent LD. We therefore propose the interpretation of prioritized candidate druggable targets with the nuance that their prioritization may reflect the importance of the underlying pathway of which they are a member. Therefore, many prioritized targets may reflect the same underlying pathway, and indeed many drugs may target multiple genes on the same pathway.
In conclusion, our study supports the idea that gene-level statistics are more suitable for gene-based inference than traditional SNP-based approaches, enabling researchers to identify a greater number of disease-associated druggable genes. The discrepancies between large scale GWAS and more targeted experimental investigations raise important questions. Many of the genes identified through our enhanced gene-based tests possess strong biological relevance within well-established pathways, suggesting they should be detectable through GWAS. We assert that many genes validated in experimental contexts confer protection or risk for diseases via multiple intricate pathways, making their detection in GWAS inversely proportional to the complexity of these pathways and level of causality involved. Additionally, a disease-associated gene containing multiple causal SNPs is generally more likely to be detected through gene-based association testing than by the lead SNP approach. While our integration of xQTLs in gene-based testing offers insights into biological plausibility for specific disease-gene associations, it is essential to consider simultaneous gene associations with other phenotypes when selecting candidate druggable targets. For example, we show in Supplementary Fig. 63 that RIPK1 expression in cerebellum is negatively correlated with AD, indicating a potentially protective effect, whereas the corresponding genetic correlation with coronary artery disease (CAD) is positive. Therefore, future research must consider that pharmacologically targeting such a gene may reduce risk of one disease while simultaneously increasing the risk of another. The development of methods which can identify these targets is ongoing (86), though it is still in its early stages.
Methods
Data sources for GWAS and drug-target interactions
We used publicly available summary statistics from genome-wide association studies (GWASs) for all gene discovery and estimated ancestry-specific linkage disequilibrium (LD) from the 1000 Genomes Phase 323 reference panels using PLINKv1.987. Disease GWAS summary statistics included those for Alzheimer’s disease (AD36; n = 487 K EUR), schizophrenia (SCZ88; n = 320 K EUR), major depressive disorder (MDD89; n = 500 K EUR), amyotrophic lateral sclerosis (ALS90; n = 138 K EUR), and Type 2 diabetes (T2D91; n = 752 K EUR, n = 129 K AFR, n = 247 K EAS, n = 42 K SAS, n = 73 K HIS). We also used expression quantitative trait loci (eQTL) summary statistics from GTEx v892 for cortex, frontal cortex, hippocampus, cerebellum, and spinal cord tissues in predominantly European populations. All analyses therefore used single nucleotide polymorphisms (SNPs) with minor allele frequency greater than ~0.1%. Our Data availability statement provides the repositories from which all GWAS summary data used in the main text were downloaded. We defined our list of 3,369 ‘druggable genes’ as those with drug-target interactions by joining lists of interactions from ChEMBL24, BindingDB25, and GtoPdb26 to their mapped molecules in DrugBank27.
Single-nuclei RNA-sequencing (snRNA-seq) data
We used individual-level snRNA-seq data from the Religious Orders Study and Rush Memory and Aging Project (ROSMAP37) to compare gene expression in the prefrontal cortex across the following broad cell type groups: epithelial, fibroblast, glial, immune, neuron, and vascular. Epithelial cells included choroid plexus epithelial cells and ependymal cells; glial cells included astrocytes, microglia, oligodendrocytes, and oligodendrocyte precursor cells; immune cells included choroid plexus macrophages and T-cells; neurons included excitatory and inhibitory neurons; vascular cells included ependymal cells, pericytes, and smooth muscle cells.
Multi-omics validation data
The results presented in Fig. 5a used reference data from multiple sources. These included The Alzheimer’s Cell Atlas93 for items 1-3 (see Fig. 5a key), the Genome-wide Positioning Systems platform for Alzheimer’s disease43 for items 4-7, manual literature searches and verification in the Open Targets Platform94, Target Illumination GWAS Analytics95, and DisGeNET96 databases for item 8, the Alzheimer’s Disease Sequencing Project Gene Verification Committee97 for item 9, and GTEx v892 for item 10.
Gene-based association test (GenT)
Our Gene-based association Test (GenT) is a test of association between a gene-specific set of SNPs and a disease phenotype. The primary advantage of GenT over similar existing gene-based tests is that its null distribution is shown under transformation to be a Gamma distribution with matched first two moments when LD structure is known98,99 (see Supplementary Note 1). As a result, it does not rely on complex numerical approximation (Pascal11; MAGMA12), simulation (VEGAS14), permutation (MAGMA12, LDAK15), or an assumed heritability model (LDAK15). This makes GenT both theoretically supported and free of the demanding computational burden that repeated sampling imposes. The null distribution of GenT when LD is estimated is approximated using a moment method which relies only on unbiased estimation of the LD matrix corresponding to the set of SNPs used for each gene. Let \({\hat{{{{\boldsymbol{\beta }}}}}^{{{{\mathcal{l}}}}}}={({\hat{\beta }_{1}^{{{{\mathcal{l}}}}}},\ldots,{\hat{\beta }_{{m}^{{{{\mathcal{l}}}}}}^{{{{\mathcal{l}}}}}})}^{\top }\) represent the vector of marginal association estimates corresponding to the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene and disease phenotype for which there are \({m}^{{{{\mathcal{l}}}}}\) SNPs used. GenT tests \({H}_{0}^{{{{\mathcal{l}}}}}:{{{{\boldsymbol{\beta }}}}}^{{{{\mathcal{l}}}}}={{{\boldsymbol{0}}}}\) vs \({H}_{1}^{{{{\mathcal{l}}}}}:{{{{\boldsymbol{\beta }}}}}^{{{{\mathcal{l}}}}}\ne {{{\boldsymbol{0}}}}\) using the statistic \({S}_{G}^{{{{\mathcal{l}}}}}={\sum }_{j=1}^{{m}^{{{{\mathcal{l}}}}}}{T}_{j}^{{{{\mathcal{l}}}}}\) where \({T}_{j}^{{{{\mathcal{l}}}}}\) is the standard chi-squared statistic used to test \({H}_{0j}^{{{{\mathcal{l}}}}}:{\beta }_{j}^{{{{\mathcal{l}}}}}=0\) in GWAS. Under \({H}_{0j}^{{{{\mathcal{l}}}}}\), \({T}_{j}\sim {\chi }^{2}\left(1\right)\) which implies under \({H}_{0}^{{{{\mathcal{l}}}}}\) that \({S}_{G}^{{{{\mathcal{l}}}}}\) can be very well approximated by \(\Gamma \left({{{\rm{\alpha }}}},{{{\rm{\xi }}}}\right)\) as \({m}^{{{{\mathcal{l}}}}}\) grows98,99, \(E\left({S}_{G}^{{{{\mathcal{l}}}}},|,{H}_{0}^{{{{\mathcal{l}}}}}\right)={m}^{{{{\mathcal{l}}}}}\), and \({Var}\left({S}_{G}^{{{{\mathcal{l}}}}},|,{H}_{0}^{{{{\mathcal{l}}}}}\right)=2{{{\rm{tr}}}}({{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}{{{{\bf{R}}}}}^{{{{\mathcal{l}}}}})\) where \({{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}={{{\rm{Corr}}}}({\hat{{{{\boldsymbol{\beta }}}}}^{{{{\mathcal{l}}}}}})\) is the \({m}^{{{{\mathcal{l}}}}}\times {m}^{{{{\mathcal{l}}}}}\) matrix of LD correlations between SNPs corresponding to the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene. This implies \(\xi={m}^{{{{\mathcal{l}}}}}/{{{\rm{tr}}}}\left(2{{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}{{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}\right)\) and \(\alpha={m}^{{{{\mathcal{l}}}}}\xi\). We provide additional details about this distributional result and extensive simulations to verify its precision in Supplementary Note 1. In practice, we assign all SNPs within 50Kb of the start and end positions of the gene to it as in ref. 14. Supplementary Note 15 discusses assignment of SNPs to gene-specific sets based on xQTL effect sizes. Supplementary Figs. 31 and 32 show results of these analyses using hippocampal eQTL effect sizes to define SNP sets in genome-wide GenT analyses with AD.
Fine-mapping multiple genes in a single locus
In standard applications of GenT and the subsequent gene-based tests that we introduce, a single test is performed for each gene. This can lead to inflation of association test statistics across the genome because multiple genes which are physically proximal to each other in local region may share SNPs in LD11. Indeed, the empirical evidence suggests that for highly polygenic traits such as T2D, the effect of LD hitchhiking between neighboring gene-specific SNP sets can be more severe than that which is encountered in SNP-level analyses. GenT and its extensions are marginal association tests in their original form and do not directly provide a way to infer driver genes in a locus. We introduce three methods to address the issue of inflation from LD and prioritization of driver genes in a locus: (i) post-hoc inflation correction, (ii) prioritization of genes in a locus based on P-value ranking and LD with surrounding genes, and (iii) fine-mapping of multiple gene-based test statistics in a single locus.
