Abstract
Behavioral tasks can be solved employing various strategies. Sometimes, different strategies result in the same observable behavior, making them latent. In this study, we infer the latent behavioral strategy used by monkeys in a working memory updating task by comparing the representational geometry of two prefrontal regions - the lateral prefrontal cortex (LPFC) and the prearcuate cortex (PAC) - with that of recurrent neural network (RNN) models trained to solve the task using different strategies. We found that neural activity patterns in both LPFC and PAC align with only one of the proposed strategies, suggesting that monkeys employ this latent strategy to perform the task. These findings open avenues for investigating the processes that lead to strategy learning and the decision-making mechanisms that determine which strategies are chosen when multiple options are available.
Similar content being viewed by others
Introduction
Organisms can respond differently to the same stimulus depending on their internal state, a capability referred to as behavioral flexibility1. Additionally, they can use various internal states to produce a similar response to a stimulus, each of which is referred to as a task strategy2,3. Different strategies are implemented by different neural dynamics4,5,6 and can be examined mechanistically using simulations4,7,8. Often, different strategies result in noticeable behavioral differences, such as variations in response accuracy3 or reaction times4. However, in some instances, different strategies may not produce any observable behavioral differences (i.e., latent strategies), and they can only be differentiated via the activity signatures in neural systems.
To date, most research of cognitive strategy focuses on the behaviorally observable ones9, while latent strategy has been largely overlooked due to the detection difficulty. Parameterized models fitted to the neural activities, on the other hand, usually only reveal the implementation mechanism of processes rather than cognitive-level strategies10,11,12,13. Only recently, a few studies showed that recurrent neural network (RNN) simulations can be used to infer which latent strategy is used by animals, when multiple strategies are available4,7,8. Here, we explored the latent strategies employed by animals in a working memory task by comparing the geometric representational properties of two prefrontal regions—the lateral prefrontal cortex (area 9/46, LPFC) and the prearcuate cortex (area 8a, PAC)—with those of RNN models explicitly trained to utilize different strategies to solve the same task.
Monkeys and RNNs were trained to solve a working memory updating task14. Human behavioral studies have shown that working memory updating tasks can be performed employing different strategies15. One possible strategy is to initially encode memories in the absence of attention, and then transfer the information to an attended format at the time of recall16 (Retrieve at Recall strategy or R@R). Alternatively, another strategy is to rehearse and update the memory online as the items are shown17 (Rehearse and Update strategy or R&U). The Retrieve at Recall strategy would presumably involve initially encoding multiple memories in orthogonal neural activity subspaces until one of them is selectively rotated for readout18. On the other hand, the Rehearse and Update strategy would presumably involve encoding both the initial and the updated memories directly onto the readout activity subspace.
In both LPFC and PAC, we observed activity patterns that were consistent with the RNNs trained to solve the task using the Rehearse and Update strategy, and were inconsistent with the RNNs trained to solve the task using the Retrieve at Recall strategy. Thus, we infer that the monkeys employ the latent Rehearse and Update strategy to solve the task.
Results
Behavioral results
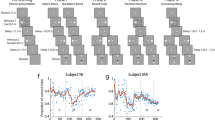
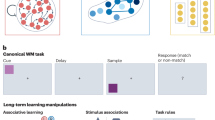
We trained RNNs and two monkeys to perform a working memory updating task (Fig. 1a). In this task, they had to report the location of the last target (red square) seen. The monkeys reported with a saccadic eye movement to the target location. For Target 1/Target 2 (T/T) trials (Fig. 1a top), monkeys had to report the location of Target 2, while for Target 1/Distractor (T/D) trials (Fig. 1a, bottom), monkeys had to report the location of Target 1. On average, monkeys reported the correct location on over 60% of trials (Monkey A: Overall: M(SD) = 0.700(0.021), T/D: M(SD) = 0.688(0.022), T/T: M(SD) = 0.711(0.032); Monkey B: Overall: M(SD) = 0.616(0.037), T/D: M(SD) = 0.626(0.034), T/T: M(SD) = 0.611(0.063)) (Supplementary Fig. 1a). To get a reward, in addition to reporting the correct location, the monkeys had to respond within 500 ms of the go-cue, and maintain fixation on the target for at least 200 ms (rewarded trials: Monkey A: Overall: M(SD) = 0.533(0.011), T/D: M(SD) = 0.506(0.018), T/T: M(SD) = 0.559(0.008); Monkey B: Overall: M(SD) = 0.563(0.036), T/D: M(SD) = 0.563(0.041), T/T: M(SD) = 0.569(0.065)) (Supplementary Fig. 1b). The monkeys’ performance tended to decrease towards the end of sessions. We speculate that this decrease is the product of fatigue or loss of motivation. Prior to this decrease in performance the animals were more attentive and motivated, leading to higher performance (Supplementary Fig. 1c–f). For the rest of the error trials, we found that that the animals were most likely guessing where to go, since they had similar likelihood of going to any of the other three erroneous targets (Supplementary Analysis 1., Supplementary Fig. 1g). For subsequent neural data analysis we only included rewarded trials (i.e., when the animals responded faster than 500 ms and fixated on the correct target for longer than 200 ms).
A Task performed by monkeys. After a 0.5 s fixation period, the Item 1 period always contained a target (Target 1, red square), presented for 0.4 s (Monkey A) or 0.3 s (Monkey B). Following a 1 s Delay 1, an Item 2 period occurred. In T/D trials, Item 2 was a distractor (green square), whereas in T/T trials, it was a new target (Target 2, red square). Item 2 was displayed for 0.4 s (Monkey A) or 0.3 s (Monkey B). After a 1 s Delay 2, the disappearance of the fixation spot served as a go cue. To receive a reward (drop of fruit juice), monkeys had to saccade to the last presented target location (Target 1 in T/D trials, Target 2 in T/T trials). B Electrode implantation sites. We chronically implanted 64 electrodes in the lateral prefrontal cortex (LPFC) (red-highlighted area) of each monkey, along with either 32 (Monkey A) or 64 (Monkey B) electrodes in the prearcuate cortex (PAC) (purple). C Input layer activity of RNNs in example T/T (left) and T/D (right) trials. The RNN input layer consisted of 9 units (4 locations × 2 types + 1 fixation). Colors indicate item locations, and line styles indicate item types (solid: target, dashed: distractor). Display durations of Item 1 and Item 2 were fixed at 0.3 s across all RNNs. D Output activity of Retrieve at Recall (R@R) RNNs in example T/T and T/D trials. The RNN output layer comprised 4 units corresponding to the possible locations. The output units of the R@R RNNs remained uninformative until Item 2 appeared. For T/T trials, the output represented the Item 2 location during Delay 2. For T/D trials, the output corresponded to the Item 1 location. E Output activity of Rehearse and Update (R&U) RNNs in example T/T and T/D trials. The R&U RNNs exhibited informative output immediately after item presentation. In T/T trials, the output corresponded to the Item 1 location during Delay 1 and switched to the Item 2 location in Delay 2. In T/D trials, the output remained at the Item 1 location across both delays.
