Abstract
Colorectal cancer liver metastasis (CRLM) is a leading cause of mortality, driven by poorly defined molecular interactions within the hepatic niche. Here, we identify a distinct population of pro-metastatic Early Growth Response 1 (Egr1)+ neutrophils that accumulate in the pre-metastatic liver. Mechanistically, we show that KIAA1199-high cancer cells secrete granulin-rich extracellular vesicles, which are internalized by hepatocytes. This uptake triggers a subset of functionally reprogrammed hepatocytes, characterized by a profound metabolic reprogramming and the suppression of peroxisome proliferator-activated receptor gamma (PPARγ) signaling, leading to increased secretion of Serum Amyloid A2 (SAA2). Hepatocyte-derived SAA2 subsequently activates Formyl Peptide Receptor 2 (FPR2) on neutrophils, stabilizing Egr1-driven transcriptional program via the PI3K-AKT pathway to enhance neutrophil survival and pro-angiogenic activity. These Egr1+ neutrophils co-localize with reprogrammed hepatocytes at the tumor-liver interface, where they promote vascular remodeling to facilitate metastatic colonization. Pharmacological restoration of PPARγ or FPR2 inhibition abrogate CRLM in preclinical models in female mice. Furthermore, a combined KIAA1199-SAA2 signature predicts liver metastasis risk in patients. Our findings delineate a KIAA1199-PPARγ/SAA2-Egr1 axis orchestrating the pre-metastatic niche and propose metabolic normalization as a preventative strategy for liver metastasis.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) represents a formidable global health challenge as the second most diagnosed malignancy and the fifth leading cause of cancer-related mortality worldwide1,2. The clinical trajectory of CRC is dramatically altered by metastatic spread, with liver metastasis occurring in over 50% of patients and serving as the predominant cause of cancer-specific mortality3. Despite advances in multimodal therapy, the prognosis for colorectal liver metastasis (CRLM) remains poor, with a 5-year survival rate of only 12–30%4. Elucidating the cellular and molecular mechanisms that govern metastatic tropism to the liver is therefore critical for identifying therapeutic targets and improving patient outcomes.
Successful metastatic colonization represents the culmination of a complex cascade that extends beyond the intrinsic properties of circulating tumor cells (CTCs). Before metastatic seeding occurs, primary tumors orchestrate the formation of a pre-metastatic niche (PMN)—a receptive microenvironment that facilitates the engraftment and outgrowth of disseminated tumor cells at distant sites5. The PMN is characterized by multiple hallmark features, including immunosuppression, chronic inflammation, extracellular matrix remodeling, and vascular alterations6. Among these, angiogenesis and vascular remodeling emerge as particularly critical early events7. Enhanced neovascularization not only provides essential oxygen and nutrients but also increases vascular permeability, thereby promoting cellular trafficking and molecular exchange within the niche and enabling tumor cell extravasation and survival8,9. However, a fundamental question remains: how do distant primary tumors transmit signals to precisely remodel the hepatic parenchyma and metabolic landscape to support these vascular and immune changes?
Emerging evidence suggests that the hepatic metabolic microenvironment plays a decisive role in PMN formation. The liver functions as a central metabolic hub, and perturbations in hepatocyte metabolism can drastically alter the local secretome, thereby influencing immune cell recruitment. Recent studies highlight tumor-associated neutrophils (TANs) as pivotal orchestrators of PMN establishment. These specialized myeloid cells are recruited to target organs through complex mechanisms involving chemokine gradients and increased vascular permeability10,11,12. Once localized within the incipient PMN, TANs promote metastasis through matrix remodeling (MMP-8/9), angiogenesis (VEGF, FGF, IL-8), immunosuppression via cooperation with other myeloid cells (TAMs, MDSCs)13, and the formation of neutrophil extracellular traps (NETs)14,15. Yet, the upstream intercellular signaling axes that link primary tumor-derived factors to hepatic metabolic reprogramming and subsequent neutrophil recruitment remain poorly defined.
KIAA1199, also known as cell migration-inducing hyaluronan-binding protein (CEMIP)16, has emerged as a key regulator of immune modulation and immune escape in CRC16,17,18. Our previous studies established that KIAA1199 is significantly upregulated in CRC tissues and strongly associated with invasion, metastasis, chemotherapeutic resistance, and poor clinical outcomes19,20,21,22,23. While KIAA1199 is known to facilitate local invasion, its capacity to remotely condition the pre-metastatic liver via secreted mediators, such as extracellular vesicles (EVs), warrants investigation.
In this work, we report a distinct subset of Egr1⁺ neutrophils that emerge within the hepatic PMN orchestrated by KIAA1199-high CRC. Mechanistically, we decipher a multi-step signaling cascade wherein KIAA1199 drives the secretion of Granulin (GRN)-rich EVs from CRC cells. Upon internalization by hepatocytes, these functional cargos induce a subset of functionally reprogrammed hepatocytes, characterized by a profound metabolic reprogramming and the suppression of PPARγ signaling. This metabolic shift directly enhances SAA2 expression and secretion. Hepatocyte-derived SAA2 engages FPR2 receptors on resident neutrophils, activating PI3K–AKT signaling that stabilizes an EGR1-driven transcriptional program promoting prolonged survival and enhanced pro-angiogenic capacity, thereby promoting vascular remodeling within PMN. By defining the KIAA1199/GRN-PPARγ/SAA2-FPR2/EGR1 axis as a central orchestrator of PMN establishment, our work provides mechanistic insight into how CRC cells remotely reprogram the hepatic metabolic and immune microenvironment to support metastatic colonization. Moreover, we identify metabolic normalization via PPARγ activation as a potential therapeutic strategy to disrupt the formation of the liver metastatic niche.
Results
KIAA1199 overexpression establishes a hepatic pre-metastatic niche driving colorectal cancer liver metastasis
Our previous studies demonstrated that elevated KIAA1199 expression is a hallmark of colorectal cancer (CRC) and is strongly associated with increased metastatic risk and poor prognosis19,20,24. We further validated these findings using both TCGA-CRC datasets and our independent clinical cohort (Fig. 1A–E, Supplementary Fig. 1A–C). Spatial transcriptomic analysis provided additional confirmation, revealing significantly higher KIAA1199 levels in tumor tissues compared to adjacent normal counterparts, in both primary colon tumors and matched liver metastases (Fig. 1F). These multi-platform validations collectively reinforce the robustness of our earlier observations. Our previous functional investigations established that primary CRC tumors with high KIAA1199 expression exhibit enhanced invasiveness and organ-metastatic potential19,20. In recent years, the pre-metastatic niche (PMN) has emerged as a pivotal contributor to metastatic colonization6. Therefore, beyond conferring intrinsic invasiveness to tumor cells, we investigated whether KIAA1199 contributes to metastatic progression by modulating the hepatic pre-metastatic microenvironment in colorectal cancer liver metastasis (CRLM).
A KIAA1199 expression in metastatic (M, n = 67 patients) versus non-metastatic (NM, n = 353 patients) CRC samples from TCGA cohort. Expression is significantly higher in the metastatic group (unpaired t test). B Kaplan–Meier cumulative incidence curve and Cox regression analysis of TCGA-CRC patients stratified by KIAA1199 expression (high: top 20%, n = 95; low: bottom 20%, n = 96). High KIAA1199 expression correlates with a significantly increased risk of disease progression (log-rank test). C Representative immunohistochemical (IHC) staining of KIAA1199 in CRC tissue microarrays (left). Corresponding MRI scans (right) track liver metastasis progression in patients from diagnosis (0 months) to 12- and 18-month follow-up (n = 104 patients). White arrows: baseline lesions; Red arrowheads: new lesions; Red dashed lines: maximum diameter of metastatic progression. Scale bars: 100 µm (IHC) and 1 cm (MRI). D Proportion of patients with distant metastasis in KIAA1199-low (n = 63 patients) versus KIAA1199-high (n = 41 patients) groups based on IHC scoring. The high-expression group showed a significantly higher metastatic rate (63% vs. 36%; Chi-square test). E Kaplan–Meier curves of cumulative liver metastasis incidence stratified by KIAA1199 expression (n = 104 patients, log-rank test). F Representative H&E staining (left) and spatial transcriptomic maps (right) showing KIAA1199 distribution in primary CRC (top) and matched CRLM (bottom) (n = 2 patients). Red dashed lines delineate tumor areas. Scale bars: 200 µm. G Schematic of the dual-tumor experimental timeline. BALB/c mice received orthotopic implantation of CT26 cells (KIAA1199 OE/KD or controls) on day 0. On day 7, wild-type CT26 cells were injected intrasplenically to mimic dissemination, followed by splenectomy 15 min later. Analysis was performed on day 21. H Representative gross liver images and H&E staining showing metastatic burden (n = 5 mice). Scale bars: 2 mm (gross) and 2 mm (histology). I–K Quantification of relative liver weight (I), number of metastatic nodules (J), and metastatic area percentage (K) (n = 5 mice, Student’s t test). L Schematic of the survival study workflow. Orthotopic implantation (KIAA1199 OE/KD or controls) was followed by intrasplenic injection on day 7. Survival was monitored until day 60. M Kaplan–Meier survival curves (n = 10 mice, log-rank test). All statistical tests are two-sided. Data are presented as mean ± s.d. Source data and exact p values are provided as a Source Data file.
To systematically examine this possibility, we developed a dual-tumor mouse model. First, orthotopic tumors were established using CRC cells engineered to express either high or low levels of KIAA1199. When the primary tumor was established but before any spontaneous metastasis occurred, we introduced a standardized liver metastasis model via spleen injection of CRC cells (Fig. 1G). This approach allowed us to evaluate how KIAA1199 expression in the primary tumor influences subsequent metastatic colonization. Strikingly, mice with KIAA1199-low orthotopic tumors showed limited metastatic burden and prolonged survival, whereas those bearing KIAA1199-high tumors exhibited markedly enhanced metastatic colonization and reduced overall survival (Fig. 1H–M, Supplementary Fig. 2A). Additionally, fluorescent labeling experiments with mCherry/GFP-tagged CRC cells demonstrated that all liver lesions originate from intrasplenic injections, not from the orthotopic primary tumor (Supplementary Fig. 2B). Collectively, these results demonstrate that KIAA1199-overexpressing primary CRC tumors promote functional pre-metastatic niche formation in the liver, thereby facilitating CRLM.