Option (i) is the most straightforward, though its post-hoc adjustment of gene-based P-values beyond that which is due only to high polygenicity may reduce power to detect disease-associated genes. Option (ii) is conceptually equivalent to the clump procedure in PLINK87 and can classify genes into one of three mutually exclusive groups: (a) lead genes, (b) clumped genes, and (c) non-significant genes. Lead genes are significant beyond a user-specified threshold and not correlated with any other genes in the locus beyond a user-defined threshold; clumped genes are significant but are correlated with at least one other gene in the locus which has a smaller gene-based test P-value. Option (iii) uses the limiting distribution of a transformation of the gene-based test statistic \({S}_{G}^{{{{\mathcal{l}}}}}={\sum }_{j=1}^{{m}^{{{{\mathcal{l}}}}}}{T}_{j}^{{{{\mathcal{l}}}}}\) from above under the gene-based null hypothesis that \(E\left({S}_{G}^{{{{\mathcal{l}}}}}\right)={m}^{{{{\mathcal{l}}}}}\):
The reason for the transformation of \({S}_{G}^{{{{\mathcal{l}}}}}\to {\ddot{S}}_{G}^{{{{\mathcal{l}}}}}\) is that the joint distribution of multiple gene-based test statistics is unknown, but the limiting distribution of them under the above transformation is multivariate normal. This result states that each transformed gene-based test statistic \({\ddot{S}}_{G}^{{{{\mathcal{l}}}}}\) follows a standard normal distribution as \({m}^{{{{\mathcal{l}}}}}\to \infty\) under \({H}_{0}^{{{{\mathcal{l}}}}}\), i.e., as the number of SNPs included in the gene-based association test increases. This property allows us to maximize the penalized joint likelihood for \(K\) correlated genes in a single locus using tools such as the Sum of Single Effects (SuSiE) model28 that will perform fine-mapping of the correlated genes. The matrix of correlations between gene-based test statistics is a result of shared LD between their SNP sets and is derived in Supplementary Note 8 for GenT, MuGenT, and xGenT. We provide additional details about fine-mapping of transformed gene-based test statistics in Supplementary Note 9, and the clumping algorithm of option (ii) described above in Supplementary Note 8.
Multiple-ancestry gene-based association test (MuGenT)
The Multiple-ancestry Gene-based association Test (MuGenT) is used to test the association between a gene and multiple traits, which could either be a single disease phenotype in multiple populations, multiple disease phenotypes in a single population, or multiple disease phenotypes in multiple populations. The null hypothesis for this test is that no SNPs in the gene set are associated with any of the tested traits; the alternative hypothesis is that at least one SNP is associated with at least one trait. Figure 3a showed that sharing of causal SNPs across traits/ancestries leads to greater power of the MuGenT test when the effects are genetically correlated, though does not meaningful improve the statistical power of MuGenT when they are not. The rationale for MuGenT and its derivation will be introduced using the context of a single disease phenotype measured in \(p\) populations. Let \({\hat{\beta }_{{jk}}^{{{{\mathcal{l}}}}}}\) represent the estimate of marginal association between the disease phenotype and jth SNP corresponding to the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene in the kth population, \({\hat{{{{\boldsymbol{\beta }}}}}_{k}^{{{{\mathcal{l}}}}}}={({\hat{\beta }_{1k}^{{{{\mathcal{l}}}}}},\ldots,{\hat{\beta }_{{m}^{{{{\mathcal{l}}}}}k}^{{{{\mathcal{l}}}}})}}^{\top }\), and \({\hat{{{{\boldsymbol{\beta }}}}}}_{j}^{{{{\mathcal{l}}}}}={({\hat{\beta }_{j1}^{{{{\mathcal{l}}}}}},\ldots,{\hat{\beta }_{{jp}}^{{{{\mathcal{l}}}}}})}^{\top }\). Also let \({{{{\bf{R}}}}}_{k}^{{{{\mathcal{l}}}}}\) represent the \({m}^{{{{\mathcal{l}}}}}\times {m}^{{{{\mathcal{l}}}}}\) matrix of LD correlations between SNPs corresponding to the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene in the \(k{{{\rm{th}}}}\) population. MuGenT tests the null hypothesis \({H}_{0}^{{{{\mathcal{l}}}}}:{\cap }_{k=1}^{p}{{{{\boldsymbol{\beta }}}}}_{k}^{{{{\mathcal{l}}}}}={{{\boldsymbol{0}}}}\) vs \({H}_{1}^{{{{\mathcal{l}}}}}:{\cup }_{k=1}^{p}{{{{\boldsymbol{\beta }}}}}_{k}^{{{{\mathcal{l}}}}}\ne {{{\boldsymbol{0}}}}\) using the statistic \({S}_{M}^{{{{\mathcal{l}}}}}={{{{\boldsymbol{1}}}}}_{p}^{\top }{{{{\bf{Z}}}}}_{{{{\mathcal{l}}}}}^{\top }{{{{\bf{Z}}}}}_{{{{\mathcal{l}}}}}{{{{\boldsymbol{1}}}}}_{p}/{m}^{{{{\mathcal{l}}}}}\) where \({{{{\bf{Z}}}}}_{{{{\mathcal{l}}}}}=({Z}_{{jk}}^{{{{\mathcal{l}}}}})\) and \({Z}_{{jk}}^{{{{\mathcal{l}}}}}\) is the Z-statistic used to test the association between the jth SNP of the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene in the kth population and the disease phenotype. Under \({H}_{0}^{{{{\mathcal{l}}}}}\), \({S}_{M}^{{{{\mathcal{l}}}}}\sim \Gamma \left(\alpha,\xi \right)\) with increasing precision as \({m}^{{{{\mathcal{l}}}}}\) grows98,99, \(E\left({S}_{M}^{{{{\mathcal{l}}}}},|,{H}_{0}^{{{{\mathcal{l}}}}}\right)=p\), \({Var}\left({S}_{M}^{{{{\mathcal{l}}}}},|,{H}_{0}^{{{{\mathcal{l}}}}}\right)={({m}^{{{{\mathcal{l}}}}})}^{-2}{{{{\boldsymbol{1}}}}}_{{p}^{2}}^{\top }{{{{{\boldsymbol{{\Upsilon }}}}}}^{{{{\mathcal{l}}}}}{{{\bf{1}}}}}_{{p}^{2}}\) (see Supplementary Note 3 for the derivation of \({{{{\boldsymbol{{\Upsilon }}}}}}^{{{{\mathcal{l}}}}}\), which is a transformed sum of all \({{{{\bf{R}}}}}_{k}^{{{{\mathcal{l}}}}}\)). This distributional relationship implies that \(\xi=p{\left({m}^{{{{\mathcal{l}}}}}\right)}^{2}/{{{{\boldsymbol{1}}}}}_{{p}^{2}}^{\top }{{{{\boldsymbol{{\Upsilon }}}}}}^{{{{\mathcal{l}}}}}{{{{\boldsymbol{1}}}}}_{{p}^{2}}\) and \(\alpha=p\xi\).
When multiple ancestries are used with MuGenT, the quantity \({{{{\boldsymbol{{\Upsilon }}}}}}^{{{{\mathcal{l}}}}}:={{{\rm{Cov}}}}\left[{{{\rm{vec}}}}\left({{{{\bf{Z}}}}}_{{{{\mathcal{l}}}}}^{\top }{{{{\bf{Z}}}}}_{{{{\mathcal{l}}}}}\right)\right]\), which is the primary component which determines the null distribution of the MuGenT test statistic, has a special patterned structure which is best understood by observing Supplementary Fig. 5. This result shows that LD covariance of the SNP set exists only within populations and not between populations either within or between individual SNPs. This greatly reduces the number of nonzero elements in \({{{{\boldsymbol{{\Upsilon }}}}}}^{{{{\mathcal{l}}}}}\), and the remaining nonzero covariance elements are summed products of within-population total LD in the SNP set between ancestry pairs. In summary, differing LD patterns between populations are handled by MuGenT since the variance of its test statistic is calculated without assuming LD patterns are the same across populations.
Multiple-ancestry gene-based test of population heterogeneity (MuGenT-PH)
The Multiple-ancestry Gene-based Test of Population Heterogenity (MuGenT-PH) test is of heterogeneity in gene associations between ancestral populations. Let \({\hat{\beta }_{{jk}}^{{{{\mathcal{l}}}}}}\) represent the estimate of marginal association between the disease phenotype and jth SNP corresponding to the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene in the kth population, \({\hat{{{{\boldsymbol{\beta }}}}}_{k}^{{{{\mathcal{l}}}}}}={({\hat{\beta }_{1k}^{{{{\mathcal{l}}}}}},\ldots,{\hat{\beta }_{{m}^{{{{\mathcal{l}}}}}k}^{{{{\mathcal{l}}}}}})}^{\top }\), and \({\hat{{{{\boldsymbol{\beta }}}}}_{j}^{{{{\mathcal{l}}}}}}={({\hat{\beta }_{j1}^{{{{\mathcal{l}}}}}},\ldots,{\hat{\beta }_{{jp}}^{{{{\mathcal{l}}}}}})}^{\top }\). The null hypothesis that MuGenT-PH tests is \({H}_{0}^{{{{\mathcal{l}}}}}:{\cap }_{k,s=1{;k}\ne s}^{p}{{{{\boldsymbol{\beta }}}}}_{k}^{{{{\mathcal{l}}}}}={{{{\boldsymbol{\beta }}}}}_{s}^{{{{\mathcal{l}}}}}\) vs \({H}_{1}^{{{{\mathcal{l}}}}}:{\cup }_{k,s=1{;k}\ne s}^{p}{{{{\boldsymbol{\beta }}}}}_{k}^{{{{\mathcal{l}}}}}\ne {{{{\boldsymbol{\beta }}}}}_{s}^{{{{\mathcal{l}}}}}\) using the statistic \({S}_{M-{PH}}^{{{{\mathcal{l}}}}}\). The statistic \({S}_{M-{PH}}^{{{{\mathcal{l}}}}}\) is the sum of \(m\) SNP-specific statistics \({T}_{j}^{{{{\mathcal{l}}}}}\) which are used to test \({H}_{0j}^{{{{\mathcal{l}}}}}:{{{{\boldsymbol{\beta }}}}}_{j}^{{{{\mathcal{l}}}}}-{p}^{-1}{{{{\boldsymbol{1}}}}}_{p}^{\top }{{{{\boldsymbol{\beta }}}}}_{j}^{{{{\mathcal{l}}}}}={{{\boldsymbol{0}}}}\) vs \({H}_{1j}^{{{{\mathcal{l}}}}}:{{{{\boldsymbol{\beta }}}}}_{j}^{{{{\mathcal{l}}}}}-{p}^{-1}{{{{\boldsymbol{1}}}}}_{p}^{\top }{{{{\boldsymbol{\beta }}}}}_{j}^{{{{\mathcal{l}}}}}\ne {{{\boldsymbol{0}}}}\), which is the null hypothesis of a standard ANOVA test for differences of group means from the grand mean. In this context, \({H}_{0j}^{{{{\mathcal{l}}}}}\) tests for population-specific differences in the disease association with SNP \(j\) from the population-averaged association. In our estimation of the grand mean in ANOVA, we used an inverse-variance weighting approach in which the kth weight for the jth SNP was equal to \({Var}{\left({\hat{\beta }}_{{jk}}\right)}^{-1}\). Under \({H}_{0j}^{{{{\mathcal{l}}}}},{T}_{j}^{{{{\mathcal{l}}}}}\sim {\chi }^{2}\left(p\right)\), \({S}_{M-{PH}}^{{{{\mathcal{l}}}}}\sim \Gamma \left(\alpha,\xi \right)\) approximately98,99, \(E\left({S}_{M-{PH}}^{{{{\mathcal{l}}}}},|,{H}_{0}^{{{{\mathcal{l}}}}}\right)={m}^{{{{\mathcal{l}}}}}p\), and \({Var}\left({S}_{M-{PH}}^{{{{\mathcal{l}}}}},|,{H}_{0}^{{{{\mathcal{l}}}}}\right):={\upsilon }^{{{{\mathcal{l}}}}}\) (see Supplementary Note 4 for the derivation of \({\upsilon }^{{{{\mathcal{l}}}}}\)). These results imply that \(\xi={m}^{{{{\mathcal{l}}}}}p/\upsilon\) and \(\alpha={\left({m}^{{{{\mathcal{l}}}}}p\right)}^{2}/{\upsilon }^{{{{\mathcal{l}}}}}\).
