Abstract
Foundation models require massive amounts of training data, which are often costly to obtain in materials science. Meanwhile, long-established physical knowledge such as molecular force fields and geometric analysis provides direct guidance for material behavior, but remains insufficiently leveraged. Here we demonstrate that expert knowledge can directly supervise pre-training and substantially reduce data requirements. A set of potential energy surface (PES) basis functions, which encode guest-host interaction energetics, is developed as unified descriptors for different guest molecules. A multi-modal architecture is designed to fuse information from both material structure and PES. Pre-training is achieved by learning comprehensive geometric features spanning different spatial scales. Consequently, a foundation model for porous materials is developed under limited data regimes, named SpbNet. SpbNet is evaluated on over 50 downstream tasks, including adsorption, separation, and intrinsic properties, etc. SpbNet consistently outperforms models pre-trained on datasets nearly 20 times larger, reducing the relative errors by over 20%. In addition, SpbNet demonstrates strong generalization capabilities across both in-distribution and out-of-distribution materials, such as Metal Organic Frameworks, Covalent Organic Frameworks, and zeolites.
Similar content being viewed by others
Introduction
Foundation models have demonstrated remarkable potential across diverse domains, including chemistry1, medicine2,3,4, and meteorology5. Trained on large-scale datasets, these parameter-rich architectures exhibit exceptional versatility in addressing a wide range of downstream tasks. Most importantly, compared with conventional machine learning approaches, foundation models show superior data efficiency, enabling rapid adaptation to downstream tasks with limited training samples6. In the field of materials science, porous materials, such as metal-organic frameworks (MOFs), covalent organic frameworks (COFs), porous polymer networks (PPNs), and zeolites, possess high specific surface areas7 and tunable structural features8, conferring substantial agricultural and industrial value9,10,11,12,13,14,15. Therefore, developing a foundation model tailored for porous materials is of considerable scientific and practical importance.
However, pre-training a foundation model typically requires an extremely large amount of data, which is often prohibitively expensive or even impractical in materials science. For instance, despite decades of research, fewer than 1000 experimental zeolite structures have been reported16. To address this challenge of data scarcity, several studies have incorporated domain expertise to enhance data efficiency, either through model architecture design17 or task formulation during training18. Nevertheless, in terms of constructing larger-scale pre-trained foundation models as well as improving data efficiency on pre-trained data, the potential of expert knowledge remains largely unexplored. Here, we propose a holistic pre-training framework for porous materials that integrates expert knowledge to enhance data efficiency. The framework comprises three key components: (1) input descriptors19,20, which encode raw material structures using unbiased molecular force fields; (2) a dual-stream multi-modal architecture designed to jointly capture structural and potential energy surface (PES) information; and (3) pre-training tasks that enable the model to learn comprehensive, multi-scale geometric representations18,20.
For the descriptors, in addition to the raw material structure, we introduce a set of PES basis functions. These basis functions can be linearly combined to represent the interaction energies between the material and various guest molecules, thereby mitigating bias toward specific adsorbates. We further design a dual-stream multi-modal model architecture comprising three key components: a graph neural network (GNN) that processes raw material structures, a vision transformer that encodes PES representations, and a cross-attention fusion module that integrates structural and energetic information. This dual-stream design effectively leverages the strengths of modern GNNs and the expressive power of PES basis functions. Finally, we construct multi-scale pre-training tasks covering probes of different sizes and material structures of varying granularity. We refer to the resulting foundation model as the Structure-informed Potential energy surface Basis function Network (SpbNet). SpbNet is pre-trained on 0.1 million MOF structures. For in-distribution downstream tasks, we evaluate SpbNet across more than 40 MOF-related benchmarks, including gas adsorption, separation, and intrinsic properties such as elastic modulus. For out-of-distribution (OOD) evaluations, we assess SpbNet on eight tasks involving COFs, PPNs, and zeolites. SpbNet consistently outperforms other pre-trained models trained on over 2 million porous materials20,21 and 120 million molecular and material samples22. These results highlight that the potential of incorporating expert knowledge in the construction of foundation models remains far from fully realized, particularly in enhancing data efficiency.
Results
Overview of SpbNet
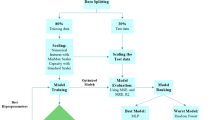
The overall architecture and training pipeline of SpbNet are illustrated in Fig. 1. The design of SpbNet comprises four major components: (1) Descriptors, encompassing both the raw material structure and the proposed PES basis functions; (2) Model architecture, corresponding to the multi-modal input design; (3) Pre-training tasks, which facilitate multi-scale geometric feature learning; and (4) Fine-tuning process, which adopts a simple multi-head-based fine-tuning strategy.
SpbNet consists of four main parts: (1) Descriptors. SpbNet adopts raw material structure and potential energy surface (PES) basis functions as input. PES basis functions are regarded as a multi-channel three-dimensional (3D) image and linearly combined through a linear layer to obtain a single-channel 3D image. (2) Model architecture. A Graph Neural Network (GNN) extracts material structure features from the raw material structure. i, j, k represent the atoms in the material. The combined PES is processed with a Vision Transformer (ViT). Structural features are incorporated into the ViT via the cross-attention mechanism. (3) Pre-training. The model is pre-trained via the prediction of global geometric features and atom-level properties including number of atoms and Signed Distance Function (SDF). A Class49 ([CLS]) token is appended to the PES patches to predict the global features via dedicated prediction heads, including Topology type (Topo), Pore Limited Diameter (PLD), Largest Cavity Diameter (LCD), Void Fraction, etc. Features of PES patches are used to predict SDF and number of atoms through separate output heads, including SDF Head and ANP (Atom Number Prediction) Head. (4) Fine-tuning. After pre-training, the model retains the backbone weights while discarding all pre-training heads. A new task-specific prediction head is added to predict the target property. Downstream tasks cover guest molecule-related properties, intrinsic material properties, and generalization to out-of-distribution material systems. Trainable parts are marked in blue background.
