Abstract
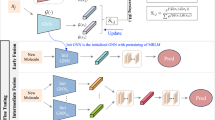
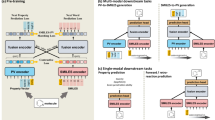
The pre-training molecular representation learning (MRL) has shown considerable potential in computer-aided drug discovery. Recently, many multimodal pre-training MRL methods have been presented, incorporating multimodal molecular data for pre-training and achieving high-accuracy predictions in downstream tasks. However, most current methods require completeness of modality for molecular data in the pre-training phase and often overlook their adaptation to real-world scenarios where, for example, molecular modalities except 2D topological graphs (2D modality) are often unavailable. In this study, we propose a multimodal pre-training MRL framework called M2UMol, which separately matches 2D modality to multiple modalities and undergoes pre-training jointly with a modality classifier. In this way, M2UMol elegantly transfers multimodal knowledge into the 2D modal encoder and allows for inputting incomplete modalities in the pre-training stage. Moreover, in downstream tasks with only the 2D modality given, M2UMol enables the precise simulation of molecular multimodal information based on the pre-trained 2D modal encoder. Comprehensive experimental results show the superior performance of M2UMol in a wide range of molecular tasks with higher efficiency in pre-training than pioneer models and demonstrate the validity of the multimodal knowledge transfer. Furthermore, we developed a user-friendly package based on M2UMol, integrating molecular representation learning, key functional group analysis, molecular multimodal retrieval, etc. It may be conveniently used in diverse fields related to drug discovery and promises to facilitate the process of developing drugs. Our code, pre-trained weights of M2UMol, and the package are available at https://github.com/Zhankun-Xiong/M2UMol.
Similar content being viewed by others
Data availability
The raw data of the pre-training dataset were sourced from the public dataset DrugBank64, available at https://go.drugbank.com/releases/latest. The processed molecular property prediction datasets are available at: http://snap.stanford.edu/gnn-pretrain/data/chem_dataset.zip. The BindingDB67 dataset is available at: https://www.bindingdb.org/bind/index.jspand the BioSNAP68 source is available at: https://github.com/kexinhuang12345/MolTrans/tree/master/dataset/BIOSNAP/full_data. The drug-drug interaction dataset is available at: https://github.com/Zhankun-Xiong/MRCGNN/tree/main/Ryu’s%20dataset. The data generated in this study have been publicly deposited to Hugging Face under https://doi.org/10.57967/hf/7153, and the data version used for this publication is available86. Source data are provided with this paper87. Source data are provided with this paper.
Code availability
The codes, pre-trained model, and the developed package are freely available at https://doi.org/10.5281/zenodo.17798744. The version used for this publication is available88.
References
Walters, W. P. & Barzilay, R. Applications of deep learning in molecule generation and molecular property prediction. Acc. Chem. Res. 54, 263–270 (2020).
Shen, J. & Nicolaou, C. A. Molecular property prediction: Recent trends in the era of artificial intelligence. Drug Discov. Today. Technol. 32, 29–36 (2019).
Ryu, J. Y., Kim, H. U. & Lee, S. Y. Deep learning improves prediction of drug-drug and drug-food interactions. Proc. Natl. Acad. Sci. USA 115, E4304–E4311 (2018).
Lin, X., Quan, Z., Wang, Z.-J., Ma, T. & Zeng, X. Kgnn: Knowledge graph neural network for drug-drug interaction prediction. In IJCAI, 380, 2739–2745 (2020).
Qiu, Y., Zhang, Y., Deng, Y., Liu, S. & Zhang, W. A comprehensive review of computational methods for drug-drug interaction detection. IEEE/ACM Trans. Comput. Biol. Bioinforma. 19, 1968–1985 (2021).
Bagherian, M. et al. Machine learning approaches and databases for prediction of drug–target interaction: a survey paper. Brief. Bioinforma. 22, 247–269 (2021).
Chen, X. et al. Drug–target interaction prediction: databases, web servers and computational models. Brief. Bioinforma. 17, 696–712 (2016).