Multiple-ancestry gene-based test of population pleiotropy (MuGenT-Pleio)
The Multiple-ancestry Gene-based Test of Pleiotropy (MuGenT-Pleio) is used to infer a nonzero association between a gene and disease phenotype in all of \(p\) populations. It can also be applied to the study of multiple phenotypes in the same population for an analogous joint inference. Let \({\hat{\beta }}_{{jk}}^{{{{\mathcal{l}}}}}\) represent the estimated marginal association between the jth SNP corresponding to the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene and disease phenotype in the kth population. MuGenT-Pleio simply tests whether any SNP in the set of \({m}^{{{{\mathcal{l}}}}}\) corresponding to the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene is associated with the disease in all populations by applying an intersection-union test (IUT) to each SNP. The null hypothesis of MuGenT-Pleio is \({H}_{0}^{{{{\mathcal{l}}}}}:{\cap }_{j=1}^{{m}^{{{{\mathcal{l}}}}}}[{\cup }_{k=1}^{p}{\beta }_{{jk}}^{{{{\mathcal{l}}}}}\notin {{{{\mathscr{R}}}}}_{{jk}}]\) vs \({H}_{1}^{{{{\mathcal{l}}}}}:{\cup }_{j=1}^{{m}^{{{{\mathcal{l}}}}}}[{\cap }_{k=1}^{p}{\beta }_{{jk}}^{{{{\mathcal{l}}}}}\in {{{{\mathscr{R}}}}}_{{jk}}]\) where \({{{{\mathscr{R}}}}}_{{jk}}\) is the rejection region corresponding to the nominal significance level \(\alpha\) used to test each of \({H}_{0{jk}}^{{{{\mathcal{l}}}}}:{\beta }_{{jk}}^{{{{\mathcal{l}}}}}=0\) vs \({H}_{1{jk}}^{{{{\mathcal{l}}}}}:{\beta }_{{jk}}^{{{{\mathcal{l}}}}}\ne 0\). Since \({m}^{{{{\mathcal{l}}}}}\) SNPs are considered to correspond to the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene, and their LD correlation matrix is generally non-diagonal and potentially different between the \(p\) populations, we correct the nominal Type I error rate using the following relationship:
where \({m}_{{eff}.}^{{{{\mathcal{l}}}}}\) is the effective number of independent SNPs in the set of \({m}^{{{{\mathcal{l}}}}}\)100,101 (see Supplementary Note 5 for the derivation of Eq. 1). It can be seen from Eq. 1 that the achieved level \(\widetilde{\alpha }\) can be less or greater than the nominal levels which define \({{{{\mathscr{R}}}}}_{{jk}}\). We correct the nominal level of the standard IUT applied to \({m}^{{{{\mathcal{l}}}}}\) SNPs and \(p\) populations by expanding or contracting the original rejection regions to form \(\widetilde{{{{\mathscr{R}}}}}\) such that \(P[{\bigcup }_{j=1}^{{m}^{{{{\mathcal{l}}}}}}({\bigcap }_{k=1}^{p}{\beta }_{{jk}}^{{{{\mathcal{l}}}}}\in \widetilde{{{{\mathscr{R}}}}})|{H}_{0}^{{{{\mathcal{l}}}}}]\approx \alpha\). We show in Supplementary Note 5 that the quantile of a chi-square distribution defining \(\widetilde{{{{\mathscr{R}}}}}\) which achieves Type I error \(\alpha\) is approximately equal to
and that this quantile is smaller or larger than the nominal \({F}_{{\chi }^{2}\left(1\right)}^{-1}\left(1-\alpha \right)\) quantile depending on the relationship between \(p\) and \({m}_{{eff}.}^{{{{\mathcal{l}}}}}\).
Multiple-ancestry gene-based test of population-selective effects (MuGenT-Sel)
The Multiple-ancestry Gene-based Test of Selective effects (MuGenT-Sel) test is used to infer that a gene is associated with a trait in a subset of populations or a subset of traits in a single population. We will introduce MuGenT-Sel in the context of two traits in a single population. MuGenT-Sel is developed for inference between two traits and can be applied pairwise to all traits/populations under consideration. Denote \({\hat{\beta }_{{jk}}^{{{{\mathcal{l}}}}}}\) as the marginal estimate of association between the jth SNP of \({m}^{{{{\mathcal{l}}}}}\) corresponding to the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene and the kth of 2 traits and denote \({Z}_{{jk}}^{{{{\mathcal{l}}}}}\) as its corresponding Z-statistic used to test \({H}_{0{jk}}^{{{{\mathcal{l}}}}}:{\beta }_{{jk}}^{{{{\mathcal{l}}}}}=0\) in GWAS. Since the goal of MuGenT-Sel is to infer a lack of gene association for one trait only, we develop this method based on the probability of gene associations belonging to one component of a mixed distribution. Note, failure to reject \({H}_{0k}^{{{{\mathcal{l}}}}}:{\cup }_{j=1}^{{m}^{{{{\mathcal{l}}}}}}{\beta }_{{jk}}^{{{{\mathcal{l}}}}}=0\) is not sufficient evidence that \({H}_{0k}^{{{{\mathcal{l}}}}}\) should be regarded as true with fixed probability given a particular model and so we cannot simply evaluate the rejection and non-rejection of each \({H}_{0k}^{{{{\mathcal{l}}}}}\) to infer gene \({{{\mathcal{l}}}}\) is only associated with one trait. The null hypothesis of MuGenT-Sel is \({H}_{0}^{{{{\mathcal{l}}}}}:({\cap }_{k=1}^{2}{{{{\boldsymbol{\beta }}}}}_{k}^{{{{\mathcal{l}}}}}\ne {{{\boldsymbol{0}}}})\cup ({\cap }_{k=1}^{2}{{{{\boldsymbol{\beta }}}}}_{k}^{{{{\mathcal{l}}}}}={{{\boldsymbol{0}}}})\) vs \({H}_{1}^{{{{\mathcal{l}}}}}:{{{{\boldsymbol{\beta }}}}}_{k}^{{{{\mathcal{l}}}}}={{{\boldsymbol{0}}}}\cap {{{{\boldsymbol{\beta }}}}}_{s}^{{{{\mathcal{l}}}}}\ne {{{\boldsymbol{0}}}}\) for \(k\ne s\), where \({{{{\boldsymbol{\beta }}}}}_{k}^{{{{\mathcal{l}}}}}=({\beta }_{{jk}}^{{{{\mathcal{l}}}}})\). Conceptually, MuGenT-Sel evaluates the probability of \({H}_{1}^{{{{\mathcal{l}}}}}\) by comparing the marginal densities of each \({{{{\boldsymbol{\beta }}}}}_{k}^{{{{\mathcal{l}}}}}\) in a non-null distribution to the marginal densities in a null distribution.