RNNs were trained until they reached >95% accuracy (R@R: M(SD) = 0.998(0.003); R&U: M(SD) = 0.998(0.013)). Of note, this >95% accuracy in the models is higher than the monkeys’ performance (60−80%), because monkeys’ errors are likely driven by factors that are not present in the models (i.e., external distractions or decreased motivation due to satiation). We presume that when the monkeys perform the task correctly, it is because they are correctly following the learned task rules, which would be comparable to a well-trained RNN model. Nevertheless, for comparison, we carried out all the analyses on networks that only reached ~70% of performance. These analyses showed that all the results were qualitatively similar to the well-trained networks (Supplementary Analysis 2, Supplementary Fig. 2).
The inputs to the RNNs could either be Target 1 followed by Target 2 (T/T, Fig. 1c, left), or Target 1 followed by Distractor (T/D, Fig. 1c, right). The R&U strategy was enforced by evaluating the output of the network during both Delays 1 and 2 (Fig. 1d). The R@R strategy was enforced by evaluating the output during Delay 2 only (Fig. 1e).
Full space decoding analysis
Overall, we recorded 89 neurons from LPFC and 129 neurons from PAC in two animals across 11 sessions. in areas 9/46 (LPFC) and 8a (PAC) (Fig. 1b). Out of the recorded neurons, 100% (129/129) of PAC cells and 94% (84/89) of LPFC cells were task responsive, and 100% (129/129) of PAC cells and 87% of LPFC cells were selective at some point during the task (from target onset to go-cue onset). Our main analyses were based on the pseudo population activities pooled from both animals (see Methods), but we also separately analysed individual animals in the real sessions (Supplementary Figs. 3−7). A summary of all statistical analyses can be found in Supplementary Tables 1−3.
To get a rough estimate of how information was coded in the models and cortical regions, we carried out a cross-temporal decoding analysis (Fig. 2). We applied principal component analysis (PCA) to denoise and reduce the dimensionality of data (see Methods), and we defined the space consisting of the first 15 PCs in each pseudo population and RNN, which counted for at least 90% of total variances, as the full space (ΣEVR PC1-15: R@R: M(SD) = 1.000(0.000), R&U: M(SD) = 1.000(0.000); LPFC: M(SD) = 0.908(0.005), PAC: M(SD) = 0.920(0.004)). We evaluated the stability of the code during late Delay 1 and 2 (i.e., the last 500 ms of each delay) (Fig. 2b). To do this, we calculated the ratio of the decoding accuracy for time-displaced decoders (i.e., off diagonal) and time-specific decoders (i.e., diagonal). A value of 1 indicates a stable code that does not change across time, while a value lower than 1 reflects a dynamic code. For each model and brain region, we compared the stability ratio distributions to corresponding baseline distributions created by label-shuffled data as well as between Delay 1 and Delay 2 (see “Methods”), with statistical significance and effect size (Hedge’s g, see Methods) measured. During Delay 2, both regions and both models showed stable codes (R@R: M(SD) = 0.999(0.003), p = 0.740; R&U: M(SD) = 0.998(0.006), p = 0.640; LPFC: M(SD) = 0.996(0.016), p = 0.730; PAC: M(SD) = 0.995(0.014), p = 0.710) (Fig. 2b, light colors). However, while for the LPFC, PAC and R&U RNNs the code during Delay 1 was equally as stable as during Delay 2, that of the R@R RNNs was more dynamic (Code Stability Ratio: R@R: MD = −0.092, p < 0.001, g = −1.863; R&U: MD = −0.001, p = 0.436, g = −0.114; LPFC: MD = −0.001, p = 0.550, g = −0.086; PAC: MD = 0.000, p = 0.766, g = 0.033) (Fig. 2b). To compare the similarities between RNN models and biological observations, we performed pairwise Kolmogorov−Smirnov (K−S) tests (D) and calculated the Kullback-Leibler (KL) divergence (DKL) as well as the probability of overlap (p) between both models and both cortical regions to quantify the divergence (D and DKL) or similarity (p) between the two cumulative distribution functions (smaller D implies more similarity). We found that both brain regions were closer to the R&U than the R@R model (R@R-LPFC: D = 0.870, DKL = 12.644, p = 0.109; R@R-PAC: D = 0.870, DKL = 12.859, p = 0.119; R&U-LPFC: D = 0.420, DKL = 1.108, p = 0.990; R&U-PAC: D = 0.380, DKL = 0.920, p = 0.990).
A Cross-temporal decodability. Mean decoding accuracy of Item 1 and Item 2 locations from the full state space of R@R (upper left) and R&U (upper right) RNNs, as well as LPFC (lower left) and PAC (lower right) populations. Item 1 (I1), Item 2 (I2), and Go cue onsets are indicated by dotted lines. White outlines indicate significantly above-baseline decodability (p < 0.05). B Stability ratio distributions. Distributions of target location stability during Delay 1 (dark colors) and Delay 2 (light colors) in R@R (yellow) and R&U (blue) RNNs and LPFC (green) and PAC (purple) populations. C Code morphing ratio distributions of Item 1 location code across delays in T/D trials, with significance levels compared to corresponding baseline distributions (gray) indicated. In B, C the mean and median values are marked by triangles and white lines, respectively. Significance of the stability ratio difference between Delay 1 and Delay 2 is indicated (***p < 0.001; **p < 0.01; *p < 0.05; n.s.: p > 0.05). Single monkey analyses can be found in Supplementary Fig. 3a−c and Supplementary Fig. 4a−c.
Second, in T/D trials, we assessed whether the code for the target location changed between Delays 1 and 2 (Fig. 2c). We calculated the degree of code morphing as logarithm of the ratio between the mean within-delay decodability and the cross-delay decodability (see Methods). A low morphing ratio indicates a time-invariant code19,20,21. Likewise, we compared the distributions against thelabel-shuffled baselines to test for statistical significance. Target code was generalizable between Delay 1 and 2 in the LPFC, PAC, and R&U RNNs, while it morphed significantly in the R@R RNNs (Code Morphing Ratio: R@R: M(SD) = 0.939(0.174), p = 0.010; R&U: M(SD) = 0.004(0.011), p = 0.812; LPFC: M(SD) = 0.022(0.022), p = 0.673; PAC: M(SD) = 0.050(0.026), p = 0.485). The pairwise K−S tests, KL divergence, and overlap on the code morphing ratio distributions also showed that both brain regions were closer to the R&U than the R@R model (R@R-LPFC: D = 1.000, DKL = 23.113, p = 0.010; R@R-PAC: D = 1.000, DKL = 23.087, p = 0.010; R&U-LPFC: D = 0.630, DKL = 1.230, p = 0.990; R&U-PAC: D = 0.880, DKL = 2.328, p = 0.257).
