Abstract
As natural hosts of avian influenza viruses, wild birds pose an increasing threat to public health. Here, using surveillance data from wild bird infections across the United States (2022-2025), we show that HPAI-H5 transmission exhibits strong interspecific variation, seasonality, and spatial heterogeneity linked to migratory flyways. Phylogeographic analysis reveals that viral genotypes evolve from early, limited transmission along single migratory routes to a nationwide dispersal pattern spanning multiple migratory flyways. Anseriformes exhibits the highest number of infections but the lowest transmission risk, whereas Strigiformes demonstrates the greatest transmission risk. The HPAI-H5 transmission in wild birds along migratory flyways exhibits significant spatial heterogeneity and is associated with bird migration. Meteorological conditions are correlated with outbreak timing and may inform early warning efforts; however, these relationships are nonlinear. These findings provide a foundation for risk assessment, early warning systems, and integrated management of avian influenza in wild bird populations.
Similar content being viewed by others
Introduction
Avian influenza is a disease caused by avian influenza viruses that primarily spreads among birds but can sometimes undergo cross-species transmission to mammals, including humans, making it a zoonotic disease of wide concern1. Highly pathogenic avian influenza (HPAI), particularly the highly pathogenic H5N1 clade 2.3.4.4b, has caused unprecedented outbreaks in wild birds and poultry globally since 2020 and continues to pose potential pandemic risks to public health2. Wild birds, especially long-distance migratory birds, have long been natural hosts of avian influenza and are considered key hosts and vectors for the virus to achieve intercontinental transmission and maintain an endemic circulation. Their complex migration routes, seasonal aggregations, and ecological interactions with multiple species provide ideal conditions for the continuous evolution, reassortment, and geographic spread of the virus3. The United States is located at the intersection of multiple major global migratory bird flyways (Pacific, Central, Mississippi, and Atlantic Flyways) and has diverse ecosystems and high avian biodiversity, making it a critical region for studying the transmission risks of avian influenza in wild birds4. In March 2024, Texas reported the world’s first case of cattle infection with highly pathogenic H5 avian influenza, which rapidly spread to multiple states across the United States, causing intercontinental transmission. Genetic evolutionary analysis suggests that the infecting virus likely originated from wild birds, was transmitted from wild birds to cattle, and then spread between herds through cattle movement5. In November 2025, Washington State confirmed a case of H5N5 infection resulting in death, the world’s first fatal human case caused by this H5 subtype avian influenza virus. The state health department detected the H5N5 avian influenza virus in the patient’s backyard and concluded that poultry or wild birds were likely the source of the infection6. These findings underscore the urgent need for in-depth research on the transmission mechanisms of avian influenza among wild birds.
Although numerous studies have confirmed the predominant role of migratory birds in the intercontinental transmission of HPAI viruses through metagenomic analysis7,8,9, there is a lack of systematic understanding of the spatiotemporal dynamic evolution and heterogeneous driving factors of the current global epidemic caused by H5N1 clade 2.3.4.4b. First, the role of host heterogeneity is far more complex than previously understood. The traditional transmission concept centered on Anseriformes is being challenged by the reality of highly efficient transmission and high pathogenicity of the virus in nontraditional hosts such as raptors and mammals10. It is essential to quantify the transmission risk of avian influenza in different categories of birds and analyze their roles in the transmission chain. Second, the underlying patterns of spatial heterogeneity are unclear. While existing studies have confirmed viral transmission along North America’s four migratory flyways, the evolutionary patterns of key genotypes within the currently circulating clades are insufficiently explored11. Furthermore, whether each flyway develops distinct spatiotemporal hotspot dynamics due to its unique ecological and geographical characteristics constitutes a critical unanswered question for achieving regionalized precision in prevention and control strategies. According to the universal theoretical framework of climate and disease, climate-related disasters are associated with infectious disease risk12. Although the impacts of extreme climatic events (e.g., droughts and heatwaves) on bird migration have been widely reported, the specific ecological mechanisms by which such events may drive the transmission of highly pathogenic avian influenza in wild birds remain unclear13. Therefore, drawing on transmission ecology theory, it is useful to prioritize key meteorological factors, such as drought indices and temperature, as covariates and to employ advanced machine-learning models to disentangle these complex relationships.
This study integrates viral genomics, epidemiological modeling, spatiotemporal heterogeneity analysis, and machine-learning approaches to provide a systematic, multidimensional assessment of HPAI-H5 outbreaks in wild birds in the United States. The results show that viral genotypes evolve from early, limited spread along a single flyway to a diffusion pattern spanning multiple migratory flyways, and this pattern is associated with migratory bird movements. Although Anseriformes account for the largest number of infections and exhibit marked seasonality, they show the lowest transmission risk, whereas Strigiformes display the highest transmission risk. HPAI-H5 activity in wild birds across the United States exhibits a pronounced autumn–winter peak, and transmission hotspot regions shift with bird migration; within each flyway, transmission risk differs between migratory and non-migratory birds. Random forest models indicate nonlinear relationships between meteorological factors and infection risk in wild birds, reflecting heterogeneous impacts of climate change on avian influenza virus circulation. Collectively, these findings provide new insights into the complex transmission ecology of HPAI-H5 among wild bird populations across North America’s four major flyways, and offer data-driven evidence and a theoretical basis to support risk assessment and scientifically targeted prevention and control efforts.