Where \({{{{\bf{R}}}}}_{k}^{{{{\mathcal{l}}}}}\) is the LD matrix corresponding to the \(m\) SNPs assigned to the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene for the kth trait/population, the ‘null density’ for trait \(k\) is \(g\left({{{{\bf{z}}}}}_{k}^{{{{\mathcal{l}}}}};{{{{\bf{R}}}}}_{k}^{{{{\mathcal{l}}}}}\right)\), where \({{{{\bf{z}}}}}_{k}^{{{{\mathcal{l}}}}}=({Z}_{{jk}}^{{{{\mathcal{l}}}}})\) and \(g\left(\cdot ;{{{{\bf{R}}}}}_{k}^{{{{\mathcal{l}}}}}\right)\) is the density of a gamma distribution parameterized by \({{{{\bf{R}}}}}_{k}^{{{{\mathcal{l}}}}}\). The ‘non-null’ density for trait \(k\) is \(g\left({{{{\bf{z}}}}}_{k}^{{{{\mathcal{l}}}}};{{{{\bf{R}}}}}_{k}^{{{{\mathcal{l}}}}},{{{{\boldsymbol{\gamma }}}}}_{k}^{{{{\mathcal{l}}}}}\right)\), where \({{{{\boldsymbol{\gamma }}}}}_{k}^{{{{\mathcal{l}}}}}\) is a mean parameter vector for \({{{{\bf{z}}}}}_{k}^{{{{\mathcal{l}}}}}\) which has undergone LASSO shrinkage and maximizes the marginal penalized multivariate normal likelihood of \({{{{\bf{z}}}}}_{k}^{{{{\mathcal{l}}}}}\). We begin by calculating \({p}_{k}^{{{{\mathcal{l}}}}}=g\left({{{{\bf{z}}}}}_{k}^{{{{\mathcal{l}}}}};{{{{\bf{R}}}}}_{k}^{{{{\mathcal{l}}}}},{{{{\boldsymbol{\gamma }}}}}_{k}^{{{{\mathcal{l}}}}}\right){\left[g\left({{{{\bf{z}}}}}_{k}^{{{{\mathcal{l}}}}};{{{{\bf{R}}}}}_{k}^{{{{\mathcal{l}}}}},{{{{\boldsymbol{\gamma }}}}}_{k}^{{{{\mathcal{l}}}}}\right)+g\left({{{{\bf{z}}}}}_{k}^{{{{\mathcal{l}}}}};{{{{\bf{R}}}}}_{k}^{{{{\mathcal{l}}}}}\right)\right]}^{-1}\), which is the probability of \({{{{\bf{z}}}}}_{k}^{{{{\mathcal{l}}}}}\) belonging in the non-null distribution given \(\left({{{{\bf{R}}}}}_{k}^{{{{\mathcal{l}}}}},{{{{\boldsymbol{\gamma }}}}}_{k}^{{{{\mathcal{l}}}}}\right)\). Secondly, we transform \({p}_{k}^{{{{\mathcal{l}}}}}\) using a shrinkage operator and instead consider \({\pi }_{k}^{{{{\mathcal{l}}}}}=d\left({p}_{k}^{{{{\mathcal{l}}}}}\right)={\left(2{p}_{k}^{{{{\mathcal{l}}}}}-1\right)}_{+}\). The purpose of \(d\left(\cdot \right)\) is to transform probabilities of ½ to 0, the rationale being that genes which truly belong in their null distribution will only on average be assigned probability ½ of belonging in it, but we intend for their probability of belonging in the non-null distribution to be 0 in this case. We then calculate the probability the gene belongs in the non-null distribution for only one trait as \({\pi }_{1}^{{{{\mathcal{l}}}}}+{\pi }_{2}^{{{{\mathcal{l}}}}}-{2\pi }_{1}^{{{{\mathcal{l}}}}}{\pi }_{2}^{{{{\mathcal{l}}}}}\) and return it as the MuGenT-Sel result.
Gene-based test integrating xQTL information (xGenT)
The xQTL Gene-based association Test (xGenT) can be used to infer that SNPs in a gene-specific set share both disease and xQTL signals. xGenT performs this task by including a weighting matrix in the standard GenT test. This weighting matrix is proportional to the xQTL effect sizes, so when none are truly present the xGenT statistic will not be large. When there are both strong xQTL and disease GWAS signals in absolute magnitude for one or more SNPs, the xGenT statistic will be large. Let \({\hat{{{{\boldsymbol{\beta }}}}}^{{{{\mathcal{l}}}}}}={({\hat{\beta }_{1}^{{{{\mathcal{l}}}}}},\ldots,{\hat{\beta }}_{{m}^{{{{\mathcal{l}}}}}}^{{{{\mathcal{l}}}}})}^{\top }\) and \({\hat{{{{\boldsymbol{\nu }}}}}}_{k}^{{{{\mathcal{l}}}}}={\left({\hat{\nu }}_{1k}^{{{{\mathcal{l}}}}},\ldots,{\hat{\nu }}_{{m}^{{{{\mathcal{l}}}}}k}^{{{{\mathcal{l}}}}}\right)}^{\top }\) represent the \({m}^{{{{\mathcal{l}}}}}\)-length vectors of estimated marginal associations between the \(m\) SNPs corresponding to the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene and the disease phenotype and the kth gene product (e.g., gene expression, protein abundance), respectively. Since there are potentially many gene products and/or tissues which researchers may want to consider as contexts for a potentially causal effect, we let \(k=1,\ldots,p\), define the set \({{{\mathscr{X}}}}\) to represent them, and combine all xQTL association estimates into the \({m}^{{{{\mathcal{l}}}}}\times p\) matrix \({\hat{{{{\bf{N}}}}}}_{{{{\mathcal{l}}}}{{{\mathscr{X}}}}}=({\hat{{{{\boldsymbol{\nu }}}}}}_{k})\). Examples may include gene expression in multiple tissues, gene expression and protein abundance in a single tissue, or their combination. Let \({{{{\boldsymbol{\theta }}}}}_{{{{\mathcal{l}}}}{{{\mathscr{X}}}}}\) represent the \(p\)-length vector of total/marginal causal effects which gene products in the set \({{{\mathscr{X}}}}\) have on the disease phenotype. The null hypothesis that xGenT tests is \({H}_{0}^{{{{\mathcal{l}}}}{{{\mathscr{X}}}}}:{{{{{\bf{N}}}}}_{{{{\mathcal{l}}}}{{{\mathscr{X}}}}}{{{\boldsymbol{\theta }}}}}_{{{{\mathcal{l}}}}{{{\mathscr{X}}}}}={{{\boldsymbol{0}}}}\) vs \({H}_{1}^{{{{\mathcal{l}}}}{{{\mathscr{X}}}}}:{{{{{\bf{N}}}}}_{{{{\mathcal{l}}}}{{{\mathscr{X}}}}}{{{\boldsymbol{\theta }}}}}_{{{{\mathcal{l}}}}{{{\mathscr{X}}}}}{{{\boldsymbol{\ne }}}}{{{\boldsymbol{0}}}}\) using the statistic \({S}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}}={{{{\bf{z}}}}}_{{{{\mathcal{l}}}}}^{\top }{{{{\bf{L}}}}}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}}{{{{\bf{z}}}}}_{{{{\mathcal{l}}}}}\) where \({{{{\bf{z}}}}}_{{{{\mathcal{l}}}}}=[{\hat{\beta }_{j}^{{{{\mathcal{l}}}}}}/{{{\rm{SE}}}}({\hat{\beta }_{j}^{{{{\mathcal{l}}}}}})],\) and \({{{{\bf{L}}}}}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}}={\sum }_{k=1}^{p}{\hat{{{{\boldsymbol{\nu }}}}}}_{k}^{{{{\mathcal{l}}}}}{\left({\hat{{{{\boldsymbol{\nu }}}}}}_{k}^{{{{\mathcal{l}}}}}\right)}^{\top }\). It follows that \(E\left({S}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}},|,{H}_{0}^{{{{\mathcal{l}}}}},{{{{\bf{L}}}}}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}}\right)={m}^{{{{\mathcal{l}}}}}\) and \({Var}\left({S}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}},|,{H}_{0}^{{{{\mathcal{l}}}}},{{{{\bf{L}}}}}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}}\right)=2{{{\rm{tr}}}}\left({{{{\bf{L}}}}}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}}{{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}{{{{\bf{L}}}}}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}}{{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}\right)\) where \({{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}\) is the \({m}^{{{{\mathcal{l}}}}}\times {m}^{{{{\mathcal{l}}}}}\) matrix of LD correlations between the \({m}^{{{{\mathcal{l}}}}}\) SNPs corresponding to the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene. This implies that \({S}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}}|{H}_{0}^{{{{\mathcal{l}}}}},{{{{\bf{L}}}}}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}}\sim \Gamma \left(\alpha,\xi \right)\) approximately98,99 where \(\xi={m}^{{{{\mathcal{l}}}}}/{{{\rm{tr}}}}\left(2{{{{\bf{L}}}}}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}}{{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}{{{{\bf{L}}}}}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}}{{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}\right)\) and \(\alpha={m}^{{{{\mathcal{l}}}}}\xi\). Though the causal effects are neither known nor estimable without performing MR, under the assumption of no horizontal pleiotropy, \({{{{\boldsymbol{\beta }}}}}_{{{{\mathcal{l}}}}}={{{{{\bf{N}}}}}_{{{{\mathcal{l}}}}{{{\mathscr{X}}}}}{{{\boldsymbol{\theta }}}}}_{{{{\mathcal{l}}}}{{{\mathscr{X}}}}}\). This implies that under this assumption \({S}_{{{{\mathscr{X}}}}}^{{{{\mathcal{l}}}}}\) tests total/marginal causality since \(E({\hat{\beta }_{j}^{{{{\mathcal{l}}}}}}{\hat{\nu }_{{jk}}^{{{{\mathcal{l}}}}}})=E({\hat{\beta }}_{j}^{{{{\mathcal{l}}}}})E({\hat{\nu }}_{{jk}}^{{{{\mathcal{l}}}}})={({\nu }_{{jk}}^{{{{\mathcal{l}}}}})}^{2}{\theta }_{{{{\mathcal{l}}}}k}\Rightarrow E({S}_{{xG}}^{{{{\mathcal{l}}}}}|{H}_{0}^{{{{\mathcal{l}}}}})\propto {{{{\boldsymbol{\theta }}}}}_{{{{\mathcal{l}}}}{{{\mathscr{X}}}}}\) (see Supplementary Note 6 for additional details).
However, like GenT and MuGenT, xGenT is primarily intended to be used as a screening tool and not to evaluate causality. In the previous subsection, we stated that the xGenT test statistic could be proportional to a total/marginal causal effect of (e.g.,) gene expression on disease risk only when no horizontal pleiotropy is present. xGenT in its standard form cannot evaluate the extent to which horizontal pleiotropy is present since it is a single-gene screening tool which evaluates association evidence and so should not alone be used to infer causality. A rejected xGenT null hypothesis can generally be interpreted to mean that there is a shared signal between disease SNPs and xQTLs in a gene-specific set, though stricter analytical tools such as MR, transcriptome-wide association study (TWAS), and colocalization should be used if the goal of the analysis is to make a causal inference regarding the relationship between a gene product and disease risk. We demonstrate this sequential testing strategy in Fig. 5b, in which we first screened the genome using xGenT then performed follow-up MR analyses on genes with supporting evidence from xGenT. These follow-up analyses provided supporting evidence of potential causality for many genes prioritized by xGenT, but still xGenT alone should not be used to make a causal inference in practice.