Geometry of item-encoding subspaces
Decoding is a valid, but coarse, way of assessing the geometry of the representations in neural networks. To get a more fine-grained assessment of the representational geometry, we analyzed the principal angles and alignment of the activity subspaces that encode target information in the two delay periods (see “Methods”). If two subspaces are identical, then we expect them to have an angle of zero and a perfect alignment, while if they are different (e.g., rotated), then we expect them to have either angles or misalignments larger than zero (Fig. 3a). We also analyzed the transferability of item codes between subspaces (the performance of decoders trained on one subspace and tested on the other), as we expected them to be transferable between identical subspaces, but not so between rotated subspaces. Again, the distributions of all measures were compared to corresponding baseline distributions for statistical significance test (see “Methods”).
A Schematic of 2D subspaces with varying geometric relationships. Left: subspaces that are coplanar, and with aligned representational configurations. Middle: subspaces that are aligned but non-coplanar. Right: subspaces that are coplanar but misaligned. Subspaces can also be non-coplanar and misaligned (not shown). B Distributions of coplanarity (cosine of principal angles) between I1D1 T/T and I2D2 T/T subspaces in R@R (yellow) and R&U (blue) RNNs, as well as LPFC (green) and PAC (purple) populations. C Distributions of subspace alignment (cosine of minimal rotational angles) between I1D1T/T and I2D2T/T subspaces, with significance levels compared to corresponding baseline distributions indicated. D Distributions of code transferability ratios between I1D1T/T and I2D2T/T subspaces, with significance levels compared to corresponding baseline distributions indicated. In B−D the mean and median values are marked by triangles and white lines. Significance relative to baseline distributions (gray) is indicated (***p < 0.001; **p < 0.01; *p < 0.05; n.s.: p > 0.05). Single monkey analyses can be found in Supplementary Figs. 3d and 4d.
In T/T trials, we estimated the activity subspace that encodes item 1 location during the last 500 ms of Delay 1 (I1D1 T/T) and the subspace that encodes item 2 location during the last 500 ms of Delay 2 (I2D2 T/T). We found that these subspaces in LPFC, PAC and R&U RNNs were coplanar (cosine of the principal angle was not significantly smaller than 1), while the R@R RNNs they were not coplanar (Fig. 3b) (Coplanarity: R@R: M(SD) = 0.446(0.471), p = 0.030; R&U: M(SD) = 0.804(0.493), p = 0.380; LPFC: M(SD) = 0.789(0.194), p = 1.000; PAC: M(SD) = 0.931(0.057), p = 1.000). Similarly, the subspaces in LPFC, PAC and R&U RNNs were aligned (cosine of misalignment angle was not significantly smaller than 1), while the R@R RNNs were not aligned (Fig. 3c) (Alignment: R@R: M(SD) = 0.839(0.400), p = 0.040; R&U: M(SD) = 0.940(0.253), p = 0.970; LPFC: M(SD) = 0.994(0.010), p = 1.000; PAC: M(SD) = 0.998(0.003), p = 1.000). Finally, we found the code transferability ratio between I1D1 T/T and I2D2 T/T was significantly lower than baseline in R@R RNNs, but not in R&U RNNs, LPFC, and PAC (Fig. 3d) (Code Transferability Ratio: R@R: M(SD) = 0.453(0.108), p < 0.001; R&U: M(SD) = 0.782(0.172), p = 0.200; LPFC: M(SD) = 0.889(0.179), p = 0.930; PAC: M(SD) = 0.816(0.149), p = 0.750). The pairwise K−S tests,KL divergence and overlap on the distributions of subspace coplanarity, subspace alignment and code transferability ratio also showed that both brain regions were closer to the R&U than the R@R model (Subspace coplanarity: R@R-LPFC: D = 0.470, DKL = 2.106, p = 0.663; R@R-PAC: D = 0.750, DKL = 14.058, p = 0.287; R&U-LPFC: D = 0.450, DKL = 1.556, p = 0.713; R&U-PAC: D = 0.250, DKL = 4.596, p = 0.802. Subspace alignment: R@R-LPFC: D = 0.520, DKL = 16.069, p = 0.515; R@R-PAC: D = 0.650, DKL = 20.878, p = 0.376; R&U-LPFC: D = 0.340, DKL = 11.028, p = 0.871; R&U-PAC: D = 0.170, DKL = 19.331, p = 0.822. Code transferability ratio: R@R-LPFC: D = 0.910, DKL = 2.940, p = 0.119; R@R-PAC: D = 0.860, DKL = 2.473, p = 0.188; R&U-LPFC: D = 0.260, DKL = 0.181, p = 0.871; R&U-PAC: D = 0.150, DKL = 0.052, p = 0.822). These results indicate that in T/T trials, the subspace that encodes Target 1 information during Delay 1 is equivalent to the subspace that encodes Target 2 information during Delay 2 in both brain regions, as well as in the R&U model, while in the R@R model they are different.
Neural trajectories in the “readout” subspace
The analyses above show that there is a common representational subspace in T/T trials during Delay 1 and Delay 2 for the PAC, LPF, and R&U models, but not for the R@R models. To determine whether the same subspace is also utilized during Delay 2 in T/D trials, we compared the same metrics (coplanarity, alignment, and code transferability) between the representational subspaces that encode the reported item information during Delay 2 in T/T and T/D trials. We found that both brain regions and both models had a common representational subspace during Delay 2 in both trial types (Supplementary Fig. 8). Thus, a single activity subspace encodes target location information during late Delay 2 in both regions and models, akin to the “readout” subspace reported by Panicello and Buschman (2021)18.
In Fig. 2, we used decoding analyses on the full activity space to show that the R&U model and both PAC and LPFC had similar code stability and code morphing properties, while the R@R model behaved differently. Decoding on projections of activity onto the readout subspace led to equivalent conclusions (Supplementary Fig. 9). To get a deeper understanding of the representational properties of the brain regions and models, we analyzed the trajectories of the average population activity projected onto the readout subspace (Fig. 4a). In particular, we focused on the amount of drift observed in the population trajectories between Delay 1 and Delay 2 (Fig. 4b and Supplementary Fig. 10). We found that the drift between delays in the R@R model was similar in T/T and T/D trials (their ratio was close to 1) (Fig. 4b). On the other hand, the drift between delays in the R&U model, PAC and LPFC was primarily present in T/T trials, and almost absent in T/D trials (their ratio was close to 0) (Fig. 4b) (Drift Ratio: R@R: M(SD) = 0.834(0.154), p = 1.000; R&U: M(SD) = 0.166(0.099), p < 0.001; LPFC: M(SD) = 0.296(0.094), p < 0.001; PAC: M(SD) = 0.183(0.053), p < 0.001). The pairwise K−S tests, KL divergence, and overlap on the drift ratio distributions also showed that both brain regions were closer to the R&U than the R@R model (R@R-LPFC: D = 0.980, DKL = 18.309, p = 0.030; R@R-PAC: D = 1.000, DKL = 22.705, p = 0.010; R&U-LPFC: D = 0.600, DKL = 0.930, p = 0.644; R&U-PAC: D = 0.280, DKL = 0.861, p = 0.683). These observations confirm that the R@R model employs different representational subspaces during Delay 1 and Delay 2, consistent with an initial representation in a subspace (unattended format) during Delay 1 that is rotated to a “readout” subspace (attended format) during Delay 218, while the R&U model employs the same subspace in both delays, consistent with the direct loading onto the “readout” subspace during Delay 1. Both prefrontal regions, PAC and LPFC, show a consistent alignment with the R&U model and consistent differences with the R@R model. Thus, our data strongly support the conclusion that the monkeys employed the R&U strategy to solve the task.
