Abstract
Biodiversity loss driven by climate change and human activities poses a critical global challenge. Population restoration and reintroduction programs are essential for mitigating this threat to endangered species, yet their outcomes often remain unpredictable due to poorly understood success factors, such as the inevitable inbreeding during bottleneck events. The conservation programme of the crested ibis (Nipponia nippon) marks a successful example where the population rose from just seven individuals to over 9000 in the past four decades. By developing an individual-based model that simulates the restoration process and incorporates species-specific demography and inbreeding data, we conclude that this successful restoration is largely deterministic, as our results closely mirror empirical recovery time and population-level inbreeding coefficients. To establish general guidelines for reintroduction programs, we compare how inbreeding depression influences the recovery success of two reintroduction strategies involving small founder populations. Our simulations reveal that the ‘firework’ approach (one-source translocations) outperforms the ‘sequential’ (serial translocations) approach in restoration effectiveness. Furthermore, by expanding analyses over a broad demographic space, we demonstrate that the net effect of inbreeding varies with species-specific demography and highlight the importance of considering their interaction when interpreting conservation outcomes and designing future reintroduction programs.
Similar content being viewed by others
Introduction
More than one in four species assessed by the IUCN is threatened with extinction, and biodiversity is expected to decline further as climate change and other anthropomorphic threats intensify1. To reverse this trend, effective conservation programmes are urgently needed2. Predicting the likelihood of population recovery from ecological threats, setting up more protected areas, and providing optimal species reintroduction strategies are therefore central to these efforts3,4,5,6,7,8,9,10,11. Despite the evident need, population restoration and reintroduction outcomes vary widely across species4,12,13,14,15,16, and the causes of failure are often not well understood15,17,18. In-depth analyses of empirical conservation outcomes can help identify drivers of success and failure, offering broader insights that can be applied to conservation efforts of other threatened species4,6,18,19.
Population bottlenecks can result from ecological catastrophes, yet are also common in reintroduction programmes, which inevitably begin with small populations due to financial constraints, conservation uncertainties, or to minimise negative impacts on the source population13,16,20. The primary goals of such programmes are to help endangered species maintain genetic diversity by establishing multiple self-sustaining populations or restoring species’ historical geographic ranges6,20,21. However, unavoidable inbreeding in small populations during early reintroduction phases often leads to reduced genetic diversity, which raises looming concerns about long-term conservation success4,17. Inbreeding is therefore a likely contributor to the variable outcomes of restoration efforts.
Inbreeding depression—the reduced fitness of a population due to mating between genetically close-related individuals—has been a central focus of conservation biology since its inception22, and is regarded as one of the most cryptic threats to population viability3,23,24. In wild animals, inbreeding is known to result in abnormal embryo development25,26, reduced fecundity23, survival rates3,27,28, and recruitment probability29,30, which ultimately threaten population viability through a reduced population growth rate19,31. These effects stem from the increased occurrence of partially recessive deleterious alleles in the homozygous state through identity-by-descent, as well as the accumulation of such alleles due to genetic drift, which invariably accompany inbreeding in small, endangered populations. Even in species that have recovered from bottlenecks, genome-wide sequencing studies have revealed that considerable inbreeding persists3,19,32,33. This raises concerns that the detrimental effects of inbreeding could undermine the viability of current steadily growing populations, or hinder the success of future reintroduction programmes19,23,32,34. A remarkable conservation success is the recovery of the endangered crested ibis (Nipponia nippon) in East Asia. This long-lived species faced a severe bottleneck in the last century due to habitat loss and pollution35. Fortunately, its population has been officially announced to have rebounded from just seven wild individuals to over 9000 (wild and captive) individuals in the past four decades, thanks to extensive conservation efforts in China36,37. As with many restoration programmes, however, it remains unclear how and why this species overcame the expected negative consequences of inbreeding, and what lessons can be learned to guide conservation efforts for other endangered species.
The crested ibis is thus an ideal empirical example for which a more extensive analysis of its conservation can be of great use—especially given that long-lived species are thought to be particularly vulnerable to extinction during bottleneck events19,23. For instance, their slow reproductive pace may hinder them from rapidly recovering from a small population size4,19. Despite this, the populations of some long-lived critically endangered species appear to have successfully recovered from past bottleneck events12,13,19,23,24,38. This has partially been attributed to the idea that repeated reproduction over a long lifespan can help buffer the harmful effects of inbreeding at the population level, thereby aiding in successful restoration23,24,39. In other words, lifetime fecundity can be substantial for species that have a slow birth rate yet continue to breed over decades. However, the influences of demography, such as fecundity and mortality rates, are often overlooked in predictions of how inbreeding impacts population viability3,6,19,28.
Therefore, in this study, we create an individual-based model incorporating both empirical inbreeding depression data and demographic rates from the crested ibis population restoration. Our first aim is to investigate to what extent such a theoretical model (Fig. 1A) can make predictions that align with empirical conservation outcomes and to understand whether the crested ibis restoration success was achieved by chance or is largely deterministic. We then conduct two additional mutually independent model analyses, as examples of broader lessons we can learn by applying this simulation framework to inform conservation practice. First, we use the same ibis restoration model to implement two idealised reintroduction approaches and compare their effectiveness (Fig. 1B): ‘firework’ and ‘sequential’. The ‘firework’ approach involves simultaneously establishing multiple independent populations using founders taken from a successfully restored source population (analogous to sparks radiating outward from a firework). In addition to the one-time release of founders to establish isolated populations, we also simulate a variant in which immigrants are regularly translocated from the source population after the initial reintroduction. The ‘sequential’ approach involves establishing new populations in a stepwise manner, where founders are taken from newly recovered populations, resembling a chain of translocation events extending over time. We choose to study these idealised reintroduction approaches because they are the basic building blocks that can be combined to produce more complex and diverse reintroduction programmes used in conservation practice. This comparison aims to provide general reintroduction recommendations for endangered species that require careful management for population expansion. Lastly, we investigate more generally how demography and inbreeding interact to impact population viability in restoration programmes. To do so, we compare the simulation results with and without inbreeding depression in a large demographic parameter space beyond that of the crested ibis.