Pre-metastatic niche formation characterized by Egr1+ neutrophil subset infiltration
To delineate the mechanisms by which high KIAA1199 expression in primary CRC tumors remodels the hepatic microenvironment, we established paired orthotopic mouse models with high and low KIAA1199 expression. Analysis at fourteen days post-implantation, when no metastases were detectable by MRI, revealed a significant increase in liver-infiltrating neutrophils in KIAA1199-high mice compared to controls consistent with our previous findings (Fig. 2A, B; Supplementary Fig. 3A–E)22. Single-cell RNA sequencing followed by unsupervised clustering analysis identified a distinct neutrophil subset, designated Neu-C2 (Fig. 2C; Supplementary Fig. 4A–E). Pseudotime trajectory analysis positioned Neu-C2 at a terminal differentiation stage, and gene set enrichment analysis revealed significant upregulation of programs governing cell adhesion, angiogenesis, and cell cycle regulation (Fig. 2D, E), collectively indicating a specialized pro-metastatic phenotype.
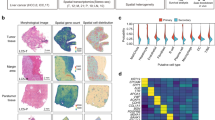
A Schematic of the pre-metastatic niche model. BALB/c mice were implanted orthotopically with CT26 (Vector or KIAA1199-OE) cells. On day 14 (pre-metastatic stage), livers were imaged by MRI and harvested for microenvironmental analyses. Scale bars: 2 mm (gross) and 2 mm (MRI). B scRNA-seq analysis showing tSNE projections (left) and cell type proportions (right) of hepatic immune and non-immune cells. Red dashed lines: neutrophils; Blue dashed lines: hepatocytes (n = 4 mice). C Subclustering of hepatic neutrophils identifies five distinct subpopulations (Neu-C0 to Neu-C4). Stacked bar plot shows their distribution. (n = 4 mice). D Pseudotime trajectory analysis (Monocle) of neutrophil subclusters (n = 4 mice). E Gene set variation analysis (GSVA) of biological pathways across neutrophil pseudotime trajectories (n = 4 mice). F Western blot analysis of neutrophils isolated from livers on day 14 following orthotopic CT26 implantation (n = 3 independent experiments). G Flow cytometry quantification of Egr1⁺ neutrophils in livers of BALB/c and C57BL/6 mice bearing Vector or KIAA1199-OE tumors (n = 3 mice, Student’s t test). H. Multiplex immunohistochemistry (mIHC) of liver sections showing representative Egr1⁺ neutrophils (white arrowheads) (n = 3 mice). Scale bars: 100 µm (left), 20 µm (right). I t-SNE projections from external scRNA-seq datasets showing Egr1⁺ neutrophil distribution in patients (Low vs. High KIAA1199) and tissue types (peritumoral liver tissue (Liver_P) vs. liver metastases (Liver_T)). Red dashed lines highlight the Egr1⁺ subset (n = 24 patients). J Spatial transcriptomic analysis of CRLM patients. Left: Egr1⁺ neutrophil distribution. Middle: Angiogenesis gene signature enrichment. Right: Spatial distance (k-nearest neighbor) from Egr1⁺ neutrophils to defined tissue zones. Red dashed lines: tumor boundary. K Representative H&E and mIHC images of mouse (top, n = 2 mice) and human (bottom, n = 2 patients) CRLM tissues. Markers: DAPI (blue), EGR1 (red), LY6G (green). Yellow arrowheads: co-localized cells. Scale bars: 100 µm. L Tube formation assay. HUVECs were co-cultured with conditioned media from dHL-60 cells transfected with EGR1 or Vector (n = 3 independent experiments; Scale bars: 20 µm.). M Schematic summary: Egr1+ neutrophils localize to the tumor microenvironment to promote neovascularization. FACS sequential gating strategies are shown in Supplementary Fig. 5G. All statistical tests are two-sided. Data are presented as mean ± s.d. Source data and exact p values are provided as a Source Data file.
To identify upstream transcriptional regulators of this neutrophil subset, we applied pySCENIC (Single-Cell rEgulatory Network Inference and Clustering) to derive the top transcription factors (TFs) linked to Neu-C2 and intersected these with differentially expressed genes (DEGs) (Supplementary Fig. 5A, B). Early Growth Response 1 (EGR1) emerged as a key regulator, being specifically upregulated in liver-infiltrating neutrophils from KIAA1199-high mice (Fig. 2F; Supplementary Fig. 5C, D). To validate the functional significance of EGR1, we performed gain-of-function experiments in differentiated HL-60 cells (dHL60, a human neutrophil-like cell line) and in co-culture systems (containing CRC cells, AML12 hepatocytes and neutrophils), simulating the hepatic microenvironment where neutrophils reside in vivo. EGR1 overexpression in dHL-60 cells and KIAA1199-driven EGR1 expression in neutrophils both recapitulated the Neu-C2 transcriptomic signature, supporting their identification as Egr1⁺ neutrophils (Supplementary Fig. 5E, F).
In further validation studies, hepatic accumulation of Egr1⁺ neutrophils increased significantly in KIAA1199-high tumor-bearing mice and in patients with CRLM (Fig. 2G–I; Supplementary Fig. 5G; Supplementary Fig. 6A–F). Notably, in metastatic lesions Egr1⁺ neutrophils localized predominantly to peritumoral regions rather than tumor cores and spatially overlapped with areas of active angiogenesis (Fig. 2J, K)25. Functionally, conditioned medium from Egr1⁺ neutrophils directly promoted endothelial tube formation in vitro (Fig. 2L; Supplementary Fig. 6G)26. Collectively, these data demonstrate that high KIAA1199-expressing CRC establishes a pro-metastatic liver niche by recruiting and programming EGR1-driven neutrophils, which in turn facilitate CRLM through enhanced angiogenesis (Fig. 2M).
Functionally reprogrammed hepatocytes orchestrate Egr1⁺ neutrophil positioning at the tumor-liver interface via SAA2-FPR2 axis
To trace the origin of Egr1⁺ neutrophils, we conducted systematic temporal and spatial profiling across multiple tissues. We observed that Egr1⁺ neutrophils progressively accumulated specifically in the liver of mice bearing KIAA1199-overexpressing tumors, while their frequency remained consistently low in all peripheral sites including bone marrow, spleen, and peripheral blood (Fig. 3A, B; Supplementary Fig. 7A–C). Given their terminally differentiated status (Fig. 2D), these data suggest that Egr1⁺ neutrophils are likely reprogrammed locally in the liver rather than recruited from the circulation.
A Flow cytometric quantification of Egr1⁺ neutrophils across five tissue compartments (tumor, LN, blood, spleen, liver) on day 14 (n = 3 mice, Student’s t test). B Time-course analysis of Egr1⁺ neutrophil frequency in the liver over 18 days (n = 3 mice per time point, two-way ANOVA). C UMAP visualization and re-clustering of hepatocytes into 13 subpopulations (Hepa-C0 to Hepa-C12). Bar plot shows proportional distribution. (n = 4 mice). D Pseudotime enrichment analysis of hepatocyte subpopulations (n = 4 mice). E. Spatial transcriptomic maps of two CRLM patients showing the localization of Hepa-C11 (functionally reprogrammed hepatocytes) and EMT signatures. Right: Spatial proximity (k-distance) analysis of Hepa-C11 to tumor borders, EMT regions, and Egr1⁺ neutrophils. F mIHC showing co-localization of SAA2 (cyan), EGR1 (red), and LY6G (green) in the liver of KIAA1199-high mice (n = 2 mice). Yellow arrowheads: Egr1⁺ neutrophils. Scale bars: 100 µm. G mIHC showing spatial distribution of Egr1⁺ neutrophils, SAA2, and CD31⁺ vasculature in human CRLM samples across tumor core, transitory, and leading edge regions (n = 2 patients). Scale bars: 100 µm. H–J In vivo validation. Mice bearing orthotopic tumors were treated with WRW4 (FPR2 antagonist, 2 mg/kg/day) or recombinant SAA2 (1 μg/kg/day). Livers were harvested on day 14 for neutrophil isolation. I Western blot of isolated neutrophils (n = 3 independent experiments). (J) Flow cytometry of hepatic Egr1+ neutrophils (n = 3 mice, Student’s t test). K Schematic of therapeutic intervention in the dual-tumor model. Mice received WRW4 (2 mg/kg/day) treatment. L Representative gross liver and H&E images at day 21 (n = 4 mice). Scale bars: 2 mm (gross) and 2 mm (histology). M, N Quantification of liver metastatic area (M) and nodule number (N) (n = 4 mice, Student’s t test). O. Kaplan–Meier survival curves over 60 days (n = 10 mice, log-rank test). FACS sequential gating strategies are shown in Supplementary Fig. 5G. Where applicable, all statistical tests are two-sided. Data are presented as mean ± s.d. Source data and exact p values are provided as a Source Data file.
Cell-cell communication analysis revealed significantly enhanced hepatocyte-neutrophil interactions in the KIAA1199-high group compared to controls, accompanied by notable shifts in hepatocyte cluster composition (Fig. 2B; Supplementary Fig. 8A). These findings implicate hepatocyte-derived signals in shaping neutrophil phenotype and support a model of liver-driven in situ reprogramming of Egr1⁺ neutrophils. Consistent with the spatial and functional features of Egr1⁺ neutrophils, we identified a distinct hepatocyte subpopulation—Hepa-C11—present almost exclusively in mice bearing KIAA1199-overexpressing orthotopic CRC tumors (Fig. 3C; Supplementary Fig. 8B–D). Pseudotime trajectory analysis positioned this cluster at an intermediate stage along the developmental trajectory, while copy number variation (CNV) analysis revealed genomic instability patterns characteristic of high malignant potential, prompting us to designate this cluster as “functionally reprogrammed hepatocytes” (Supplementary Fig. 9A, B). Functionally, functionally reprogrammed hepatocytes exhibited a transcriptomic profile significantly enriched in angiogenesis, cell adhesion, and immunoregulatory pathways, closely mirroring the pro-tumor characteristics of Egr1⁺ neutrophils (Fig. 3D). Ligand-receptor interaction analysis identified strong SAA2–FPR2 signaling between functionally reprogrammed hepatocytes and Egr1⁺ neutrophils (Supplementary Fig. 9C–G). Spatial transcriptomic and multiplex immunofluorescence analyses of CRLM patient samples revealed that these two cell populations co-localized specifically at the tumor-liver interface, overlapping with regions exhibiting epithelial-mesenchymal transition (EMT) markers. This spatial relationship was consistently observed in both human CRLM specimens and mouse models (Fig. 3E–G, Supplementary Fig. 9H, Supplementary Fig. 10A–D).