Descriptors
Descriptors play a pivotal role in materials informatics. Although recent studies have increasingly employed raw structural information (i.e., atomic numbers and positions) as input, expert-designed descriptors continue to demonstrate clear advantages in specific tasks19,20,23. In particular, PES are highly suitable for tasks involving guest molecules—the primary application domain of porous materials19,20. This suitability arises because PES directly characterizes the interaction energy between a material and a guest molecule, which governs the latter’s behavior. However, PES profiles differ substantially across guest species (see Supplementary Fig. 1 for details), rendering molecule-specific PES representations unsuitable as universal inputs for a foundation model. To construct a generalizable interaction energy descriptor, we propose a set of PES basis functions that describe the interaction energy between porous materials and various guest molecules. The potential energy of any guest molecule ϕ at position r can be expressed as a linear combination of 21 basis function terms with molecule-specific coefficients Coe(ϕ), comprising 13 terms of Pauli exchange-repulsion energy, 7 terms of London dispersion energy, and 1 term of Coulombic energy:
Detailed formulations and derivations are provided in Section “Derivation of potential energy surface basis functions”. The 21 terms (Epauli, Elondon, and Ecoulomb) serve as the input features of SpbNet. These 21 basis functions are evaluated at predefined sampling points across the lattice, resulting in a 21-channel three-dimensional image that spans the entire periodic cell. The 21 channels are subsequently linearly combined through a linear layer to generate a single-channel image representing a universal PES, which can then be processed using vision-based methods. Further implementation details are described in Sections “Representing PES basis functions via multi-channel 3D image” and “Linearly combining PES basis functions via residual connection”. Importantly, this set of basis functions is free from bias toward any specific guest molecule, making it a suitable and transferable input representation for a universal foundation model.
Model architecture
Model architecture represents another critical component of a foundation model. It must be capable of capturing essential structural features from the input, such as bond lengths, bond angles, and other local geometrical descriptors. Given the multi-modal nature of SpbNet’s inputs, the architecture must effectively integrate heterogeneous representations. As illustrated in Fig. 1, the main architecture of SpbNet consists of a GNN for processing material structures24 and a vision transformer25 for encoding the linearly combined PES. To fuse information from the structural and energetic modalities, we employ a cross-attention mechanism commonly adopted in multi-modal architectures26,27. Following the design in ref. 25, an additional learnable [CLS] token is appended to the PES patch sequence. The feature extracted from the [CLS] token is subsequently passed through a prediction head (see Section “Training strategy” for details) to infer the target property.
To enable simultaneous prediction of multiple material properties during pre-training, we construct separate task-specific heads (multi-head design). During fine-tuning, only the backbone parameters are retained, while all pre-training heads are discarded and replaced with randomly initialized heads for downstream task. Compared with single-stream architectures, where the same layers and weights are used to process features from different modalities, the dual-stream design allows distinct sets of layers (self-attention and cross-attention layers28) to specialize in modality-specific representations26,27. This separation not only preserves the diversity of structural and energetic features but also facilitates flexible substitution of the structural encoder, thereby leveraging advances in modern GNN architectures.
Pre-training
Pre-training strategy constitutes one of the most critical components of a foundation model. In this work, we focus on training the network to learn well-established geometric analysis of porous materials. A series of multi-scale pre-training tasks are designed. At the material level, the model is pre-trained to predict (1) topology type (e.g., pcu, fcu, etc. The model is trained to classify the material into a specific topology in the pre-defined topology type set20), (2) pore limiting diameter (PLD), (3) largest cavity diameter (LCD), (4) accessible surface area (ASA), and (5) accessible volume fraction (VF). To mitigate bias toward specific guest molecules, ASA and VF are computed using probes of varying sizes29. At the atomic level, the model is trained to predict (1) the number of atoms within each PES patch, and (2) the signed distance function (SDF) at the patch center. The SDF is also computed with probes of different sizes. Calculation details and the differences in geometric features obtained using various probe sizes are provided in Supplementary Text 1 and Supplementary Fig. 2. These pre-training tasks comprehensively leverage the accumulated domain knowledge of geometric characterization in porous materials, thereby providing a robust and domain-informed pre-training process. The correlation between these geometric features and downstream tasks can be found in Supplementary Figs. 3–9.
Pre-training results
Figure 2 presents the pre-training performance of SpbNet. As shown, for macroscopic geometric features such as VF and LCD, SpbNet achieves an R2 score of 0.9 and a Pearson correlation coefficient of 0.95. When the probe size decreases from 2.0 to 0.5, the accuracy of VF remains largely unchanged, whereas the accuracy of ASA decreases. This observation is consistent with Supplementary Fig. 7, which shows that correlations among ASAs computed using different probe sizes are more complex than those of VF. The visualization of the topology type classification task can be found in Supplementary Fig. 10, with the accuracy of 94.9%.
a Pearson correlation coefficient of geometric features. b Coefficient of Determination (R2) score of geometric features. SpbNet is trained to predict different geometric features: (1) Accessible Surface Area (ASA) of different probes, (2) Void Fraction (VF) of different probes, (3) Pore Limited Diameter (PLD), and (4) Largest Cavity Diameter (LCD). Both ASA and VF are calculated using probes of 0.5 Å and 2.0 Å. Thus there are totally 6 different geometric features. These results are calculated from 100 Metal Organic Frameworks (MOFs) randomly selected in the pre-training validation set. c t-distributed Stochastic Neighbor Embedding (t-SNE) plot of Potential Energy Surface (PES) patches' feature colored by the contained number of atoms. From the lower left to the upper right, the number of atoms gradually increases. d t-SNE plot of PES patches' feature colored by Signed Distance Function (SDF). From the top left to the bottom right, the SDF gradually increases. For both t-SNE plots, PES patches are from 100 MOFs randomly selected from the validation set of the pre-training process. Source data are provided as a Source Data file.
For the fine-grained SDF and atom-number prediction tasks, the t-SNE visualizations are shown in Fig. 2c, d. The PES patch embeddings exhibit distinct trends as the SDF and atom numbers vary, suggesting that the model effectively learns to predict both properties simultaneously. The R2 scores for atom-number and SDF prediction tasks are 0.911 and 0.957, respectively (see Supplementary Fig. 11 for visualization). As shown in Fig. 2c, most PES patches contain only a few atoms, reflecting SpbNet’s high spatial resolution in decomposing PES into fine patches. In Fig. 2d, most patch centers display positive SDF values, indicating that the porous regions occupy the majority of the MOF volume.
Collectively, these findings demonstrate that SpbNet effectively learns well-defined geometric features through pre-training. The multi-head architecture further allows the model to capture diverse types of features simultaneously without mutual interference, even when identical properties are calculated by different scales of probes. Through such comprehensive pre-training objectives, SpbNet develops generalizable geometric representations that are transferable to downstream applications such as adsorption, facilitating rapid and data-efficient adaptation. Although predicting these properties is relatively straightforward, the model does not inherently acquire such intermediate features when trained solely on downstream task labels. Incorporating these properties as auxiliary objectives in multi-task training also enhances downstream accuracy (Supplementary Table 2), underscoring their value in guiding the model toward more informative structural representations.