Todeschini, R. & Consonni, V. Handbook of molecular descriptors (John Wiley & Sons, 2008).
Skoraczyński, G. et al. Predicting the outcomes of organic reactions via machine learning: are current descriptors sufficient? Sci. Rep. 7, 3582 (2017).
Lu, C. et al. Molecular property prediction: a multilevel quantum interactions modeling perspective. In Proceedings of the Thirty-Third AAAI Conference on Artificial Intelligence and Thirty-First Innovative Applications of Artificial Intelligence Conference and Ninth AAAI Symposium on Educational Advances in Artificial Intelligence, 1052–1060 (2019).
Wang, Z. et al. Advanced graph and sequence neural networks for molecular property prediction and drug discovery. Bioinformatics 38, 2579–2586 (2022).
Bjerrum, E. J. Smiles enumeration as data augmentation for neural network modeling of molecules. arXiv preprint arXiv:1703.07076 (2017).
Quan, Z. et al. A system for learning atoms based on long short-term memory recurrent neural networks. In 2018 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 728–733 (IEEE, 2018).
Weininger, D. Smiles, a chemical language and information system. 1. introduction to methodology and encoding rules. J. Chem. Inf. Comput. Sci. 28, 31–36 (1988).
Gilmer, J., Schoenholz, S. S., Riley, P. F., Vinyals, O. & Dahl, G. E. Neural message passing for quantum chemistry. In International conference on machine learning, 1263–1272 (PMLR, 2017).
Xiong, Z. et al. Pushing the boundaries of molecular representation for drug discovery with the graph attention mechanism. J. Med. Chem. 63, 8749–8760 (2019).
Schütt, K. et al. Schnet: A continuous-filter convolutional neural network for modeling quantum interactions. Advances in neural information processing systems, 30, (2017).
Gasteiger, J., Groß, J. & Günnemann, S. Directional message passing for molecular graphs. In International Conference on Learning Representations (2020).
Hu, Z., Dong, Y., Wang, K., Chang, K.-W. & Sun, Y. Gpt-gnn: Generative pre-training of graph neural networks. In Proceedings of the 26th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, 1857–1867 (2020).
Hu, W. et al. Strategies for pre-training graph neural networks. In International Conference on Learning Representations (2020).
Zaidi, S. et al. Pre-training via denoising for molecular property prediction. In International Conference on Learning Representations (2023).
Chithrananda, S., Grand, G. & Ramsundar, B. Chemberta: large-scale self-supervised pretraining for molecular property prediction. arXiv preprint arXiv:2010.09885 (2020).
Ross, J. et al. Large-scale chemical language representations capture molecular structure and properties. Nat. Mach. Intell. 4, 1256–1264 (2022).
Wang, Y., Wang, J., Cao, Z. & Barati Farimani, A. Molecular contrastive learning of representations via graph neural networks. Nat. Mach. Intell. 4, 279–287 (2022).
You, Y. et al. Graph contrastive learning with augmentations. Adv. neural Inf. Process. Syst. 33, 5812–5823 (2020).
You, Y., Chen, T., Shen, Y. & Wang, Z. Graph contrastive learning automated. In International Conference on Machine Learning, 12121–12132 (PMLR, 2021).
Stärk, H. et al. 3d infomax improves gnns for molecular property prediction. In International Conference on Machine Learning, 20479–20502 (PMLR, 2022).
Liu, S. et al. Multi-modal molecule structure–text model for text-based retrieval and editing. Nat. Mach. Intell. 5, 1447–1457 (2023).
Su, B. et al. A molecular multimodal foundation model associating molecule graphs with natural language. arXiv preprint arXiv:2209.05481 (2022).
Liu, S. et al. Pre-training molecular graph representation with 3d geometry. In International Conference on Learning Representations (2021).
Zeng, Z., Yao, Y., Liu, Z. & Sun, M. A deep-learning system bridging molecule structure and biomedical text with comprehension comparable to human professionals. Nat. Commun. 13, 862 (2022).