In all modelling scenarios, population restoration begins with a small founder population. The breeding cycle consists of pair formation or maintenance, reproduction, and survival stages. The inbreeding coefficients (F) of a breeding pair’s offspring were calculated by tracking the pedigree (exemplified by the cartoon, see ‘Methods’ for details). Birds of different colours represent different sexes. In the main model, inbreeding was considered to only negatively impact hatching success based on the empirical situation in crested ibises. Fledging success and nest survival (marked by asterisks in the diagram) are included in the crested ibis population restoration model (corresponding results in Fig. 2) and the reintroduction model variants (corresponding results in Fig. 3), but not for the analysis that explores broader parameter ranges (corresponding results in Figs. 4 and 5). This diagram was self-designed, and the ibis silhouettes it contains are used under a Creative Commons Attribution 4.0 license (CC BY 4.0) from Creazilla (https://creazilla.com/media/silhouette/2248/ibis).
Results
Model overview
The simulation model begins with a bottlenecked population consisting of a few individuals. These individuals mate randomly to form breeding pairs that remain together for subsequent breeding seasons (Fig. 1). Each pair reproduces once per year. Their reproductive success is determined by the species’ clutch size, hatching success (influenced by the offspring’s inbreeding coefficients), and fledging success of the hatchlings. An individual’s inbreeding coefficient is calculated by tracking a pedigree matrix that contains all individuals that have ever appeared in the population (see ‘Methods’ and Supplementary Methods for details). After reproduction, individuals survive based on stage-specific probabilities (juvenile or adult survival rates). If both partners of a breeding pair survive, they will maintain the pair bond and breed again in the next season. If one partner dies or reaches the maximum reproductive age, the other individual randomly pairs with an unpaired adult of the opposite sex. This generates a population of overlapping generations. Simulations continue until the population reaches a threshold size or goes extinct.
Model predictions align well with the empirical crested ibis population restoration
To investigate the empirical restoration of the crested ibis, we incorporated species-specific biological information into the model, from empirical data (see the Supplementary Notes and the online repository40). The information includes kinship of the founder population, the negative effect of the inbreeding coefficient on hatching success (derived from empirical pedigree data analysis), clutch size, fledging success, nest survival, and stage-specific mortality rates (for details see Methods and Table1). Of 300 simulations, 291 replicates (97%) resulted in successful population recovery, reaching a threshold population size of 9000 (Fig 2A, B, Fig. S2), suggesting that the empirical restoration success of ibis restoration is largely deterministic. Furthermore, the time required for successful restoration among simulation replicates (median: 44 years, mean ± sd: 46.5 ± 10.0 years) is close to the empirical restoration time of 41 years (grey dashed lines in Fig. 2A, Fig. S2).
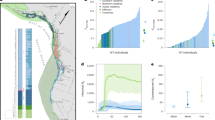
A The distribution of the time a population took to achieve successful restoration (in green, N = 291 replicates) or go extinct (in purple, N = 9 replicates) out of 300 simulation replicates. The grey dashed line represents the empirical time (41 years) the crested ibis population took to recover from seven individuals to over 9000. B Simulated population dynamics (plotted in the log scale of base 10) started from the empirical bottlenecked scenario with seven founders (i.e., two breeding pairs and three juveniles from one of the pairs). The log-transformed size of the founder population (seven individuals) is shown by the solid dot. The 100 trajectories presented here include all 9 replicates that ended in extinction and 91 randomly selected replicates that led to successful restoration (see Fig. S2 for trajectories of all 300 replicates). C The trajectory of population-level distribution of inbreeding coefficient among individuals in one randomly selected simulation replicate. The green solid line and the coloured ribbons represent the population average inbreeding coefficient and its standard deviation over simulation time.
Among simulations that resulted in successful restoration (Fig. 2B, green lines), most populations entered exponential growth (i.e., a straight line on the log scale) after short initial fluctuations. In contrast, populations in unsuccessful restorations (Fig. 2B, purple lines) remained small, never entered steady exponential growth, and eventually decreased to extinction—often rapidly (median: 8 years, mean ± sd : 15.8 ± 20.7 years). During restoration, the population’s average inbreeding coefficient first increases until reaching a plateau (Fig. 2C). The initial increase of the population-level inbreeding coefficient resulted from unavoidable inbreeding caused by the small size of the founder population. Once the population survives this initial phase, fewer inbred individuals can produce more offspring, contributing to faster population growth. Also, as population size increases, mating with close kin becomes less likely, leading to a decline in the variance of inbreeding coefficients among individuals. Among the successful replicates, the median population-level inbreeding coefficient in the final restoration year (0.19) was similar to the empirical estimate in the sampled population (0.17, 95% CI: 0.155–0.175, N = 321, grey dashed line in Fig. S3A).
The effect of inbreeding on hatching success was a central assumption of the crested ibis population restoration model. While this relationship was estimated from empirical data (see ‘Methods’), we also performed a sensitivity analysis to test model robustness. Varying the parameter value (s0) by ±50% did not substantially change the results (median population restoration time ranged from 40 to 50 years, and restoration success rate ranged from 95% to 99%, Fig. S4). More notable differences occurred when the value of s0 was doubled, leading to a decrease in restoration success (to below 90%) and longer restoration time (median: 58 years), although this does not affect the average inbreeding level in the successfully restored populations (Fig. S4). We also varied the relatedness between the adult founders, ranging from full siblings, half-siblings, to unrelated. All scenarios showed high restoration success rates (93.7–97.3%), though simulated restoration times suggest that the empirical founders were likely to have ranged from half-siblings to unrelated (Fig. S5). When inbreeding was assumed to not only impact hatching success but also reduce the survival of juveniles or adults, restoration success declined, but the population-level inbreeding coefficient when reaching restoration remained similar (Fig. S6).
In the main model, we implemented hard selection (i.e., mortality rates were independent of population density) for simplicity. However, soft selection is known to potentially hinder population recovery by negatively affecting individual fitness at population density41. To test whether this influences our model results, we additionally investigated how density regulation affects simulation outcomes. The results were robust to soft selection, as density dependence primarily slowed population growth during later stages of recovery, when the population size became large, but did not reduce the probability of successful restoration (Figs. S7 and S8, corresponding to two different ways of density regulation). Intuitively, this is because restoration success hinges on the early stage, which determines whether a population can enter stable exponential growth, whereas density effects only become important later.
The alignment between simulation results and the empirical data (i.e., restoration time and population-level inbreeding coefficients) and the robustness of the model suggest that our model provides a good fit for the real-world restoration dynamics of the crested ibis population.