To validate the functional importance of this cellular interaction, we co-cultured hepatocytes with KIAA1199-overexpressing CRC cells, which led to robust upregulation and secretion of SAA2. This phenomenon was also observed in vivo in orthotopic CRC models (Supplementary Fig. 10E–I). Importantly, recombinant SAA2 was sufficient to induce the generation of Egr1⁺ neutrophils in liver both in vitro and in vivo rather than peripheral blood, spleen, primary tumors, lymph nodes. Whereas pharmacological blockade of FPR2 using the selective FPR2 antagonist WRW4 abrogated this effect, thereby confirming the functional importance of the SAA2–FPR2 axis in promoting the local acquisition of the Egr1⁺ neutrophil phenotype (Fig. 3H–J; Supplementary Fig. 10J–M). In dual-tumor models, inhibition of SAA2–FPR2 signaling using WRW4 significantly reduced liver metastasis and prolonged survival, without affecting the growth of the orthotopic primary tumor (Fig. 3K–O; Supplementary Fig. 11A–D).
Collectively, these findings demonstrate that functionally reprogrammed hepatocytes orchestrate the positioning and phenotypic activation of Egr1⁺ neutrophils via SAA2–FPR2 signaling axis at the tumor–liver interface. Furthermore, our results identify FPR2 as a promising therapeutic target for preventing colorectal cancer liver metastasis.
KIAA1199-driven vesicular granulin elicits hepatocyte reprogramming and SAA2 upregulation via PPARγ suppression
To elucidate the molecular mechanisms underlying SAA2 upregulation in functionally reprogrammed hepatocytes, we performed comprehensive transcriptomic analysis with ranked gene expression profiling and revealed marked suppression of the PPAR signaling pathway within this specific hepatocyte subpopulation (Fig. 4A, B). Given the central role of PPARs (PPARα, PPARβ, PPARγ) in hepatic metabolic regulation, we conducted targeted metabolomic analysis, which demonstrated a significant shift from lipid synthesis/storage toward malignancy-associated metabolic pathway, including enhanced glycolysis and glutaminolysis. This metabolic reprogramming supports increased energy production and stress adaptation—a hallmark of malignant transformation (Fig. 4C). Among PPAR members, PPARγ was the most significantly downregulated in functionally reprogrammed hepatocytes (Fig. 4D). To investigate the functional relationship between PPARγ and SAA2, we activated PPARγ using rosiglitazone (Rog), a selective agonistt. This intervention significantly reduced SAA2 expression and protein secretion, suggesting a negative regulatory relationship (Fig. 4E; Supplementary Fig. 12A–C). This inverse relationship was further supported by their spatial co-localization in CRLM liver tissue and by transcriptional data from functionally reprogrammed hepatocytes (Fig. 4F).
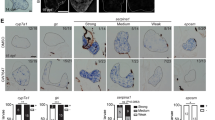
A Gene set enrichment analysis (GSEA) showing significant downregulation of the PPAR signaling pathway in Hepa-C11 hepatocytes. Enrichment score (ES), normalized enrichment score (NES), P value, and false discovery rate (FDR, Benjamini–Hochberg corrected) are indicated (n = 4 mice). B Scaled heatmap of key PPAR pathway–related genes across hepatocyte subclusters (Hepa-C1 to Hepa-C12), with cell numbers shown above each column (n = 4 mice). C Heatmap of metabolic pathway activities across hepatocyte subclusters (Hepa-C1 to Hepa-C12). Color scale represents relative pathway activity, and the red box highlights the distinctive metabolic pattern of Hepa-C11 (n = 4 mice). D Western blot of PPAR isoforms (α, β/δ, γ) in fractionated AML12 hepatocytes following co-culture with CRC cells (Vector vs. KIAA1199) (n = 3 independent experiments). E Western blot analysis of PPARγ and SAA2 in liver tissues from orthotopic tumor-bearing mice treated with or without rosiglitazone (Rog, 10 μg/kg/day) and sacrificed on day 14 (n = 3 independent experiments). F Spatial transcriptomic maps showing SAA2 (left) and PPARG (right) expression in two colorectal liver metastasis (CRLM) samples. Red dashed lines delineate tumor boundaries. G Predicted PPARγ binding motifs (P1 and P2) within the mouse Saa2 promoter region (−100 bp to +2000 bp). H Dual-luciferase reporter assays in AML12 cells using wild-type or mutant Saa2 promoter constructs. Mutation of the P2 site abolished PPARγ-mediated repression of Saa2 (n = 3 independent experiments, Student’s t test). I ChIP-qPCR confirming PPARγ binding to the Saa2 promoter in AML12 cells (n = 3 independent experiments, Student’s t test). J Schematic of the triple co-culture system (CRC + AML12 + Neutrophils) treated with Rog (10 μM). K ELISA quantification of SAA2 in co-culture supernatants (n = 3 mice, Student’s t test). L–M Flow cytometry quantification of Egr1+ neutrophils generated in the co-culture system using C57BL/6 (L) and BALB/c (M) neutrophils (n = 3 mice, Student’s t test). N Schematic of Rog treatment in the dual-tumor mouse model. O Representative gross liver and H&E images (n = 4 mice). Scale bars: 2 mm (gross) and 2 mm (histology). P–R Quantification of liver metastatic area (P), metastatic nodules (Q), and liver weight (R; n = 4 mice, Student’s t test). S Kaplan–Meier survival curves comparing KIAA1199 and KIAA1199+Rog groups (n = 10 mice, log-rank test). FACS gating strategies are shown in Supplementary Fig. 5G. Where applicable, all statistical tests are two-sided. Data are presented as mean ± s.d. Source data and exact p values are provided as a Source Data file.
To determine whether PPARγ directly regulates SAA2 transcription, we conducted bioinformatic analysis using UCSC Genome Browser and JASPAR, identifying two evolutionarily conserved PPARγ-binding motifs within the SAA2 promoter region (–2,000 to +100 bp) (Fig. 4G). To test the functional significance of these binding sites, we performed dual-luciferase reporter assays, which confirmed that PPARγ suppresses SAA2 promoter activity. Specifically, overexpression of PPARγ significantly reduced wild-type promoter-driven luciferase activity, whereas the SAA2-MT2 construct (mutation at binding site 2) abolished this repression. In contrast, SAA2-MT1 (binding site 1 mutant) retained PPARγ responsiveness, indicating that site 2 is essential for PPARγ-mediated transcriptional inhibition (Fig. 4H). ChIP-qPCR further demonstrated the direct binding of PPARγ to the SAA2 promoter in hepatocytes, confirming the specificity of this regulatory axis (Fig. 4I). Functionally, activation of PPARγ by rosiglitazone not only suppressed SAA2 production but also reduced the generation of Egr1⁺ neutrophils in vitro and in vivo (Fig. 4J–M; Supplementary Fig. 12D, E). To evaluate the therapeutic potential of targeting this pathway, we treated orthotopic CRC models with rosiglitazone (Rog). This intervention significantly attenuated KIAA1199-driven liver metastasis and prolonged overall survival (Fig. 4N–S; Supplementary Fig. 12F).
To decipher the intercellular signaling axis bridging KIAA1199-high CRC cells and hepatocytes, we performed integrated proteomic and transcriptomic analyses. This multi-omics approach implicated KIAA1199 as a potent driver of extracellular vesicle (EV) biogenesis and secretion (Supplementary Fig. 13A). Consistent with this, quantitative characterization of purified EVs confirmed that KIAA1199 overexpression significantly enhances EV release. Nanoparticle Tracking Analysis (NTA) demonstrated a marked increase in particle concentration in the KIAA1199-high group compared to the low-expression group (Supplementary Fig. 13B), a finding further corroborated by the elevated abundance of the key ESCRT biogenesis marker TSG101 (Supplementary Fig. 13C). Morphologically, while EVs from both groups maintained a standard exosomal size range (30-150 nm), the KIAA1199-high group exhibited a subtle shift towards a smaller average diameter, potentially reflecting a distinct sub-population of vesicles preferentially secreted upon KIAA1199 induction. Critically, the internalization of these KIAA1199-high CRC-derived EVs by hepatocytes proved essential for inducing the “functionally reprogrammed” phenotype (Supplementary Fig. 13D, E). Through cargo profiling, we identified granulin (GRN) as a pivotal downstream effector preferentially enriched in KIAA1199-induced EVs (Supplementary Fig. 13F–H). Strikingly, these GRN-laden EVs were efficiently taken up by hepatocytes and were sufficient to recapitulate the transcriptional and metabolic rewiring observed in functionally reprogrammed hepatocytes (Supplementary Fig. 13I–K).
Collectively, our study delineates a metastatic cascade wherein KIAA1199 remodels the secretome of CRC cells, promoting the release of GRN-rich EVs. The uptake of these vesicles by hepatocytes instigates a metabolic reprogramming characterized by PPARγ suppression and consequent SAA2 upregulation. This signaling axis ultimately fosters a pro-metastatic microenvironment. Furthermore, our data highlight the therapeutic promise of the PPARγ agonist rosiglitazone, which effectively disrupts this KIAA1199-GRN-PPARγ-SAA2 axis, thereby normalizing hepatic metabolism and preventing the formation of the pre-metastatic niche in CRLM.
Liver-derived Egr1⁺ neutrophils promote metastasis through extended survival and VEGFA secretion via PI3K/AKT/EGR1 circuit
To directly evaluate the pro-tumorigenic activity of Egr1⁺ neutrophils, we established a functional metastasis model in which we co-injected Egr1⁺ neutrophils with CRC cells via intrasplenic injection (Fig. 5A). Compared to control neutrophils, Egr1⁺ neutrophils markedly accelerated CRLM formation and reduced overall survival (Fig. 5B–E; Supplementary Fig. 14A). To elucidate the molecular mechanisms underlying this enhanced pro-metastatic activity, we performed gene set enrichment analysis (GSEA), which revealed significant activation of NF-κB, PI3K–AKT, and MAPK pathways in Egr1⁺ neutrophils. (Fig. 5F). Through systematic phosphoproteomic screening of neutrophils in the KIAA1199 co-culture system, we identified PI3K-AKT signaling as the primary response pathway activated following SAA2-FPR2 engagement (Fig. 5G). In time-course experiments, we observed that the proportion of Egr1⁺ neutrophils progressively increased in culture, accompanied by elevated Ki67 expression and reduced apoptotic signatures, indicating significantly prolonged survival compared to Egr1⁻ neutrophils (Fig. 5H–J). This extended lifespan represents a critical functional advantage, as conventional neutrophils typically survive only 1-2 days. Moreover, Egr1⁺ neutrophils displayed markedly increased VEGFA expression and secretion (Fig. 5K–N; Supplementary Fig. 14B, C), and blocking VEGFA (using a VEGFA–neutralizing antibody) markedly impairs the pro-angiogenic activity of Egr1⁺ neutrophils (Supplementary Fig. 14D–F), implicating VEGFA as a key effector of their enhanced pro-angiogenic capacity. Importantly, pharmacological inhibition of PI3K–AKT signaling with Wortmannin abrogated both the survival advantage and pro-angiogenic functions of Egr1⁺ neutrophils. These findings confirm that liver-derived Egr1⁺ neutrophils, upon receiving SAA2–FPR2 signals, extend their lifespan and promote angiogenesis via the PI3K/AKT/EGR1 circuit (Fig. 5O–R).