Performance on in-distribution metal-organic-frameworks
Gas adsorption stands as one of the most critical application scenarios for porous materials9,30. We conduct experiments on two representative datasets: hMOF31,32,33,34 (comprising ~130,000 hypothetical MOFs) and CoREMOF35,36 (containing ~14,000 experimentally synthesized MOFs). The Mean Absolute Error (MAE) is adopted as the performance metric. SpbNet is compared with CGCNN37, MOFormer38, and MOFTransformer20,21. Detailed architectures of the baseline models can be found in Supplementary Text 2. Beyond typical industrial gases such as CO2 and CH4, we also incorporate (1) noble gases (including Kr, Xe) and (2) other hydrocarbons (including propylene, propyne, and para-xylene (p-xylene)).
SpbNet demonstrates superior performance across different gases (see Fig. 3a and Supplementary Data 1). Specifically, for CO2 adsorption, SpbNet achieves a substantial reduction in MAE by 29.1%. Notably, even when predicting CH4 adsorption (the probe used to calculate the PES), SpbNet still maintains a remarkable improvement, reducing the MAE from 0.248 to 0.175, an improvement of 29.4%. These results unequivocally demonstrate that the enhanced generalization capabilities do not compromise accuracy for specific guest molecules. For other hydrocarbons, SpbNet reduces the relative error for C3H6 and p-xylene by 23.9% and 9.6%, respectively (as shown in the right panel of Fig. 3a). Furthermore, SpbNet exhibits promising performance for noble gases such as Kr and Xe, with MAE decreases of 11.0% and 17.8%, respectively. Additional quantitative results are provided in Supplementary Data 1.
a Gas adsorption. CO2, CH4, propylene, and paraxylene. b Gas separation. CH4/N2, N2/O2, C3H6/C3H8. c Adsorption heat. d Henry’s constant. e Intrinsic properties. Elastic modulus (KVRH) (where VRH represents Voigt–Reuss–Hill techniques39), shear modulus (GVRH), decomposition temperature, and heat capacity. SpbNet is compared to CGCNN37, MOFormer38, PMTransformer20,21 For the hMOF31 --34 and CoREMOF35,36 datasets with sufficient data, we randomly selected 5000 samples for training and 1000 samples for test and validation. For other datasets, we split the training, validation, and test sets using a ratio of 8:1:1. Detailed metrics, data sources, and training data volume are provided in Supplementary Data 1 and 2. Mean absolute errors (MAE) are annotated above each subplot. Source data are provided as a Source Data file.
Beyond gas adsorption, SpbNet is versatile and applicable to three distinct categories of tasks: (1) multi-guest molecule-related tasks (e.g., gas separation); (2) single-guest molecule-related tasks (e.g., adsorption heat and Henry’s constant); and (3) material intrinsic properties (e.g., thermal decomposition temperature).
For gas separation, as illustrated in Fig. 3b, SpbNet significantly improves accuracy for CH4/N2 and N2/O2 separation, reducing the MAE by 32.3% and 21.0%, respectively. Even for chemically similar gases like propane and propylene, SpbNet achieves an 8.03% improvement. For single-guest molecule-related tasks (Fig. 3c, d), we assessed: (1) heat of adsorption; (2) Henry’s constant; and (3) gas diffusion. In the context of CO2 and CH4 adsorption heat, SpbNet reduces the relative error by 0.75% and 20.9%. Regarding the Henry’s constant for propylene and CH4, SpbNet demonstrates substantial enhancements, reducing the MAE by 17.2% and 41.0%. Additionally, we evaluated SpbNet’s performance in gas diffusion. Further quantitative results are detailed in Supplementary Data 1.
Aside from guest molecule-related properties, SpbNet is also capable of predicting intrinsic material properties (Fig. 3e). Four different intrinsic properties are examined: (1) mechanical stability (including elastic modulus and shear modulus); (2) thermal stability; and (3) heat capacity. For mechanical properties, tasks are conducted to predict the bulk elastic modulus (KVRH) and shear modulus (GVRH), where VRH represents Voigt–Reuss–Hill techniques39. SpbNet achieves higher accuracy, reducing the MAE by 11.1% and 13.7%, respectively. Regarding thermally relevant properties, SpbNet demonstrates comparable performance in the prediction of thermal decomposition temperature, reducing the MAE by 2.34%. Although the heat capacity data is sparse, consisting of only a few hundred data points, the four compared models perform well, maintaining an R2 score above 0.7 (see Supplementary Data 1).
Furthermore, we compare SpbNet with other universal models designed for molecules and general materials in Supplementary Table 1 and Supplementary Text 3. Notably, even when benchmarked against modern foundation models pre-trained on 209 million data points40 and 120 million data points22, SpbNet achieves the best performance. Nevertheless, it must be emphasized that SpbNet is primarily trained to capture the macroscopic geometric structure of the material, which is closely related to the behavior of guest molecules within the material pores. Consequently, SpbNet primarily leverages its structure encoder to infer other properties, such as electronic structure, which may not be its main focus.
Out-of-distribution generalization
Except for performance on in-distribution downstream datasets, another critical evaluation metric for a foundation model is its transferability to OOD datasets. In this section, we evaluate SpbNet’s performance on other porous material systems, specifically COFs, PPNs, and zeolites. The results are presented in Fig. 4. As shown in Fig. 4a, b, although the model has never encountered COF or zeolite structures during the pre-training stage, SpbNet accurately predicts the geometric properties of both material classes. This confirms that the pre-trained SpbNet possesses inherent generalization capability across different porous systems. The model exhibits substantially better generalization on COFs than on zeolites, owing to the closer structural similarity between MOFs and COFs (see Supplementary Figs. 12 and 13 for the visualization of the features). Interestingly, despite MOFs and zeolites showing comparable statistical properties, such as pore volume and atomic density (Supplementary Fig. 14), the model generalizes poorly to zeolites. This is primarily because zeolites are composed predominantly of Si, O, and Al atoms, leading to distinct chemical environments. In contrast, the model retains strong generalization on COFs even when there exist notable differences from MOFs in both atomic density and pore volume.
a Accuracy of geometric features prediction for COFs. b Accuracy of geometric features prediction for zeolites. SpbNet trained on MOF-only pre-training dataset is directly evaluated on the COFs and zeolites. 100 COFs and zeolites are randomly selected from the test set of the benchmark dataset. c Prediction error distributions (box plots) on COFs. d Prediction error distributions (box plots) on PPNs. e Prediction error distributions (box plots) on zeolites. For each porous material, we randomly selected 5000 samples for fine-tuning, and then validated and tested SpbNet on 1000 samples. Mean absolute errors (MAE) are annotated above each subplot. Source data are provided as a Source Data file.