Liu, S., Du, W., Ma, Z.-M., Guo, H. & Tang, J. A group symmetric stochastic differential equation model for molecule multi-modal pretraining. In International Conference on Machine Learning, 21497–21526 (PMLR, 2023).
Luo, Y., Yang, K., Hong, M., Liu, X. & Nie, Z. Molfm: A multimodal molecular foundation model. arXiv preprint arXiv:2307.09484 (2023).
Zhu, Y. et al. Featurizations matter: a multiview contrastive learning approach to molecular pretraining. In ICML 2022 2nd AI for Science Workshop (2022).
Bai, P., Miljković, F., John, B. & Lu, H. Interpretable bilinear attention network with domain adaptation improves drug–target prediction. Nat. Mach. Intell. 5, 126–136 (2023).
Van der Maaten, L. & Hinton, G. Visualizing data using t-sne. J. Mach. Learning Res.9, 2579–2605 (2008).
Long, Q., Wang, M. & Li, L. Generative imagination elevates machine translation. In Proceedings of the 2021 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, 5738–5748 (2021).
Xia, J. et al. Mole-bert: Rethinking pre-training graph neural networks for molecules. In The Eleventh International Conference on Learning Representations (2022).
Zang, X., Zhao, X. & Tang, B. Hierarchical molecular graph self-supervised learning for property prediction. Commun. Chem. 6, 34 (2023).
McCune, D. F., Gaivin, R. J., Rorabaugh, B. R. & Perez, D. M. Bulk is a determinant of oxymetazoline affinity for the α1a-adrenergic receptor. Receptors Channels 10, 109–116 (2004).
Wang, T. & Isola, P. Understanding contrastive representation learning through alignment and uniformity on the hypersphere. In International Conference on Machine Learning, 9929–9939 (PMLR, 2020).
Fang, Y. et al. Knowledge graph-enhanced molecular contrastive learning with functional prompt. Nat. Mach. Intel. 5, 542–553 (2023).
BOTEV, Z., GROTOWSKI, J. & KROESE, D. Kernel density estimation via diffusion. Ann. Stat. 38, 2916–2957 (2010).
Chen, M. et al. Dilirank: the largest reference drug list ranked by the risk for developing drug-induced liver injury in humans. Drug Discov. Today 21, 648–653 (2016).
Goldberg, B. & Stern, A. The mechanism of oxidative hemolysis produced by phenylhydrazine. Mol. Pharmacol. 13, 832–839 (1977).
García Marín, I. D. et al. New compounds from heterocyclic amines scaffold with multitarget inhibitory activity on aβ aggregation, ache, and bace1 in the alzheimer disease. Plos one 17, e0269129 (2022).
Di, L. & Kerns, E. H. Drug-like properties: concepts, structure design and methods from ADME to toxicity optimization (Academic Press, 2015).
Black, W. et al. From indomethacin to a selective cox-2 inhibitor: development of indolalkanoic acids as potent and selective cyclooxygenase-2 inhibitors. Bioorg. Med. Chem. Lett. 6, 725–730 (1996).
Bavry, A. A. et al. Harmful effects of nsaids among patients with hypertension and coronary artery disease. Am. J. Med. 124, 614–620 (2011).
Banik, M., Gopi, S. P., Ganguly, S. & Desiraju, G. R. Cocrystal and salt forms of furosemide: solubility and diffusion variations. Cryst. Growth Des. 16, 5418–5428 (2016).
Goud, N. R. et al. Novel furosemide cocrystals and selection of high solubility drug forms. J. Pharm. Sci. 101, 664–680 (2012).
Harriss, B. I., Vella-Zarb, L., Wilson, C. & Evans, I. R. Furosemide cocrystals: Structures, hydrogen bonding, and implications for properties. Cryst. growth Des. 14, 783–791 (2014).
Karaytuğ, M. O. et al. Piperazine derivatives with potent drug moiety as efficient acetylcholinesterase, butyrylcholinesterase, and glutathione s-transferase inhibitors. J. Biochem. Mol. Toxicol. 37, e23259 (2023).
Deshler, L. & Zuman, P. Polarographic reduction of aldehydes and ketones: Part xviii. ethacrynic acid. Analytica Chim. Acta 73, 337–354 (1974).