A Population trajectories projected onto the readout subspace. Trajectories of state projections from the end of Delay 1 (circles) to the end of Delay 2 (stars)(green: [I1, I2] = [Loc 2, Loc 4]; orange: [I1, I2] = [Loc 4, Loc 2]) in T/T (solid lines, solid dots) and T/D (dashed lines, hollow dots) trials. Targets in the right hemifield (locations 2 and 4) were selected because they are contralateral to the electrode implantation site - left hemisphere. For the same plots using all 4 locations, see Supplementary Fig. 10. Data are shown for example, RNNs and neural populations. The approximate representational neighbourhood for the different locations during Delay 2 is highlighted with the large translucent ellipses, for reference only (green: Loc 4; orange: Loc 2). For the R@R network, we highlight the region that contains the projections of all target locations during Delay 1 (gray ellipse). B Distributions of drift ratios for the location pairs in R@R (yellow) and R&U (blue) RNNs, as well as LPFC (green) and PAC (purple) populations. Mean and median values are marked by triangles and white lines. Significance relative to corresponding baseline distributions (gray) is indicated (***p < 0.001; **p < 0.01; *p < 0.05; n.s.: p > 0.05). Single monkey analyses can be found in Supplementary Figs. 3e and 4e.
Discussion
Certain tasks can be solved employing different strategies that lead to identical responses to stimuli, in which case the strategy is latent. While previous studies have used behavioral and neural measurements to infer observable strategies4,5,15,16,17,22, to date, we have not been able to infer latent strategies. Since, by definition, latent strategies cannot be discerned based on behavior, instead we need to rely on differences in neural signals across strategies. In this study, we inferred the latent strategy employed by monkeys by comparing the geometry of their neural representations with the geometry of the representation of RNNs trained to solve the same task employing two different strategies (Supplementary Fig. 12).
The PAC is known for playing a role in motor preparation and execution23 rather than working memory maintenance24. On the other hand, the LPFC plays a role in working memory maintenance18,25. Target information replacement occurs earlier in PAC than in LPFC14, suggesting that while the animals may be able to report the updated location very quickly if necessary, updating working memory could take longer. In line with this prediction, behavioral observations in humans have shown that responses in tasks that require updating a single item were usually very fast and automatic26, while the explicit retrieval of target and removal of unrelated memory triggered by a retro-cue tends to take longer27.
Despite the different functions and timing of signals in LPFC and PAC, they shared qualitative similarities in all our metrics. Both regions exhibited sustained stable activities during the late delay periods, and target information was encoded and updated on a “readout” space throughout the trial in both regions. It is important to highlight that when we use the term “strategy,” we are referring to the animal’s approach to solving the task, rather than a mechanism specific to a single brain region. We assume that brain regions coordinate to implement this animal-level strategy. Therefore, while individual brain regions may have distinct functional roles within the organism, their activity should collectively align with the strategy the animal is employing. Thus, the objective of testing the PAC and LPFC separately is not to suggest any neural redundancy in the PFC. Rather, we argue that the activity observed in both regions, despite their differences14, is more consistent with the animal employing the rehearse and update strategy, rather than the retrieve at recall strategy.
Human studies have shown that the running memory span task can be solved using different strategies. In this task, a list of elements is presented in succession for an unpredictable number of items, after which the subjects are asked to recall the last n (~10) items presented28. One strategy that can be employed involves the unattended maintenance of the items, and, when the list ends, attempting to recall as many items as possible from this unattended storage16. We refer to this strategy as the Retrieve at Recall (R@R) strategy. Panichello and colleagues (2021) used neural data analysis to show that a similar strategy can be used to solve a retro-cue task18. The authors trained monkeys to retrospectively select one item from a set of items held in short-term memory. They showed that prior to attentional selection, memory items were represented in independent subspaces of neural activity in prefrontal cortex, but after selection, these representations were transformed to a “readout” subspace, which was used to guide behavior18. Furthermore, Piwek and colleagues (2023) found that this neural transformation emerged naturally in RNNs trained to perform the same task29. Our memory updating task can also be solved using this strategy, as shown here using RNNs (Fig. 1d).
Another strategy that can be employed to solve the running memory span task involves keeping track of the last few items, by dropping older ones and adding new ones in an updating process17. We refer to this strategy as the Rehearse and Update (R&U) strategy. Our task can be solved using this strategy, as shown here using RNNs (Fig. 1e). The prefrontal cortex plays an important role in working memory rehearsal25,30, and updating14,31,32,33. Here, we showed that monkey prefrontal activity is consistent with the R&U strategy, and inconsistent with the R@R strategy. Thus, using neural signals, we were able to infer the latent strategy employed by the monkeys to perform the task. However, this should not be interpreted as a claim that the R&U strategy is the only strategy that monkeys can use, since humans have been shown to employ either strategy under different task conditions22. Furthermore, while it has been suggested that monkeys and humans employ different strategies to solve working memory tasks3, this discrepancy may be explained by different stages of learning, rather than differences in strategies in both species7.
Several previous studies have characterized the neural mechanisms implementing working memory maintenance, including stable coding supported by sustained activity11, activity-silent mechanisms supported by short-term plasticity34,35, and dynamic coding36,37. Furthermore, these memories can be encoded in different formats, depending on whether they are attended to or not18,29. The strategies considered in this study (R@R and R&U) may be implemented by one or a combination of these mechanisms. However, it is not our goal to study the specific neural implementation of working memory per se. Rather, our goal is to compare the representational geometries of RNNs and brain regions to infer which strategy was being used by the monkeys. Relatedly, our study does not test the activity-silent hypothesis. The R@R RNNs solve the task without any short-term plasticity, and information about Target 1 location is maintained during Delay 1 in the activity of the neurons (Fig. 2A). The precise format of this Delay 1 activity is not controlled for: in some R@R RNN simulations the code was dynamic, and in some it was stable (see the variability of Delay 1 code stability in Fig. 2B).
Although this is an important step in understanding how strategies are implemented, our findings do not explain why animals prefer one strategy over another. To address this, we would need to investigate the learning process that leads to the selection of a specific strategy, which may itself involve latent factors38,39. Furthermore, when organisms learn that multiple strategies can be used to effectively solve a task, we must explore the decision-making process, both conscious and unconscious40, that determines why one strategy is chosen over the others41.