Further OOD evaluation is conducted on downstream tasks. For two gas adsorption tasks (CH4, 298 K, 65 bar and 5.8 bar) on COFs41 (Fig. 4c left), SpbNet reduces the MAE by 27.6% and 34.4%, respectively. For PPNs42 (CH4, 298 K, 65 bar and 1 bar) (Fig. 4d), the improvements reach 32.5% and 74.7%. Furthermore, for COFs’ adsorption heat and Henry’s constant43 (CO2, 298 K), SpbNet reduces the MAE by 17.7% and 13.2%. On the zeolite dataset44 (CH4, 298 K), the corresponding improvements reach 39.7% and 40.8% (Fig. 4e). These findings demonstrate that transferring SpbNet to new porous material systems does not lead to a significant degradation of its accuracy advantage, confirming the generalization ability of the pre-trained model to support fine-tuning of downstream tasks across diverse material classes.
Label efficiency
Foundation models possess broad application prospects due to their ability to achieve strong performance with minimal training data, significantly expanding their application scope compared to models trained from scratch. We evaluate the label efficiency of SpbNet on the gas adsorption and diffusion tasks, varying the number of training samples from 5000 to 20,000. The results are illustrated in Fig. 5. With the same pre-training data volume (100 K data), SpbNet consistently outperforms MOFTransformer20,21. Furthermore, even when compared to MOFTransformer pre-trained on 1.9 million data (~19 times the amount used for SpbNet), SpbNet still maintains superior performance on the gas adsorption task. Only for the gas diffusion task, when MOFTransformer is fine-tuned on approximately three times the amount of downstream task data (15 K data), the performance of SpbNet and MOFTransformer becomes comparable. For gas adsorption, both SpbNet and MOFTransformer significantly benefit from increasing the size of the training dataset. However, for gas diffusion, the effect of increasing the data volume is less pronounced, indicating that diffusion is an inherently more challenging task than adsorption. These results underscore the high data efficiency of SpbNet. SpbNet not only obtains sufficient pre-training capabilities from a relatively small dataset (100 K) but also demonstrates rapid adaptability to downstream tasks.
a Performance of models under different volume of fine-tuning data for H2 Uptake. b Performance of models under different volume of fine-tuning data for H2 Diffusion. Models are fine-tuned on 5 K, 10 K, 15 K, and 20 K data, which are randomly selected from the whole training set. Figures demonstrate the Mean Absolute Error (MAE) of different models under different fine-tuning data volume. SpbNet is compared to MOFTransformer20 (mt) and PMTransformer21 (pm). SpbNet is pre-trained on 100 thousand MOFs. MOFTransformer is pre-trained on on MOF datasets of varying size (100 thousand, 500 thousand, 1 million). PMTransformer is pre-trained on 1.9 million porous materials. Part of results are from the figure of MOFTransformer20. Source data are provided as a Source Data file.
Discussion
To fundamentally understand the underlying mechanism of the pre-training tasks, we performed ablation studies. We considered four different versions of SpbNet: (1) Trained from scratch (i.e., without any pre-training process and with randomly initialized weights). (2) Pre-training utilizing only global geometric features. (3) Pre-training using global geometric features and the atom number prediction task. (4) Pre-training using global geometric features, the atom number prediction task, and the SDF prediction task.
Feature space analysis
We first evaluate the feature space extracted by SpbNet. Figure 6 a demonstrates the averaged cosine similarity between the global [CLS] token and other local PES tokens. When the model is trained from scratch, the feature of the [CLS] token rapidly aligns with other tokens as the network deepens. This is primarily due to the absence of a token-scale loss constraint, where the inherent low-pass filtering property of the attention mechanism causes the features of all tokens to become overly similar, a phenomenon known as over-smoothing45. In contrast, when fine-tuned from pre-trained weights, the average similarity remains at a relatively low level, indicating that each energy patch maintains more localized information compared to the global feature. We further discovered that one crucial factor controlling this effect is the size of the pre-training data (see Supplementary Fig. 15 for the comparison between large and small pre-training dataset). When the data volume is sufficiently large, even with only material-level supervision signals, each token can still retain local information; conversely, over-smoothing is prone to occur when the data volume is small.
a Average cosine similarity between the class ([CLS]) token49 and potential energy surface (PES) tokens. Cosine similarities are computed between the added [CLS] token and all PES patch tokens, averaged over 100 randomly selected samples from the pre-training dataset. Four model variants are evaluated: (1) without pre-training (scratch); (2) pre-trained with global geometric (Geo) features (geo); (3) with Geo features and Atom Number Prediction (ANP) (geo+anp); and (4) with Geo features, ANP, and Signed Distance Function (SDF) prediction (geo+anp+sdf). Solid lines denote mean values, shaded regions denote standard deviation. b Effective rank of PES tokens. The effective rank46 characterizes the intrinsic working dimensionality of the feature space. It decreases from lower to higher layers, implying progressive compression of structural information into compact semantic representations56,57. ANP and SDF tasks increase the effective rank, indicating strengthened local feature learning. Solid lines and shaded regions respectively represent mean and standard deviation. c Prediction error distributions (box plots) of different SpbNet variants. Mean absolute errors (MAE) are annotated above the subplots. Global geometric pre-training greatly improves performance, while ANP and SDF further enhance accuracy. Removing basis functions (Wobasis) reduces accuracy. Source data are provided as a Source Data file.
These results indicate that the behavior of the [CLS] token in fine-tuning and training-from-scratch is essentially different. After the model is pre-trained, the model already learns to compress the structure information into a low-rank feature space. In this fine-tuning phase, the [CLS] token learns to aggregate this information to make prediction. When the model is trained from scratch, the lower layers still maintain local information of each token. However, the global feature of the [CLS] token quickly spreads to other tokens and erases almost all local information of PES tokens. At this point, the network degenerates to a simple MLP.
Effect of task combination on feature rank
To observe the impact of different pre-training tasks, Fig. 6b illustrates the effective ranks under various pre-training task combinations. Similar to the Principal Component Analysis method, the effective rank quantifies the actual dimensionality that significantly contributes to the high-dimensional feature space46. From the lower to the higher layers, the effective rank gradually decreases, suggesting that the feature of each PES token is compressed into a low-dimensional semantic space. During the model’s fine-tuning process, these extracted semantics are leveraged for making robust predictions. Both the atom number prediction task and the SDF prediction task enhance the effective rank throughout this process, enabling the model to utilize more layers to extract richer local information. The local information is also maintained after fine-tuning, as evidenced by the linear probing experiments shown in Supplementary Figs. 16 and 17.