Litwin, A., Adams, L. E., Zimmer, H. & Hess, E. V. Immunologic effects of hydralazine in hypertensive patients. Arthritis Rheumatism J. Am. Coll. Rheumatol. 24, 1074–1077 (1981).
Main, B. G. & Tucker, H. 3 recent advances in β-adrenergic blocking agents. Prog. Med. Chem. 22, 121–164 (1985).
Shrivastav, P. S., Buha, S. M. & Sanyal, M. Detection and quantitation of β-blockers in plasma and urine. Bioanalysis 2, 263–276 (2010).
Rampášek, L. et al. Recipe for a general, powerful, scalable graph transformer. Adv. Neural Inf. Process. Syst. 35, 14501–14515 (2022).
Wang, L., Liu, Y., Lin, Y., Liu, H. & Ji, S. Comenet: Towards complete and efficient message passing for 3d molecular graphs. In Koyejo, S. et al. (eds.) Advances in Neural Information Processing Systems, vol. 35, 650–664 (Curran Associates, Inc., 2022).
Gu, Y. et al. Domain-specific language model pretraining for biomedical natural language processing. ACM Trans. Comput. Healthc. (HEALTH) 3, 1–23 (2021).
Chen, T., Kornblith, S., Norouzi, M. & Hinton, G. A simple framework for contrastive learning of visual representations. In International conference on machine learning, 1597–1607 (PMLR, 2020).
Ganin, Y. & Lempitsky, V. Unsupervised domain adaptation by backpropagation. In International conference on machine learning, 1180–1189 (PMLR, 2015).
Wang, H. et al. Multi-modal learning with missing modality via shared-specific feature modelling. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 15878–15887 (2023).
Wishart, D. S. et al. Drugbank 5.0: a major update to the drugbank database for 2018. Nucleic acids Res. 46, D1074–D1082 (2018).
Wu, Z. et al. Moleculenet: A benchmark for molecular machine learning. Chem. Sci. 9, 513–530 (2018).
Xiong, Z. et al. Multi-relational contrastive learning graph neural network for drug-drug interaction event prediction. In Proceedings of the Thirty-Seventh AAAI Conference on Artificial Intelligence. 5339–5347 (2023).
Gilson, M. K. et al. Bindingdb in 2015: A public database for medicinal chemistry, computational chemistry and systems pharmacology. Nucleic acids Res. 44, D1045–D1053 (2016).
Zitnik, M., Sosic, R. & Leskovec, J. Biosnap datasets: Stanford biomedical network dataset collection. http://snap.stanford.edu/biodata Cited by 5 (2018).
Yu, Q. et al. Multimodal molecular pretraining via modality blending. In The Twelfth International Conference on Learning Representations.
Hamilton, W., Ying, Z. & Leskovec, J. Inductive representation learning on large graphs. Advances in neural information processing systems 30, (2017).
Sun, F.-Y., Hoffmann, J., Verma, V. & Tang, J. Infograph: Unsupervised and semi-supervised graph-level representation learning via mutual information maximization. In International Conference on Learning Representations (2020).
Xu, M., Wang, H., Ni, B., Guo, H. & Tang, J. Self-supervised graph-level representation learning with local and global structure. In International Conference on Machine Learning, 11548–11558 (PMLR, 2021).
Rong, Y. et al. Self-supervised graph transformer on large-scale molecular data. Adv. Neural Inf. Process. Syst. 33, 12559–12571 (2020).
Yu, H., Zhao, S. & Shi, J. STNN-DDI: A substructure-aware tensor neural network to predict drug-drug interactions. Brief. Bioinforma. 23, bbac209 (2022).
Nyamabo, A. K., Yu, H. & Shi, J.-Y. SSI-DDI: Substructure-substructure interactions for drug-drug interaction prediction. Brief. Bioinf. https://doi.org/10.1093/bib/bbab133 (2021).