Methods
Subjects and surgical procedures
Two adult male macaques (Macaca fascicularis) were used in this experiment: Monkey A (age 12) and Monkey B (age 12). All animal procedures were approved by, and conducted in compliance with, the standard of the National University of Singapore Institutional Animal Care and Use Committee (NUS IACUC #R18-0295). Procedures also conformed to the recommendations described in Guidelines for the Care and Use of Mammals in Neuroscience and Behavioral Research (National Academies Press, 2003). Each animal was first implanted with a titanium head-post (Crist Instruments, MD, USA) before arrays of intracortical microelectrodes (MicroProbes, MD, USA) were implanted in multiple regions of the left frontal cortex. In Monkey A, two arrays were placed over the dorsal and ventral aspect of the LPFC (Area 9/46), one array was placed over PAC (Area 8A), and one array was placed over the pMA (not included in the analyses here), with 32 electrodes in each. In Monkey B, two arrays of 32 electrodes were placed over the PAC, and two arrays of 32 electrodes each were placed over the LPFC (as shown in Fig. 1b). The arrays consisted of platinum-iridium wires with 200 µm separation, 1−5.5 mm long and with 0.5 MΩ of impedance, arranged in 8 × 4 grids. For arrays positioned in the pre-arcuate region (PAC), we used standard electrical microstimulation to confirm that saccades could be elicited with low currents in monkey A (however, we could not perform this procedure in monkey B).
Recording techniques
The neural signals in both monkeys were recorded using a 128-channel Grapevine recording system (Ripple Neuro, UT, USA) at 30 kHz sampling rate. The wideband signals were bandpass-filtered between 300 and 3000 Hz, and spikes were detected on each channel separately using an automated sorting algorithm based on Hidden Markov modelling42. We recorded the eye positions of each subject using the Eyelink 2000 (SR Research Ltd, ON, CA) on another standalone computer. We designed and ran the behavioral tasks using the Psychopy in Python43 on a third computer connected to the recording computer using parallel ports for event mark synchronization during recording.
Behavioral tasks
Both monkeys performed a 2-item delayed saccade task. Each trial began with a 0.5 s fixation period, during which the monkey fixated on a white central dot. A first stimulus (Item 1, I1) was then presented at one of four possible locations on the corners of a 3 × 3 grid (10° visual angle) for a fixed duration (Monkey A: 0.4 s; Monkey B: 0.3 s), followed by a 1 s delay (D1) where the monkey maintained fixation. A second stimulus (Item 2, I2) was then presented at one of the three remaining locations for the same duration. I1 was always a task-relevant target (red square), while I2 was either a task-irrelevant distractor (T/D) or a new target (T/T), with equal probability. The target (red) and distractor (green) were identical in shape and size. After a second 1 s delay (D2), the fixation dot disappeared, signalling the monkey to make a saccadic response to the location of the most recent target (I1 in T/D trials; I2 in T/T trials). In other words, the animal needed to make an eye movement to the location of I1 when I2 was a distractor, or to the location of I2 when it was a new target. A juice reward was delivered for correct responses within 0.5 s after the go cue. I1 and I2 never appeared at the same location within a trial, yielding 24 trial conditions based on I1 × I2 locations (4 × 3) and I2 types (2).
Behavioral performance analysis
The behavioural accuracy was first calculated separately in each real session as the proportion of correct trials of each type among all trials of that type, and then averaged across all sessions. For example, the accuracy of T/T trials in session i is
where \({N}_{i}^{T/T,{correct}}\) and \({N}_{i}^{T/T}\) represent the number of correct and total T/T trials in session i, respectively, and the mean accuracy across n sessions is
To measure the accuracy as a function of session stages, we calculated the accuracy over bins of 100 trials and slide at steps of 20 trials from the beginning of each session to the end. For the conservative calculation of accuracy, we counted only the rewarded trials as correct and the remaining to be incorrect. For the liberal calculation of accuracy, we included both the rewarded trials and trials with correct responded locations but failed to either 1. maintain fixation at the location for at least 200 ms or 2. produce saccadic response within 500 ms post-go-cue.
Neuron firing rates and preprocessing
We recorded a total of 89 neurons in LPFC (Monkey A: 17, Monkey B: 72) and 129 neurons in PAC (Monkey A: 3, Monkey B: 126) across 11 sessions (Monkey A: 3, Monkey B: 8). The number of LPFC cells in each session ranged between 3 and 14 (M(SD) = 8.091(3.145)), and the number of PAC cells in each session ranged between 0 to 21 (M(SD) = 11.727(7.669)). All cells were used to build pseudo populations. Single-neuron firing rates were computed using a 50 ms sliding window (10 ms overlap). Firing rates were z-scored relative to the mean and standard deviation of pre-I1 baseline activity (0.2 s) across all trials20. Since I1 and I2 display durations differed between Monkey A (0.4 s) and Monkey B (0.3 s), we removed the middle 0.1 s of each delay period in Monkey A for pooled analyses to ensure temporal alignment.
A cell was considered responsive if it had firing rates significantly different from baseline (before the first stimulus onset) in one or more 50 ms time bins (10 ms overlap) at any point during the trial (from target onset to go-cue onset). Firing rates were time-averaged within each bin, and statistical significance was tested using 1-sample t-test with a statistical significance threshold of p < 0.001 with two-sided alternative hypotheses. Using the same method, we visualized the proportion of responsive neurons among all recorded neurons as a function of time in each session of each animal in the extended analysis (see Supplementary Figs. 6a and 7a).
We also analysed the selectivity of single neurons to the location of I1 and I2. We averaged neuron activities into 50 ms time bins for each trial and performed one-way ANOVA on the I1 and I2 locations at each bin. Neurons with p < 0.001 were considered location selective at the bin. In the extended analysis (Supplementary Figs. 6b and 7b), we visualized the proportion of I1 and I2 selective neurons as a function of time in each session of each animal.
Percentage of Explained Variance (PEV) analysis
To estimate how much information of the task variables (i.e., I1 and I2 locations) was coded by the activity of each single neuron, we calculated the percentage of explained variance (PEV, or ω2). ω2 is defined by
where df is the degree of freedom, and
We visualized the PEV of all single neurons as well as the averaged PEV from neurons recorded in each session in the Supplementary Figs. 6c, d and 7c, d.
Pseudo population
We created pseudo populations by sampling equal numbers of trials per condition from each session of both monkeys and pooling these trials together, as if these neurons responded together in each pseudo trial. We created 100 pseudo populations separately for the LPFC (89 neurons) and the PAC (129 neurons). The selectivity of the neurons was as reported in our previous work14. Only correct trials were included in the pseudo population. All neural data analyses were conducted on the pseudo populations.