Results
Spatial dissemination of HPAI-H5 genotypes in the United States
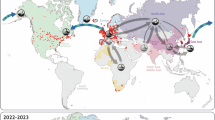
A total of 3867 valid gene sequences of HPAI-H5 subtype viruses from wild birds in the United States were screened from the GISAID database from January 2022 to April 2025. All sequences belonged to the clade 2.3.4.4b evolutionary lineage. In terms of subtype composition, H5N1 virus sequences constituted the vast majority (n = 3855), with an additional 12 sequences being H5N5 viruses. The wild bird hosts were classified into different orders on the basis of biological taxonomy, and a Sankey diagram (Supplementary Fig. 1) was constructed to display the associations and flow patterns of various genotypes between different years and bird orders. The results revealed significant interannual variations in the number of submitted samples. The number of sequences submitted reached its peak in 2022 (n = 1934), while the lowest number of sequences was recorded in 2024 (n = 371). In terms of host distribution, Accipitriformes was the primary host group for viral infection (n = 1622), followed by Anseriformes (n = 747) and Strigiformes (n = 422). The dominant viral genotypes and their host preferences exhibited clear annual evolutionary patterns. The greatest viral genotype diversity was detected in 2022, with multiple cocirculating genotypes; however, the data from 2025 revealed that D1.1 (n = 717) became the absolute dominant genotype, with its transmission mainly concentrated in Accipitriformes (n = 389). Based on the eight genotypes with the largest number of sequences during the study period, discrete phylogeographic reconstructions were performed (Fig. 1). The results showed that the spatial dissemination of HPAI-H5 genotypes across the United States was highly consistent with the major migratory flyways, although distinct diffusion patterns were observed among different genotypes. A1 and A2 genotypes showed transmission primarily concentrated along the Atlantic Flyway, with multiple migration events occurring among northeastern coastal states and relatively limited westward spread. In contrast, B1-lineage viruses (B1.1 and B1.3) exhibited a much broader connectivity pattern, spanning both the Atlantic and Mississippi Flyways and forming dense East–Midwest transmission networks and frequent bidirectional exchange between coastal and inland states. The B2.1 genotype displayed a pronounced inland focal structure, with transmission centered in North Dakota and Minnesota, consistent with a pattern of spread primarily within the Mississippi Flyway. Among all lineages, the B3 (B3.2 and B3.6) and D1.1 genotypes exhibited the widest geographic dissemination, forming extensive and multidirectional transmission networks that spanned the entire United States. The two B3 genotypes generated dense transmission links connecting the Great Lakes region, the central plains, the Gulf Coast and several western states, whereas D1.1, despite also producing wide-ranging movements, maintained a stronger and more stable presence in the Northeast. The spatial scale and orientation of these diffusion patterns indicate that their spread is not consistent with a single migratory flyway; instead, the nationwide distribution of these genotypes reflects the combined influence of the Atlantic, Mississippi, Central, and Pacific Flyways.
The spatial spread of viral genotypes closely aligns with major migratory bird flyways. A The different migratory flyways of migratory birds in the United States. B–I The reconstructed Markov jump events for each genotype (A1, A2, B1.1, B1.3, B2.1, B3.2, B3.6, D1.1). Red circles represent the number of sequences collected in each state. Black arrows denote state-to-state viral migration events, and line thickness corresponds to the frequency of inferred transitions. The base map is generated using the R package (rnaturalearth) with data from Natural Earth.
Notably, Washington State has detected dozens of avian influenza cases in wild birds, waterfowl, and backyard poultry over the past few weeks, which aligns with the surge in outbreak activity nationwide. Genetic evolutionary analysis revealed that the virus from the world’s first fatal human H5N5 case reported in Washington State in November 2025 was highly homologous to H5N5 sequences from wild bird sources (Supplementary Fig. 2). The human isolate presented as a distinct terminal branch nested within contemporaneous wild bird sequences, indicating a zoonotic spillover event. The human-derived sequence did not exhibit significant topological changes or fundamental positional alterations that typically signify adaptive evolution of the virus toward mammalian hosts. The close phylogenetic relationship with the great black-backed gull isolate suggests that the patient may have had direct or indirect contact with infected wild birds. This finding underscores the necessity and urgency of strengthening avian influenza surveillance in wild birds.
Analysis of the transmission risk of different wild bird categories
From January 2022 to April 2025, a total of 12,808 wild birds infected with the H5 subtype of avian influenza were collected. In accordance with the biological classification of the orders, the different species of wild birds were categorized into different orders (Fig. 2A). The results revealed that the top five orders of infected wild birds in the United States were Anseriformes (9040 birds), Accipitriformes (1953 birds), Strigiformes (445 birds), Passeriformes (405 birds), and Charadriiformes (395 birds). Further analysis was conducted to plot the monthly proportion of infections for different orders of birds (Fig. 2B). Infections in Anseriformes exhibited a clear seasonal pattern, with the highest proportion of infections occurring in autumn and winter each year. The transmission capacity of different orders was quantified using the basic reproduction number (R0). The results revealed (Fig. 2C) that Strigiformes had the strongest transmission capacity, with a basic reproduction number of 3.164 (95% CI: 2.711–3.726), whereas Anseriformes had the weakest transmission capacity, with a basic reproduction number of 0.992 (95% CI: 0.990–0.994). In addition, the basic reproduction number for Accipitriformes was 1.655 (95% CI: 1.571–1.748), that for Passeriformes was 1.029 (95% CI: 1.019–1.039), and that for Charadriiformes was 1.279 (95% CI: 1.198–1.369).
Transmission capacity varies among different orders of wild birds. A Logarithmic graph of HPAI-H5 infection numbers among wild birds of different orders. B The composition ratio of infections in the top five orders by infection numbers for each month. C The basic reproduction number and its 95% confidence interval for the top five orders (N = 12,238) by infection numbers. Points indicate point estimates of the basic reproduction number; lines indicate 95% confidence intervals.
Time series decomposition and seasonal variation in different orders
A time series chart of the monthly infections for the top five orders of wild birds with the greatest numbers of infections was constructed, with a smoothing curve added to visually display the trend of changes (Fig. 3). During the entire study period, avian influenza outbreaks among wild birds in the United States clearly exhibited regular patterns. Among them, the pattern for Anseriformes (migratory birds) was consistent with the overall infection pattern, while the infection patterns for other orders were not as apparent.
The epidemiological trends of HPAI-H5 differ among wild bird orders. The bars represent the number of infections in wild birds, and the lines represent the smoothed curves fitted using the GAM model and the Loess regression model. A Overall situation, B Anseriformes, C Accipitriformes, D Strigiformes, E Passeriformes, and F Charadriiformes. A, B were fitted using the GAM model, while C–F were fitted using the Loess regression model.
The Seasonal-Trend decomposition using LOESS (STL) results show that avian influenza outbreaks among wild birds exhibited clear trends and seasonality (Supplementary Fig. 3). Avian influenza outbreaks among wild birds occur annually, with a seasonal span of six months. Infections in wild bird orders, except for Passeriformes, have shown a continuous downward trend, while Passeriformes were more severely affected in 2024. The seasonality of infections in Anseriformes aligned with the overall seasonal pattern of wild birds.
The time series was divided into the four seasons of the year, with spring and autumn being the migration seasons for migratory birds in the United States, and a seasonal boxplot was created (Fig. 4). During autumn and winter, the number of infections was twice that during spring and five times that during summer. The results of the Kruskal–Wallis test conducted between different seasons revealed that the overall infection count had a chi-square value of 14.66 (P < 0.05), indicating significant differences in infection numbers between different seasons. Among them, autumn had the highest number of infections, with a median and interquartile range of 658 (196, 868), whereas summer had the lowest, at 39 (14, 137). Pairwise comparisons between different seasons were conducted using Dunn’s test with Bonferroni correction. The results revealed significant differences in infection numbers between autumn and summer and between winter and summer, whereas the differences between the other seasons were not significant. Further analysis of wild birds of different orders revealed that Anseriformes had significant differences in H5 infection numbers between spring and autumn, spring and winter, summer and autumn, and summer and winter. However, the differences between seasons were not statistically significant for Accipitriformes, Passeriformes, Charadriiformes, or Strigiformes.