Nyamabo, A. K., Yu, H., Liu, Z. & Shi, J.-Y. Drug-drug interaction prediction with learnable size-adaptive molecular substructures. Brief. Bioinf. https://doi.org/10.1093/bib/bbab441 (2021).
Yang, Z., Zhong, W., Lv, Q. & Chen, C. Y.-C. Learning size-adaptive molecular substructures for explainable drug-drug interaction prediction by substructure-aware graph neural network. Chem. Sci. 13, 8693–8703 (2022).
Zhu, X., Shen, Y. & Lu, W. Molecular Substructure-Aware Network for Drug-Drug Interaction Prediction. In Proceedings of the 31st ACM International Conference on Information & Knowledge Management, 4757–4761 (2022).
Li, Z. et al. DSN-DDI: an accurate and generalized framework for drug-drug interaction prediction by dual-view representation learning. Briefings Bioinformatics 24. https://doi.org/10.1093/bib/bbac597 (2023).
He, H., Chen, G. & Yu-Chian Chen, C. 3DGT-DDI: 3D graph and text based neural network for drug-drug interaction prediction. Briefings in Bioinformatics 23, https://doi.org/10.1093/bib/bbac134, https://academic.oup.com/bib/article-pdf/23/3/bbac134/43745041/bbac134.pdf (2022).
Cortes, C. & Vapnik, V. Support-vector networks. Mach. Learn. 20, 273–297 (1995).
Ho, T. K. Random decision forests. In Proceedings of 3rd international conference on document analysis and recognition, vol. 1, 278–282 (IEEE, 1995).
Lee, I., Keum, J. & Nam, H. Deepconv-dti: Prediction of drug-target interactions via deep learning with convolution on protein sequences. PLoS Comput. Biol. 15, e1007129 (2019).
Nguyen, T. et al. Graphdta: Predicting drug–target binding affinity with graph neural networks. Bioinformatics 37, 1140–1147 (2021).
Huang, K., Xiao, C., Glass, L. M. & Sun, J. Moltrans: molecular interaction transformer for drug–target interaction prediction. Bioinformatics 37, 830–836 (2021).
Xiong, Z. Data of multi-to-uni modal knowledge transfer pre-training for molecular representation learning: V1.0, https://doi.org/10.57967/hf/7153 (2025).
Xiong, Z. Source data files for paper “multi-to-uni modal knowledge transfer pre-training for molecular representation learning. https://doi.org/10.5281/zenodo.18219572 (2026).
Xiong, Z. Zhankun-xiong/m2umol: M2umol, https://doi.org/10.5281/zenodo.17798744 (2025).
Irwin, R., Dimitriadis, S., He, J. & Bjerrum, E. J. Chemformer: A pre-trained transformer for computational chemistry. Mach. Learn. Sci. Technol. 3, 015022 (2022).
Yang, M., Chen, T., Liu, Y.-X. & Huang, L. Visualizing set relationships: Evenn’s comprehensive approach to venn diagrams. Imeta 3, e184 (2024).
Acknowledgements
W.Z. is supported by the National Natural Science Foundation of China (62372204, 62072206), National Administration of Traditional Chinese Medicine Science and Technology Project (No. GZY-KJS-2025-003), Huazhong Agricultural University Scientific & Technological Self-innovation Foundation and Fundamental Research Funds for the Central Universities (2662024SZ006). S.L. is supported by the National Natural Science Foundation of China (62472191). P.Z. is not funded by any of the funders.
Author information
Authors and Affiliations
Contributions
Z.X., Z.W., and F.H. contributed equally. Z.X. conceived the research project. Z.X. developed the primary method and code. Z.W. analyzed the baselines in the paper. M.Q., S.F., and L.Y. assisted in analyzing the effectiveness and interpretability of the method. Z.X., Z.W., F.H., P.Z., and W.Z. wrote the paper. All authors, including X.Z. and S.L., read and commented on the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Prayag Tiwari, and the other anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xiong, Z., Wang, Z., Huang, F. et al. Multi-to-uni modal knowledge transfer pre-training for molecular representation learning. Nat Commun (2026). https://doi.org/10.1038/s41467-026-69302-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-69302-6