A Schematic of the adoptive transfer model. Neutrophils were pre-treated ex vivo with rSAA2 (20 μg/mL, 12 h) to induce the Egr1⁺ phenotype, then co-injected intrasplenically with CT26 cells (1:1). Splenectomy was performed 15 min later, and mice were analyzed on day 14. B Representative liver and H&E images from Neu (Egr1⁻ neutrophils) and Egr1⁺ Neu groups. Red arrowheads: metastases (n = 5 mice, Student’s t test). Scale bars: 2 mm (gross) and 2 mm (histology). C, D Quantification of metastatic nodules (C) and metastatic area (D) (n = 5 mice, Student’s t test). E Kaplan–Meier survival curves comparing Neu and Egr1⁺ Neu groups (n = 10 mice, log-rank test). F GSEA showing enrichment of NFκB, PI3K-AKT, and MAPK pathways in Egr1+ neutrophils (FDR, Benjamini–Hochberg corrected; n = 4 mice). G Phospho-proteomic Western blot screening of neutrophils from triple co-culture, identifying PI3K-AKT activation. Heatmap summarizes relative band intensities (n = 3 independent experiments). H Liver-infiltrating neutrophils were isolated and cultured ex vivo with rSAA2 (20 μg/mL) for 0–16 h. Flow cytometry quantified Egr1⁺ neutrophils over time (n = 3 mice, two-way ANOVA). I Cell death analysis (viability dye) over time (n = 3 independent experiments, two-way ANOVA). J Ki67 proliferation assay (n = 5 mice, Student’s t test). K, L ELISA of VEGFA in supernatants from triple co-cultures (n = 3 independent experiments, Student’s t test). M. Western blot analysis of dHL-60 cells and liver-infiltrating neutrophils from C57BL/6 and BALB/c mice treated with rSAA2 (20 μg/mL) with or without the FPR2 antagonist WRW4 (10 μM). Heatmap shows densitometric quantification (n = 3 independent experiments). N ELISA of VEGFA secretion under the conditions in (M) (n = 3 independent experiments, Student’s t test). O Western blot analysis of dHL-60 cells treated with rSAA2 with or without the PI3K inhibitor Wortmannin (100 nM) (n = 3 independent experiments). P ELISA of VEGFA secretion under the conditions in (O) (n = 3 independent experiments, Student’s t test). Q Western blot of cleaved caspase-3 expression (n = 3 independent experiments, Student’s t test). R Time-course cell death analysis of Egr1⁻ and Egr1⁺ neutrophils stimulated with rSAA2, with or without Wortmannin (n = 3 independent experiments, two-way ANOVA). FACS gating strategies are shown in Supplementary Fig. 5G. Where applicable, all statistical tests are two-sided. Data are presented as mean ± s.d. Source data and exact p values are provided as a Source Data file.
Collectively, these findings establish a mechanistic link whereby functionally reprogrammed hepatocytes orchestrate the functional activation of Egr1⁺ neutrophils might through FPR2–PI3K/AKT/EGR1 signaling axis, thereby driving CRLM through prolonged neutrophil survival and enhanced angiogenesis.
Egr1⁺ neutrophils drive angiogenesis to accelerate KIAA1199-mediated CRLM
Our functional studies revealed that Egr1⁺ neutrophils possess potent pro-angiogenic activity that is mechanistically linked to their pro-metastatic function (Fig. 6A, B). To investigate this relationship in vivo, we examined vascular patterns in KIAA1199-overexpressing orthotopic models. These analyses revealed significantly increased hepatic neovascularization that spatially co-localized with Egr1⁺ neutrophils infiltration (Fig. 6C). This same pattern—enhanced angiogenesis with elevated Egr1⁺ neutrophil infiltration—was consistently observed in KIAA1199-driven CRLM mice and in human CRLM specimens. Quantitative spatial analysis demonstrated that both cell populations predominantly localized to the tumor–liver interface rather than intratumorally regions (Fig. 6D, E). These consistent spatial relationships strongly implicate Egr1⁺ neutrophils in driving angiogenesis-dependent promotion of CRLM.
A Subcutaneous tumor growth in mice co-injected with CRC cells and either Egr1⁺ or control neutrophils. Tumor volume was monitored for 21 days. Inset shows representative tumors at endpoint (n = 5 mice, two-way ANOVA). B CD31 immunofluorescence (green) and vascular density quantification in subcutaneous tumors. Yellow arrowheads indicate endothelial structures (n = 5 mice, Student’s t test). Scale bars: 100 µm. C mIHC of pre-metastatic livers (day 14) showing spatial association of Egr1+ neutrophils (white arrowheads) and CD31⁺ vessels (yellow arrowheads). Scale bars: 100 µm. D, E Co-localization analysis of Egr1⁺ neutrophils and CD31⁺ vasculature in mouse (D, n = 2 mice) and human (E, n = 2 patients) CRLM samples, highlighting the tumor boundary. Scale bars: 100 µm. F, I SVEC4-10 endothelial cell viability (CCK-8) (n = 3 independent experiments, Student’s t test). G, J Endothelial functional assays using supernatants from triple co-cultures treated with Rog or WRW4 (n = 3 independent experiments, Student’s t test). H, K Endothelial migration (scratch wound assay) (n = 3 independent experiments, Student’s t test). L Representative tube formation images (HUVEC and SVEC4-10) treated with neutrophil-conditioned media (n = 3 independent experiments). Scale bars: 20 µm. M, N Quantification of vessel numbers (M) and junction points (N) (n = 3 independent experiments, Student’s t test). Where applicable, all statistical tests are two-sided. Data are presented as mean ± s.d. Source data and exact p values are provided as a Source Data file.
To directly assess the functional significance of this relationship, we collected conditioned media from Egr1⁺ neutrophils generated under multiple experimental conditions (co-culture systems containing CRC cells and AML12 hepatocytes) and tested their effects on endothelial cell function. These conditioned media significantly enhanced endothelial cell proliferation, migration, and tube formation compared to media from control neutrophils. Critically, pharmacological intervention at two key points in our identified pathway—either activating PPARγ with rosiglitazone or blocking FPR2 with WRW4—significantly abrogated these pro-angiogenic effects (Fig. 6F–N; Supplementary Fig. 15A–D; Supplementary Fig. 16A–J). In light of our previous data demonstrating that Egr1⁺ neutrophils upregulate and secrete VEGFA through the PI3K/AKT/EGR1 circuit (Fig. 5), blocking VEGFA (using a VEGFA–neutralizing antibody) dramatically attenuates the angiogenic response resulted by the conditioned media from KIAA1199-driven Egr1⁺ neutrophils (Supplementary Fig. 16K–M). These results support a comprehensive model in which KIAA1199-induced functionally reprogrammed hepatocytes potentiate the angiogenic program of Egr1⁺ neutrophils via PPARγ/SAA2/FPR2 signaling axis, thereby accelerating CRLM through enhanced neovascularization at the tumor-liver interface.
KIAA1199-SAA2 Signature Enables Precise Risk Stratification for Colorectal Cancer Liver Metastasis
To translate our mechanistic insights into clinically actionable biomarkers, we collected paired primary tumor tissues and matched serum samples from CRC patients in Union Cohort #1 (n = 94). All samples were obtained during routine clinical care. Demographic variables including age and sex were recorded at the time of sample collection but were not used as covariates in the subsequent analyses. Correlation analysis demonstrated serum KIAA1199 levels accurately reflect intertumoral expression (Pearson’s r = 0.79, p < 0.001), establishing its validity as a surrogate biomarker (Supplementary Fig. 17A). Consistent with our hypothesis, serum KIAA1199 levels positively correlated with serum SAA2 concentrations (Fig. 7A). Importantly, both markers were significantly elevated in patients with liver metastasis compared with non-metastatic cases (Fig. 7B). Moreover, high serum KIAA1199 was associated with an increased risk of CRLM (Fig. 7C), in line with our earlier observation that intratumoral KIAA1199 predicts metastatic potential. Similarly, elevated SAA2 was independently associated with heightened metastatic risk (Fig. 7D). Notably, patients with concurrent high expression of KIAA1199 and SAA2 exhibited a substantially greater risk of developing liver metastasis compared to patients with high expression of either marker alone (Fig. 7E), underscoring the clinical value of these two factors as combined predictors of CRLM. Based on these findings, we developed a KIAA1199–SAA2 combined predictive model for risk stratification. In receiver operating characteristic (ROC) curve analysis, the combined model consistently outperformed either KIAA1199 alone or SAA2 alone in predicting liver metastasis (Fig. 7F; Supplementary Fig. 17B). Decision curve analysis further confirmed that the combined model provided greater net clinical benefit across a wide range of threshold probabilities (Fig. 7G). To facilitate clinical implementation, we developed a nomogram incorporating both biomarkers to estimate the probability of CRLM at 6, 12, and 24 months (Fig. 7H). The model demonstrated excellent calibration, with predicted probabilities closely matching observed outcomes. Prospective validation in Union Cohort #2 (n = 5), with longitudinal imaging follow-up, confirmed the predictive power of the model (Fig. 7I). External validation in an independent Union Cohort #3 (n = 70) further established the robustness and reproducibility of the model across diverse patient populations (Fig. 7J).