Significant differences in the seasonality of avian influenza epidemics are observed among different wild bird orders. The X axis represents different seasons, and the Y axis represents the total number of HPAI-H5 infections in wild birds. Pairwise comparisons were performed using Dunn’s test (two-sided) with Bonferroni correction for multiple comparisons. P values shown above the box plots are adjusted P values; values in red indicate statistically significant differences. A Overall situation (N = 12,808); B Anseriformes (N = 9040); C Accipitriformes (N = 1953); D Strigiformes (N = 445); E Passeriformes (N = 405); F Charadriiformes (N = 395). The lower and upper boundaries of the box (interquartile) represent the 25th and the 75th percentile, respectively.
Seasonal analysis of avian influenza in wild birds across different migratory flyways
In the bar chart showing the number of wild birds infected with H5 avian influenza across different migratory flyways by month (Supplementary Fig. 4), wild bird infections across different flyways were shown to exhibit distinct patterns of fluctuation and trends. The STL decomposition results (Supplementary Fig. 5) revealed that wild bird infections along the Pacific Flyway and Atlantic Flyway tended to decrease, whereas those along the Central Flyway tended to decrease starting in mid-2022, with the trend gradually stabilizing thereafter. The Mississippi Flyway exhibited a pattern of initial decrease in wild bird infections followed by a gradual increase. The birds along the four migratory flyways in the United States were categorized into migratory and non-migratory species, and box plots of avian influenza infections across different flyways and seasons were generated, with differential analysis performed (Fig. 5).
The transmission of avian influenza in wild birds across different migratory flyways exhibits distinct seasonal characteristics. The X axis represents different seasons, and the Y axis represents the total number of HPAI-H5 infections in wild birds. Pairwise comparisons were performed using Dunn’s test (two-sided) with Bonferroni correction for multiple comparisons. P values shown above the box plots are adjusted P values; values in red indicate statistically significant differences. A–C Pacific Flyway (A, without distinguishing between migratory and non-migratory birds (N = 3089); B Migratory birds (N = 2757); C Non-migratory birds (N = 332)). D–F Central Flyway (D, without distinguishing between migratory and non-migratory birds (N = 2030); E Migratory birds (N = 1831); F Non-migratory birds (N = 199)). G–I Mississippi Flyway (G, without distinguishing between migratory and non-migratory birds (N = 3667); H Migratory birds (N = 3316); I Non-migratory birds (N = 351)). J–L Atlantic Flyway (J, without distinguishing between migratory and non-migratory birds (N = 3711); K Migratory birds (N = 3099); L Non-migratory birds (N = 612)). The lower and upper boundaries of the box (interquartile) represent the 25th and the 75th percentile, respectively.
Along the Pacific Flyway, overall wild bird infections significantly differed across seasons (chi-square value: 11.45, P < 0.05). Pairwise comparisons revealed differences in infection numbers between spring and autumn. For migratory birds along this flyway, there were also significant differences in infection numbers between spring and autumn, while non-migratory birds showed no statistically significant differences in infection numbers across different seasons. Along the Central Flyway, there were no statistically significant differences in H5 avian influenza infection numbers among wild birds across different seasons, whether considering the overall population or migratory and non-migratory birds separately. Along the Mississippi Flyway, there were no statistically significant differences in overall avian influenza infections or non-migratory bird avian influenza infections across different seasons. However, among migratory birds, there were differences in avian influenza infections across seasons (chi-square value: 9.02, P = 0.03). Pairwise comparisons indicated that this difference was primarily driven by different infection numbers between summer and autumn. Along the Atlantic Flyway, overall wild bird infections significantly differed across seasons (chi-square value: 9.33, P = 0.03). Pairwise comparisons revealed differences in infection numbers between summer and winter. For migratory birds along this flyway, there were also significant differences in infection numbers between spring and winter as well as between summer and winter, while non-migratory birds showed no statistically significant differences in infection numbers across different seasons.
Analysis of transmission risk among wild birds across different migratory flyways
The transmission capacity of H5 subtype avian influenza was evaluated for overall wild bird infections and migratory and non-migratory wild bird infections across different migratory flyways, and quantified using the basic reproduction number (Fig. 6). Along the Pacific Flyway, the basic reproduction number for overall H5 subtype avian influenza was 1.009 (95% CI: 1.005–1.012); for migratory birds it was 1.010 (95% CI: 1.007–1.014), and for non-migratory birds it was 1.011 (95% CI: 1.002–1.020). Because the confidence intervals for different bird types overlapped, there were no differences in transmission capacity among different categories of wild birds along this flyway. Along the Central Flyway, there was no difference between the overall H5 subtype avian influenza transmission capacity (R0: 1.099, 95% CI: 1.081–1.116) and the transmission capacity of migratory birds (R0: 1.106, 95% CI: 1.087–1.125), but both were greater than the transmission capacity of non-migratory birds (R0: 1.028, 95% CI: 1.005–1.052). Along the Mississippi Flyway, non-migratory birds had the strongest H5 subtype avian influenza transmission capacity (R0: 6.414, 95% CI: 4.972–8.398), which was much greater than the transmission capacity of migratory birds (R0: 1.071, 95% CI: 1.057–1.085). Along the Atlantic Flyway, the overall H5 subtype avian influenza basic reproduction number was less than 1 (R0: 0.996, 95% CI: 0.995–0.998), with non-migratory birds having the highest basic reproduction number at 1.112 (95% CI: 1.086–1.138) and migratory birds having a basic reproduction number of 1.002 (95% CI: 1.000–1.003).
Avian influenza transmission capacity in wild birds differs across migratory flyways. The X axis represents the basic reproduction number values, where the scale is logarithmically transformed. The Y axis represents different categories of wild birds. A Pacific Flyway (N = 3089); B Central Flyway (N = 2030); C Mississippi Flyway (N = 3667); D Atlantic Flyway (N = 3711). Points indicate point estimates of the basic reproduction number (R₀); lines indicate 95% confidence intervals.
Spatiotemporal distribution analysis of HPAI-H5 avian influenza in wild birds
Spatiotemporal distribution maps of H5 avian influenza in wild birds across the United States in different years and seasons were created (Supplementary Fig. 6). Throughout the study period, all states reported wild bird infection cases, with Florida having the highest number of infected wild birds at 782. In 2022, Minnesota had the most infections (587 birds); in 2023, California had the most infections (213 birds); in 2024, Florida had the most infections (201 birds); and in 2025, Maine had the most infections (169 birds). The infection situation exhibited a decreasing trend year by year, and the states with severe infections were all located in coastal areas of the United States. Spatial clustering analysis was conducted for all birds in the United States and for migratory and non-migratory birds across different seasons, and the results are shown in Table 1.