Defining disease-associated druggable genes in novel loci
For T2D, we had access to ancestry-specific and trans-ancestry GWAS summary statistics. To define the set of previously detected T2D-associated genes from existing GWAS, we began by identifying lead SNPs present in the trans-ancestry GWAS. To do this, we used the --clump procedure in PLINKv1.987 five separate times with each of the ancestry-specific LD reference panels from the 1000 G Phase 323. This provided five sets of lead SNPs in the trans-ancestry T2D GWAS, from which we formed the union set of and regarded all genes within \(\pm\)1 Mb of any of these lead SNPs are previously detected T2D-associated genes. For SCZ, ALS, and MDD, we followed the same procedure but used European LD reference panels. For AD, we started with the complete list of genes identified in ref. 36 –the largest AD GWAS to date–and regarded the set of previously known AD genes as those within \(\pm\)1 Mb of any genes identified by ref. 36. We used the published list of identified genes because the publicly available set of GWAS summary statistics from36 contains less genes identified at the level of genome-wide significance than those reported in the published set.
To provide supporting evidence of association for druggable genes we detected in novel loci, we performed fine-mapping of SNPs in each gene-specific set using the sum of single effects (SuSiE) method28. SuSiE intends to construct sets of SNPs which have fixed probability of containing a causal SNP, and we performed it using the susieR package in R28,102,103. SuSiE used ancestry-specific LD reference panels from 1000 Genomes Phase 323 and included all SNPs within \(\pm\)100Kb of the midpoint of each gene by base pair position. All raw LD matrices, denoted \({{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}\) for the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene, were regularized by \({{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}\leftarrow 0.99\times {{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}+0.01\times {{{{\bf{I}}}}}^{{{{\mathcal{l}}}}}\). All SuSiE parameters were fixed at their default values. SuSiE results are interpreted as supporting evidence because the null hypothesis of no association between any SNP in the gene-specific SNP set and the disease – i.e., the null hypothesis which GenT tests – is tested using a completely different form of inference. These results are not interpreted as replications because both SuSiE and gene-based testing use some of the same data.
Estimating the T2D between-population genetic correlation
We estimated the pairwise genetic correlations between EUR, AFR, EAS, SAS, and HIS populations on their measured Type 2 diabetes using the Popcorn method33. We used LD reference panels from the 1000 Genomes Phase 323 European (matched to the EUR T2D cohort), African (matched to AFR), East and South Asian (matched to EAS and SAS, respectively), and Admixed American (matched to HIS) populations. The full code we used to perform these analyses is available at https://github.com/noahlorinczcomi/gent_analysis.
Estimating the number of participants needed to detect SYK
We estimated the number of additional participants needed to detect the association between SYK and AD in the publicly available AD GWAS summary statistics from36 in \(n\) total participants. We assumed detection of SYK by the lead SNP rs10512201, which respectively had estimated effect size and standard error of -0.439 \((\hat{\beta })\) and 0.0103 \(\left(s\right)\) in the GWAS summary statistics. We assumed \(s\propto {n}^{-1/2}\) and transformed \(\hat{\beta }\) using \(g\) accordingly such that \(g(\hat{\beta })\sim N(g[\beta ],{n}^{-1})\), defined the genome-wide significance threshold for the statistic \(g{(\hat{\beta })}^{2}n\) as \(29.71\approx Q={F}_{{\chi }_{1}^{2}}^{-1}\)(1-5E-8) and searched a grid of non-centrality parameters \({\lambda }_{k}=g{\left(\beta \right)}^{2}{n}_{k}\) defined by hypothetical sample size \({n}_{k}\) until \(1-{F}_{{\chi }_{1,\lambda }^{2}}\left(Q\right)\) was equal at least to 0.9, or 90% power to detect the lead SYK SNP at the level of genome-wide significance. The R code used to perform this analysis is available at our Github.
Simulations
We performed simulations to confirm the null distributions of GenT, MuGenT, MuGenT-PH, MuGenT-Pleio, MuGenT-Sel, and xGenT were correctly specified and present most of these results in the Supplementary Note 14. Primary results are presented in the top panels of Figs. 2 to 4. With the exception of head-to-head GenT vs MAGMA simulations performed in Supplementary Note 14, we describe the general procedure for generating all other simulated data here and present all details of the simulation procedures in the Supplementary Note 14. All R code used to reproduce our simulation results is available from our Github repository: (https://github.com/noahlorinczcomi/gent_analysis). Since all of our methods are gene-based, we generated GWAS summary statistics which corresponded to a single gene over at least 1000 replications. We generated all marginal SNP association estimates by transforming their corresponding joint association estimates, which were generated by a multivariate normal distribution with mean vector \({{{{\bf{b}}}}}^{{{{\mathcal{l}}}}}\) and \({\left(n{{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}\right)}^{-1}\) for the \({{{\mathcal{l}}}}{{{\rm{th}}}}\) gene, each of which varied in dimension by the specific simulation setup. The vector \({{{{\bf{b}}}}}^{{{{\mathcal{l}}}}}\) is the vector of true joint associations of the SNPs with the trait and was in all cases made to be sparse. Its nonzero elements were selected randomly and, in the case of simulations with MuGenT, were made to be the same across populations. The matrix \({{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}\) was the LD matrix and for many results presented in the main text had a first-order autoregressive structure with correlation parameter equal to 0.9, which we denote as AR1(0.9) in Figs. 2–4. This means that two simulated SNPs separated by \(k\) base pairs had LD correlation of 0.9k.
We also tested a compound symmetry structure of the simulated LD matrices and present those results in the Supplementary Note 12, which are consistent with all simulation results presented in the main text. In Supplementary Note 12, we compare the Type I error rate and power of GenT to those of existing methods such as VEGAS14, GATES104, ACAT105, and exset106. These results demonstrate that on average GenT has a Type I error which is on average better controlled at nominal levels than GATES, VEGAS, and ACAT, and exhibits similar Type I error control to exset across a range of simulation scenarios (Supplementary Fig. 17). GenT has power to detect gene-level associations that is comparable to exset and VEGAS, which on average was less than the power of GATES and ACAT when power was low. However, we note that these two methods have slightly inflated Type I error and so a comparison of their power to that of GenT, VEGAS, and exset it not fair. On average, GenT completes in 0.1 sec for a single gene with 500 SNPs.
In the MuGenT simulation results presented in Fig. 3, we did not assume all LD matrices to be of the same structure across populations. The parameter \(n\) was the GWAS sample size(s). After fixing \(({{{{\bf{b}}}}}^{{{{\mathcal{l}}}}},{{{{\bf{R}}}}}^{{{{\mathcal{l}}}}})\), we generated \({\hat{{{{\bf{b}}}}}^{{{{\mathcal{l}}}}}}\), and in Fig. 2\({\hat{{{{\bf{R}}}}}}^{{{{\mathcal{l}}}}}\), then the marginal SNP association estimates \({\hat{{{{\boldsymbol{\beta }}}}}^{{{{\mathcal{l}}}}}}\) as \({\hat{{{{\boldsymbol{\beta }}}}}^{{{{\mathcal{l}}}}}}={{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}{\hat{{{{\bf{b}}}}}^{{{{\mathcal{l}}}}}}\) and their corresponding variances from \({\chi }_{n-1}^{2}/({n}^{2}-n)\) where \({\chi }_{a}^{2}\) is a central chi-squared distributed variate with \(a\) degrees of freedom. This process assumes the genotypes and phenotype are each standardized to have mean 0 and unit variance. Marginal Z-statistics for each SNP were then calculated using the quotient of the estimated marginal effect sizes and their corresponding standard errors. False and true positive rates were determined by calculating the proportion of test results which passed the nominal 5% significance threshold across all simulation replicates. When we estimated the Type I error rate of GenT when the number of individuals in the LD reference panel was finite, we randomly drew \({\hat{{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}}\) from a Wishart distribution parameterized by \({n}_{{{{\rm{ref}}}}.}\) and \({{{{\bf{R}}}}}^{{{{\mathcal{l}}}}}\), respectively the LD reference panel sample size and matrix of true LD correlations and transformed it to be a correlation matrix.
We restate that GenT and its extensions are marginal gene-based association tests which use SNP sets formed from SNPs in and around each gene body. One test is performed for each gene, and pairs of gene-specific SNP sets may contain SNPs which are in LD with each other. We show in Supplementary Note 8 that this induces correlation between gene-based test statistics which are in the same loci. The effect of shared LD between SNP sets on inflation may be more severe than that which is encountered in SNP-level analyses. Two comments on inflation are worth considering: (i) inflation is not necessarily indicative of inflated Type I error, (ii) for a fixed number of causal genes, the degree of observed genome-wide inflation is proportional to the degree to which genes are localized within gene-dense loci across the genome. Point (i) holds because a gene-based association test is of a marginal null hypothesis, the rejection of which is not incorrect if the gene is non-causal but in LD with a causal gene. Point (ii) In our simulations of Type I error described above, we are evaluating the frequency with which the gene-based null hypothesis for a single gene is incorrectly rejected at a fixed Type I error rate. These simulations are not intended to evaluate the extent to which GenT and its extensions can identify causal genes in a locus. We introduced the gene-based fine-mapping method above and show its simulation performance in Supplementary Note 9.
Simulations performed to make a head-to-head comparison of GenT to MAGMA were specially designed to be compatible with the Functional Mapping and Annotation of Genome-Wide Association Studies (FUMA) web tool107 and the current MAGMA v1.10 software using default parameters12. These simulations are completely described in Supplementary Note 14. These simulations generated data using estimated LD matrices from the 1000 Genomes Phase 3 European reference panel23. Both GenT and MAGMA formed SNP-gene pairs by assigning all SNPs which were available in the LD reference and within 50Kb of gene start/end positions (hg19) to gene-specific sets. Simulations evaluated Type I and Type II error (power) by generating data for 18,160 genes genome-wide a single time. We then calculated the proportions of rejected gene-based null hypotheses across the genome. In power simulations, we fixed the power of the GenT test for each gene at 0.50 and compared MAGMA power to it (cf. Supplementary Note 14).