Population reintroduction success depends crucially on the origin and number of founders
Having shown that our theoretical model fits well with the empirical restoration of the crested ibis, we conducted further model analyses to draw broader conclusions. Our first analysis using the theoretical model includes model variants corresponding to two idealised reintroduction strategies, which we term ‘firework’ and ‘sequential’ approaches. These strategies differ in the source of founders used to establish newly introduced populations. In the ‘firework’ approach, founders of the newly introduced populations were taken from the original source population, after it was successfully restored. The new populations were either established solely by these founders (Fig. 3A1) or also with additional immigrants translocated from the source population at regular intervals (Fig. 3A2). In the ‘sequential’ approach, the founders of the first round of reintroduction also came from the restored source population, but subsequent reintroduction rounds drew founders from the most recently restored population in a stepwise, chain-like fashion. For clarity and ease of comparison, only a few representative trajectories of population reintroduction are shown in Fig. 3; however, our simulations explored a wide range of scenarios, varying the number of unrelated founder pairs from three to 13 for each reintroduction approach, with 50 independent replicates for each scenario (Fig. S9).
One randomly selected simulation replicate from each different scenario is used to demonstrate the modelling outcomes. The solid lines represent the trajectory of the population-average inbreeding coefficient (F) of the populations; the coloured ribbons depict the standard deviation of F over the course of simulations. A Simulated trajectories where individuals are reintroduced simultaneously following either the standard ‘firework’ approach (A1, with no immigrants) or its variant (A2, with immigrants) in the scenario with three or 11 founder pairs. The simulation trajectory of the source population is coloured in grey, and the trajectories of reintroduced populations were in orange (for the standard ‘firework’) and pink (for the ‘firework’ with immigrants). The triangles represent the introduction of initial founders; the pink and purple dots in (A2) show time points when groups of immigrants (either three pairs or 11 pairs, consistent with the number of initial founders) are translocated from the source to the growing reintroduced population. B Simulated trajectories where individuals are reintroduced following the ‘sequential’ approach with three or 11 founder pairs. The dots indicate the mean F of the founders at the beginning of each reintroduction round. The simulation with three founder pairs collapsed at the beginning of the fourth round, while the one with eleven founder pairs achieved all ten rounds of restoration.
With the ‘firework’ reintroduction approach, increasing the number of founders resulted in lower inbreeding levels in the reintroduced population, regardless of whether immigrants were added (Fig. 3A1 and A2; compare the trajectories with three and 11 founder pairs). However, adding immigrants further slowed the accumulation of population-level inbreeding coefficients (Fig. 3A1 and A2; compare trajectories with the same number of founder pairs) and accelerated growth of the reintroduced population (Fig. S10). For example, a scenario with three initial founder pairs supplemented by the repeated addition of three immigrant pairs from the source population across five rounds (Fig. 3A2) yielded comparable population growth dynamics to a one-time reintroduction of 11 founder pairs (Fig. 3A1). These two scenarios showed similar final-year population average inbreeding coefficients (0.21 vs. 0.21, Fig. 3A1 and A2) and population sizes (285 vs. 280 individuals, Fig. S10).
In population reintroductions following the ‘sequential’ approach, there is a notable accumulation of the inbreeding coefficient over consecutive rounds (Fig. 3B). This approach resulted in more frequent population reintroduction failures, especially with a small number of founders (Fig. S9B). For instance, with only three founder pairs, there is a faster increase in the population-level inbreeding coefficient, resulting in sudden population collapse at the beginning of the fourth reintroduction round (Fig. 3B). In contrast, introducing 11 founder pairs allowed the population to complete all ten rounds of reintroduction, despite the continuous accumulation of inbreeding in each consecutively reintroduced population. Overall, the full programme of ten reintroduction rounds under the ‘sequential’ approach was only possible when each reintroduction round involved more than seven founder pairs (Fig. S9).
Our simulations showed that, for both reintroduction approaches, increasing the number of founders leads to faster population growth and mitigates the accumulation of inbreeding coefficients over reintroduction steps. As founder numbers increased, the restoration effectiveness gap between the two approaches narrowed. Nonetheless, the ‘firework’ approach generally outperformed the ‘sequential’ approach by yielding higher success probability, shorter restoration times of newly established populations, and requiring fewer founder pairs to achieve successful reintroduction.
The net effect of inbreeding depression on population restoration depends on demography
We further analysed the theoretical model to explore how inbreeding and demography interact to influence population viability under a broader demographic parameter space. Specifically, we varied clutch size, juvenile mortality rate, and adult mortality rate within ranges informed by demographic parameters of 41 bird species (Table S1). For each parameter combination, we ran simulations both with and without inbreeding depression, reducing hatching success. This comparison allowed us to disentangle the interactions between demographic parameters and inbreeding depression in shaping population viability after a bottleneck event.
From a full analysis across these parameter combinations, we can categorise species into three types (markers I–III in Fig. 4A–C) based on how demographic rates and inbreeding jointly affect population viability. Type I species (of which the crested ibis is an example) have favourable demographic rates (e.g., low juvenile and adult mortality rates). For these species, inbreeding depression has a negligible impact on the restoration success after a bottleneck (Fig. 4C), as they show a high recovery probability regardless (Fig. 4A, B). In contrast, type III species have unfavourable demographic rates that alone lead to low recovery probabilities (Fig. 4A, B), making population extinction during bottleneck events almost inevitable regardless of inbreeding depression (Fig. 4C). Type II species occupy an intermediate zone: they have demographic rates that result in a good chance of population restoration when inbreeding is assumed to be neutral (Fig. 4A), but restoration is unlikely when inbreeding reduces reproductive fitness (Fig. 4B, C). All three types are reflected by the distribution of the 41 species in the full simulation analysis (Fig. 5), and variation in clutch size introduces further complexity. Specifically, small clutch sizes considerably hinder recovery in species with high adult mortality, whereas in species with larger clutches, population viability is more constrained by high juvenile mortality. Although the model assumes a uniform effect of inbreeding depression across species, the net impact of inbreeding on population recovery from a bottleneck varies considerably depending on the demographic context. Given the large variation in avian demography (Table S1), our model results highlight the necessity to consider the interactions between a species’ demography and the impact of inbreeding depression when evaluating the likelihood of population recovery from bottleneck events.
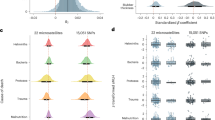
The outcomes were drawn from simulations that started with a bottlenecked founder population of three pairs and a scenario where the clutch size was set to three eggs. The probability value in each cell was calculated based on 100 independent simulation replicates. The heatmaps in A and B show the probability of population restoration under broad ranges of juvenile and adult mortality rates when the effect of inbreeding depression is respectively absent (s0 = 0) or present (s0 = 1). The colour scale is consistent in both panels. C The net effect of inbreeding depression on the probability of population restoration. Each cell in the grid represents the restoration probability difference between the corresponding cells in (A) and the present (B). The white area presents a parameter space where restoration is not possible regardless of the presence or absence of inbreeding depression. The labelled dots I–III in the panels represent hypothetical species whose restoration probabilities are influenced by the combined effect of inbreeding depression and demographic parameters in different ways (see main text for details).