Models
We trained artificial recurrent neural networks (RNNs) to simulate neural activity under different latent strategies in trials of 60 time steps (I1/I2 display = 6 time steps, D1/D2 = 20 time steps, post-go-cue = 8 time steps). Each RNN comprised 9 input units (4 locations × 2 types + 1 fixation), 64 recurrent units, and 4 output units (4 locations). The recurrent dynamics followed:
And the output dynamics can be described as:
where It, rt and Yt denote the input, recurrent, and output unit activities at time t, and Win, Wrec, Wout represent the input, recurrent and output weights respectively. σ represents the noise at the recurrent layer and was drawn from N(0, 0.1). Recurrent weights were orthogonally initialized, while input and output weights followed Xavier-normal initialization. We optimized Win, Wrec, Wout using stochastic gradient descent with NAdam. The loss was computed as the mean squared error (MSE) of the outputs over the target time window:
For Retrieve at Recall (R@R) RNNs, during Delay 1 all output units were expected to have equal activity (0.25), and during Delay 2 the output of the choice item (I1 in T/D trials, I2 in T/T trials) was expected to be 1, while the non-choice items remained at 0. For Rehearse and Update (R&U) RNNs, the output corresponded to I1’s location during Delay 1 and switched to the choice item’s location during Delay 2. 100 RNNs were trained per strategy, each with different weight initializations.
Principal Component Analysis (PCA)
We applied principal component analysis (PCA) to each pseudo-population to reduce dimensionality before decoding, using the scikit-learn library44. To preserve variance across conditions, we computed mean firing rates per condition, averaging over Delay 1 and Delay 2. The resulting activity matrix X (Nconditions (24) × Nneurons) was used to fit a PCA model, which was then applied to the full time series for dimensionality reduction. We defined the full space of the transformed data as the projections on the first 15 PCs, which counted for at least 90% of the total explained variance (EVR) on average in both regions. The same procedure was applied to the hidden unit activities of task-optimized RNNs.
Decoding Analysis
Cross-temporal Decodability
To measure the decodable stimulus location information across the population of recorded neurons, we used linear discriminant analysis (LDA) based on the algorithm from scikit-learn44 to predict the locations of Items 1 and 2 (Fig. 2 and Supplementary Figs. 2a–4a, 5b, 6e–7e, 9a). Decoding analyses were conducted on neural data from 0.2 s before Item 1 onset to the go cue. A 50 ms non-overlapping smoothing window was applied before training and testing decoders. PCA-based dimensionality reduction was applied before decoding, and analyses were performed on projected activities. The same approach was used for RNN hidden unit activities. For full-space decodability (Fig. 2), we used projections on the top 15 PCs to train and test decoders. For readout subspace decodability (Supplementary Fig. 9a), we used state projections on the readout subspace.
Within each pseudo population and RNN, we randomly sampled half of the trials for training and the other half for testing, iterating 100 times. At each iteration, a decoder was trained using training set activities at one time bin to predict item locations and tested on the test set activities across all time bins. Decoding performance was averaged over 100 iterations to minimize variability from set-splitting. We tested decodability significance at each time bin using a 95th-percentile test (see Statistical Tests), comparing decoder accuracy to baseline distributions generated from data with shuffled location labels (shuffled 100 times per pseudo-population and RNN). We repeated this process until all time bins were trained and tested.
Code stability and code morphing
To quantify information coding stability, we computed the stability ratio, defined as the mean within-time decodability (trained and tested at the same time bin) divided by the mean cross-time decodability (trained and tested at different time bins) across selected time windows21. More specifically, cross-time decodability was calculated at every possible pair of train-test bins that were different from each other, in other words, the off-diagonal bins in cross-temporal decoding plot, while the within-time decodability was the on-diagonal bins, where decoders were trained and tested at the same bin. These decoder performances were averaged across bins and trial type conditions for stability ratio calculation. We calculated this ratio separately for late Delay 1 (last 0.5 s) and late Delay 2, followed by statistical comparisons (see Statistical Tests).
The population code of the target may change after the presentation of a distracting stimulus (i.e., code morphing)19. Here, we quantified the code morphing ratio in the T/D trials as the logarithm of the ratio of the mean cross-temporal decodability trained and tested in late Delay 1 over the decodability trained in late Delay 1 and tested in late Delay 2 (and vice versa). A low code morphing ratio indicates a time-invariant code. Baseline distributions were created by decoders trained with label-shuffled data, iterated 100 times, and averaged within each pseudo-population and RNN.
Subspace analysis
Subspace estimation
Low-dimensional geometries can be estimated from the population based on various methods18,20,29,44. We estimated low-dimensional subspaces using methods adapted from Panichello and Buschman (2021)18 and Piwek et al. (2023)29 to obtain 2D planes best fitting the representations of Item 1 and Item 2 during early (first 0.5 s of) and late (last 0.5 s of) Delay 1 and Delay 2. For each pseudo population, we first grouped the trials by conditions (Nconditions = Nlocation combinations (12) × Ntype (2)). We then averaged the neural population states by conditions and projected the Nconditions (24) × Nneurons data to the axes of the top three principal components (PCs). On average, the sum of explained variances by these three PCs reached about 50% of total variances in LPFC and over 60% in PAC as well as in R@R and R&U RNNs (ΣEVRPC1-3: R@R: M(SD) = 0.995(0.010), R&U: M(SD) = 0.994(0.007); LPFC: M(SD) = 0.499(0.016), PAC: M(SD) = 0.611(0.013)).
Within each time window, we first averaged the activities across all timepoints, returning us an Nconditions (24) × NPCs (3) matrix. For item-specific subspaces (Fig. 3 and Supplementary Fig. 8), we separated the matrix by the type condition (T/T and T/D) for analysis clarity, each with a 12 × 3 matrix. Within each type, we estimated the mean representation by averaging the rows corresponding to conditions with the same location of each item: for example, the representation of I1 presented at location 1 was calculated as the mean of all conditions with I1-location = 1, regardless of the location of I2. This returned us an Nlocations (4) × NPCs (3) matrix for each item. Next, we conducted a secondary PCA on this matrix and extracted the first two PCs. These two eigenvectors corresponded to the directions of maximum variances by locations, hence were used to define the best-fitting 2D plane of the locational representation. Finally, we projected the states from the 3-PC space onto these 2D planes and used these projections for subsequent analyses.
The “readout” subspaces (Fig. 4) were estimated based on the same approach, except that we did not separate between trial types, and the mean representation was calculated by averaging the rows with the same location of the final choice item (i.e., I1 for T/D and I2 for T/T). Also, instead of estimating at separate time windows, we only used activities during late D2 to estimate the “readout” axes, as stable outputs were observed during this period in both models and brain regions (as shown in Fig. 2).
Geometry analysis
With the subspace estimated using the approach described above, we then examined the geometric relationships between subspace pairs, specifically the coplanarity and the degree of alignment of the representational configurations between subspaces. Metrics were calculated using methods adapted from Panichello and Buschman (2021)18 and Piwek et al. (2023)29. For coplanarity, we measured the cosine of the principal angle between the normal vectors of the two planes. We applied a correction to the directions of the plane-defining vectors based on the representational configurations to guarantee vectors from two different subspaces were under a common frame of reference, as described below29. This was because the original directions of PC axes generated by the PCA algorithm are ambiguous with respect to their sign, which would result in inconsistent signs of the normal vectors of the 2D planes and interfere with the principal angle calculations. To avoid this problem, we corrected the directions of the eigenvectors by first rotating PC1 vector to be positively collinear (i.e., parallel and in the same direction) to a predefined directional side of the location configurations, creating a PC1corrected, and then rotating PC2 vector by the shortest angular distance to be orthogonal to PC1corrected. We then used PC1corrected and PC2corrected as the plane-defining vectors as well as for normal vector calculations. Planes with mirror-reflected configurations of one another would lead to angles greater than 90° (cos(θ) <0), whereas the planes of rigid rotation of each other would have principal angles restricted smaller than 90° (cos(θ) > 0).