Mendelian randomization
We performed univariable Mendelian Randomization treating bulk gene expression or protein abundance in select brain tissues as the exposure and AD risk as the outcome and presented the results in Fig. 4f and Fig. 5b. IVs in each MR analysis were selected as the cis-xQTLs with P-values less than \(5\times {10}^{-8}\). We used the vectors of marginal association Z-statistics as inputs to MR as in ref. 108 and we controlled for LD between eQTL IVs by pre-transforming each Z-statistic vector to remove it. Let \(({{{{\bf{z}}}}}_{{jk}}^{x},{{{{\bf{z}}}}}_{{jk}}^{y})\) represent the tuple of exposure \((x)\) and outcome \((y)\) marginal, untransformed Z-statistic vectors for the jkth exposure-AD pair.
We estimated LD between the corresponding \({m}_{{jk}}\) SNPs using the 505 European samples in the 1000 Genomes Phase 3 reference panel23 and denote the estimated matrix as \({\hat{{{{\bf{R}}}}}}_{{jk}}\). Pre-transforming \(({{{{\bf{z}}}}}_{{jk}}^{x},{{{{\bf{z}}}}}_{{jk}}^{y})\) to remove LD meant multiplying \(({{{{\bf{z}}}}}_{{jk}}^{x},{{{{\bf{z}}}}}_{{jk}}^{y})\) by \({{{\bf{F}}}}\) where \({{{\bf{F}}}}\propto {{{\bf{G}}}}\) and \({{{\bf{GG}}}}={\hat{{{{\bf{R}}}}}}_{{jk}}^{-1}\). Kollo & van Rosen109 proved that \({{{\rm{E}}}}({\hat{{{{\bf{R}}}}}}_{{jk}}^{-1})\approx {m}_{{jk}}{(505-{m}_{{jk}}-1)}^{-1}{{{{\bf{R}}}}}_{{jk}}^{-1}={c}_{{jk}}{{{{\bf{R}}}}}_{{jk}}^{-1}\) and so we used \({{{\bf{F}}}}={\alpha }_{{jk}}{{{\bf{G}}}}\) to remove LD structure where \({\alpha }_{{jk}}=\max (0.5,{c}_{{jk}})\). We used the MRBEE method to estimate causal effects, which controls for bias from horizontal pleiotropy and weak instruments108. Supplementary Note 16 presents additional MR results for the xGenT-prioritized genes displayed in Fig. 5b. These results compared the causal estimates made by MRBEE to those from 11 other MR methods in the MendelianRandomization R package (v0.8.0110). These results suggest overall agreement between MRBEE and other methods which address MR assumptions using different tools.
NTRK1 experiments
We targeted NTRK1 with a selective NTRK1 protein (TrKA) inhibitor, GW441756111, in human iPSC-derived neurons. GW441756 was purchased from Selleckchem.com (Cat #S2891) with a purity of 99.53%, which selectively targets TrKA but not TrKB, TrKC, or others112. Cells were cultured in a 37 °C, 5% CO2 humidified incubator. To generate mature neurons, AD patient iPSCs (IUGB269.1 and IUGB416) from NCRAD (National Centralized Repository for Alzheimer’s Disease) were amplified and maintained in mTeSR medium (STEMCELL Technologies). The whole process for deafferenting iPSCs to mature neurons took 35 days. At day 0, iPSCs were dissociated with accutase into single cells, suspended with neural induction medium (STEMCELL Technologies) plus ROCK inhibitor (Y−27632, STEMCELL Technologies), and plated onto Matrigel coated plates with a density of 0.5 × 105 cells/cm2. At day 6 and 12, cells were passaged with accutase into single cells, suspended with neural induction medium plus ROCK inhibitor, and replated onto Matrigel coated plates with a density of 0.5 × 105 cells/cm2. At day 18, cells were passaged with accutase into single cells, suspended with neural induction medium plus ROCK inhibitor, and replated onto Matrigel coated plates with a density of 1.0 × 105 cells/cm2.
From day1−18, cell medium was changed to fresh neural induction medium without ROCK inhibitor every day for neuronal progenitor cell induction. From day 19-24, cell medium was changed to fresh forebrain neuron differentiation medium (STEMCELL Technologies) every day for neuronal precursor differentiation. At day 25, cells were dissociated with accutase into single cells, suspended with forebrain neuron maturation medium (STEMCELL Technologies), and plated onto Matrigel coated plates with a density of 1.5 × 105 cells/cm2. Cells were matured into a mixed population of excitatory and inhibitory forebrain-type neurons for 10 days with medium changed every two days. Patient iPSC-derived neurons in 12-well plates were treated with the NTRK1 inhibitor (GW441756) for 24 h with the indicated doses (0, 1, 10, and 20 μM in DMSO).
Neurofibrillary tangles (NFTs) are intracellular aggregates of hyperphosphorylated Tau proteins and are hallmarks of AD and other neurodegeneration diseases113. Patient iPSC-derived neurons display considerable level of phosphorylated Tau114. Here we used Western blot to evaluate the effect of NTRK1 inhibitor (GW441756) on Tau phosphorylation. Cells were lyzed in N-PER buffer (Thermo Fisher Scientific) supplemented with Protease/Phosphatase Inhibitor Cocktail (Thermo Fisher Scientific). Total protein was collected, and protein concentration were measured with Pierce BCA Protein Assay Kit (Thermo Fisher Scientific). 10ug protein were loaded into Mini-PROTEAN TGX Precast Gels (Bio-Rad) for Western blot analysis with chemiluminescence system. Primary antibodies for Western blot were as follows: rabbit anti-p-Tau 217 (Cell Signaling Technology, #51625S), 1:1000; rabbit anti-p-Tau 181 (Cell Signaling Technology, #12885S), 1:1000; rabbit anti-Tau (Cell Signaling Technology, #46687S), 1:1000; mouse anti-β-Actin (Proteintech, #66009−1-Ig), 1:1000. Primary antibodies were diluted in PBST/5% nonfat dry milk (PBST: PBS + 0.1% Tween 20) and incubated overnight at 4 °C. Secondary antibodies were diluted in PBST/5% milk and incubated for 30 min at room temperature. Western blot signals were detected using West Pico or West Femto (Thermo Fisher Scientific) ECL substrate. Band intensity was quantified using NIH ImageJ software and used to indicate protein levels.
Ethics
Our study complied with all relevant regulations regarding the use of human study participants and received approval from the Institutional Review Board (IRB) at the Cleveland Clinic Foundation, Cleveland, OH and was conducted in accordance to the criteria set by the Declaration of Helsinki.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Supporting Data are available from Zenodo [https://doi.org/10.5281/zenodo.18121206]. We provided gene-based association test results from GenT for AD, ALS, MDD, SCZ in Supporting Data 1-4; multi-ancestry gene-based association testing with MuGenT for T2D in Supporting Data 5; gene-based population heterogeneity testing with MuGenT-PH for T2D in Supporting Data 6; brain eQTL- and pQTL-integrated gene-based association testing for AD in Supporting Data 7 and 8, respectively; and all fine-mapped SNPs for AD, ALS, MDD, SCZ, and all T2D populations in Supporting Data 9−17. We created a web application (https://nlorinczcomi.shinyapps.io/gent/) to interactively query and download all gene-based association test and fine-mapping results with GenT and SuSiE for the 32 phenotypes listed in Supporting Data 18. Fine-mapped GenT results for five complex diseases are provided in Supporting Data 19-23. In our main analyses, GWAS summary statistics were downloaded from public repositories. AD GWAS summary statistics are under accession code GCST90027158; GWAS summary statistics for SCZ are available on Figshare [https://doi.org/10.6084/m9.figshare.19426775]; GWAS summary statistics for ALS are available under accession code GCST90027164; GWAS summary statistics for MDD are available on https://datashare.ed.ac.uk/handle/10283/3203; GWAS summary statistics for T2D are available from https://www.diagram-consortium.org/downloads.html (accession label: “Multi-ancestry GWAS meta-analysis summary statistics”). All other repositories from which we downloaded GWAS summary statistics for phenotypes whose GenT results are presented at (https://nlorinczcomi.shinyapps.io/gent/) are available in Supporting Data 18. GWAS summary statistics for xQTL in cortex, cerebellum, frontal cortex, hippocampus, and spinal cord tissues were downloaded from the ‘eQTL Tissue-Specific All SNP Gene Associations’ section of (https://gtexportal.org/home/downloads/adult-gtex/qtl) (v8). ROSMAP data are available at the AD Knowledge portal under restricted access: https://adknowledgeportal.synapse.org. Supplementary Figs. 1–33 provide additional simulation and real data analysis examples to further demonstrate and evaluate the performance of our gene-based testing approaches. Supplementary Figs. 34–37 provide additional diagnostic results for MuGenT testing in T2D. Supplementary Figs. 38–51 display locus-specific plots for a set of genes which were significant in xGenT testing with AD. Supplementary Figs. 52–68 display their local genetic correlations across multiple phenotypes and GTEx v8 tissues, and Supplementary Figs. 69–84 display the distributions of their eQTL effect sizes across all GTEx tissues. Supplementary Fig. 85 demonstrates between-ancestry heterogeneity of LD patterns in the KIF11 locus. Supplementary Fig. 86 displays a cross-tissue network of local genetic correlations in the RIPK2 locus. Supplementary Fig. 87 shows cell-type specific expression of three candidate AD-associated genes using ROSMAP data and identified with xGenT, and Supplementary Fig. 88 shows a set of protein-protein interactions involving NTRK1. Source Data are provided with this article. Source data are provided with this paper.
Code availability
All code used to generate our simulation and real data analysis results is available at (https://github.com/noahlorinczcomi/gent_analysis) and is deposited in Zenodo [https://doi.org/10.5281/zenodo.17449888]. We have created an R package for researchers to apply GenT, MuGenT, MuGenT-PH, MuGenT-Pleio, MuGenT-Sel, and xGenT to their own data which is available from (https://github.com/noahlorinczcomi/gent), where tutorials are available. The R package we used for data analysis is available from Zenodo [https://doi.org/10.5281/zenodo.17449854]. We made all summary gene-based association testing and fine-mapping results from analyses from 32 phenotypes available for query and download from (https://nlorinczcomi.shinyapps.io/gent).