Forty-one avian species (in black dots, see Table S1 for the full Latin name63) are mapped in the panels according to their demographic rates (i.e., juvenile and adult mortality rates and clutch size) to predict possible effects of inbreeding depression on restoration of a bottlenecked population under our model assumptions. The coloured gradient, ranging from light to dark orange, represents an increasing net effect of inbreeding depression on the probability of population restoration. Population restoration was not possible in the white areas.
Discussion
We developed an individual-based model inspired by the empirical restoration of the crested ibis, and performed two additional analyses to draw broader conclusions from this remarkable conservation success. Our crested ibis population restoration model provides a good fit to the empirical recovery of the species, as demonstrated through the agreement between simulation results and empirical data in terms of population restoration time and average inbreeding coefficient. On this basis, we further adjusted and analysed the model to compare the outcomes of two applied reintroduction strategies and found the ‘firework’ approach to be more effective than the ‘sequential’ approach. The former approach generally requires fewer founders and leads to a higher restoration probability, shorter restoration time, and lower inbreeding levels. We subsequently analysed the model to investigate beyond the demographic parameters of the crested ibis and demonstrate that a bottlenecked population’s chance of recovery depends on the interplay between its vulnerability to inbreeding depression and its species-specific demography.
Inbreeding can pose a significant extinction risk to bottlenecked populations, such as during the early stages of reintroduction schemes. Our simulations show that the ‘firework’ approach outperforms the ‘sequential’ approach by mitigating the accumulative effect of inbreeding that occurs through sequential reintroduction rounds. There is empirical evidence for the success of ‘firework’ schemes, as seen in the white-tailed deer (Odocoileus virginianus) and bearded vultures (Gypaetus barbatus), with reintroduced populations maintaining steady growth over time13,38. However, this approach is not guaranteed to always lead to success, as demonstrated by the conservation project of the Seychelles Magpie-robin (Copsychus sechellarum), which was successful in some translocation attempts but failed in others15. The varied mortality risks induced by local ecological threats are proposed to play a role in this species, reflecting the importance of demography in determining restoration success. When incorporating immigrant translocations, the ‘firework’ approach more effectively limits the accumulation of inbreeding. A supporting example is the white-tailed eagle (Haliaeetus albicilla) reintroduction in western Scotland, where a second release in 1990 followed early signs of population decline after the initial 1970s release and helped successfully establish a breeding population42,43. The effect of continuous large-scale translocations on sustaining the source population warrants caution when applying ‘firework’ reintroductions in practice.
The disadvantage of the ‘sequential’ approach revealed in our study aligned with another theoretical model44, which showed a dramatic increase in extinction rate after the second (compared to the first) bottleneck event. Empirically, sequential bottlenecked populations have been found to suffer from reduced genetic diversity, such as several second-order bottlenecked populations of the New Zealand saddleback (Philesturnus carunculatus rufusater) and the dice snakes (Natrix tessellata) illegally introduced into the lakes north of the Alps in Switzerland45,46,47. This limitation, however, can be mitigated by releasing a larger number of founders from previously successfully reintroduced populations. In practice, consistently translocating individuals from the source population is often more difficult than from reintroduced populations. Therefore, given constraints from policy regulations, limited time or financial resources16,48,49, the ‘sequential’ approach can sometimes have a practical advantage over the ‘firework’ approach.
Reintroduction programmes often combine the two approaches. For example, the reintroduction of the Alpine ibex (Capra ibex) started with the ‘firework’ approach to establish the first three wild populations in Switzerland12. After that, some secondary reintroductions continued to follow the ‘firework’ approach, while others adopted the ‘sequential’ approach with founders taken from a single reintroduced population. Genetic analysis, aligned with our model conclusion, showed a stronger loss of genetic diversity in the secondary reintroduced populations following the latter approach. Our work underscores that theoretical analysis can help clarify ambiguous outcomes in empirical reintroduction programmes and make predictions to optimise conservation strategies.
From a practical conservation perspective, our simulation outcomes suggest that inbreeding-induced restoration failure is most likely to occur during the early phase of population establishment. In successful cases, both the population restoration and reintroduction models exhibit similar dynamic patterns of the population-level inbreeding coefficient (F). The rapid initial increase in F indicates that severe inbreeding occurs due to the small population size. The average F later reaches a plateau once the population overcomes the first hurdle, accompanied by converging individual variation of F over the course of time. This trend indicates that the population has entered a positive eco-evolutionary vortex50. The reduction of individual variation in inbreeding levels in the population is mainly caused by diminishing chances of mating with close kin in an expanding population. This vortex also contributes to the exponential growth in the simulations, aligned with population dynamics in seven watershed subpopulations of crested ibises51. However, such rapid recovery has not occurred in the Floreana mockingbird Mimus trifasciatus and some other crested ibis subpopulations51,52, probably due to factors including habitat quality (related to density-dependent soft selection) or external threats, such as predation risks and disease outbreaks. Our simulation findings provide insights for conservation evaluation, especially for the recovery state of closed populations or captive restoration programmes.
Our model also helps to further understand the effects of inbreeding on population viability in general. It is often puzzling that many populations show steady growth despite being marked by known negative effects of inbreeding on reproductive fitness and decreased genetic diversity23,32,33,34. Taylor et al.23 emphasised that current population growth is not a reliable criterion for predicting population viability, as genetic load created by inbreeding in the spotted kiwi (Apteryx owenii) has resulted in reduced hatching success, which was predicted to eventually break down the population. Similarly, despite rapid population recovery, concerns arose after genetic analyses confirmed high levels of inbreeding in the endangered black-faced spoonbill (Platalea minor)32. Results from our model suggest that considerable inbreeding does not necessarily hinder populations from recovering from a bottleneck, especially for species with high survival and reproductive rates. However, even with little or no impact of inbreeding depression, populations with unfavourable demographic rates, such as low survival and/or reproduction, can still face high extinction risks, as for example seen in the continuous failed restoration of the great bustard (Otis tarda), given its generally low survival rates14. In the case of the crested ibis, along with remarkable conservation efforts, a combination of relatively low cost of inbreeding and repeated reproduction attempts enabled by the species’ longevity synergistically contributed to its rapid population recovery. Furthermore, the negative effects of high mortality rates and inbreeding depression on population recovery can sometimes be compensated for by large clutch sizes, showing the complex roles of different demographic traits in influencing population viability. In general, our results highlight that the impact of inbreeding on population viability is dependent on the species’ demography, and thus both aspects should be carefully considered when predicting population viability or attempting reintroduction schemes.