For alignment, we measured the cosine of the minimal rotation angle, minimizing the distances between corresponding representational points on the two subspaces. To calculate the rotation angle, we first forced the two planes to be coplanar. Next, we estimated the representational configuration from each plane by averaging the states by item locations and then projected the configuration points on corresponding planes, giving us two Nlocations (4) × 2 configuration matrices. We then applied an orthogonal Procrustes analysis on these two matrices, which returned an orthogonal matrix R that could rotate one matrix to most closely match the other, with the R(0,0) as the cosine of the minimal rotation angle.
For both coplanarity and alignment, the baseline hypothesis was the full equivalence between subspaces; in other words, we tested whether the two subspaces were significantly non-coplanar and misaligned. To create the baseline distributions, we first split the data into a training set and a test set and separately estimated subspaces. We then calculated the coplanarity and alignment between the training and test set subspaces at each item and time bin of interest, and finally took the mean of them. The rationale was that a given subspace is perfectly coplanar and aligned to itself, and subspaces estimated from different subsets of the same pseudo-session or recurrent neural network (RNN) should be highly similar, with some variability due to differences between training and test sets. For example, for the comparison between I1D1 and I2D2, we averaged coplanarity and alignment between I1D1train vs. I1D1test and I2D2train vs. I2D2test. For each pseudo population and RNN, we repeated this process 100 times and took their average. The resulting parallel distributions were then compared to observed values, and empirical p-values were calculated using the 95th-percentile test (see Statistical Tests), and a significant p value indicated consistent deviation from the equivalent hypothesis.
Code transferability ratio
In addition to the geometry analysis, we also examined the code transferability between subspaces. Specifically, we trained LDA decoders with the state projections on one subspace and tested with the projections on the other subspace. The codes were expected to be transferable if the two subspaces were equivalent. Given the noisy nature of neural data, the decoder performances in the LPFC and PAC were generally worse than in the RNNs. To make the quantifications comparable between neural data and RNNs, we calculated the ratio of cross-subspace decodability (i.e., code transferability) over the within-subspace decodability (i.e., trained and tested with the projection on the same subspace), and a high ratio (close to 1) indicates the decodability to be at a similar level cross- and within-subspace. The baseline distributions were created by training the cross- and within-subspace decoders on data with shuffled location labels, and for each pseudo population and RNN, we repeated the process 100 times and took the average.
Drift ratio
To quantify the dynamics of population activity projection onto the readout subspace, for each condition (I1-I2 Combination × Type) we calculated the Euclidean distance between the mean projection points during late (last 0.5 s of) Delay 1 and during late Delay 2 and scaled to the range from −1 to 1. We then calculated the drift ratio by dividing the drift distance in T/D trials by the drift distance in T/T trials. A low ratio would indicate the readout projections to be changed only when information updates were required. The baseline distributions were created by estimating the distances from data with shuffled location labels, and we repeated this process 100 times and took the average for each pseudo population and RNN.
Significance tests
For tests compared to baselines, we considered two distributions to be significantly different from baseline if the 95th percentile range of the test distribution did not overlap with the baseline distribution45,46. The p value was calculated as:
where X represents the number of overlapping points and N represents the total number of points in the baseline distribution.
For tests comparing between two test distributions, we pooled the data together and randomly re-assigned them to create two new distributions and calculated the mean difference. We iterated this process for 1000 times to create a null distribution. The p value was then calculated as the proportion of points in the null distribution that are larger or smaller (depending on the test direction of interest) than the true mean difference between the test distributions. We additionally calculated the effect size of the comparisons using the measure of Hedge’s g as:
where n and s refer to the number of data points and standard deviations of each distribution46,47.
Kolmogorov-Smirnov (K–S) test
To statistically compare the distributions between RNN models and biological regions, we applied Kolmogorov-Smirnov (K–S) tests on the primary quantification measures between all model-region pairs using Scipy47. The K–S test assesses whether two empirical distributions vary from each other based on their cumulative distribution functions, with a K–S statistic (D) indicating the distance between the two cumulative distribution functions (the smaller the closer) and a p-value (with two-sided alternative hypothesis) indicating the statistical significance.
Kullback–Leibler (KL) Divergence test
An alternative index we used to compare the similarities between the RNN models and biological regions is the KL divergence, which quantifies the dissimilarity between a model-generated probability distribution function and the empirical observation. It is a common measure of the loss of a model in machine learning. Greater KL divergence indicates more differences between the two distributions. Specifically, the KL divergence (DKL) of a probability distribution Q from a reference probability distribution P is calculated as
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are available at https://doi.org/10.6084/m9.figshare.28530998.v1 and https://doi.org/10.6084/m9.figshare.28530935.v1. Source data are provided with this paper.
Code availability
A code package for performing the cross-temporal decoding is available at https://github.com/YQian2333/Inferring-WM-Strategies.
References
Mante, V., Sussillo, D., Shenoy, K. V. & Newsome, W. T. Context-dependent computation by recurrent dynamics in prefrontal cortex. Nature 503, 78–84 (2013).
Purcell, B. A. & Kiani, R. Hierarchical decision processes that operate over distinct timescales underlie choice and changes in strategy. Proc. Natl. Acad. Sci. USA 113, E4531–E4540 (2016).
Wittig, J. H., Morgan, B., Masseau, E. & Richmond, B. J. Humans and monkeys use different strategies to solve the same short-term memory tasks. Learn. Mem. 23, 644–647 (2016).
Fascianelli, V. et al. Neural representational geometries reflect behavioral differences in monkeys and recurrent neural networks. Nat. Commun. 15, 6479 (2024).
Genovesio, A., Brasted, P. J., Mitz, A. R. & Wise, S. P. Prefrontal cortex activity related to abstract response strategies. Neuron 47, 307–320 (2005).
Venkatraman, V., Payne, J. W., Bettman, J. R., Luce, M. F. & Huettel, S. A. Separate neural mechanisms underlie choices and strategic preferences in risky decision making. Neuron 62, 593–602 (2009).
Tsuda, B., Richmond, B. J. & Sejnowski, T. J. Exploring strategy differences between humans and monkeys with recurrent neural networks. PLoS Comput. Biol. 19, e1011618 (2023).
Ji-An, L., Benna, M. K. & Mattar, M. G. Discovering cognitive strategies with tiny recurrent neural networks. Nature 644, 993–1001 (2025).