Task-specific performance
Figure 6c illustrates the ablation study results on six selected downstream tasks. The inclusion of token-level pre-training tasks yields performance improvements. Notably, the SDF prediction task is particularly beneficial for gas separation, which may be attributed to its role in capturing the molecular sieving effect. The detailed surface shape information provided by the SDF assists the model in accurately determining whether a guest molecule can easily pass through or be retained by the material’s pores. Conversely, the atom number prediction task produced a certain negative effect on the zeolite tasks. This negative transfer may be due to the significant structural differences between zeolites and MOFs, as illustrated by the t-SNE plots of different materials in Supplementary Figs. 12 and 13. These combined results decisively demonstrate the effectiveness and mechanism of the atom number and SDF prediction tasks in enhancing SpbNet’s representation learning capability. In addition, removing basis functions also reduces the model accuracy. The effect of basis functions is relatively limited on CH4-related tasks, which can be attributed to that the CH4 molecule is adopted as the probe to calculate the PES without basis functions.
Limitations
In this work, we present SpbNet, an expertise-guided supervised pre-training framework for foundation models of porous materials. It achieves high data efficiency and strong performance across a range of guest-molecule and geometry-related downstream tasks. The model leverages unbiased descriptors derived from molecular force fields and adopts a supervised pre-training paradigm centered on geometric representations. Crucially, SpbNet integrates a broad and physically grounded set of pre-training tasks designed to improve generalization beyond any single downstream objective. This approach is particularly suited to porous materials. The key functions of porous materials, such as adsorption, separation, and storage, are governed by host-guest interactions dictated by pore geometry.
SpbNet combines three components within a unified framework: the PES as an input descriptor that encodes guest-host interaction energetics; a multi-modal architecture that couples PES-based and structural representations; and a geometry-oriented pre-training objective targeting features correlated with macroscopic metrics like adsorption capacity and separation ratio. Together, these components provide both local energetic and global geometric perspectives, forming a comprehensive pre-training scheme that enhances generalization and data efficiency under limited data availability.
Nevertheless, the strong task specificity of this supervised strategy may constrain its generalization compared with broader self-supervised learning (SSL) approaches. For tasks in which the underlying physics is weakly linked to geometric features, such as electronic structure prediction, the benefits of PES-based or geometry-driven supervision diminish. In this case, the model instead relies on latent representations from its structural encoder and the adaptability of its multi-modal design.
Overall, these findings suggest that under data-scarce conditions, carefully designed input descriptors, tailored architectures, and broad supervised pre-training tasks explicitly aligned with downstream objectives can substantially enhance generalization and achieve superior data efficiency compared with mainstream unsupervised approaches.
Methods
Derivation of potential energy surface basis functions
The first crucial step in our approach is to establish a universal PES representation suitable for diverse guest molecules. A feasible solution is to define a set of PES basis functions, \(\{{E}_{i}({{{\bf{r}}}})\}\), based on the position vector r. The potential energy E of any guest molecule ϕ at position r can then be expressed as a linear combination of these basis functions:
Here, Ei(r) are the basis functions, and Coei(ϕ) represents the coefficient unique to the guest molecule ϕ. For simplicity, we initially consider the case of a single atomic probe. Additionally, we focus only on the major non-bonded interactions.
Molecular force field and PES components
We assume that the non-bonded interaction potential energy primarily consists of three components: the Pauli exchange mutual repulsion potential (Epauli), the London dispersion potential (Elondon), and the Coulomb potential (Ecoulomb). The interaction energy between atom i (in the MOF) and atom j (the probe) is defined by the following standard force field terms47:
The parameters ϵ and σ denote the well depth and the van der Waals radius, respectively, which depend on the specific (pseudo) atoms. q represents the partial charge, and \(k=\frac{1}{4\pi {\epsilon }_{0}}\) is the electrostatic force constant.
For any atom i in the material, the Lennard–Jones (LJ) component of the van der Waals potential, V(rij) = Epauli(i, j) + Elondon(i, j), simplifies to:
The cross-parameters σij and ϵij are calculated using the Lorentz–Berthelot mixing rules:
Let j be the reference probe used for calculation and ϕ be the new probe atom in the actual task. Assuming the distance riϕ = rij = r, and substituting the mixing rules into the LJ potential for both probes:
If we were to calculate the ratio Vϕ/Vj, the result would be a complex function f(σϕ, σi, r) dependent on the distance r and the specific material atom i. Since the total potential energy for a material M is the summation Σi∈MV, the complex dependence of Vϕ/Vj on i and r prevents a simple linear correlation between different probes (ΣVϕ and ΣVj). This necessitates a decomposition into universal basis functions.
Construction of universal basis functions
Our goal is to find a set of basis functions E such that the ratio of the total potential energy of two probes satisfies ∀ ϕ, j: Eϕ/Ej = g(ϕ, j), where the coefficient g(ϕ, j) is independent of the material atom σi and the position r. This proportionality must hold even after summing over all atoms in the material structure (∑Eϕ/∑Ej = g(ϕ, j)), which is possible only if Eϕ/Ej is independent of i and r.
The required basis functions can be constructed by applying the binomial theorem to the Pauli repulsion and London dispersion terms, leveraging the Lorentz–Berthelot mixing rules. We analyze the London dispersion potential L as an example (riϕ = rij = r):
The term inside the first set of parentheses depends on the material atom i and position r, defining the basis function. The term inside the second set of parentheses depends only on the probes ϕ and j, defining the coefficient. Since the coefficient is independent of i and r, we can sum over all atoms i ∈ M, and the total potential energy remains a linear combination with the same coefficients.
Thus, for a specified exponent kl, the London basis function (summed over material atoms) is:
Final PES basis functions and implementation
The Pauli repulsion term is derived similarly with an exponent of 12, and the form of the electrostatic potential (which is Ecoulomb ∝ r−1 for a single charge qj = 1) is already a simple basis. Using the UFF molecular force field parameters47, the complete set of PES basis functions is:
where σϕ = σj + Δσ. Any (pseudo) atom j can serve as the reference probe, yielding a universally applicable set of basis functions. In our implementation, we specifically use the methane (a pseudo-atom representing methane) probe to calculate these basis functions. It should be noted that these functions describe the PES of a single (pseudo) atom. Further refinement for more complex guest molecules containing multiple atoms is reserved for future work.
Additionally, considering that many common guest molecules (e.g., CH4, N2, H2) are non-polar, their interactions are less affected by the Coulomb term. Introducing partial charges can potentially introduce a bias against specific guest molecules during the pre-training stage. Therefore, the partial charges and the corresponding Ecoulomb potential are not considered in practice, leaving a total of 13 + 7 = 20 basis terms.