References
Claussnitzer, M. et al. A brief history of human disease genetics. Nature 577, 179–189 (2020).
Uffelmann, E. et al. Genome-wide association studies. Nat. Rev. Meth. Prim. 1, 59 (2021).
Tam, V. et al. Benefits and limitations of genome-wide association studies. Nat. Rev. Genet. 20, 467–484 (2019).
Gusev, A. et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat. Genet. 48, 245–252 (2016).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
Farh, K. K. H. et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature 518, 337–343 (2015).
Reay, W. R. et al. Advancing the use of genome-wide association studies for drug repurposing. Nat. Rev. Genet. 22, 658–671 (2021).
Lago, S. G. et al. The druggable schizophrenia genome: from repurposing opportunities to unexplored drug targets. NPJ Gen. Med. 7, 25 (2022).
Qi, H. et al. New “drugs and targets” in the GWAS era of bipolar disorder. Bipolar Disord. 25, 410–421 (2023).
Huang, H. et al. Gene-based tests of association. PLoS Genet 7, e1002177 (2011).
Lamparter, D. et al. Fast and rigorous computation of gene and pathway scores from SNP-based summary statistics. PLoS Comput. Biol. 12, e1004714 (2016).
de Leeuw, C. A. et al. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Vsevolozhskaya, O. A. et al. DOT: Gene-set analysis by combining decorrelated association statistics. PLoS Comput. Biol. 16, e1007819 (2020).
Liu, J. Z. et al. A versatile gene-based test for genome-wide association studies. Am. J. Hum. Genet. 87, 139–145 (2010).
Berrandou, T. E. et al. LDAK-GBAT: fast and powerful gene-based association testing using summary statistics. Am. J. Hum. Genet. 110, 23–29 (2023).
Zhang, W. et al. Partitioning gene-based variance of complex traits by gene score regression. PLoS One 15, e0237657 (2020).
Hof, J. P. et al. LDAK-KVIK performs fast and powerful mixed-model association analysis of quantitative and binary phenotypes. Nat. Genet 57, 2116–2123 (2025).
Hecker, J. et al. Reporting correct p values in VEGAS analyses. Twin Res. Hum. Genet. 20, 257–259 (2017).
Yurko, R. et al. H-MAGMA, inheriting a shaky statistical foundation, yields excess false positives. Ann. Hum. Genet. 85, 97–100 (2021).
Barbeira, A. N. et al. Exploring the phenotypic consequences of tissue specific gene expression variation inferred from GWAS summary statistics. Nat. Comms. 9, 1825 (2018).
Gerring, Z. F. et al. A gene co-expression network-based analysis of multiple brain tissues reveals novel genes and molecular pathways underlying major depression. PLoS Genet 15, e1008245 (2019).
Gkatzionis, A. et al. Statistical methods for cis-Mendelian randomization with two-sample summary-level data. Genet. Epidemiol. 47, 3–25 (2023).
1000 Genomes Project Consortium. A global reference for human genetic variation. Nature. 526, 68 (2015).
Gaulton, A. et al. The ChEMBL database in 2017. NAR 45, D945–D954 (2017).
Gilson, M. K. et al. BindingDB in 2015: a public database for medicinal chemistry, computational chemistry and systems pharmacology. Nucleic Acids Res 44, D1045–D1053 (2016).
Harding, S. D. et al. The IUPHAR/BPS guide to PHARMACOLOGY in 2022: curating pharmacology for COVID−19, malaria and antibacterials. Nucleic Acids Res 50, D1282–D1294 (2022).
Wishart, D. S. et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res 46, D1074–D1082 (2018).
Zou, Y. et al. Fine-mapping from summary data with the “Sum of Single Effects” model. PLoS Genet 18, e1010299 (2022).
Kosoy, R. et al. Genetics of the human microglia regulome refines Alzheimer’s disease risk loci. Nat. Genet. 54, 1145–1154 (2022).
Phan, K. et al. Multiple pathways of lipid dysregulation in amyotrophic lateral sclerosis. Brain Comms 5, 340 (2023).
Zhen, L. et al. EphB2 deficiency induces depression-like behaviors and memory impairment: involvement of NMDA 2B receptor dependent signaling. Front. Pharmacol. 9, 862 (2018).
Choi, S. J. et al. A schizophrenia-related deletion leads to KCNQ2-dependent abnormal dopaminergic modulation of prefrontal cortical interneuron activity. Cereb. Cortex. 28, 2175–2191 (2018).
Brown, B. C. et al. Transethnic genetic-correlation estimates from summary statistics. Am. J. Hum. Genet. 99, 76–88 (2016).
Ortiz, R. et al. Type 2 diabetes and cardiometabolic risk may be associated with increase in DNA methylation of FKBP5. Clin. Epigenet. 10, 1–8 (2018).
Huang, Q. Y. et al. Cdc42: a novel regulator of insulin secretion and diabetes-associated diseases. Int. J. Mol. Sci. 20, 179 (2019).
Bellenguez, C. et al. New insights into the genetic etiology of Alzheimer’s disease and related dementias. Nat. Genet. 54, 412–436 (2022).
Bennett, D. A. et al. Religious orders study and rush memory and aging project. J. Alz. Dis. 64, S161–S189 (2018).
Nagele, R. G. et al. Contribution of glial cells to the development of amyloid plaques in Alzheimer’s disease. Neurobiol. Aging 25, 663–674 (2004).
Uddin, M. S. et al. Glial cells in Alzheimer’s disease: from neuropathological changes to therapeutic implications. Ageing Res. Rev. 78, 101622 (2022).
The UniProt Consortium. UniProt: the Universal Protein Knowledgebase in 2023. Nucleic Acids Res. 51, 523–531 (2023).
Savas, J. N. et al. Amyloid accumulation drives proteome-wide alterations in mouse models of Alzheimer’s disease-like pathology. Cell Rep. 21, 2614–2627 (2017).
Magistri, M. et al. Transcriptomics profiling of Alzheimer’s disease reveal neurovascular defects, altered amyloid-β homeostasis, and deregulated expression of long noncoding RNAs. J. Alzheimer’s. Dis. 48, 647–665 (2015).
Zhou, Y. et al. AlzGPS: a genome-wide positioning systems platform to catalyze multi-omics for Alzheimer’s drug discovery. Alzheimer’s. Res. Ther. 13, 1–13 (2021).
Liu, K. et al. Human peripheral blood-derived mesenchymal stem cells with NTRK1 over-expression enhance repairing capability in a rat model of Parkinson’s disease. Cytotech 70, 1291–1299 (2018).
Cozza, A. et al. SNPs in neurotrophin system genes and Alzheimer’s disease in an Italian population. J. Alzheimer’s. Dis. 15, 61–70 (2008).
Hall, B. E. et al. Transcriptome analysis of rheumatoid arthritis uncovers genes linked to inflammation-induced pain. Sci. Rep. 14, 25893 (2024).
de Frutos Lucas, J. et al. Investigating neurotrophin genetics and hippocampal volume. bioRxiv https://doi.org/10.1101/2022.01.01.474700 (2022).
Yang, K. et al. NTRK1 knockdown induces mouse cognitive impairment and hippocampal neuronal damage through mitophagy suppression via inactivating the AMPK/ULK1/FUNDC1 pathway. Cell Death Discov. 9, 404–419 (2023).
Raney, B. J. et al. The UCSC genome browser database: 2024 update. Nucleic Acids Res 52, 1082–1088 (2024).
Maccioni, R. B. et al. The protein kinase cdk5: structural aspects, roles in neurogenesis and involvement in Alzheimer’s pathology. Eur. J. Biochem. 268, 1518–1527 (2001).
Liu, S. L. et al. The role of Cdk5 in Alzheimer’s disease. Mol. Neurobiol. 53, 4328–4342 (2016).
Wehrman, T. et al. Structural and mechanistic insights into nerve growth factor interactions with the TrkA and p75 receptors. Neuron 53, 25–38 (2007).
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Sondka, Z. et al. The COSMIC Cancer Gene Census: describing genetic dysfunction across all human cancers. Nat. Rev. Cancers 18, 696–705 (2018).
Indo, Y. Molecular basis of congenital insensitivity to pain with anhidrosis (CIPA): mutations and polymorphisms in TRKA (NTRK1) gene encoding the receptor tyrosine kinase for nerve growth factor. Hum. Mutat. 18, 462–471 (2001).
Sjöstedt, E. et al. An atlas of the protein-coding genes in the human, pig, and mouse brain. Science. 367, eaay5947 (2020).
Wang, S. et al. TREM2 drives microglia response to amyloid-β via SYK-dependent and-independent pathways. Cell 185, 4153–4169 (2022).
Hampel, H. et al. The amyloid-β pathway in Alzheimer’s disease. Mol. Psychiat. 26, 5481–5503 (2021).
Davies, G. et al. Genetic contributions to variation in general cognitive function: a meta-analysis of genome-wide association studies in the CHARGE consortium (N= 53,949). Mol. Psychiat. 20, 183–192 (2015).
Demenais, F. et al. Multiancestry association study identifies new asthma risk loci that colocalize with immune-cell enhancer marks. Nat. Genet. 50, 42–53 (2018).
Allaire, P. et al. Genetic and clinical determinants of telomere length. HGG Advances. 4, 100201 (2023).
Pickrell, J. K. et al. Detection and interpretation of shared genetic influences on 42 human traits. Nat. Genet. 48, 709–717 (2016).
Sakaue, S. et al. A cross-population atlas of genetic associations for 220 human phenotypes. Nat. Genet. 53, 1415–1424 (2021).
Richard, S. et al. Thyroid hormone action during GABAergic neuron maturation: The quest for mechanisms. Front. Endocrin. 14, 1–13 (2023).