Overall, wild birds nationwide and migratory wild birds exhibited spatial clustering (the non-random geographical distribution of H5 infections, where the proximity of high-prevalence locations exceeds what would be expected by chance) in spring, summer, and autumn, while non-migratory wild birds exhibited spatial clustering year-round and in autumn. The local indicators of spatial association (LISA) cluster map and cold spot-hotspot analysis map (Fig. 7) revealed that in spring, infections in overall birds and migratory birds formed hotspot areas (statistically significant high-value areas) in North Dakota, South Dakota, Minnesota, and Wisconsin, indicating high-high clustering. In summer, the hotspot areas shifted to the northwest, appearing in Washington, Oregon, and Idaho. In addition, Alaska showed high-low clustering phenomena in both spring and summer. In winter, for overall and migratory birds, Kansas showed high-low clustering phenomena, whereas non-migratory bird infections exhibited high-low clustering in Texas.
The spatiotemporal distribution of highly pathogenic avian influenza in wild birds is heterogeneous. A Annual analysis (A1–A3 LISA clustering for all wild birds, migratory birds, and non-migratory birds. A4–A6 Cold spot-hotspot analysis for all wild birds, migratory birds, and non-migratory birds). B–E Seasonal analysis for spring, summer, autumn, and winter, respectively, with the same structure as the annual analysis.
Spatiotemporal distribution analysis of HPAI-H5 avian influenza along migratory flyways
Further spatiotemporal analysis of H5 avian influenza in wild birds across major migratory flyways revealed substantial differences in the timing and geographic distribution of infections among the flyways. With respect to the Pacific Flyway, the global spatial autocorrelation results indicate that different types of wild birds did not exhibit significant spatial clustering across different seasons. Analysis of local spatial clustering patterns (LISA and Getis-Ord Gi*) revealed significant local clustering and seasonal dynamics in the geographical distribution of viral infections (Supplementary Figs. 7 and 8). During spring, Montana was identified as exhibiting high clustering, with Montana, Washington, and Idaho constituting hotspot areas for viral infections, whereas non-migratory bird infections in California and Arizona represented the primary cold spot areas. During summer, viral hotspot areas underwent spatial shifts, with California becoming the primary hotspot area, while Oregon and Montana served as infection hotspots for migratory birds. In autumn, the spring hotspot areas for non-migratory birds transformed into cold spot areas, and during winter, Washington became the hotspot area for non-migratory bird infections. In the Central Flyway, non-migratory birds significantly differed in terms of the Moran’s I index for the entire year and autumn. LISA clustering maps (Supplementary Fig. 9) and cold spot-hotspot maps (Supplementary Fig. 10) indicate that hotspot areas were driven primarily by non-migratory birds, resulting in the formation of clustered regions covering Wyoming, Colorado, and New Mexico. During winter, infections in all birds and migratory birds along this flyway exhibited hotspot areas in Wyoming and New Mexico. On the Mississippi Flyway, the spread of H5 avian influenza virus exhibited strong spatial autocorrelation. Global Moran’s I index analysis revealed that avian influenza infections in total birds and migratory birds exhibited significant spatial clustering patterns during spring, summer, and autumn. In addition, migratory birds demonstrated significant spatial clustering at the annual scale. For non-migratory birds, their spatial clustering effects were primarily concentrated in spring and autumn. According to the LISA clustering map (Supplementary Fig. 11) and cold spot-hotspot map (Fig. 8), Wisconsin exhibited that both total birds and migratory birds formed stable high-high clustering areas, indicating that this state is a key hotspot for viral transmission. During spring, the southern regions of this flyway were all cold spot areas, whereas Minnesota in the northern part of the route was identified as a hotspot area for both migratory and non-migratory bird infections. At the beginning of summer, the infection hotspots further concentrated northwards and were primarily distributed in the northern regions of the flyway. During autumn, Minnesota, Iowa, and Wisconsin were hotspot areas for total and migratory bird infections, whereas Mississippi and Alabama were hotspot areas for non-migratory birds. By winter, the bird infection hotspot areas all shifted to the southern regions of this flyway. On the Atlantic Flyway, migratory birds showed significant spatial clustering during summer, whereas non-migratory birds exhibited significant spatial clustering annually as well as during summer and autumn. According to the LISA clustering map (Supplementary Fig. 12) and cold spot-hotspot map (Supplementary Fig. 13), annual-scale analysis confirmed that Georgia and Florida are two key viral transmission core areas in the southern part of this flyway, constituting persistent infection hotspots. During summer, migratory bird infection hotspot areas were in the north, whereas non-migratory bird infection hotspot areas were in the south, indicating a clear north‒south spatial differentiation. During autumn and winter, non-migratory bird infection hotspot areas remained in Florida and Georgia, whereas migratory birds exhibited no hotspot areas.
Avian influenza transmission along the Mississippi Flyway exhibits significant spatial autocorrelation. A Annual analysis (A1–A3 Cold spot-hotspot analysis of all wild birds, migratory birds, and non-migratory birds). B–E Seasonal analysis for spring, summer, autumn, and winter, respectively, with the same structure as the annual analysis.
Analysis of meteorological factors based on Random Forest and partial dependence
A Random Forest regression model was established with wild bird infection counts as the dependent variable and selected meteorological factors as independent variables. The three hyperparameters of the Random Forest model were optimized: ntree (number of trees), maxnodes (maximum depth of individual trees), and mtry (number of variables randomly selected at each split). The optimal parameter combination selected was ntree=500, maxnodes=32, and mtry=5. Tenfold cross-validation was performed to obtain the optimal model, and the coefficient of determination was calculated to be 0.87. On the basis of the variable importance calculated from the Random Forest model, higher values indicate greater variable importance (Supplementary Table 1). Based on the current dataset, the variables ranked from highest to lowest impact were: Palmer Drought Severity Index, Palmer Modified Drought Index, average temperature, precipitation, Palmer Z-Index, and cooling degree days.
Partial dependence and individual conditional expectation (ICE) with 500 simulations revealed a nonlinear relationship between infection counts and meteorological factors (Fig. 9). Both the average temperature and the number of cooling degree days monotonically decreased with the increasing number of infections. When the number of cooling degree days exceeded 100 days, the number of wild bird infections exhibited little change as the values increased. The Palmer Drought Severity Index and Palmer Modified Drought Index exhibited an inverted U-shaped relationship with the number of infections, with the highest number of infections occurring when the Palmer Drought Severity Index or Palmer Modified Drought Index was approximately −3.3, followed by a rapid decrease in wild bird infections as the values increased. The Palmer Z-Index and precipitation exhibited a U-shaped relationship with the number of wild bird infections, with the lowest number of wild bird infections occurring when the Palmer Z-Index was between −3 and 0 or when precipitation was ~2.8 inches. Beyond this range, the number of wild bird infections gradually increased as the values either increased or decreased.