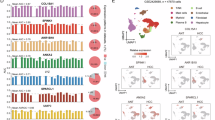
A Correlation between serum KIAA1199 and SAA2 protein levels (n = 94 patients, Spearman correlation). B Serum KIAA1199 and SAA2 levels in non-metastatic (NM) vs. metastatic (M) patients (NM, n = 47 patients; M, n = 47 patients; Student’s t test). C, D Kaplan–Meier analysis of liver metastasis-free survival stratified by serum KIAA1199 (C) and SAA2 (D) (high vs. low; n = 94 patients). E Kaplan–Meier analysis stratified by the combined serum KIAA1199 and SAA2 signature (High-High vs. Low-Low vs. Mixed; n = 94 patients, log-rank test). F Time-dependent ROC curves comparing prediction accuracy of KIAA1199 alone, SAA2 alone, and the combined model in Union Cohort #1 (n = 94 patients). G Decision curve analysis (DCA) showing net clinical benefit. H Nomogram integrating KIAA1199 and SAA2 for predicting CRLM probability at 6, 12, and 24 months. Three representative cases (Patients 1–3) are highlighted. I Prospective validation in Union Cohort #2 (n = 5 patients), showing longitudinal imaging of Patients 1–3 from initial diagnosis to recurrence detection. J External validation in Union Cohort #3 (n = 70): Kaplan–Meier curves (stratified by single or combined markers) and ROC analysis. K Schematic illustration of the proposed mechanism. High KIAA1199 expression in colorectal cancer induces a pre-metastatic niche characterized by enrichment of functionally reprogrammed hepatocytes and Egr1⁺ neutrophils at the tumor–liver interface. Mechanistically, KIAA1199-high CRC–derived GRN⁺ extracellular vesicles are taken up by hepatocytes, leading to suppression of PPARγ signaling, metabolic reprogramming, and increased SAA2 expression and secretion. SAA2 activates neutrophils via FPR2, promoting in situ differentiation into Egr1⁺ neutrophils. These neutrophils activate the PI3K–AKT pathway, suppress caspase-3-mediated apoptosis to extend survival, and induce VEGFA expression to enhance angiogenesis, collectively facilitating tumor colonization and metastatic progression. CRLM, colorectal cancer liver metastasis; PMN, pre-metastatic niche; CPM, clinical prediction model; FR-hHepatocytes, functionally reprogrammed human hepatocytes. Where applicable, all statistical tests are two-sided. Data are presented as mean ± s.d. Source data and exact p values are provided as a Source Data file.
Collectively, these results establish a stable and reproducible KIAA1199–SAA2 predictive model with strong performance for CRLM risk stratification across multiple independent cohorts. By providing a clinically practical framework for identifying high-risk patients who may benefit from intensified surveillance and early intervention, this model represents a promising avenue for clinical translation of our mechanistic findings.
Discussion
In this study, we identified a distinct population of Egr1⁺ neutrophils that specifically emerge within the pre-metastatic liver microenvironment shaped by KIAA1199-high colorectal tumors. Through the integration of differential gene signatures with transcription factor inference analysis, we identified EGR1 as the dominant regulator of this subset, as its overexpression in neutrophils was sufficient to recapitulate the distinctive transcriptional and functional programs of these cells. These findings establish EGR1 as a central coordinator of the neutrophils transcriptional reprogramming and, in turn, define Egr1⁺ neutrophils as a discrete functional entity. EGR1, a prototypical immediate early gene rapidly induced by diverse stimuli including growth factors and cytokines, has established roles in proliferation, differentiation, and vascular biology27. Consequently, it is frequently highly expressed in tumor tissues—particularly in metastatic or highly proliferative tumor cells—as also observed in our multicolor immunofluorescence analyses28,29.
Although neutrophils have traditionally been regarded as terminally differentiated and short-lived effectors, recent single-cell studies have revealed their remarkable transcriptional plasticity along a continuous “neutrotime” trajectory30. EGR1 is a pleiotropic transcription factor often upregulated in proliferating cancer cells. Within this framework, the observed EGR1 expression in metastatic CRC cells may reflect the highly proliferative and invasive state within the metastatic niche. Our data position EGR1 as a master regulator that drives neutrophils toward a specialized pro-angiogenic and pro-metastatic state. This finding aligns with prior work showing EGR1 functions as an active transcriptional regulator in tumor-associated neutrophils in non-small cell lung cancer31. Our results extend this concept by demonstrating that Egr1⁺ neutrophils in the pre-metastatic liver niche display prolonged survival and enhanced pro-angiogenic capacity, directly linking this phenotype to metastatic progression. We propose that EGR1 functions analogously to FOXP3 in regulatory T cells—a master regulator that fundamentally reprograms cell identity and function32. This perspective challenges the long-standing paradigm of neutrophils as homogeneous, short-lived effectors and redefines them as transcription factor-driven specialists with diverse functional states.
A key conceptual advance of our study is the elucidation of the upstream origin of Egr1⁺ neutrophils. In contrast to the prevailing paradigm that tumor-associated neutrophils are predominantly recruited from the circulation through chemokine gradients33,34, we demonstrate that Egr1⁺ neutrophils arise in situ within the inflamed hepatic parenchyma. Specifically, we identified a subset of functionally reprogrammed hepatocytes driven by KIAA1199-derived extracellular vesicles that suppress PPARγ signaling and upregulate SAA2, thereby “educating” local neutrophils into an EGR1-driven transcriptional state. This finding represents a significant conceptual shift from earlier reports emphasizing systemic neutrophil recruitment35,36, and instead highlights local hepatocyte–neutrophil crosstalk as the decisive driver of neutrophil specialization. Mechanistically, PPARγ, initially characterized as a regulator of lipid metabolism, functions as a critical orchestrator of metabolic and inflammatory homeostasis in hepatocytes37. Acting as a repressive transcription factor, PPARγ restrains pro-inflammatory programs and maintains hepatocytes in a quiescent state. Our data reveal that PPARγ directly represses SAA2 transcription, establishing a regulatory axis that links hepatocyte metabolic rewiring to neutrophil reprogramming. Within this context, SAA2 emerges as a pivotal molecular bridge between hepatocyte metabolism and the inflammatory microenvironment. Previous studies demonstrated that PPARγ agonists suppress SAA expression, whereas inhibition of PPARγ enhances its release37,38,39. Our results extend this paradigm by establishing the functional consequences of this regulation—specifically, the generation of Egr1⁺ neutrophils that drive pre-metastatic niche formation.
A key feature of our study is the spatial characterization of Egr1⁺ neutrophils within the pre-metastatic liver microenvironment. Using multiplex immunofluorescence and spatially resolved transcriptomics, we observed a striking enrichment and co-localization of both functionally reprogrammed hepatocytes and Egr1⁺ neutrophils at the tumor-parenchyma interface in established liver metastases. This finding indicates that these cells function not only in the initiation of metastasis but also as pioneering effectors at the invasive margin once metastasis is established. This spatial organization is functionally significant, as both cell populations express transcriptional programs linked to epithelial–mesenchymal transition, angiogenesis, and extracellular matrix remodeling. And Egr1⁺ neutrophils play a primary and independent role in driving pro-angiogenic remodeling within the tumor microenvironment. Our findings are conceptually distinct from previous reports that emphasized hepatocyte-derived serum amyloid A as a systemic mediator of pre-metastatic niche formation40 or described an “invasive zone” enriched in stromal and immune remodeling features35. However, those studies focused largely on STAT3-dependent systemic SAA secretion without resolving the specific cellular interactions at the invasive front. By contrast, our data reveal a localized mechanism where functionally reprogrammed hepatocytes suppress PPARγ signaling and upregulate SAA2 expression, thereby directly reprogramming adjacent neutrophils into an Egr1⁺pro-angiogenic state. This local hepatocyte-neutrophil axis integrates metabolic reprogramming and inflammation, providing a mechanistic explanation for how the CRLM microenvironment is established at the invasive margin. The translational implications of these findings are substantial. Our results suggest that circulating SAA2 and KIAA1199 levels may serve as predictive biomarkers of CRLM risk. Moreover, therapeutic interventions targeting this axis, particularly the PPARγ-SAA2-Egr1⁺ neutrophil pathway, could provide strategies for metastatic prevention by disrupting this specialized microenvironmental niche. It is noteworthy that, although rosiglitazone is a well-established PPARγ agonist, its systemic use has been associated with cardiovascular safety concerns, leading to prescribing restrictions in some countries41,42. In this study, rosiglitazone was employed solely as a short-term mechanistic tool, administered at a low dose (10 μg/kg/day) for a limited duration. Future studies should explore safer and more hepatocyte-targeted strategies to modulate PPARγ activity, to maintain therapeutic relevance while ensuring an appropriate safety margin.
Type 2 diabetes (T2D) and hyperglycemia are recognized risk factors for both the incidence and progression of CRC43. In our study, the PPARγ agonist rosiglitazone effectively blocked KIAA1199-mediated suppression of PPARγ signaling and attenuated CRLM, underscoring the therapeutic potential of targeting this pathway. This finding has intriguing implications for understanding the relationship between metabolic disease and cancer progression. Epidemiological studies support this notion, showing that patients with T2D have a higher risk of CRC, while mechanistic evidence indicates that hyperglycemia promotes CRLM by mobilizing monocytes via the CCL3–CCR1 axis44. Conversely, metformin-based regimens have been associated with improved outcomes in CRLM patients45, further reinforcing the concept that metabolic disease intersects with hepatic immune–metabolic reprogramming. Although our clinical dataset did not reveal a statistically significant association between diabetes and CRLM—likely due to limited sample size and the complex interplay of metabolic comorbidities—the biological mechanisms and preliminary clinical signals warrant further investigation. Modulating PPARγ activity in diabetic patients may provide dual benefits: restoring systemic glucose homeostasis while simultaneously disrupting the hepatocyte–neutrophil axis that supports CRLM. This perspective highlights an emerging paradigm in precision oncology—the potential to repurpose established metabolic drugs based on mechanistic understanding of the intersection between metabolism and cancer biology. By linking PPARγ agonists with disruption of the pre-metastatic niche, our findings suggest approaches for preventing metastasis in high-risk CRC patients.