Daw, N. D., Gershman, S. J., Seymour, B., Dayan, P. & Dolan, R. J. Model-based influences on humans’ choices and striatal prediction errors. Neuron 69, 1204–1215 (2011).
Trübutschek, D. et al. A theory of working memory without consciousness or sustained activity. eLife 6, e23871 (2017).
Wimmer, K., Nykamp, D. Q., Constantinidis, C. & Compte, A. Bump attractor dynamics in prefrontal cortex explains behavioral precision in spatial working memory. Nat. Neurosci. 17, 431–439 (2014).
Churchland, A. K. et al. Variance as a signature of neural computations during decision making. Neuron 69, 818–831 (2011).
Stine, G. M., Zylberberg, A., Ditterich, J. & Shadlen, M. N. Differentiating between integration and non-integration strategies in perceptual decision making. eLife 9, e55365 (2020).
Qian, Y., Herikstad, R. & Libedinsky, C. Working memory updating in the macaque lateral prefrontal cortex. J. Neurosci. 45, e1770242024 (2025).
Hockey, R. Rate of presentation in running memory and direct manipulation of input-processing strategies. Q. J. Exp. Psychol. 25, 104–111 (1973).
Cowan, N. et al. On the capacity of attention: its estimation and its role in working memory and cognitive aptitudes. Cognit. Psychol. 51, 42–100 (2005).
Postle, B. R. Context in verbal short-term memory. Mem. Cognit. 31, 1198–1207 (2003).
Panichello, M. F. & Buschman, T. J. Shared mechanisms underlie the control of working memory and attention. Nature 4, 601–605 (2021).
Parthasarathy, A. et al. Mixed selectivity morphs population codes in prefrontal cortex. Nat. Neurosci. 20, 1770–1779 (2017).
Parthasarathy, A. et al. Time-invariant working memory representations in the presence of code-morphing in the lateral prefrontal cortex. Nat. Commun. 10, 1–11 (2019).
Tan, P. K., Tang, C., Herikstad, R., Pillay, A. & Libedinsky, C. Distinct lateral prefrontal regions are organized in an anterior-posterior functional gradient. J. Neurosci. Off. J. Soc. Neurosci. 43, 6564–6572 (2023).
Bunting, M., Cowan, N. & Scott Saults, J. How does running memory span work? Q. J. Exp. Psychol. 59, 1691–1700 (2006).
Bruce, C. J. & Goldberg, M. E. Primate frontal eye fields. I. Single neurons discharging before saccades. J. Neurophysiol. 53, 603–635 (1985).
Jonikaitis, D., Noudoost, B., & Moore, T. Dissociating the contributions of frontal eye field activity to spatial working memory and motor preparation. J. Neurosci. 43, 8681–8689 (2023).
Funahashi, S., Bruce, C. J. & Goldman-Rakic, P. S. Mnemonic coding of visual space in the monkey’s dorsolateral prefrontal cortex. J. Neurophysiol. 61, 331–349 (1989).
Kessler, Y., Zilberman, N. & Kvitelashvili, S. Updating, fast and slow: items, but not item-context bindings, are quickly updated into working memory as part of response selection. J. Cogn. 6, 11 (2023).
Tortajada, M. et al. Decoding load or selection in visuospatial working memory? Psychophysiology 61, e14636 (2024).
Pollack, I., Johnson, L. B. & Knaff, P. R. Running memory span. J. Exp. Psychol. 57, 137–146 (1959).
Piwek, E. P., Stokes, M. G. & Summerfield, C. A recurrent neural network model of prefrontal brain activity during a working memory task. PLoS Comput. Biol. 19, e1011555 (2023).
Miller, E. K., Erickson, C. A. & Desimone, R. Neural mechanisms of visual working memory in prefrontal cortex of the macaque. J Neurosci 16, 5154–5167 (1996).
D’Ardenne, K. et al. Role of prefrontal cortex and the midbrain dopamine system in working memory updating. Proc. Natl. Acad. Sci. USA 109, 19900–19909 (2012).
Kim, H., Smolker, H. R., Smith, L. L., Banich, M. T. & Lewis-Peacock, J. A. Changes to information in working memory depend on distinct removal operations. Nat. Commun. 11, 6239 (2020).
Nir-Cohen, G., Kessler, Y. & Egner, T. Neural substrates of working memory updating. J. Cogn. Neurosci. 32, 2285–2302 (2020).
Lundqvist, M. et al. Gamma and beta bursts underlie working memory. Neuron 90, 152–164 (2016).
Mongillo, G., Barak, O. & Tsodyks, M. Synaptic theory of working memory. Science 319, 1543–1546 (2008).
Meyers, E. M., Freedman, D. J., Kreiman, G., Miller, E. K. & Poggio, T. Dynamic population coding of category information in inferior temporal and prefrontal cortex. J. Neurophysiol. 100, 1407–1419 (2008).
Stokes, M. G. et al. Dynamic coding for cognitive control in prefrontal cortex. Neuron 78, 364–375 (2013).
Kuchibhotla, K. V. et al. Dissociating task acquisition from expression during learning reveals latent knowledge. Nat. Commun. 10, 2151 (2019).
Drieu, C. et al. Rapid emergence of latent knowledge in the sensory cortex drives learning. Nature 641, 960–970 (2025).
Cary, M. & Reder, L. M. Metacognition in strategy selection. in Metacognition: Process, Function and Use 63–77 (Kluwer Academic Publishers, 2002).
Wichary, S. & Smolen, T. Neural underpinnings of decision strategy selection: a review and a theoretical model. Front. Neurosci. 10, 500 (2016).
Peirce, J. et al. PsychoPy2: experiments in behavior made easy. Behav. Res. Methods 51, 195–203 (2019).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Tang, C., Herikstad, R., Parthasarathy, A., Libedinsky, C. & Yen, S. C. Minimally dependent activity subspaces for working memory and motor preparation in the lateral prefrontal cortex. eLife 9, 1–23 (2020).
Ojala, M. & Garriga, G. C. Permutation tests for studying classifier performance. In Proc. Ninth IEEE International Conference on Data Mining 908–913 https://doi.org/10.1109/ICDM.2009.108 (IEEE, 2009).
Hedges, L. V. Distribution theory for Glass’s estimator of effect size and related estimators. J. Educ. Stat. 6, 107–128 (1981).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Acknowledgements
This work was supported by a grant by the Ministry of Education of Singapore (MOE-T2EP30121-0010) to C.L. We would like to thank Clement Lim for his help with the animal work, and Shih-Cheng Yen for useful discussions.
Author information
Authors and Affiliations
Contributions
R.H. and C.L. conceived the project and designed the experiments. R.H. performed the experiments and recorded data. Y.Q. analysed the data and performed modelling. Y.Q., R.H., and C.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Qian, Y., Herikstad, R. & Libedinsky, C. Inferring latent behavioral strategy from the representational geometry of prefrontal cortex activity. Nat Commun 17, 2850 (2026). https://doi.org/10.1038/s41467-026-69380-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69380-6