There is a nonlinear association between meteorological factors and avian influenza. The X axis represents the values of the corresponding meteorological factors, and the Y axis represents the number of wild bird infections. PDSI is the Palmer Drought Severity Index, PMDI is the Palmer Modified Drought Index, and PZI is the Palmer Z-Index.
Discussion
Since 2022, the global spread of the H5N1 2.3.4.4b clade highly pathogenic avian influenza virus has increased worldwide, with high prevalence among various wild bird species and efficient infection of multiple mammalian species, including seals, sea lions, cetaceans, and cattle14. The United States’ first report of this avian influenza virus clade being transmitted from wild birds to cattle and causing intercontinental spread has attracted global attention5. The results of this study also revealed that the outbreak from 2022 to 2025 was primarily dominated by the clade 2.3.4.4b evolutionary lineage, which is highly consistent with the persistent outbreak situation caused by this viral clade globally15. From a genotypic perspective, the viral population in wild birds in the United States is evolving from a high diversity towards a single dominant genotype. The transition from multiple genotypes cocirculating in 2022 to the single D1.1 genotype dominating by 2025 suggests that this genotype may have acquired significant fitness advantages or stronger adaptation to key hosts, with Anseriformes, Accipitriformes, and Strigiformes being the three bird orders that require the most attention in avian influenza prevention and control. Notably, the increased prevalence of the D1.1 genotype was concurrent with the shift of Accipitriformes becoming the primary host group. This temporal overlap is consistent with a potential ecological or evolutionary association, suggesting that the D1.1 genotype may be characterized by traits that facilitate its circulation within raptor populations. However, potential sampling bias must be considered when interpreting these findings. Phylogeographic reconstruction further revealed a significant association between the spatial transmission patterns of different genotypes within the United States and migratory bird flyways16. The transmission pathways of early genotypes such as A1, A2, and B2.1 were relatively limited, spreading along single migratory flyways—either the Atlantic or Mississippi—indicating their relatively narrow host range or ecological adaptability. In contrast, viruses of the B1-lineage (B1.1 and B1.3) have demonstrated cross-flyway transmission capacity, capable of establishing bidirectional connections between coastal and inland host populations. Most notably, the D1.1 genotype has exhibited an unprecedented nationwide dispersal pattern, with its transmission network spanning all four major migratory flyways. This finding not only aligns with the epidemiological trend of D1.1 becoming the dominant genotype in 2025, but also suggests that this genotype may have acquired a broader host spectrum, stronger environmental adaptability, or more efficient inter-regional transmission risk. This evolutionary trajectory from localized to nationwide distribution and from single-flyway to multi-flyway transmission may reflect the virus’s gradual optimization of its transmission strategies during adaptation to the North American wild bird ecosystem. Further experimental research is urgently needed to elucidate the molecular mechanisms underlying the emergence of the D1.1 genotype and its public health implications.
The occurrence of avian influenza in wild birds in the United States shows clear seasonality, with more severe outbreaks in autumn and winter and a lower severity in summer, which is consistent with observations from countries such as Japan and Bangladesh17,18. H5 outbreaks occur annually in wild birds in the United States, with peaks typically in autumn, lasting ~6 months. Migratory birds are primarily dominated by Anseriformes, and their infection dynamics are strongly associated with the overall seasonal pattern of HPAI-H5 outbreaks in the United States. This pattern is highly consistent with their large-scale, flocking migration and wintering behaviors19. Wild bird avian influenza infections are most common in the United States during autumn, which is related to the autumn migration of migratory birds. Greater attention should be given to avian influenza prevention and control during autumn migration than spring migration. The R0 value for Anseriformes being less than 1 suggests that although Anseriformes constitute the main body of the outbreak, the virus has not formed efficient, sustained transmission chains within their populations. Therefore, Anseriformes may play more of a role as the primary viral reservoir and long-distance vector in the outbreak, which also explains why large-scale highly pathogenic avian influenza outbreaks have not occurred despite such massive migration, and why the HPAI-H5 positive rate is very low in migratory bird avian influenza surveillance. The R0 values for raptors such as Strigiformes and Accipitriformes were significantly greater than 1, with Strigiformes exhibiting the highest transmission potential. This finding suggests that although the total number of infections in these species is far lower than that in Anseriformes, once the virus enters their populations, it may have the capacity to trigger efficient local transmission. Their position at the top of the food chain makes them key nodes for viral spillover from aquatic environments to terrestrial environments, and even to mammals20,21. Seasonal decomposition also revealed that the infection patterns of these raptors do not display strong seasonality associated with large-scale migration, suggesting that their infections may be more associated with local, opportunistic predatory behaviors. The avian influenza outbreak in wild birds in the United States was relatively severe in 2022 but showed a gradual declining trend over time. This pattern of change may be attributed to the synergistic effects of multiple factors, including the establishment of natural immunity in wild bird populations, changes in viral virulence, alterations in ecological environmental factors, and the implementation of relevant prevention and control measures. Assessment of the transmission risk among wild birds on different migratory routes revealed that on the Mississippi and Atlantic Flyways, the transmission risk of resident birds far exceeds that of migratory birds. Particularly along the Mississippi Flyway, the notably high R0 values among non-migratory birds suggest that the presence of resident populations is consistent with the formation of highly efficient and sustained viral transmission “engines,” which may serve as localized reservoirs for the outbreak.