As demonstrated here, theoretical models offer a powerful and feasible approach for analyses of empirical population restorations and to be able to draw broader conclusions from them, information that itself can be used to guide future conservation efforts. Whilst our current model is tailored toward a crested ibis population, we encourage future models to use inbreeding depression and demographic data from other species to draw more specific population restoration predictions. For instance, in alignment with empirical evidence from the crested ibis, we here simulate that inbreeding negatively affects embryo development, but in other species, inbreeding depression may affect different aspects of fitness. Other factors can also be incorporated into such models, either to make more species-specific predictions or to gain a more general understanding of the range of factors that can influence population viability53. For example, inbreeding-induced genetic purging5,54 or Allee effects55,56 could both affect the restoration success of small populations. Allee effects have been found to vary across populations of the crested ibis depending on local geographic features and habitat quality56, highlighting that incorporating spatial structure and environmental heterogeneity into future models can also be informative. Despite the simplified assumptions, our simulations across a broad demographic space reveal a crucial but often overlooked role of demography in shaping population viability, which can potentially signal species vulnerability under current model premises. We encourage empirical studies to collect more quantitative data on species-specific inbreeding effects and demographic information, in combination with theoretical analyses such as the one presented here, that together can provide useful insights into the development of successful conservation and population restoration programmes.
Methods
Ethics declarations
All fieldwork was conducted under the guidance of the Ethics Committee of the Institute of Zoology, Chinese Academy of Sciences. The collection of pedigree information and captivity data was permitted by Dongzhai National Nature Reserve, and carried out in accordance with the guidelines of the Institutional Animal Care and Use Committee of the Institute of Zoology, Chinese Academy of Sciences.
Study system
The crested ibis is currently categorised as an endangered species in the IUCN Red List of threatened species57. It was declared locally extinct in Russia, Korea and Japan in the last century, and was rediscovered in 1981 in a mountain village of Yang County, Shaanxi Province, China. At the time, the wild population had only seven individuals, including two breeding pairs, one of which had three nestlings58. Since then, extensive conservation efforts—both in situ and ex situ—have been undertaken. As of 2022, the crested ibis has officially been reported by the State Council of China to have recovered to a population size exceeding 9000 individuals over the past 41 years36.
Quantification of the inbreeding effect
The empirical dataset used in this study was collected from the Crested Ibis Breeding Centre at Dongzhai National Nature Reserve (DNNR) in Henan Province, China. A pedigree was constructed based on recorded breeding partnerships and parental information for 321 captive-bred crested ibises from 2008 to 2023 using the R package ‘pedantics’59. Individual inbreeding coefficients were calculated from this pedigree using the same package, yielding a population mean of 0.17 (95% CI: 0.155-0.175). We used mixed linear models to quantify the effects of inbreeding on fitness during early-life stages—specifically fertilisation, hatching, and fledging (see Supplementary Notes for details of the statistics). The effects of inbreeding on juvenile and adult survival were not estimated due to the lack of such information for birds living in captivity. Instead, we used simulations to explore the potential effects of inbreeding on mortality rates at different life stages and evaluate how this affects population restoration dynamics (Fig. S6).
Field data collection for estimating demographic parameters
Breeding biology information was collected from the wild crested ibis population derived from the 122 individuals released at Dongzhai National Nature Reserve since 2014. We monitored breeding activity throughout the breeding season from February to June during 2021–2023. The crested ibis nests were searched over 549.66 km2 of mountainous terrain. Most nests were found in Masson pine (Pinus massoniana), with an average nest height of 12.58 ± 1.70 m (N = 33 nests). Nests were checked every other day via binoculars or drones (DJI Mini 4 Pro) to identify clutch size, the number of hatchlings (30 days after clutch completion) and the number of fledglings (42 days after hatching). Daily survival rate (DSR) was estimated using the RMark package in R60,61. A total of 40 nests were continuously monitored in the wild. Those with incomplete records were excluded from downstream analyses. Juveniles (22 individuals) and adults (40 individuals) were tracked with GPS devices (HQBG3621L, from Hunan Global Xinshi Technology Co., Ltd.).
Demographic parameters used in the model include clutch size, nest survival rate, hatching success, fledging success, and the mortality rates of juveniles and adults. The wild individuals originated from released offspring of the captive population living in the same nature reserve, and thus are likely to share similar fitness effects of inbreeding as the captive population. Details on how these demographic parameters were estimated are provided in the Supplementary Note.
Calculating inbreeding coefficients from a simulated pedigree
The inbreeding coefficient F measures the level of inbreeding, which is the probability that two alleles at a given locus in an individual are identical by descent from a common ancestor. This can be estimated by analysing the pedigree to track the kinship of all common ancestors of an individual’s parents. We calculated inbreeding coefficients by constructing a numerator relationship matrix (A) using the tabular method62. Specifically, all individuals in the pedigree were numbered from 1 to N, and the N × N matrix A contains the pairwise relationship coefficients between any two individuals. The diagonal element corresponding to a focal individual i is given by \({a}_{i,i}=1+0.5{a}_{{i}_{p},{i}_{q}}\), where ip and iq are the parents of the focal individual i, located in the p-th and q-th row of the matrix; and \({a}_{{i}_{p},{i}_{q}}\) is thus the relationship coefficient between them (Supplementary Methods). The off-diagonal entries in the lower triangle of A are computed recursively as \({a}_{i,j}=0.5({a}_{i,{j}_{p}}+{a}_{i,{j}_{q}})\), where jp and jq are the parents of individual j. Following this method, all elements of A can be generated through pedigree tracking. The inbreeding coefficient of individual i is then derived from its corresponding diagonal element in A by Fi = ai,i − 1. We programmed the pedigree matrix and the calculation of F in C++ (version 22) using a memory-efficient algorithm that significantly reduced the storage demands for the matrix. The algorithm pseudo-code is provided in the Supplementary Methods, and the C++ code is accessible in the Zenodo repository40.