Representing PES basis functions via multi-channel 3D image
For a specific material, the crystal structure is first expanded into a supercell to ensure that the minimum length of the lattice vector exceeds 8Å. Coordinates are then uniformly sampled within the supercell to construct a 24 × 24 × 24 PES grid (13,824 sampling points in total). For each coordinate r, the corresponding n terms of the PES basis functions are computed based on the formulas in Section “Derivation of potential energy surface basis functions”. The resulting data structure is a tensor of shape 24 × 24 × 24 × n, where n is the number basis functions. This data structure is naturally interpreted as a three-dimensional (3D) n-channel image with a resolution of 24 × 24 × 24, where each sampling point serves as a voxel and each of the n basis function terms corresponds to a distinct channel.
Linearly combining PES basis functions via residual connection
In Eq. (9), while a direct polynomial expansion of \({({\sigma }_{i}+{\sigma }_{\phi })}^{6}\) is mathematically feasible, we note the potential benefit of employing the difference substitution σϕ = σj + Δσ. The term in the basis expansion that is independent of Δσ is the one corresponding to the original London dispersion potential energy of the reference probe j (similarly for the Pauli term). This decomposition allows the final PES to be conceptually viewed as the original potential energy of probe j (specifically, j = CH4 is used to calculate the basis functions) plus a series of correction terms. This is analogous to the residual connection paradigm in deep learning48, which is known to enhance training stability and facilitate the training of deeper or more complex networks. SpbNet adopts this practice by fixing the coefficient of the reference PES (probe j) to 1, while allowing the coefficients of the other basis functions to be learnable.
Consequently, the PES basis functions are categorized into: (1) Original PES Term: A single-channel 3D image, shaped 24 × 24 × 24 × 1, which represents the PES of the reference probe j (CH4) and acts as the identity mapping. (2) Correction Terms: Composed of the remaining 18 channels (Pauli and London terms excluding the reference term), shaped 24 × 24 × 24 × 18. These correction terms are transformed into a single channel via a learnable linear layer (with weight shape (18,1) and no bias) and subsequently added to the Original PES Term to obtain the final universal PES. This specific implementation of the residual PES was found to be critical for performance. Direct implementation where all basis function coefficients (including that of the original CH4 PES) were learnable resulted in significant performance loss, performing even worse than simply using the original CH4 PES without any basis function expansion.
Model architecture
The architecture of SpbNet is designed around a cross-attention mechanism28 integrated with a structure encoder. Cross-attention is widely used in multi-modal learning to enable effective fusion of heterogeneous information26,27. In SpbNet, this mechanism facilitates the interaction between structural features of porous materials and the PES representations.
SpbNet consists of two main components: a structure encoder and a PES decoder. For the structure encoder, we adopt the PaiNN architecture24 to generate atom-level embeddings following the implementation in ref. 20. All hyperparameters remain consistent with the original PaiNN settings. Since each atom possesses an individual feature vector, the representation of an entire structure can be viewed as an unordered set of atomic embeddings.
For the PES decoder, we employ a Vision Transformer (ViT)25. The PES grid of size 24 × 24 × 24 × 1 (see Section “Linearly combining PES basis functions via residual connection”) is divided into cubic patches with a patch size of 3, yielding (24/3)3 = 512 patches in total. Each 3 × 3 × 3 × 1 patch is treated as a token, transforming the PES into an ordered sequence of patch tokens. This token sequence is processed by a 36-layer Transformer decoder28, where cross-attention layers inject the structural embeddings from the encoder into the PES token stream. The resulting feature of each patch is used to predict atomic quantities within that region.
A learnable [CLS] token49 is prepended to the PES token sequence, and its final feature is passed through two linear layers followed by a \(\tanh\) activation to predict both pre-training objectives (e.g., void fraction and topology) and downstream properties (e.g., gas adsorption). The hidden feature dimension is set to 512, and the model employs eight attention heads.
Training strategy
We follow the standard training paradigm commonly adopted in Transformer-based models. During the pre-training stage, the framework consists of (1) a linear layer that combines the PES basis functions (see Section “Linearly combining PES basis functions via residual connection”), (2) the structure encoder, and (3) the PES Transformer decoder as the backbone, with a single linear layer serving as the prediction head. An additional [CLS] token is appended to the sequence of PES patch tokens. The extracted feature of the [CLS] token is subsequently processed by a pooling layer composed of a single linear layer followed by a \(\tanh\) activation function. Each pre-training task is associated with an independent prediction head, implemented as a single linear layer to predict the corresponding target property (e.g., topology type, PLD, LCD, etc.). All trainable components share the same training schedule. The learning rate increases from 10−5 to 10−4 over the first 15 epochs and then decays to 10−5 following a cosine schedule. The AdamW optimizer is used with a weight decay of 0.25. The model is pre-trained for 40 epochs, and the checkpoint with the lowest validation loss is selected for fine-tuning.
In the fine-tuning phase, all backbone weights are retained, while the prediction head (including the pooling layer and task-specific linear layer) is replaced. The pooling layer is re-initialized, and a new downstream task-specific head is added. The same learning rate schedule is used as in the pre-training phase: the learning rate increases linearly from 0 to 10−4 during the first 2 epochs and then decays to 0 following the cosine function. AdamW is again employed as the optimizer, with the weight decay set to 0.1. The model is fine-tuned for 60 epochs, and the checkpoint achieving the lowest validation loss is used for inference. Notably, a smaller weight decay is adopted in fine-tuning, as it is empirically found to yield better performance.
Datasets
SpbNet uses different datasets as benchmarks. Except for bandgap, thermal stability, heat capacity, mechanical properties and datasets for COFs, zeolites and PPNs, other benchmarks use three public datasets: (1) hMOF31,32,33,34, (2) CoREMOF36, and (3) Tobacco50. hMOF31,32,33,34 is a commonly used dataset, containing over 130,000 hypothetical MOF materials. The hMOF dataset includes data on the adsorption capacity of seven gases (CO2, CH4, N2, H2, Kr and Xe) and adsorption heat data of four gases (CO2, CH4, N2, H2). It should be noted that the structural topologies in the hMOF dataset are relatively limited. Another, more challenging and reliable dataset is CoREMOF35,36. The CoREMOF dataset includes about 14,000 MOF structures, coming from experimental data, Cambridge Structural Database, and Web of Science searches. Thus, the CoREMOF dataset mainly includes experimental structures (instead of hypothetical structures as in hMOF and Tobacco). Compared with hMOF, CoREMOF has richer topologies and more complex structures. A large part of the experiments in this paper uses the CoREMOF dataset, such as adsorption, separation, etc. (see Supplementary Data 2). In addition, Tobacco50 is also a hypothetical dataset. Unlike hMOF, Tobacco uses a topology-guided method, thus the topologies are richer. The mechanical stability benchmark39 uses a part of the Tobacco dataset. In addition to these datasets, there are also some datasets used for specific works, such as heat capacity. For all datasets, we try to select ~5000 for training and 1000 for test and validation. For smaller datasets with less than 7000 samples, the training-validation-test data are randomly split into 8: 1: 1. All datasets, sources, training data volume, and tasks are listed in Supplementary Data 2. For pre-training, SpbNet is pre-trained on 95,370 structures, which are randomly sampled from the data provided by MOFTransformer20.