From a spatial distribution perspective, the distribution of H5 in wild birds in the United States exhibits apparent spatiotemporal heterogeneity, with hotspot areas varying across different seasons. Importantly, hotspots identified in this analysis should be interpreted in light of the underlying model uncertainty and potential surveillance biases. Research has revealed that for non-migratory birds, autumn is the primary season for spatial clustering of avian influenza infections. On a national scale, this study revealed a macropattern in which viral hotspots migrate from north-central regions in spring to northwestern regions in summer and autumn, providing suggestive spatial evidence for the association between viral transmission and migratory bird flyways. Spring hotspots are concentrated in states such as North Dakota, South Dakota, and Minnesota, which highly coincide with the geographic location of North America’s most important waterfowl breeding area—the Prairie Pothole Region22. Large numbers of migratory birds congregate here for breeding, greatly increasing the density and risk of viral transmission, thus forming the core of spring infections. The subsequent shift of hotspots to the northwest may reflect the post-breeding dispersal of birds, as well as the early stages of the autumn southwards migration of some species along the Pacific Flyway. This finding suggests that the transmission of avian influenza among wild birds across the United States is characterized by a structured spatial pattern that is closely associated with the seasonal life cycles of migratory birds. Spatial analysis of the current dataset suggests that the four major migratory flyways in the United States are not homogeneous transmission corridors, but rather, each has unique spatiotemporal distribution characteristics. The Mississippi Flyway exhibits a classic north‒south migration pattern, with its hotspots showing significant north-south movement according to seasons, highly consistent with the migratory rhythms and pathways of migratory birds. The Atlantic Flyway presents a unique north-south differentiation pattern. In summer, migratory bird infections are associated with northern hotspots, whereas non-migratory birds are associated with an independent transmission core in the south. By autumn and winter, as migratory birds migrate south or depart, only persistent hotspots maintained by non-migratory bird populations in Georgia and Florida remain. These findings align with the conclusion that non-migratory birds have higher R0 values along this flyway, collectively revealing that southern resident bird populations are the transmission core of the epidemic along this flyway. The transmission pattern of the Central Flyway is dominated by non-migratory birds supplemented with migratory birds. Its stable core hotspot areas are maintained year-round by non-migratory bird populations in Wyoming, Colorado, and New Mexico. The arrival of winter migratory birds does not alter this core pattern but rather adds additional transmission risk, which explains why winter migratory birds also form hotspots in this region. The Pacific Flyway presents a scattered, localized transmission pattern, with insignificant global spatial autocorrelation coexisting with the seasonal emergence of local hotspots, suggesting that transmission along this flyway may be more influenced by complex terrain and more diverse host communities, making it difficult for viral transmission to form large-scale aggregations.
Long-term climate change and extreme weather events affect the migration timing, migratory species, and the adaptability of migratory birds to highly pathogenic avian influenza viruses23. Therefore, analyzing meteorological factors in relation to avian influenza in migratory birds could help elucidate the potential effects of climate change on the transmission of the virus among migratory birds. Previous research in the Poyang Lake region of China revealed a decreasing trend between precipitation and avian influenza positivity rates; however, the present study exhibited a U-shaped relationship. Moreover, unlike the decreasing trend observed in this study, the relationship between temperature and avian influenza positivity rates in the Poyang Lake region exhibited an inverted U-shape. This discrepancy may be due to differences in the relationship between temperature changes and avian influenza prevalence along different migratory bird flyways3. We also found that the number of avian influenza positive cases decreased as the number of cooling degree days increased, which may be linked to the behavior of wild birds. During colder weather, birds may reduce their foraging activities and interactions. Furthermore, analysis of the current dataset suggests a pronounced nonlinear association between drought severity and the number of avian influenza infections in wild birds. In this study, three drought severity indicators reflecting different conditions were used to quantify their relationships with avian influenza in wild birds. This approach revealed the potential effects of influencing factors at different critical thresholds, providing valuable insights to help relevant authorities implement more effective and scientifically informed prevention and control measures for avian influenza in wild birds. This study also had several limitations. On the one hand, although the most comprehensive monitoring data have been used, owing to potentially insufficient monitoring efforts for wild birds in certain states or regions, the actual number of infection cases in these areas may be underestimated, which could have a certain impact on the geographical distribution and epidemic patterns of the disease. On the other hand, while factors such as temperature and precipitation were considered, there are still other influencing factors that have not been considered.
While our study focuses on the United States, the constructed analytical framework—integrating phylogeography, transmission risk estimation, spatial clustering, and machine learning-based meteorological analysis—demonstrates procedural generalizability. This multidimensional analytical pipeline effectively disentangles complex host-environment-virus interactions and could be adapted for surveillance systems in other regions (e.g., Europe or Asia), provided that sufficient surveillance data are available. However, the ecological findings derived from this framework exhibit significant context specificity. Biologically driven patterns, such as high transmission risk in raptors and the role of Anseriformes as long-distance vectors, likely represent broad ecological traits that are applicable to other migratory flyways. In contrast, specific spatial and meteorological determinants are inherently regional. For instance, the specific inverted U-shaped relationship between the Palmer Drought Severity Index and infection risk reflects the unique hydroclimatic conditions of North America. This contrasts with previous findings from Poyang Lake in China, highlighting the heterogeneity of environmental drivers across different climate zones, and suggests that interpretations for other regions should be made with caution. Furthermore, the identified spatial hotspots (e.g., the ‘Prairie Pothole Region’) are intrinsically tied to North America’s unique geography and might not be directly generalized to other geographical contexts. Therefore, while this methodological approach serves as a universal tool for early warning, specific risk thresholds and spatial hotspots must be recalibrated when applying the framework to new ecological contexts to obtain context-specific conclusions.
Methods
Date collection
In this study, HPAI-H5 virus genome sequences obtained from wild bird samples from the United States from January 2022 to April 2025 from the Global Initiative on Sharing All Influenza Data (GISAID) database were screened24. The data entries included in the analysis contained detailed metadata, primarily including sample collection date, sequence accession number, virus subtype, genotype, host species, and viral evolutionary lineage. Data on the number of wild birds infected with H5 subtype avian influenza from January 2022 to April 2025 were collected from the United States Department of Agriculture (USDA). The data include the date of wild bird sampling, the state where sampling occurred, and the types of wild birds25. Meteorological data were obtained from the National Centers for Environmental Information (NCEI), including average temperature (°F), precipitation (inch), cooling degree days (°Df), Palmer Drought Severity Index, Palmer Modified Drought Index, and Palmer Z-Index for the entire United States from January 2022 to April 2025. These data can be directly downloaded from the website and represent monthly average data weighted by land area or population. The specific algorithms can be found in the official documents from the NCEI26. When wild bird data from different orders and months were classified and statistically analyzed, if no H5 samples were detected for a particular order of wild birds, the count was recorded as 0; otherwise, calculations were performed on the basis of the original existing data. However, wild birds with unknown sampling dates that existed in the original data were removed. During the study period, there were no missing values in the meteorological data. On the basis of bird information from “Birds of the World”27, infected wild birds were classified into two categories: migratory birds and non-migratory birds. With respect to wild bird data for which bird names were ambiguous and difficult to determine, these entries were excluded from the analysis of migratory and non-migratory birds.