Model simulation details for crested ibis restoration
We parameterised the individual-based model based on the breeding system and demographic data of the crested ibis. Simulations were initiated with a small population of seven individuals from two families—two breeding pairs, one of which had three nestlings—resembling the population state when the species was rediscovered in the wild in 198135. Because the kinship between the four adult founders was unknown, we explored different assumptions about their relatedness (i.e., full siblings, half siblings, and unrelated; Fig. S5). Pedigree analysis of one sampled population revealed that inbreeding primarily reduced brood hatching success, with no significant influence on fertilisation success or nestling survival (relevant analyses are provided in the Supplementary Note and the Zenodo repository40). We therefore model only the effect of inbreeding depression on hatching success here, but consider the effect of inbreeding depression on different life stages in the Supplementary Method and Fig. S6. We modelled the average number of hatchlings V per brood as a function of clutch size C and inbreeding-dependent hatching success, as follows:
where H0 is the baseline hatching success, s0 quantifies the negative effect of inbreeding depression, and F is the inbreeding coefficient. We performed a sensitivity analysis to explore how the parameterisation of s0 affects the simulated outcomes of population recovery (Fig. S4).
The number of fledglings was drawn from a Poisson distribution with a mean value of VR0, where R0 represents the fledging success of the hatchlings. We also incorporated the empirical nest survival rate Rn, which reflects the probability a nest survives the breeding season without complete reproductive failure due to damage or predation. As crested ibises remain in the juvenile stage without breeding for two years, we modelled mortality rates across three life stages based on empirical data: first-year juvenile (j1), second-year juvenile (j2), and adult (a). The values of all parameters are listed in Fig. 1. The survivorship of juveniles and adults in the population after a breeding season is determined by sampling a Bernoulli distribution with their corresponding life-stage-dependent mortality rates as the mean. Survivors enter a new breeding cycle, except for newborns and first-year juveniles. Simulations were terminated once the population reached 9000 individuals to compare the simulated population recovery with the empirical restoration outcomes. We here assumed no carrying capacity for population growth, but density-dependence as a type of soft selection could potentially hinder population recovery by negatively affecting individual fitness41. We therefore explored the effects of how density regulation can affect population recovery in the Supplementary Method, Fig. S7 and Fig. S8. All simulation code was written in C++ and is available in the Zenodo repository40. Simulation and empirical data were analysed and visualised in R 4.2.2.
Model analysis 1: comparing the ‘firework’ and ‘sequential’ reintroduction approaches
Based on the crested ibis model, we developed a reintroduction analysis that includes two model variants to compare the conservation effectiveness of two applied reintroduction strategies, the ‘firework’ approach and the ‘sequential’ approach. In each independent simulation of the ‘firework’ approach, we first generated a source population containing over 1000 individuals using the core crested ibis model. From this population, a fixed number of founder pairs were randomly sampled ten times to initialise ten independent reintroduction events. Each reintroduction event was terminated when the population either recovered to over 1000 individuals or went extinct. We tested different founding population sizes, ranging from three to 13 pairs, and simulated each scenario ten times.
To evaluate the impact of immigration on reintroduction outcomes, we ran additional simulations by simultaneously translocating groups of founders following two alternative processes: (1) translocating founders to establish a new population with no subsequent immigrants (same as the ‘firework’ simulations described above); (2) translocating founders to establish a new population and adding the same numbers of new immigrant pairs from the source population to the new population every three years for five rounds (i.e., in years 3, 6, 9, 12 and 15). We simulated scenarios with three and 11 pairs of reintroduced founders for these two alternatives, which led to four new populations being established simultaneously. The simulations were terminated once any of the four reintroduced populations reached 1000 individuals. Each combination was run for ten replicates.
In the ‘sequential’ approach, founders were always taken from the most recently recovered population that had reached 1000 individuals, creating a series of reintroductions. The number of founder pairs also ranged from three to 13. Each reintroduction proceeded sequentially: a new round began after the previous reintroduced population had recovered. Simulations were terminated either after 10 rounds of successful reintroduction or when a newly reintroduced population went extinct. We ran 50 independent simulation replicates for each number of founder pairs.
Model analysis 2: evaluating the more general impact of inbreeding on species-specific demography
To evaluate the net effects of inbreeding depression on the probability of population recovery in a bottleneck scenario, we implemented two model variants: hatching success declines with inbreeding (s0 = 1), and inbreeding has no effect on hatching success (s0 = 0). To generalise beyond the crested ibis, we excluded ibis-specific parameters such as nest survival and fledging success, and varied other demographic parameters across wider ranges: the mortality rate of juveniles (\({m}_{{p}_{j}}\)) and adults (\({m}_{{p}_{a}}\)), and clutch size (Cp) (Table 1). Specifically, we explored 11 values of \({m}_{{p}_{a}}\), 17 values of \({m}_{{p}_{j}}\), and six values of Cp, with full factorial combinations, and each unique combination was simulated with 100 independent replicates. Simulations began from a small, unrelated founding population consisting of three unrelated pairs. All individuals older than one year were allowed to breed. The linear correlation between inbreeding coefficients and hatching success was kept the same as in the crested ibis population restoration model, as relevant data for other species were unavailable. Nest failure and hatchling death were also excluded for simplicity and generality. To contextualise the model outcomes, we mapped 41 avian species with available stage-dependent mortality data from the COMADRE Database (2024) (Available from: https://www.compadre-db.org/[15 July 2024, Version 4.23.3.1]) to the simulation outcomes to provide implications of varied inbreeding effects on species with diverse demographic rates under our model assumptions. From all available avian data in COMADRE, we filtered species to ensure they were not too dissimilar in life-history to the crested ibis, such that we can explore parameter space beyond that of our focal species while ensuring that the model remains a biologically appropriate tool to do so. Specifically, we included only species with lifespans exceeding 10 years and average clutch sizes between one and six. As a larger clutch size contributes to improved restoration success (see Fig. 5), we rounded the average clutch size up to the next integer (rather than the closest integer) for all the species mapped to the simulation outcomes, to be more conservative in attributing the extinction risk to the effects of inbreeding and juvenile and adult mortality.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All simulation outcomes and empirical data used for producing the results in this study have been deposited in the Zenodo repository: https://doi.org/10.5281/zenodo.18049101.
Code availability
All simulation and analysis codes supporting the findings of this study are accessible on Zenodo: https://doi.org/10.5281/zenodo.18049101.
References
IUCN, The IUCN Red List of Threatened Species. Version 2024-2. 2024. Accessed on 01.10.2024. https://www.iucnredlist.org.
Conference of the Parties to the Convention on Biological Diversity (COP15), Kunming–Montreal Global Biodiversity Framework. Decision 15/4 adopted 19 December 2022, UN, symbol CBD/COP/DEC/15/4 (2022).
Kardos, M. et al. Inbreeding depression explains killer whale population dynamics. Nat. Ecol. Evol. 7, 675–686 (2023).
Jamieson, I. G. Founder effects, inbreeding, and loss of genetic diversity in four avian reintroduction programs. Conserv. Biol. 25, 115–123 (2011).