Jones, P. et al. Child developmental risk factors for adult schizophrenia in the British 1946 birth cohort. Lancet 344, 1398–1402 (1994).
Murray, R. M. et al. 30 years on: how the neurodevelopmental hypothesis of schizophrenia morphed into the developmental risk factor model of psychosis. Schiz. Bull. 43, 1190–1196 (2017).
Janoutová, J. et al. Epidemiology and risk factors of schizophrenia. Neuroendo. Lett. 37, 1–8 (2016).
Eyles, D. W. How do established developmental risk-factors for schizophrenia change the way the brain develops? Transl. Psychiat. 11, 158 (2021).
Pereira, M. J. et al. FKBP5 expression in human adipose tissue increases following dexamethasone exposure and is associated with insulin resistance. Metabolism 63, 1198–1208 (2014).
Sidibeh, C. O. et al. FKBP5 expression in human adipose tissue: potential role in glucose and lipid metabolism, adipogenesis and type 2 diabetes. Endocrine 62, 116–128 (2018).
Agam, G. et al. The Role of FKBPs in Complex Disorders: Neuropsychiatric Diseases, Cancer, and Type 2 Diabetes Mellitus. Cells 13, 801 (2024).
Nielsen, J. B. et al. Loss-of-function genomic variants highlight potential therapeutic targets for cardiovascular disease. Nat. Comms. 11, 6417 (2020).
Warren, H. R. et al. Genome-wide association analysis identifies novel blood pressure loci and offers biological insights into cardiovascular risk. Nat. Genet. 49, 403–415 (2017).
Saunders, G. R. et al. Genetic diversity fuels gene discovery for tobacco and alcohol use. Nature 612, 720–724 (2022).
Leng, K. et al. CRISPRi screens in human iPSC-derived astrocytes elucidate regulators of distinct inflammatory reactive states. Nat. Neuro. 25, 1528–1542 (2022).
Belongie, K. J. et al. Identification of novel biomarkers to monitor β-cell function and enable early detection of type 2 diabetes risk. PLoS One 12, e0182932 (2017).
Kipp, A. et al. Glomeruli proteome analysis reveals early differences between pre-existing and de-novo type 2 diabetes in human renal allografts. Nephrol. Dial. Transplant. 36, 143–003 (2021).
Alam, C. M. et al. M. Keratin 8 modulates β-cell stress responses and normoglycaemia. J. Cell Sci. 126, 5635–5644 (2013).
Lopez-Ramirez, O. et al. The role of acid sensing ion channels in the cardiovascular function. Front. Physiol. 14, 1194948 (2023).
Newberry, E. P. et al. Impaired hepatic very low-density lipoprotein secretion promotes tumorigenesis and is accelerated with Fabp1 deletion. Am. J. Pathol. 194, 958–974 (2024).
Tian, R. et al. CRISPR interference-based platform for multimodal genetic screens in human iPSC-derived neurons. Neuron 104, 239–255 (2019).
Gomez-Isla, T. et al. Connections and Cognitive Impairment In Alzheimer’s Disease. In Connections, cognition and Alzheimer’s disease. Springer Berlin Heidelberg 49–166 (1997).
Heneka, M. T. et al. Neuroinflammation in Alzheimer’s disease. Lancet Neurol. 14, 388–405 (2015).
Mendes, T. K. et al. Investigating the role of RIO protein kinases in Caenorhabditis elegans. PLoS One 10, e0117444 (2015).
Kaur, D. et al. Activation of microglia and astrocytes: a roadway to neuroinflammation and Alzheimer’s disease. Inflammopharmacology 27, 663–677 (2019).
Lorincz-Comi, N. J. et al. Bayesian inference of genetic pleiotropy identifies drug targets and repurposable medicines for human complex diseases. medRxiv https://doi.org/10.1101/2025.03.17.25324106 (2025).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, s13742–015 (2015).
Trubetskoy, V. et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature 604, 502–508 (2022).
Howard, D. M. et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat. Neurosci. 22, 343–352 (2019).
Van Rheenen, W. et al. Common and rare variant association analyses in amyotrophic lateral sclerosis identify 15 risk loci with distinct genetic architectures and neuron-specific biology. Nat. Genet. 53, 1636–1648 (2021).
Suzuki, K. et al. Genetic drivers of heterogeneity in type 2 diabetes pathophysiology. Nature 627, 347–357 (2024).
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Zhou, Y. et al. The Alzheimer’s Cell Atlas (TACA): A single-cell molecular map for translational therapeutics accelerator in Alzheimer’s disease. Alzheimer’s. Dement 8, e12350 (2022).
Ochoa, D. et al. The next-generation Open Targets Platform: reimagined, redesigned, rebuilt. Nucleic Acids Res 51, D1353–D1359 (2023).
Yang, J. J. et al. TIGA: target illumination GWAS analytics. Bioinformatics 37, 3865–3873 (2021).
Piñero, J. et al. DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res 45, D833–D839 (2016).
Greenfest-Allen, E. et al. NIAGADS Alzheimer’s GenomicsDB: A resource for exploring Alzheimer’s disease genetic and genomic knowledge. Alzheimer’s. Dement 20, 1123–1136 (2024).
Covo, S. et al. A novel single-gamma approximation to the sum of independent gamma variables, and a generalization to infinitely divisible distributions. Elec. J. Stat. 8, 894–926 (2014).
Stewart, T. et al. A simple approximation to the convolution of gamma distributions. Cent. Discussion Pap. 70, 1–26 (2007).
Jiang, L. et al. Powerful and robust inference of complex phenotypes’ causal genes with dependent expression quantitative loci by a median-based Mendelian randomization. Am. J. Hum. Genet. 109, 838–856 (2022).
Galwey, N. W. A new measure of the effective number of tests, a practical tool for comparing families of non-independent significance tests. Genet. Epidemiol. 33, 559–568 (2009).
Wang, G. et al. A simple new approach to variable selection in regression, with application to genetic fine mapping. JRSSB: Stat. Method. 82, 1273–1300 (2020).
R Core Team. *R: A language and environment for statistical computing*. R Foundation for Statistical Computing. https://www.R-project.org/ (2023).
Li, M. X. et al. GATES: a rapid and powerful gene-based association test using extended Simes procedure. Am. J. Hum. Genet. 88, 283–293 (2011).
Liu, Y. et al. ACAT: a fast and powerful p value combination method for rare-variant analysis in sequencing studies. Am. J. Hum. Genet. 104, 410–421 (2019).
Lorincz-Comi, N. Exact set-based hypothesis testing of correlated normal statistics using a distributional equivalence in a high-dimensional space. https://github.com/noahlorinczcomi/exset (2025).
Watanabe, K. et al. Functional mapping and annotation of genetic associations with FUMA. Nat. Comms. 8, 1826–1837 (2017).
Lorincz-Comi, N. et al. MRBEE: A bias-corrected multivariable Mendelian randomization method. HGG Advances. 5, 100290 (2024).
Kollo, T. et al. Advanced Multivariate Statistics With Matrices. Dordrecht: Springer Netherlands (2005).
Yavorska, O. O. et al. MendelianRandomization: an R package for performing Mendelian randomization analyses using summarized data. Int. J. Epidemiol. 46, 1734–1739 (2017).
Wood, E. R. et al. Discovery and in vitro evaluation of potent TrkA kinase inhibitors: oxindole and aza-oxindoles. Bioorg. Med. Chem. Lett. 14, 953–957 (2004).
Selleck Chemicals. GW441756, Catalog No. S2891. https://www.selleckchem.com/products/gw-441756.html#s_ref (2025).
Parra Bravo, C. et al. Cellular and pathological functions of tau. Nat. Rev. Mol. Cell. Biol. 25, 845–864 (2024).
Gohel, D. et al. Sildenafil as a candidate drug for Alzheimer’s disease: real-world patient data observation and mechanistic observations from patient-induced pluripotent stem cell-derived neurons. J. Alzheimers Dis. 98, 643–657 (2024).
Acknowledgements
This work was supported by the National Institute on Aging (NIA) under Award Number U01AG073323, R01AG066707, R01AG084250, R01AG076448, R01AG082118, R01AG092462, R01AG092591, RF1AG082211, and R33AG083003, the National Institute of Neurological Disorders and Stroke (NINDS) under Award Number RF1NS133812, National Institutes of Health (NIH) under Award Number 1OT2OD038083-01, the Alzheimer’s Association award (ALZDISCOVERY−1051936), the Alzheimer’s Drug Discovery Foundation (ADDF), and Dr. Keyhan and Dr. Jafar Mobasseri Endowed Chair for Innovative Research to F.C. This work was supported in part by the Rebecca E. Barchas, MD, Professorship in Translational Psychiatry, the Valor Foundation, Project 19PABH134580006-AHA/Allen Initiative in Brain Health and Cognitive Impairment, the Elizabeth Ring Mather & William Gwinn Mather Fund, S. Livingston Samuel Mather Trust, and the Louis Stokes VA Medical Center resources and facilities to A.A.P. This work was supported in part by National Heart, Lung, and Blood Institute (NHLBI) under Award Number P01HL158501 to M.K.C. and F.C. This work was also supported by Global Center for Pathogen and Human Health Research Postdoc Fellowship.
Author information
Authors and Affiliations
Contributions
N.L.C. and F.C. conceived the study design. N.L.C. developed the methodology, performed the formal analysis, and wrote the original draft. Y.Z., J.Z., and W.M. created a visualization. W.S., I.R.P., and X.C. performed experiments. F.C. supervised all activities, provided all resources, and reviewed and edited the manuscript. Y.H., A.P., J.B., J.H., and M.C. edited the manuscript. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
All authors declare no competing financial and non-financial interests.
Peer review
Peer review information
Nature Communications thanks Junhao Wen, Chunyu Liu and the other anonymous reviewer for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lorincz-Comi, N., Song, W., Chen, X. et al. Combining xQTL and genome-wide association studies from diverse populations improves druggable gene discovery. Nat Commun 17, 2801 (2026). https://doi.org/10.1038/s41467-026-69236-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69236-z