Statistical analysis
Spatial dissemination of HPAI-H5 genotypes in the United States
The spatial dissemination of HPAI-H5 genotypes in the United States was analyzed using a discrete phylogeographic framework implemented in BEAST (v1.10.5). Whole-genome sequences collected from 2022 to 2025 were aligned with MAFFT and screened using IQ-TREE (v2.1.4) before genotype-specific datasets were constructed (Supplementary Method 1). In addition, based on information from the U.S. Fish & Wildlife Service and the United States Department of Agriculture websites25,28, migratory flyways in the United States were classified into four routes—the Pacific, Central, Mississippi, and Atlantic flyways—to illustrate the status of different migratory pathways. In November 2025, a fatal case of H5N5 infection occurred in the United States. To understand the association between this fatal case and virus infections in wild birds, HA gene sequences of H5N5 isolates from the United States between 2022 and 2025 were collected and aligned using MAFFT, followed by manual trimming to remove low-quality regions. Maximum likelihood phylogenetic inference was performed in IQ-TREE2 using the optimal substitution model selected by ModelFinder, with branch support assessed through 1000 ultrafast bootstrap replicates. The resulting maximum likelihood tree was visualized and color-annotated in iTOL. Branch lengths represent the number of nucleotide substitutions per site.
Analysis of serial interval and transmission risk
The serial interval refers to the time interval between the onset of symptoms in a primary case and the onset of symptoms in a secondary case. It follows a certain distribution, and the generation time distribution is a critical step in calculating the basic reproduction number using statistical models. The basic reproduction number is an important metric in epidemiology used to measure the transmission capacity of an infectious disease. It refers to the average number of individuals directly infected by a single infectious individual during their entire infectious period in a completely susceptible population29. In this study, the sample collection date of wild birds was used as a proxy indicator for symptom onset time, representing the time point when infection was detected, which can effectively reflect the actual timing of an epidemiological event. On the basis of the sampling time data, corresponding epidemic curves were constructed, and the est.GT function in the “R0” package was used to estimate the serial interval distribution of the wild birds. On this basis, the package’s function (est.R0.EG) was further applied to estimate the basic reproduction number and its 95% confidence interval for different bird orders and different flyways using an exponential distribution30.
Generalized additive model and loess regression model
A generalized additive model (GAM) is used to reflect the nonlinear relationships between variables. By estimating the influence of each independent variable using individual smooth spline functions, the GAM allows independent nonlinear modeling of the effect of each independent variable while maintaining the linear interpretability of the overall model. The GAM consists of different additive terms, each of which is estimated using a single smooth spline function to explain how the dependent variable changes with the independent variables31. Locally estimated scatterplot smoothing (LOESS) is a nonparametric regression method primarily used to capture trends in data through smooth curves. It combines the concepts of locally weighted regression and polynomial fitting, making it suitable for exploring data with nonlinear relationships32. In this study, GAM and LOESS regression models were used to smooth the original time series data of the top five wild birds with the greatest number of infections in the United States (Supplementary Method 2).
STL decomposition
STL decomposition is a time-series decomposition method based on local weighted regression. It smooths the time series through local weighted regression and iteratively decomposes the time series into three components: trend, seasonality, and residual33. The basic formula is as follows:
where \({Y}_{t}\) is the observed value of the time series, \({T}_{t}\) is the trend component, \({S}_{t}\) is the seasonal component, and \({{R}}_{t}\) is the residual component. In this study, STL decomposition was used to analyze the H5 avian influenza data of the top five wild bird orders with the highest infection numbers in the United States to determine whether there were any seasonality or trends. Similarly, STL decomposition was performed on avian influenza data for wild birds along the different flyways. In this study, March to May was set as spring, June to August as summer, September to November as autumn, and December to February of the following year as winter. Box plots were used to compare and analyze the differences in H5 avian influenza counts among wild birds of different orders across different seasons (Supplementary Method 3).
Spatial autocorrelation analysis
Spatial autocorrelation is a statistical method used to measure the distribution characteristics and mutual relationships of spatial data (Supplementary Method 4). The local Moran’s I index is used to reveal local indicators of spatial association. When the local Moran’s I index is significant, four types of spatial correlation states exist: high-high aggregation areas, high-low aggregation areas, low-high aggregation areas, and low-low aggregation areas34. Hotspot analysis is used to study the clustering patterns and locations of high- or low-value features in local spaces, and the \({G}_{i}^{*}\) statistic for each feature in the dataset is calculated to determine the state of spatial clustering. In this study, R 4.5.0 was used to create spatial distribution maps for different periods and seasons and to generate spatial clustering maps and cold spot-hotspot analysis maps to calculate Moran’s I index for H5 avian influenza infections in migratory and non-migratory birds across the United States and different flyways.
Random Forest algorithm and partial dependence analysis
The Random Forest algorithm is a classic machine learning algorithm that uses the bagging method to extract multiple sample sets (Supplementary Method 5). Partial dependence represents the result of individual conditional expectations converging to the mean, reflecting the overall effect of the variable, while ICE reflects the relationship for each individual observation3. In this study, Random Forest regression was used to determine the relationships between meteorological factors and the number of wild bird infections. Monthly meteorological data for the United States were used as the input layer of the Random Forest model, while the monthly number of wild bird infections nationwide served as the output layer to train the model. Grid search and cross-validation were used to identify optimal parameters, and the final model was fitted on the basis of these optimal parameters (Supplementary Method 5).
Ethical approval
As this study did not involve patients or the public, no formal ethical evaluation or informed consent was required.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The authors declare that the data supporting the findings of this study are available in the paper and the Supplementary Information file. The raw data and code have been uploaded to GitHub (https://github.com/kfang0/wild-bird-analysis-and-risk-assessment). Details on how to access the previously reported H5 sequencing data analyzed in this study (including GISAID accession numbers) are provided in the Source Data file accompanying this manuscript. For further information, please contact the corresponding author. Source data are provided with this paper.
References
Xiong, W. & Zhang, Z. Influenza Virus Genomic Mutations, Host Barrier and Cross-species Transmission. Curr Genomics 26, 161–174 (2025).
Klim, H. et al. Serological analysis in humans in Malaysian Borneo suggests prior exposure to H5 avian influenza near migratory shorebird habitats. Nat. Commun. 15, 8863 (2024).
Fang, K. et al. Detection and prevalence of avian influenza epidemic in the southwest of Poyang Lake and analysis of the influence of meteorological factors. One Health 20, 101047 (2025).
Dokter, A. M. et al. Seasonal abundance and survival of North America’s migratory avifauna determined by weather radar. Nat. Ecol. Evol. 2, 1603–1609 (2018).
Nguyen, T.-Q. et al. Emergence and interstate spread of highly pathogenic avian influenza A(H5N1) in dairy cattle in the United States. Science 388, eadq0900 (2025).
Taylor, L. First ever human case of H5N5 avian flu confirmed in the US. BMJ 391, r2453 (2025).