Hedrick, P. W. A. Garcia-Dorado, Understanding inbreeding depression, purging, and genetic rescue. Trends Ecol. Evol. 31, 940–952 (2016).
Bozzuto, C., Biebach, I., Muff, S., Ives, A. R. & Keller, L. F. Inbreeding reduces long-term growth of Alpine ibex populations. Nat. Ecol. Evol. 3, 1359–1364 (2019).
Griffith, B., Scott, J. M., Carpenter, J. W. & Reed, C. Translocation as a species conservation tool: status and strategy. Science 245, 477–480 (1989).
Sarrazin, F. & Barbault, R. Reintroduction: challenges and lessons for basic ecology. Trends Ecol. Evol. 11, 474–478 (1996).
Fischer, J. & Lindenmayer, D. B. Small patches can be valuable for biodiversity conservation: two case studies on birds in southeastern Australia. Biol. Conserv. 106, 129–136 (2002).
Ewen, J. G., Armstrong, D. P., Parker, K. A. & Seddon, P. J. Reintroduction Biology: Integrating Science and Management. (John Wiley & Sons, 2012).
Jachowski, D.S., Millspaugh, J.J., Angermeier, P.L. & Slotow, R. Reintroduction of Fish and Wildlife Populations. (University of California Press, 2016).
Grossen, C., Biebach, I., Angelone-Alasaad, S., Keller, L. F. & Croll, D. Population genomics analyses of European ibex species show lower diversity and higher inbreeding in reintroduced populations. Evol. Appl. 11, 123–139 (2018).
Schaub, M., Zink, R., Beissmann, H., Sarrazin, F. & Arlettaz, R. When to end releases in reintroduction programmes: demographic rates and population viability analysis of bearded vultures in the Alps. J. Appl. Ecol. 46, 92–100 (2009).
Ashbrook, K., Taylor, A., Jane, L., Carter, I. & Székely, T. Impacts of survival and reproductive success on the long-term population viability of reintroduced great bustards UK. Oryx 50, 583–592 (2016).
Burt, A. J. et al. The history, status and trends of the endangered Seychelles Magpie-robin Copsychus sechellarum. Bird. Conserv. Int. 26, 505–523 (2016).
Wakamiya, S. M. & Roy, C. L. Use of monitoring data and population viability analysis to inform reintroduction decisions: peregrine falcons in the Midwestern United States. Biol. Conserv. 142, 1767–1776 (2009).
Ewing, S. R., Nager, R. G., Nicoll, M. A. C., Aumjaud, A., Jones, C. G. & Keller, L. F. Inbreeding and loss of genetic variation in a reintroduced population of Mauritius kestrel. Conserv. Biol. 22, 395–404 (2008).
Armstrong, D. P. & Seddon, P. J. Directions in reintroduction biology. Trends Ecol. Evol. 23, 20–25 (2008).
Velando, A., Barros, Á & Moran, P. Heterozygosity–fitness correlations in a declining seabird population. Mol. Ecol. 24, 1007–1018 (2015).
Aisya, Z., White, D. J., Thavornkanlapachai, R., Friend, J. A., Rick, K. & Mitchell, N. J. Using PVA and captive breeding to balance trade-offs in the rescue of the island dibbler onto a new island ark. Sci. Rep. 12, 11913 (2022).
Sun, L., Zhou, T., Wan, Q.-H. & Fang, S.-G. Transcriptome comparison reveals key components of nuptial plumage coloration in crested ibis. Biomolecules 10, 905 (2020).
Soulé, M.E. & Wilcox, B.A. Conservation Biology. An Evolutionary-ecological Perspective. (University of California, San Diego, 1980).
Taylor, H. R., Colbourne, R. M., Robertson, H. A., Nelson, N. J., Allendorf, F. W. & Ramstad, K. M. Cryptic inbreeding depression in a growing population of a long-lived species. Mol. Ecol. 26, 799–813 (2017).
Robinson, J. A. et al. The critically endangered vaquita is not doomed to extinction by inbreeding depression. Science 376, 635–639 (2022).
Brekke, P., Bennett, P. M., Wang, J., Pettorelli, N. & Ewen, J. G. Sensitive males: inbreeding depression in an endangered bird. Proc. R. Soc. B 277, 3677–3684 (2010).
Ahlinder, J., Giles, B. E. & García-Gil, M. R. Life stage-specific inbreeding depression in long-lived Pinaceae species depends on population connectivity. Sci. Rep. 11, 8834 (2021).
Huisman, J., Kruuk, L. E. B., Ellis, P. A., Clutton-Brock, T. & Pemberton, J. M. Inbreeding depression across the lifespan in a wild mammal population. Proc. Natl. Acad. Sci. USA 113, 3585–3590 (2016).
Hoeck, P. E. A., Wolak, M. E., Switzer, R. A., Kuehler, C. M. & Lieberman, A. A. Effects of inbreeding and parental incubation on captive breeding success in Hawaiian crows. Biol. Conserv. 184, 357–364 (2015).
Bichet, C. et al. Contrasting heterozygosity-fitness correlations across life in a long-lived seabird. Mol. Ecol. 28, 671–685 (2019).
Szulkin, M., Garant, D., McCleery, R. H. & Sheldon, B. C. Inbreeding depression along a life-history continuum in the great tit. J. Evol. Biol. 20, 1531–1543 (2007).
Kardos, M., Keller, L.F. & Funk, W.C. What can genome sequence data reveal about population viability? Mol. Ecol. 34, e17608 (2024).
Li, S.-H., Liu, Y. et al. Not out of the woods yet: signatures of the prolonged negative genetic consequences of a population bottleneck in a rapidly re-expanding wader, the black-faced spoonbill Platalea minor. Mol. Ecol. 31, 529–545 (2022).
Harrisson, K. A. et al. Lifetime fitness costs of inbreeding and being inbred in a critically endangered bird. Curr. Biol. 29, 2711–2717 (2019).
Stock, S. E. et al. Uncovering inbreeding, small populations, and strong genetic isolation in an Australian threatened frog, Litoria littlejohni. Conserv. Genet. 24, 575–588 (2023).
Shi, D. The description of the decline of the crested ibis. J. Northwest Univ. 21, 25–30 (1991).
The State Council Information Office, People’s Republic of China, Global population of crested ibises exceeds 9000. http://english.scio.gov.cn/pressroom/2022-12/06/content_78553054.htm (2022).
Ma, L. et al. Changes in the habitat preference of crested ibis Nipponia nippon) during a Period of Rapid Population Increase. Animals 11, 2626 (2021).