Our study uncovers a distinct population of Egr1⁺ neutrophils that preferentially emerges within the pre-metastatic liver microenvironment orchestrated by KIAA1199-high CRC and delineates a comprehensive metastatic signaling axis that bridges primary CRC biology with hepatic immunometabolism (Fig. 7K). Mechanistically, we identify KIAA1199 not merely as a marker of aggression, but as a master regulator of vesicular communication, driving the secretion of Granulin (GRN)-rich extracellular vesicles. The uptake of these vesicles by hepatocytes induces a functionally reprogrammed state characterized by suppression of PPARγ signaling, metabolic rewiring, and robust upregulation of SAA2 expression and secretion. This hepatocyte-derived SAA2 acts as a critical paracrine signal, engaging the FPR2-PI3K-AKT cascade to educate a unique population of Egr1⁺ neutrophils. These neutrophils, endowed with prolonged survival and pro-angiogenic capabilities, co-localize with functionally reprogrammed hepatocytes to pioneer the pre-metastatic niche. Conceptually, this work challenges the long-standing view of neutrophils as a homogeneous population of short-lived effector cells passively recruited from the circulation and instead positions them as transcription factor-driven specialists shaped by local metabolic cues. By defining the “KIAA1199-GRN-PPARγ/SAA2-FPR2-EGR1” axis, we provide a mechanistic rationale for repurposing metabolic modulators, such as thiazolidinediones (e.g., rosiglitazone), to normalize the hepatic microenvironment and prevent metastatic seeding. Ultimately, this work integrates tumor cell biology, hepatocyte metabolism, and innate immunity into a unified framework, offering both a risk stratification signature and actionable therapeutic targets to intercept the lethal progression of colorectal liver metastasis.
Methods
All human studies were conducted under protocols approved by the Medical Ethics Committee of Tongji Medical College, Huazhong University of Science and Technology (Approval No. UHCT20210271-01). Animal experiments were performed in accordance with protocols approved by the Laboratory Animal Management and Use Committee of Hubei Provincial Center for Disease Control and Prevention (Approval No. 202410259).
Patients and tissue sample preparation
Human colorectal cancer (CRC) tissue samples were prospectively collected from patients with histopathological confirmed diagnoses who received treatment at Union Hospital, Tongji Medical College, Huazhong University of Science and Technology between September 2020 and August 2025. Patient sex and age were recorded at enrollment and were based on self-report. Sex and gender information was recorded but were not included as variables in the analysis, as the study focused on molecular and cellular mechanisms rather than sex- or gender-specific differences. No financial compensation was provided to study participants. All participants provided written informed consent after receiving a comprehensive explanation of the study protocol. Residual samples are stored at Union Hospital, Tongji Medical College, Huazhong University of Science and Technology and are available from the corresponding author upon reasonable request, subject to ethical approval.
For mouse liver tissue processing, specimens from control and experimental groups were harvested under sterile conditions, mechanically minced into approximately 1 mm³ fragments, and enzymatically dissociated into single-cell suspensions using collagenase IV (1 mg/mL; Sigma-Aldrich) and DNase I (0.1 mg/mL; Sigma-Aldrich) in serum-free DMEM at 37 °C for 30 min with gentle agitation. The resulting cell suspensions were filtered through 70 μm cell strainers, subjected to erythrocyte lysis using ACK buffer, and washed twice in PBS supplemented with 0.04% BSA. Cell viability was assessed by trypan blue exclusion, with a threshold of ≥90% viability required for downstream single-cell RNA sequencing (scRNA-seq) applications. According to the approved animal ethics protocol, the maximal permitted tumor burden did not exceed 10% of body weight, and mice were monitored closely for signs of distress. In all experiments, the maximal tumor size/burden was not exceeded, and animals were euthanized at predefined humane endpoints. Criteria for euthanasia included: (i) tumor burden exceeding 10% of body weight; (ii) weight loss >20% compared to baseline; (iii) inability to access food or water; (iv) signs of severe distress (e.g., hunching, ruffled fur, lethargy, hypothermia); (v) labored breathing; (vi) persistent bleeding, ulceration, or infection at the surgical site; (vii) severe pain unrelieved by analgesics; or (viii) any other condition assessed by trained personnel in accordance with the approved protocol. Sex was considered in the study design; to avoid additional variability associated with mixed-sex populations, only female mice were used in this study.
Library construction and sequencing
Single-cell RNA-seq libraries were generated using the “BD Rhapsody Single-Cell Analysis System” following the manufacturer’s protocols. Briefly, single cells were loaded into BD Rhapsody cartridges for bead-cell pairing, barcoding, and cDNA synthesis. Libraries were prepared using BD Rhapsody cDNA Kit and mRNA Whole Transcriptome Analysis (WTA) Amplification Kit. Final libraries were sequenced on the Illumina NovaSeq 6000 (paired-end, 150 bp).
Data processing and analysis
Raw sequencing data were processed using the “BD Rhapsody pipeline (Seven Bridges Genomics platform)”. Reads were aligned to the mouse genome (mm10), and gene-cell matrices were generated. Quality control, clustering, visualization (UMAP; tSNE), and differential gene expression analyses were performed using Seurat (v4.4.0) in R. Cells with <500 or >6000 genes or >10% mitochondrial content were excluded. Differentially expressed genes (DEGs) were identified by Wilcoxon rank-sum test (adjusted P < 0.05, |log₂FC | ≥ 0.25), and functional enrichment analyses were conducted using clusterProfiler (v4.12.6).
Validation using publicly available single-cell RNA sequencing and spatial transcriptomics data
Publicly available single-cell RNA-seq datasets (GSE225857)26 and spatial transcriptomics datasets25 from colorectal cancer liver metastases (CRLM) and matched primary colorectal tumors were downloaded and processed. Raw sequencing data were re-analyzed using the standard pipeline in Seurat. Data underwent quality control, normalization (LogNormalize method), dimensional reduction (PCA), and clustering (Louvain algorithm). Cell-type annotation was performed based on canonical marker genes, and differences in the cell type composition or gene expression profiles between metastatic and primary samples were compared and visualized using t-distributed stochastic neighbor embedding (t-SNE) or uniform manifold approximation and projection (UMAP). Tumor regions were identified based on histopathology (H\&E staining images) and spatial feature plots, and the spatial distribution of target gene (e.g., KIAA1199) expression was visualized using spatial heatmaps.
Pseudotime trajectory analysis
Pseudotime trajectory analysis was performed using Monocle (v2.3.2) in R to infer cellular differentiation trajectories among neutrophil and hepatocyte subclusters. Cells were ordered along the inferred pseudotime trajectories based on the identified highly variable genes. The pseudotime progression was visualized using trajectory plots annotated with cell subclusters.
Transcription factor regulatory network analysis
Single-cell transcription factor (TF) regulatory networks were inferred using pySCENIC (v0.11.2) in Python. Raw count data were used as input for gene regulatory network inference using GENIE3, followed by cis-regulatory motif enrichment analysis using RcisTarget with the provided database (mm10 motif database). TF regulon activities were calculated with AUCell. A regulon activity heatmap was generated to visualize TF activity across neutrophil subclusters, highlighting potential master regulatory transcription factors.
Cell-cell communication analysis
Cell-cell communication analysis was performed using CellChat (v1.6.1) in R. Normalized single-cell expression data from Seurat were used as input. Default CellChat ligand-receptor interaction databases for mouse (CellChatDB.mouse) were applied. Communication probabilities between identified cell types or subclusters were calculated based on expression levels of known ligand-receptor pairs. Significant ligand-receptor interactions were visualized using network diagrams and heatmaps. Differential communication signaling pathways between control and treated groups were further analyzed and identified based on statistical significance (p < 0.05).
Single-cell copy number variation (CNV) analysis
Single-cell CNV profiles were inferred using the inferCNV (v1.20.0) R package. Briefly, normalized single-cell RNA-seq expression data were imported from Seurat, with non-tumor cells (e.g., immune or stromal cells) set as reference cells. CNV estimations were conducted across chromosomal regions using default parameters. Results are visualized as violin plots showing relative CNV signals across cell populations, facilitating identification of malignant cells and assessment of tumor heterogeneity.
Cell culture and reagents
Murine colorectal cancer cell lines MC38 (Cat# CL-0972) and CT26 (Cat# CL-0071), murine hepatocyte line AML12 (Cat# CL-0602), murine endothelial cell line SVEC4-10 (Cat# CL-0221), human umbilical vein endothelial cells (HUVEC, Cat# CL-0675), and human myeloid cell line HL-60 (Cat# CL-0110) were obtained from Procell Life Science & Technology Co., Ltd. (Wuhan, China). Cell line authentication was performed using short tandem repeat (STR) profiling, and all lines were routinely tested for mycoplasma contamination using the MycoAlert Mycoplasma Detection Kit (Lonza).
Cells were maintained in either Dulbecco’s Modified Eagle Medium (DMEM, Cat# PM150210, Procell) or RPMI-1640 medium (Cat# PM150110, Procell) supplemented with 10% heat-inactivated fetal bovine serum (FBS; Cat# 164210, Procell) and 1% penicillin-streptomycin solution (Cat# PB180120, Procell) at 37 °C in a humidified atmosphere containing 5% CO₂. HL-60 cells were differentiated into neutrophil-like cells by treatment with 1.3% DMSO for 5 days, with differentiation confirmed by CD11b expression and morphological assessment.
The following reagents were used: WRW4 (FPR2 antagonist, Cat# HY-P1119), rosiglitazone (PPARγ agonist, Cat# HY-14600), wortmannin (PI3K inhibitor, Cat# HY-10197), and tribromoethyl alcohol (anesthetic, Cat# HY-B1372) from MedChemExpress (Monmouth Junction, NJ, USA); recombinant mouse SAA2 protein (rSAA2, Cat# RPB795Mu01) and recombinant human SAA2 protein (hrSAA2, Cat# RPB795Hu01) from Cloud-Clone Corp. (Wuhan, China). Detailed information regarding antibodies used in this study is provided in Supplementary Table 1.
Orthotopic colorectal cancer and liver metastasis mouse models
Six-week-old BALB/c or C57BL/6 mice (Changsheng Biotechnology Co., Ltd., Liaoning, China) were anesthetized via intraperitoneal injection of tribromoethyl alcohol. To establish the orthotopic colorectal cancer model, approximately 1 × 10⁶ colorectal cancer cells suspended in 50 μL of phosphate-buffered saline (PBS, Cat# PB180327, Procell, China) were injected into the submucosal layer of the cecum (on day 14, prior to visible metastasis, mice underwent MRI scanning. Imaging was performed on a Bruker BioSpec system without contrast agents. T2-weighted sequences were used for optimal lesion detection. All images were independently evaluated by three experienced radiologists blinded to the experimental groups to ensure objective analysis). Seven days later, to mimic liver metastasis, 5 × 10⁶ colorectal cancer cells suspended in 100 μL of PBS were injected into the spleen (or direct co-injected with Egr1⁺ neutrophils at a 1:1 ratio, with each cell type delivered at 2.5 × 10⁶ cells). The spleen was surgically removed 15 min post-injection, and the abdominal wall was sutured. Postoperative analgesia and supportive care were administered. Tumor growth was monitored regularly, and mice were sacrificed at predetermined time points to assess primary tumor burden and the extent of liver metastases. Mice were maintained under SPF conditions with a 12-h light/dark cycle at 22 ± 2 °C and 50–60% relative humidity, with free access to food and water. All animal experiments were approved by the Laboratory Animal Management and Use Committee of Hubei Provincial Center for Disease Control and Prevention (Approval No. 202410259).