Xie, R. et al. The episodic resurgence of highly pathogenic avian influenza H5 virus. Nature 622, 810–817 (2023).
Tian, H. et al. Avian influenza H5N1 viral and bird migration networks in Asia. Proc. Natl. Acad. Sci. USA 112, 172–177 (2015).
Olsen, B. et al. Global patterns of influenza A virus in wild birds. Science 312, 384–388 (2006).
Leguia, M. et al. Highly pathogenic avian influenza A (H5N1) in marine mammals and seabirds in Peru. Nat. Commun. 14, 5489 (2023).
Youk, S. et al. H5N1 highly pathogenic avian influenza clade 2.3.4.4b in wild and domestic birds: introductions into the United States and reassortments, December 2021-April 2022. Virology 587, 109860 (2023).
Alcayna, T., Rao, V. B. & Lowe, R. Identifying the climate sensitivity of infectious diseases: a conceptual framework. Lancet Planet Health 9, 101291 (2025).
Zhang, L. B., Ma, Z. & Liu, Y. Biological traits and biome features mediate responses of terrestrial bird demography to droughts. J. Anim. Ecol. 93, 1868–1880 (2024).
Koopmans, M. P. G. et al. The panzootic spread of highly pathogenic avian influenza H5N1 sublineage 2.3.4.4b: a critical appraisal of One Health preparedness and prevention. Lancet Infect. Dis. 24, e774–e781 (2024).
Cui, P. et al. Global dissemination of H5N1 influenza viruses bearing the clade 2.3.4.4b HA gene and biologic analysis of the ones detected in China. Emerg. Microbes Infect. 11, 1693–1704 (2022).
Damodaran, L., Jaeger, A. S. & Moncla, L. H. Ecology and spread of the North American H5N1 epizootic. Nature 649, 432–441 (2026).
Islam, A. et al. Potential risk zones and climatic factors influencing the occurrence and persistence of avian influenza viruses in the environment of live bird markets in Bangladesh. One Health 17, 100644 (2023).
Fujimoto, Y. & Haga, T. Association between highly pathogenic avian influenza outbreaks and weather conditions in Japan. J. Vet. Med. Sci. 86, 1045–1051 (2024).
Fourment, M., Darling, A. E. & Holmes, E. C. The impact of migratory flyways on the spread of avian influenza virus in North America. BMC Evol Biol 17, 118 (2017).
Vidaña, B. et al. The role of birds of prey in West Nile virus epidemiology. Vaccines 8, 550 (2020).
Grear, D. A., Hall, J. S., Dusek, R. J. & Ip, H. S. Inferring epidemiologic dynamics from viral evolution: 2014–2015 Eurasian/North American highly pathogenic avian influenza viruses exceed transmission threshold, R0 = 1, in wild birds and poultry in North America. Evolut. Appl. 11, 547–557 (2018).
McKenna, O. P., Mushet, D. M., Kucia, S. R. & McCulloch-Huseby, E. C. Limited shifts in the distribution of migratory bird breeding habitat density in response to future changes in climate. Ecol. Appl. 31, e02428 (2021).
Prosser, D. J., Teitelbaum, C. S., Yin, S., Hill, N. J. & Xiao, X. Climate change impacts on bird migration and highly pathogenic avian influenza. Nat. Microbiol. 8, 2223–2225 (2023).
Khare, S. et al. GISAID’s Role in Pandemic Response. China CDC Wkly 3, 1049–1051 (2021).
Animal and Plant Health Inspection Service. https://www.aphis.usda.gov (2025).
National Centers for Environmental Information (NCEI). https://www.ncei.noaa.gov/access (2025).
Birds of the World - Comprehensive life histories for all bird species and families. https://birdsoftheworld.org/bow/home (2025).
Migratory Bird Program Administrative Flyways | U.S. Fish & Wildlife Service. https://www.fws.gov/partner/migratory-bird-program-administrative-flyways (2025).
Kim, W.-H. & Cho, S. Estimation of the basic reproduction numbers of the subtypes H5N1, H5N8, and H5N6 during the highly pathogenic avian influenza epidemic spread between farms. Front. Vet. Sci. 8, 597630 (2021).
Wallinga, J. & Lipsitch, M. How generation intervals shape the relationship between growth rates and reproductive numbers. Proc. Biol. Sci. 274, 599–604 (2007).
Wand, H., Dassaye, R., Reddy, T., Yssel, J. & Ramjee, G. Geographical-level contributions of risk factors for HIV infections using generalized additive models: results from a cohort of South African women. AIDS Care 31, 714–722 (2019).
Zimbalist, A. et al. Utilization of locally estimated scatterplot smoothing (LOESS) regression to estimate missing weights in a longitudinal cohort of breast cancer patients. Ann. Epidemiol. 104, 55–60 (2025).
Lin, L. et al. Spatiotemporal distribution of migraine in China: analyses based on Baidu index. BMC Public Health 23, 1958 (2023).
Zhang, C., Luo, L., Xu, W. & Ledwith, V. Use of local Moran’s I and GIS to identify pollution hotspots of Pb in urban soils of Galway, Ireland. Sci. Total Environ. 398, 212–221 (2008).
Acknowledgements
We would like to express our sincere gratitude to the Global Initiative on Sharing All Influenza Data (GISAID), the United States Department of Agriculture (USDA), and the National Centers for Environmental Information (NCEI) for providing open-access data. We thank Dr. Song Chao’s team from the West China School of Public Health, Sichuan University, for their assistance with model development in the early stages. We also thank Edward Hill from the University of Liverpool for offering crucial insights into the manuscript. This research was financially supported by the Self-supporting Program of Guangzhou Laboratory (GZNL2024A01004 and SRPG22-007), the National Key Research and Development Program of China (2024YFC2311404), the Fujian Guiding Science and Technology Plan Project (2023Y0004), the Science and Technology Project of Jiangxi Provincial Health Commission (202410978), the Major Science and Technological Project of Jiangxi Province (20201BBG71010), the China Postdoctoral Science Foundation (K2825002) and the Postdoctoral Fellowship Program of CPSF (GZC20250516).
Author information
Authors and Affiliations
Contributions
K.F.: conceptualization, data analysis, visualization, writing—original draft; J.H.L.: data analysis, visualization, and writing—review; H.F.Z. and J.B.: visualization and data analysis; Z.Y.Z., W.T.S., and T.M.C.: conceptualization, project administration, writing—review and editing. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Dae-Sung Yoo, and the other, anonymous, reviewer for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fang, K., Li, J., Zhao, H. et al. Assessing HPAI-H5 transmission risk across wild bird migratory flyways in the United States. Nat Commun 17, 2524 (2026). https://doi.org/10.1038/s41467-026-69344-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69344-w