Kekkonen, J., Wikström, M. & Brommer, J. E. Heterozygosity in an isolated population of a large mammal founded by four individuals is predicted by an individual-based genetic model. PLoS ONE 7, e43482 (2012).
Slate, J., Kruuk, L. E. B., Marshall, T. C., Pemberton, J. M. & Clutton-Brock, T. H. Inbreeding depression influences lifetime breeding success in a wild population of red deer (Cervus elaphus). Proc. R. Soc. B 267, 1657–1662 (2000).
Zheng, J. Dataset: Inbreeding and demography interact to impact population recovery from bottlenecks. Zenodo. https://doi.org/10.5281/zenodo.15813368 (2025).
Bell, D. A., Kovach, R. P., Robinson, Z. L., Whiteley, A. R. & Reed, T. The ecological causes and consequences of hard and soft selection. Ecol. Lett. 24, 1505–1521 (2021).
Green, R. E., Pienkowski, M. W. & Love, J. A. Long-term viability of the re-introduced population of the white-tailed eagle Haliaeetus albicilla in Scotland. J. Appl. Ecol. 33, 357–368 (1996).
Evans, R. J. et al. Growth and demography of a re-introduced population of White-tailed Eagles Haliaeetus albicilla. Ibis 151, 244–254 (2009).
Thevenon, S. & Couvet, D. The impact of inbreeding depression on population survival depending on demographic parameters. Anim. Conserv. 5, 53–60 (2002).
Lambert, D. M. et al. Serial population bottlenecks and genetic variation: translocated populations of the New Zealand saddleback (Philesturnus carunculatus rufusater). Conserv. Genet. 6, 1–14 (2005).
Parker, K. A., Anderson, M. J., Jenkins, P. F. & Brunton, D. H. The effects of translocation-induced isolation and fragmentation on the cultural evolution of bird song. Ecol. Lett. 15, 778–785 (2012).
Gautschi, B., Widmer, A., Joshi, J. & Koella, J. C. Increased frequency of scale anomalies and loss of genetic variation in serially bottlenecked populations of the dice snake, Natrix tessellata. Conserv. Genet. 3, 235–245 (2002).
Sutton, A. E. Leadership and management influences the outcome of wildlife reintroduction programs: findings from the Sea Eagle Recovery Project. PeerJ 3, e1012 (2015).
Sutherland, W. J. et al. Conserv. Lett. 3, 229–235 (2010).
Nabutanyi, P. & Wittmann, M. J. Models for eco-evolutionary extinction vortices under balancing selection. Am. Nat. 197, 336–350 (2021).
Li, X. et al. Understanding recovery is as important as understanding decline: The case of the crested ibis in China. Land 11, 1817 (2022).
Bozzuto, C., Hoeck, P. E. A., Bagheri, H. C. & Keller, L. F. Modelling different reintroduction strategies for the critically endangered Floreana mockingbird. Anim. Conserv. 20, 144–154 (2017).
Hernández, F., Janzen, T. & Lavretsky, P. simRestore: a decision-making tool for adaptive management of the native genetic status of wild populations. Mol. Ecol. Resour. 24, e13892 (2024).
Deredec, A. & Courchamp, F. Importance of the Allee effect for reintroductions. Ecoscience 14, 440–451 (2007).
Armstrong, D. P. & Wittmer, H. U. Incorporating Allee effects into reintroduction strategies. Ecol. Res. 26, 687–695 (2011).
Li, M. et al. Impact of Allee effects on the establishment of reintroduction populations of endangered species: The case of the Crested Ibis. Glob. Ecol. Conserv. 35, e02103 (2022).
BirdLife International, Nipponia nippon The IUCN Red List of Threatened Species, 2025-1, https://doi.org/10.2305/IUCN.UK.2018-2.RLTS.T22697548A132069229.en (2025).
Liu, Y. Z. Rediscovery of crested ibis Nipponia nippon in Qinling Mountain. Chin. J. Zool. 27, 237 (1981).
Morrissey, M. B. & Wilson, A. J. Pedantics: an R package for pedigree-based genetic simulation and pedigree manipulation, characterization and viewing. Mol. Ecol. Resour. 10, 711–719 (2010).
Laake, J. R. Mark: an R interface for analysis of capture-recapture data with MARK. AFSC Processed Rep., Alaska Fish. Sci. Cent., NOAA, Natl. Mar. Fish. Serv., Seattle, WA (2013).
R Core Team, R: A Language and Environment for Statistical Computing. (R Foundation for Statistical Computing, Vienna, Austria, 2024).
Emik, L. O. & Terrill, C. E. Systematic procedures for calculating inbreeding coefficients. J. Hered. 40, 51–55 (1949).
Gill, F., Donsker, D. & Rasmussen, P. IOC World Bird List (v15.1). https://www.worldbirdnames.org/new/bow/ (2025).
Acknowledgements
We acknowledge funding from the National Natural Science Foundation of China (32301290 to J.Z., 32570596 to D.W., 32125005 to X.Z.), the International Exchange and Talent Recruitment Programme of the Postdoctoral Management Committee (339478 to J.Z.), the Institute of Zoology, Chinese Academy of Sciences (2023IOZ0104 and 2024IOZ0107) and National Key Program of Research and Development, Ministry of Science and Technology of China (2023YFF1304800) to D.W., International Partnership Programme of Chinese Academy of Sciences for Grand Challenges (073GJHZ2023091GC), Joint Research Unit of Chinese Academy of Sciences (JRU CAS:152111ZYLH20250004) to X.Z., and the Swiss National Science Foundation (211549) to X-Y.L.R. We thank Dongzhai National Nature Reserve for providing us with the conservation data of the captive crested ibis and the research permit for this species. We thank the Centre for Information Technology of the University of Groningen for their support and for providing access to the Hábrók high-performance computing cluster. We also thank Stefano Canessa for insightful discussions.
Author information
Authors and Affiliations
Contributions
J.Z. designed and implemented the model, produced the main results, and wrote the first draft of the manuscript; E.R-B. collected and analysed species demographic data; T. J. participated in model design and implementation; Z.Z., X.Z., D.W. initiated project conceptualisation, provided empirical data; X-Y.L.R. supervised model development and data visualisation. All authors contributed to the paper writing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks George Olah, Lukas Keller and Marty Kardos for their contribution to the peer review of this work. A peer review file is available.”
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zheng, J., Rees-Baylis, E., Janzen, T. et al. Inbreeding and demography interact to impact the recovery of a bottlenecked crested ibis population. Nat Commun 17, 2785 (2026). https://doi.org/10.1038/s41467-026-69278-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69278-3