Data availability
The dataset of hMOF, CoREMOF, Tobacco is from https://mof.tech.northwestern.edu(ref. 51). For other porous materials, benchmark datasets are from https://figshare.com/articles/dataset/Benchmark_data/24884001. The heat capacity data is from https://archive.materialscloud.org/record/2022.53(ref. 52). The data of mechanical stability is from https://figshare.com/articles/dataset/Neural_zip/24316339(ref. 39). All used pre-trained material structures are from https://figshare.com/articles/dataset/1_million_hypothetical_MOFs/21810147(ref. 20). All the benchmark datasets used in this study have been deposited in the Figshare database under accession code https://figshare.com/articles/dataset/Benchmark_Dataset/28943996?file=54263099. Pre-trained weight has been deposited in the Zenodo database under accession code https://doi.org/10.5281/zenodo.1785514453. Source data are provided as a Source Data file. The Source Data generated in this study has been deposited in the Zenodo database under accession code https://doi.org/10.5281/zenodo.1785649154. Source data are provided with this paper.
Code availability
The code for data preprocessing, pre-training and fine-tuning is available at https://github.com/tyvanzou/spbnetunder the MIT LICENSE. The source code has also been deposited to Zenodo55.
References
Ross, J. et al. Large-scale chemical language representations capture molecular structure and properties. Nat. Mach. Intell. 4, 1256–1264 (2022).
Singhal, K. et al. Large language models encode clinical knowledge. Nature 620, 172–180 (2023).
Yu, M. et al. Deep learning large-scale drug discovery and repurposing. Nat. Comput. Sci. 4, 600–614 (2024).
Schäfer, R. et al. Overcoming data scarcity in biomedical imaging with a foundational multi-task model. Nat. Comput. Sci. 4, 495–509 (2024).
Bi, K., Xie, L., Zhang, H., Chen, X., Gu, X. & Tian, Q. Accurate medium-range global weather forecasting with 3d neural networks. Nature 619, 533–538 (2023).
Zhou, Y. et al. A foundation model for generalizable disease detection from retinal images. Nature 622, 156–163 (2023).
Deng, H. et al. Large-pore apertures in a series of metal-organic frameworks. Science 336, 1018–1023 (2012).
Wang, C., Liu, D. & Lin, W. Metal–organic frameworks as a tunable platform for designing functional molecular materials. J. Am. Chem. Soc. 135, 13222–13234 (2013).
Snyder, B. E. et al. A ligand insertion mechanism for cooperative NH3 capture in metal–organic frameworks. Nature 613, 287–291 (2023).
Nugent, P. et al. Porous materials with optimal adsorption thermodynamics and kinetics for co2 separation. Nature 495, 80–84 (2013).
Datta, S. J. et al. CO2 capture from humid flue gases and humid atmosphere using a microporous coppersilicate. Science 350, 302–306 (2015).
Zhao, X., Wang, Y., Li, D.-S., Bu, X. & Feng, P. Metal–organic frameworks for separation. Adv. Mater. 30, 1705189 (2018).
Yang, S. et al. Selectivity and direct visualization of carbon dioxide and sulfur dioxide in a decorated porous host. Nat. Chem. 4, 887–894 (2012).
Zhou, S. et al. Asymmetric pore windows in mof membranes for natural gas valorization. Nature 606, 706–712 (2022).
Jiang, Z. et al. Filling metal–organic framework mesopores with TiO2 for CO2 photoreduction. Nature 586, 549–554 (2020).
Blatov, V. A., Blatova, O. A., Daeyaert, F. & Deem, M. W. Nanoporous materials with predicted zeolite topologies. RSC Adv. 10, 17760–17767 (2020).
Batzner, S. et al. E (3)-equivariant graph neural networks for data-efficient and accurate interatomic potentials. Nat. Commun. 13, 2453 (2022).
Cui, J. et al. Direct prediction of gas adsorption via spatial atom interaction learning. Nat. Commun. 14, 7043 (2023).
Bucior, B. J. et al. Energy-based descriptors to rapidly predict hydrogen storage in metal–organic frameworks. Mol. Syst. Des. Eng. 4, 162–174 (2019).
Kang, Y., Park, H., Smit, B. & Kim, J. A multi-modal pre-training transformer for universal transfer learning in metal–organic frameworks. Nat. Mach. Intell. 5, 309–318 (2023).
Park, H., Kang, Y. & Kim, J. Enhancing structure–property relationships in porous materials through transfer learning and cross-material few-shot learning. ACS Appl. Mater. Interfaces 15, 56375–56385 (2023).
Shoghi, N. et al. From molecules to materials: pre-training large generalizable models for atomic property prediction. In Proc. International Conference on Learning Representations, 17183–17212 (ICLR, 2024).
Shi, K. et al. Two-dimensional energy histograms as features for machine learning to predict adsorption in diverse nanoporous materials. J. Chem. Theory Comput. 19, 4568–4583 (2023).
Schütt, K., Unke, O., Gastegger, M. Equivariant message passing for the prediction of tensorial properties and molecular spectra. In Proc. International Conference on Machine Learning, 9377–9388 (PMLR, 2021).
Dosovitskiy, A. et al. An image is worth 16×16 words: transformers for image recognition at scale. In Proc. International Conference on Learning Representations, (ICLR, 2021).
Hertz, A. et al. Prompt-to-prompt image editing with cross-attention control. In Proc. International Conference on Learning Representations, (ICLR, 2023).
Lee, K.-H., Chen, X., Hua, G., Hu, H. & He, X. Stacked cross attention for image-text matching. In Proc. European Conference on Computer Vision 201–216 (Springer Cham, 2018).
Vaswani, A. et al. Attention is all you need. In Proc. Advances in Neural Information Processing Systems 30 (Curran Associates, Inc., 2017).
Willems, T. F., Rycroft, C. H., Kazi, M., Meza, J. C. & Haranczyk, M. Algorithms and tools for high-throughput geometry-based analysis of crystalline porous materials. Microporous Mesoporous Mater. 149, 134–141 (2012).
Kim, E. J. et al. Cooperative carbon capture and steam regeneration with tetraamine-appended metal–organic frameworks. Science 369, 392–396 (2020).