Isolation of hepatic neutrophils
Mice were anesthetized and perfused via the portal vein with phosphate-buffered saline (PBS) to flush the liver. The liver was rapidly excised, minced, and gently dissociated through a 70 μm cell strainer to obtain a single-cell suspension. Red blood cells were lysed using red blood cell lysis buffer (Cat# C3702, Beyotime, China). The remaining cells were washed twice with PBS and centrifuged at 1000 × g for 10 min to collect the cell pellet.
Cells were resuspended in fluorescence-activated cell sorting (FACS) buffer, composed of PBS supplemented with 0.5% fetal bovine serum (FBS) and 2 mM EDTA (Cat# PB180620, Procell, China). Hepatic neutrophils were then isolated using the MojoSort™ Mouse Neutrophil Isolation Kit (Cat# 480058, BioLegend, San Diego, USA) according to the manufacturer’s protocol. Purified neutrophils were used for downstream analyses.
Flow cytometry analysis
Approximately 1 × 10⁶ cells were washed once with FACS buffer. Cells were then incubated with an appropriate mixture of fluorochrome-conjugated antibodies in the dark at room temperature for 30 min. After incubation, cells were washed twice with FACS buffer and centrifuged at 300 × g for 5 min each time. Finally, cells were resuspended in 100 μL of FACS buffer. Flow cytometric analysis was performed using a flow cytometer (Beckman Coulter). For each sample, a minimum of 2 × 10⁴ events was acquired. Data were analyzed using FlowJo software (version 10).
Immunohistochemistry (IHC)
Freshly harvested mouse liver and orthotopic colorectal tumor tissues were fixed in 4% paraformaldehyde (Cat# P0099, Beyotime, China) overnight, dehydrated through a graded ethanol series, embedded in paraffin, and sectioned at a thickness of 4 μm. Tissue sections were deparaffinized with xylene, rehydrated through a graded ethanol series, and subjected to antigen retrieval by microwave heating in sodium citrate buffer (pH 6.0). Endogenous peroxidase activity was quenched using 3% hydrogen peroxide, followed by blocking of nonspecific binding with 5% bovine serum albumin (BSA). Sections were then incubated with primary antibodies overnight at 4 °C. The next day, HRP-conjugated secondary antibodies were applied and incubated at room temperature for 1 h. Immunoreactivity was visualized using DAB chromogen, and nuclei were counterstained with hematoxylin. Slides were mounted and observed under a light microscope.
Co-immunoprecipitation (Co-IP) and Western Blot Analysis
Cells or tissue samples were lysed using RIPA buffer (Cat# P0013B, Beyotime, China) supplemented with protease inhibitor cocktail (Cat# ST505, Beyotime, China) to extract total protein. Protein concentrations were determined using the BCA Protein Assay Kit (Cat# P0012S, Beyotime, China). For co-immunoprecipitation, equal amounts of protein lysates were incubated overnight at 4 °C with the target-specific antibody or an isotype-matched IgG control under gentle rotation. The following day, Protein A/G magnetic beads (Cat# HY-K0202, MedChemExpress, USA) were added and incubated for 2–4 h at 4 °C. The beads were collected using a magnetic stand and washed three times with PBS (5 min per wash). Bound proteins were eluted by boiling the beads in SDS loading buffer for 5 min.
For Western blotting, equal amounts of protein samples were subjected to SDS-PAGE (Cat# PG112, Epizyme Biotech, Shanghai, China) and transferred onto PVDF membranes (Cat# ISEQ00010, Millipore, USA). Membranes were blocked with 5% non-fat dry milk or bovine serum albumin (BSA) in TBST buffer and incubated overnight at 4 °C with primary antibodies. After washing, membranes were incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies at room temperature for 1 h. Immunoreactive bands were visualized with the ChemiDoc imaging system (Bio-Rad, USA), and band intensities were quantified using ImageJ software.
Luciferase reporter assay and chromatin immunoprecipitation (ChIP) assay
Two putative PPAR-γ binding sites within the Saa2 promoter region were identified using the JASPAR online database (https://jaspar.genereg.net/). The promoter sequence of interest was cloned into a luciferase reporter plasmid (GENE CREATE, Wuhan, China) and transfected into target cells. After 48 h of transfection, cells were lysed, and luciferase activity was measured using the Dual-Luciferase® Reporter Assay System (Promega, USA), with firefly luciferase activity normalized to Renilla luciferase activity.
For ChIP assays, approximately 1 × 10⁷ cells were processed using a commercial ChIP kit (Cat# P2078, Beyotime, China) according to the manufacturer’s instructions to assess the direct binding of transcription factors to the promoter region. Cell lysates were immunoprecipitated overnight with PPAR-γ antibody-conjugated Protein A/G magnetic beads, with normal IgG used as a negative control. The immunoprecipitated chromatin complexes were then washed, eluted, and reverse cross-linked. Recovered DNA fragments were analyzed by quantitative PCR (qPCR) to assess the enrichment of the promoter region, normalized to input chromatin.
Lentiviral infection and plasmid transfection
Cells were seeded in 6-well plates and infected with lentiviral particles at approximately 50% confluence. After 12 h of incubation, the culture medium was replaced with fresh complete medium, and cells were maintained for an additional 48 h before downstream processing. Stable CRC cell lines with KIAA1199 overexpression, KIAA1199 knockdown, and the corresponding control vectors were generated by lentiviral transduction followed by puromycin selection, with the concentration and duration applied according to the manufacturer’s instructions18. Lentiviral particles were produced using a standard packaging system by Genecopoeia (Guangzhou, China). Stable expression or knockdown efficiency was validated by quantitative PCR and immunoblotting. Newly generated stable cell lines are available from the corresponding author upon reasonable request.
For plasmid transfection, cells were plated to reach 60–70% confluency. Transfection was performed using Lipofectamine™ 2000 reagent (Thermo Fisher Scientific, USA) according to the manufacturer’s instructions. Forty-eight hours post-transfection, cells were harvested for downstream applications such as Western blotting, qPCR, or functional assays. Detailed sequence information is provided in Supplementary Tables 1 and 3. Additional information is available from the corresponding author upon reasonable request.
CCK-8, wound healing, transwell, and tube formation assays
Cell proliferation was measured using the Cell Counting Kit-8 (Cat# C0037, Beyotime, China). Cells were seeded in 96-well plates, and CCK-8 reagent was added at the indicated time points. After incubation for 1–2 h, absorbance at 450 nm was measured using a microplate reader.
For wound healing assays, confluent monolayers were scratched with a sterile pipette tip. After washing with PBS, cells were cultured in serum-free medium, and wound closure was imaged at 0 and 24 h.
Transwell migration assays were performed using 8.0 μm pore size Transwell inserts (Corning, USA). Cells suspended in serum-free medium were added to the upper chamber, while medium containing 10% FBS was placed in the lower chamber. After 24 h, migrated cells were fixed, stained, and counted.
For tube formation assays, HUVECs or SVEC4-10 cells were seeded onto Matrigel-coated 96-well plates and incubated for 4–6 h. Tube-like structures were imaged and quantified using ImageJ software.
Extracellular vesicle (EV) isolation
Exosome-enriched EVs were isolated from conditioned medium using the BeyoExo™ Enhanced Exosome Isolation Kit for Cell Culture Media (Beyotime Biotechnology, China; Cat. No. C3622), a polymer-based precipitation method, strictly adhering to the manufacturer’s protocol. Briefly, cells were cultured in medium supplemented with exosome-depleted serum or in serum-free medium to prevent contamination with exogenous vesicles. Culture supernatants were harvested and subjected to stepwise centrifugation to remove intact cells and cellular debris. The clarified supernatants were subsequently passed through a 0.22-µm filter, incubated with the precipitation reagent overnight at 4°C, and centrifuged (10,000×g, 4 °C, 30 min) to recover EV pellets. The resulting pellets were resuspended in sterile PBS for downstream structural and functional applications.
PKH26 labeling for EV uptake assays
For fluorescence tracking, purified EVs were labeled using the BeyoExo™ Exosome Labeling and Tracking Kit (PKH26) (Beyotime Biotechnology, China; Cat. No. C3637). Briefly, EVs were incubated with PKH26 working solution for 1-5 min at room temperature in the dark. The labeling reaction was immediately terminated using the provided quenching solution. Crucially, to distinguish EV uptake from free dye artifacts, labeled EVs were processed according to the manufacturer’s instructions to remove unbound dye prior to incubation with recipient cells.
Proteomic and transcriptomic profiling
Proteomic analysis
Proteomic analysis was performed by data-independent acquisition (DIA) mass spectrometry with three biological replicates per group (n = 3). Proteins were extracted using SDT buffer, digested with trypsin by the FASP method, and peptides were desalted, spiked with iRT standards, and analyzed on a Vanquish Neo nano-UHPLC system coupled to an Orbitrap Astral mass spectrometer (Thermo Fisher Scientific). MS1 scans were acquired over m/z 380–980 at 240,000 resolution, and MS2 spectra were collected using 299 DIA windows (2 m/z) with HCD at 25 eV. Data were searched against the UniProt reference proteome using trypsin with up to one missed cleavage, carbamidomethylated cysteine as a fixed modification, oxidation (M) and protein N-terminal acetylation as variable modifications, and FDR < 1% at peptide and protein levels.
Transcriptomic analysis
Transcriptomic profiling was conducted via bulk RNA sequencing of the corresponding cell pellets. Following rigorous quality control, sequencing reads were aligned to the reference genome, and gene-level expression was quantified and normalized using standard bioinformatics pipelines. Differential gene expression analysis was performed to identify transcripts significantly altered between conditions.
Statistical analysis of omics data
For both proteomic and transcriptomic datasets, statistical testing for differential abundance or expression included correction for multiple comparisons using the Benjamini–Hochberg procedure. To facilitate biological interpretation, functional enrichment analyses-including Gene Ontology (GO), KEGG pathway analysis, and Gene Set Enrichment Analysis (GSEA)-were performed on the lists of differentially regulated proteins and genes identified under the corrected significance framework.