Wilmer, C. E., Farha, O. K., Bae, Y.-S., Hupp, J. T. & Snurr, R. Q. Structure–property relationships of porous materials for carbon dioxide separation and capture. Energy Environ. Sci. 5, 9849–9856 (2012).
Wilmer, C. E. et al. Large-scale screening of hypothetical metal–organic frameworks. Nat. Chem. 4, 83–89 (2012).
Sikora, B. J., Wilmer, C. E., Greenfield, M. L. & Snurr, R. Q. Thermodynamic analysis of xe/kr selectivity in over 137000 hypothetical metal–organic frameworks. Chem. Sci. 3, 2217–2223 (2012).
Li, S., Chung, Y. G. & Snurr, R. Q. High-throughput screening of metal–organic frameworks for CO2 capture in the presence of water. Langmuir 32, 10368–10376 (2016).
Chung, Y. G. et al. Computation-ready, experimental metal–organic frameworks: A tool to enable high-throughput screening of nanoporous crystals. Chem. Mater. 26, 6185–6192 (2014).
Chung, Y. G. et al. Advances, updates, and analytics for the computation-ready, experimental metal–organic framework database: Core mof 2019. J. Chem. Eng. Data 64, 5985–5998 (2019).
Xie, T. & Grossman, J. C. Crystal graph convolutional neural networks for an accurate and interpretable prediction of material properties. Phys. Rev. Lett. 120, 145301 (2018).
Cao, Z., Magar, R., Wang, Y. & Barati Farimani, A. Moformer: self-supervised transformer model for metal–organic framework property prediction. J. Am. Chem. Soc. 145, 2958–2967 (2023).
Lee, J. et al. Optimal surrogate models for predicting the elastic moduli of metal–organic frameworks via multiscale features. Chem. Mater. 35, 10457–10475 (2023).
Zhou, G. et al. Uni-mol: a universal 3d molecular representation learning framework. In Proc. International Conference on Learning Representations, (ICLR, 2023).
Mercado, R. et al. In silico design of 2d and 3d covalent organic frameworks for methane storage applications. Chem. Mater. 30, 5069–5086 (2018).
Martin, R. L., Simon, C. M., Smit, B. & Haranczyk, M. In silico design of porous polymer networks: high-throughput screening for methane storage materials. J. Am. Chem. Soc. 136, 5006–5022 (2014).
Deeg, K. S. et al. In silico discovery of covalent organic frameworks for carbon capture. ACS Appl. Mater. Interfaces 12, 21559–21568 (2020).
Kim, B., Lee, S. & Kim, J. Inverse design of porous materials using artificial neural networks. Sci. Adv. 6, 9324 (2020).
Wang, P., Zheng, W., Chen, T. & Wang, Z. Anti-oversmoothing in deep vision transformers via the Fourier domain analysis: from theory to practice. In Proc. International Conference on Learning Representations, (ICLR, 2022).
Roy, O. & Vetterli, M. The effective rank: a measure of effective dimensionality. In Proc. 2007 15th European Signal Processing Conference pp. 606–610 (PTETiS, 2007).
Rappé, A. K., Casewit, C. J., Colwell, K., Goddard III, W. A. & Skiff, W. M. Uff, a full periodic table force field for molecular mechanics and molecular dynamics simulations. J. Am. Chem. Soc. 114, 10024–10035 (1992).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proc. IEEE Conference on Computer Vision and Pattern Recognition, 770–778 (IEEE Computer Society, 2016).
Devlin, J., Chang, M.-W., Lee, K. & Toutanova, K. Bert: pre-training of deep bidirectional transformers for language understanding. In Proc. 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, Volume 1 (long and Short Papers), 4171–4186 (Association for Computational Linguistics, 2019).
Colón, Y. J., Gómez-Gualdrón, D. A. & Snurr, R. Q. Topologically guided, automated construction of metal–organic frameworks and their evaluation for energy-related applications. Cryst. Growth Des. 17, 5801–5810 (2017).
Bobbitt, N. S. et al. Mofx-db: An online database of computational adsorption data for nanoporous materials. J. Chem. Eng. Data 68, 483–498 (2023).
Moosavi, S. M. et al. A data-science approach to predict the heat capacity of nanoporous materials. Nat. Mater. 21, 1419–1425 (2022).
Zou, J. et al. A data-efficient foundation model for porous materials based on expert-guided supervised learning. Zenodo https://doi.org/10.5281/zenodo.17855144 (2025).
Zou, J. et al. A data-efficient foundation model for porous materials based on expert-guided supervised learning. Zenodo https://doi.org/10.5281/zenodo.17856491 (2025).
Zou, J. et al. A data-efficient foundation model for porous materials based on expert-guided supervised learning. tyvanzou/spbnet https://doi.org/10.5281/zenodo.18168110 (2025).
Feng, R. et al. Rank diminishing in deep neural networks. Adv. Neural Inf. Process. Syst. 35, 33054–33065 (2022).
Patel, N.N. & Shwartz-Ziv, R. Learning to compress: local rank and information compression in deep neural networks. In Proc. Workshop on Machine Learning and Compression, NeurIPS (Openreview, 2024).
Acknowledgements
We gratefully acknowledge support for this work provided by National Key R & D Program of China (Grant No.: 2024YFC3406401 (X.L.), 2024YFE0101100 (D.Z.)), National Natural Science Foundation of China (NSFC) (Grant Nos.: 62472102 (B.Y.), 62372117 (W.T.), 623B1004 (J.Z.), 22525501 (X.L.), 22475049 (X.L.), 22088101 (D.Z.)), Natural Science Foundation of Shanghai (Grant No.: 24ZR1490400 (W.T.)), Shanghai Pilot Program for Basic Research-Fudan University (Grant No.: 22TQ004 (X.L.)), the Science and Technology Commission of Shanghai Municipality (Nos. 2024ZDSYS02 (X.L.), 25PY2600100 (D.Z.)).
Author information
Authors and Affiliations
Contributions
D.Z., B.Y., and X.L. conceived the study. X.L, W.T., and Q.L. reviewed the manuscript and provided critical revision suggestions. J.Z. conducted experiments. Z.L. collected the datasets. J.Z. and Z.L. wrote the manuscript. T.W. and Z.W. helped preprocessing the dataset. R.L. and Y.Y. helped the design of the computational chemistry part in the manuscript. All authors discussed the results and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Philipp Benner and the other anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zou, J., Lv, Z., Tan, W. et al. A data-efficient foundation model for porous materials based on expert-guided supervised learning. Nat Commun 17, 2618 (2026). https://doi.org/10.1038/s41467-026-69245-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69245-y