Statistical analysis
All statistical analyses were performed using R software (version 4.2.2) and GraphPad Prism (version 9.0). For comparisons between two groups, the Student’s t test was used when data followed a normal distribution with equal variance; otherwise, the Wilcoxon rank-sum test was applied. For comparisons among three or more groups, one-way analysis of variance (ANOVA) was used if assumptions of normality and homogeneity of variance were met; otherwise, the Kruskal–Wallis test was employed. The chi-square test was used to compare categorical variables among groups. All experiments were performed in at least three independent biological replicates. Continuous data are presented as mean ± standard deviation (SD) unless otherwise specified. P-values < 0.05 were considered statistically significant, with the following notation: *P < 0.05, **P < 0.01, ***P < 0.001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The publicly available data used in this study are available in the Gene Expression Omnibus (GEO) under accession code GSE225857, the National Omics Data Encyclopedia (NODE) under accession code OEP00001756, and The Cancer Genome Atlas (TCGA) Pan-Cancer dataset (https://www.cancer.gov/tcga). The single-cell RNA-sequencing data generated in this study have been deposited in the Genome Sequence Archive (GSA) under the accession code PRJCA046174. The proteomics and transcriptomics data generated in this study have been deposited in the iProX database under accession code IPX0014830000 and are also available at Zenodo under https://doi.org/10.5281/zenodo.18205510. The remaining data are available within the Article, Supplementary Information, or Source Data file, and/or are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
All data analyses and processing were performed using publicly available software and established packages, as detailed and cited in the Methods section. The code used in this study has been deposited in Zenodo under https://doi.org/10.5281/zenodo.18205510.
References
Siegel, R. L., Miller, K. D., Fuchs, H. E. & Jemal, A. Cancer statistics, 2022. Cancer J. Clinicians 72, 7–33 (2022).
Zheng, R. S. et al. Cancer incidence and mortality in China, 2022]. Zhonghua Zhong Liu Za Zhi 46, 221–231 (2024).
Raphael, M. J. & Karanicolas, P. J. Regional Therapy for colorectal cancer liver metastases: which modality and when? J. Clin. Oncol. 40, 2806–2817 (2022).
Datta, J., Narayan, R. R., Kemeny, N. E. & D’Angelic, M. I. Role of hepatic artery infusion chemotherapy in treatment of initially unresectable colorectal liver metastases: a review. JAMA Surg. 154, 768–776 (2019).
Patras, L., Shaashua, L., Matei, I. & Lyden, D. Immune determinants of the pre-metastatic niche. Cancer Cell 41, 546–572 (2023).
Liu, Y. & Cao, X. Characteristics and significance of the pre-metastatic niche. Cancer Cell 30, 668–681 (2016).
Wang, Y. et al. Pre-metastatic niche: formation, characteristics and therapeutic implication. Signal Transduct. Target Ther. 9, 236 (2024).
Zeng, Z. et al. Cancer-derived exosomal miR-25-3p promotes pre-metastatic niche formation by inducing vascular permeability and angiogenesis. Nat. Commun. 9, 5395 (2018).
Babak, M. V., Zalutsky, M. R. & Balyasnikova, I. V. Heterogeneity and vascular permeability of breast cancer brain metastases. Cancer Lett. 489, 174–181 (2020).
Coffelt, S. B., Wellenstein, M. D. & Visser, K. E. d. Neutrophils in cancer: neutral no more. Nat. Rev. Cancer 16, 431–446 (2016).
Park, J. et al. Cancer cells induce metastasis-supporting neutrophil extracellular DNA traps. Sci. Transl. Med. 8, 361ra138 (2016).
Mantovani, A., Cassatella, M. A., Costantini, C. & Jaillon, S. Neutrophils in the activation and regulation of innate and adaptive immunity. Nat. Rev. Immunol. 11, 519–531 (2011).
Hedrick, C. C. & Malanchi, I. Neutrophils in cancer: heterogeneous and multifaceted. Nat. Rev. Immunol. 22, 173–187 (2022).
Christoffersson, G. et al. VEGF-A recruits a proangiogenic MMP-9-delivering neutrophil subset that induces angiogenesis in transplanted hypoxic tissue. Blood 120, 4653–4662 (2012).
Yang, L. et al. DNA of neutrophil extracellular traps promotes cancer metastasis via CCDC25. Nature 583, 133–138 (2020).
Liu, J., Yan, W., Han, P. & Tian, D. The emerging role of KIAA1199 in cancer development and therapy. Biomed. Pharmacother. Biomed. Pharmacotherapie 138, 111507 (2021).
Chen, Y. et al. The role of CEMIP in tumors: an update based on cellular and molecular insights. Biomed. Pharmacother. 146, 112504 (2022).
Li, L. et al. Targeting pyroptosis reverses KIAA1199-mediated immunotherapy resistance in colorectal cancer. J Immunother. Cancer 13, e010000 (2025).
Zhao, L. et al. KIAA1199 promotes metastasis of colorectal cancer cells via microtubule destabilization regulated by a PP2A/stathmin pathway. Oncogene 38, 935–949 (2019).
Zhang, D. et al. Down-regulation of KIAA1199/CEMIP by miR-216a suppresses tumor invasion and metastasis in colorectal cancer. Int J. Cancer 140, 2298–2309 (2017).
Hua, Q. et al. CEMIP, a novel adaptor protein of OGT, promotes colorectal cancer metastasis through glutamine metabolic reprogramming via reciprocal regulation of β-catenin. Oncogene 40, 6443–6455 (2021).
Wang, H. et al. KIAA1199 drives immune suppression to promote colorectal cancer liver metastasis by modulating neutrophil infiltration. Hepatology 76, 967–981 (2022).
Zhang, B. et al. Tumor CEMIP drives immune evasion of colorectal cancer via MHC-I internalization and degradation. J. Immunother. Cancer 11, e005592 (2023).
Xu, G. et al. CEMIP, acting as a scaffold protein for bridging GRAF1 and MIB1, promotes colorectal cancer metastasis via activating CDC42/MAPK pathway. Cell Death Dis. 14, 167 (2023).
Che, L. H. et al. A single-cell atlas of liver metastases of colorectal cancer reveals reprogramming of the tumor microenvironment in response to preoperative chemotherapy. Cell Discov. 7, 80 (2021).
Wang, F. et al. Single-cell and spatial transcriptome analysis reveals the cellular heterogeneity of liver metastatic colorectal cancer. Sci. Adv. 9, eadf5464 (2023).
Wang, B. et al. The role of the transcription factor EGR1 in cancer. Front Oncol. 11, 642547 (2021).
Ji, R. C. Hypoxia and lymphangiogenesis in tumor microenvironment and metastasis. Cancer Lett. 346, 6–16 (2014).
Cheng, J. C., Chang, H. M. & Leung, P. C. Egr-1 mediates epidermal growth factor-induced downregulation of E-cadherin expression via Slug in human ovarian cancer cells. Oncogene 32, 1041–1049 (2013).
Grieshaber-Bouyer, R. et al. The neutrotime transcriptional signature defines a single continuum of neutrophils across biological compartments. Nat. Commun. 12, 2856 (2021).
Pettinella, F. et al. Surface CD52, CD84, and PTGER2 mark mature PMN-MDSCs from cancer patients and G-CSF-treated donors. Cell Rep. Med. 5, 101380 (2024).
Hori, S. FOXP3 as a master regulator of T(reg) cells. Nat. Rev. Immunol. 21, 618–619 (2021).
Bui, T. M. et al. Tissue-specific reprogramming leads to angiogenic neutrophil specialization and tumor vascularization in colorectal cancer. J. Clin. Invest. 134, e174545 (2024).
Wang, H. et al. Neutrophil extracellular traps in homeostasis and disease. Signal Transduct. Target Ther. 9, 235 (2024).
Wu, L. et al. An invasive zone in human liver cancer identified by Stereo-seq promotes hepatocyte-tumor cell crosstalk, local immunosuppression and tumor progression. Cell Res. 33, 585–603 (2023).
Ramirez, C. F. A. & Akkari, L. Myeloid cell path to malignancy: insights into liver cancer. Trends Cancer 11, 591–610 (2025).
Cordoba-Chacon, J. Loss of hepatocyte-specific PPARgamma expression ameliorates early events of steatohepatitis in mice fed the methionine and choline-deficient diet. PPAR Res. 2020, 9735083 (2020).
Francque, S. et al. Nonalcoholic steatohepatitis: the role of peroxisome proliferator-activated receptors. Nat. Rev. Gastroenterol. Hepatol. 18, 24–39 (2021).
Han, C. Y. et al. Adipocyte-derived serum amyloid A3 and hyaluronan play a role in monocyte recruitment and adhesion. Diabetes 56, 2260–2273 (2007).
Lee, J. W. et al. Hepatocytes direct the formation of a pro-metastatic niche in the liver. Nature 567, 249–252 (2019).
Harmon, G. S., Lam, M. T. & Glass, C. K. PPARs and lipid ligands in inflammation and metabolism. Chem. Rev. 111, 6321–6340 (2011).
Juurlink, D. N. Rosiglitazone and the case for safety over certainty. JAMA 304, 469–471 (2010).
Ma, Y. et al. Type 2 diabetes and risk of colorectal cancer in two large U.S. prospective cohorts. Br. J. Cancer 119, 1436–1442 (2018).
Peng, H. et al. Hyperglycemia induces an immunosuppressive microenvironment in colorectal cancer liver metastases by recruiting peripheral blood monocytes through the CCL3-CCR1 axis. J. Immunother. Cancer 13, e011310 (2025).
Rada, M., Krzywon, L., Petrillo, S., Lazaris, A. & Metrakos, P. A Retrospective study on the role of metformin in colorectal cancer liver metastases. Biomedicines 11, 731 (2023).
Acknowledgements
The authors thank the patients who participated in this study. This work is supported by the National Natural Science Foundation of China (82272711 to D.J.Z., 82272900 to L.Z., and 82573302, 82373050 to T.Z.).
Author information
Authors and Affiliations
Contributions
L.S.L., K.C., P.Y.Z., G.J.X., and J.G.Z. conducted experiments. L.S.L., K.C., and D.J.Z. wrote the manuscript. L.Z. and D.J.Z. designed research studies and analyzed data. Z.Y.L., D.D.Y., J.Z., P.F.Z., and J.H.R. acquired data. D.J.Z. and Z.T. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Samuel Huber and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, L., Zhao, L., Cao, K. et al. Hepatocytes functionally reprogrammed by KIAA1199-high colorectal cancer cells favour the accumulation of pro-metastatic Egr1+ neutrophils. Nat Commun 17, 2462 (2026). https://doi.org/10.1038/s41467-026-69250-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69250-1