Abstract
All-optical interrogation, based on high-resolution two-photon stimulation and imaging, has emerged as a potentially transformative approach in neuroscience, allowing for the simultaneous precise manipulation and monitoring of neuronal activity across various model organisms. However, the unintended excitation of light-gated ion channels such as channelrhodopsin (ChR) during two-photon calcium imaging with genetically encoded calcium indicators (GECIs) introduces artifactual neuronal perturbation and contaminates neural activity measurements. In this study, we propose an active pixel power control (APPC) approach, which dynamically adjusts the imaging laser power at each scanning pixel, to address the challenge. We aim to achieve simultaneous two-photon optogenetic manipulation and calcium imaging with a single femtosecond laser, while minimizing the cross-talk between manipulation and imaging. To study this technology’s capabilities, we applied it to the larval zebrafish brain in vivo. Our results demonstrate that the APPC approach preserves GECI signal quality while suppressing optogenetic artifacts significantly. This enhances the accuracy of neural circuit dissection and advances the precision of all-optical interrogation, offering a robust framework for probing neural circuit dynamics and causality in vivo with high fidelity, potentially across various model organisms. Importantly, this technology can be seamlessly integrated with commonly used two-photon microscope systems in laboratories worldwide.
Similar content being viewed by others
Introduction
All-optical interrogation of neuronal circuits represents a paradigm-shifting advancement in neuroscience, successfully applied to various model organisms, including Mus musculus (mice)1,2,3,4,5,6,7, Danio rerio (zebrafish)8,9, Drosophila melanogaster (fruit flies)10, and Caenorhabditis elegans (nematodes)11,12. This technique integrates genetically encoded calcium indicators (GECIs) with optogenetic actuators, such as channelrhodopsin (ChR), facilitating non-invasive, in vivo recording and manipulation of neuronal activity. Recent advancements in two-photon microscopy and holographic optogenetics enable neural ensemble recording and optogenetic control at single-cell resolution and sub-millisecond timescales2,5,6,7,13. Furthermore, closed-loop systems allow real-time extraction of neural activity to guide precise perturbations such as spike rates, timing, and synchrony3,14,15. These capabilities enable targeting of individual neurons based on functional signatures and advance mechanistic studies of circuit connectivity, plasticity, and disease-associated aberrant activity in conditions such as Alzheimer’s disease16,17 and Parkinson’s disease18,19. By elucidating neural circuit dynamics under targeted perturbation, this approach goes beyond merely correlating activity with behavior or cognitive states, establishing causal links; for example, it has been used to demonstrate how emotional states are encoded in neural attractor networks6,20,21,22,23,24. However, a major challenge emerges when a two-photon light source serves dual roles as both reader and stimulator in all-optical interrogation systems: cross-talk between these modules compromises experimental precision, and must be effectively minimized2,5,13,25.
Generally, there are three primary mechanisms underlying the cross-talk between calcium imaging and optogenetic stimulation5,25,26. Type I cross-talk refers to the contamination of calcium signals due to additional GECI signals when the optogenetic stimulation laser excites the GECI. Type II cross-talk involves unwanted excitation of ChR by the imaging laser. Lastly, Type III cross-talk occurs when densely packed neurons, with both somas and dendrites expressing ChR, lead to off-target activation during optogenetic stimulation. Strategies such as image-processing algorithms and time-multiplexed excitation to reduce calcium signal distortion9,27,28 in Type I cross-talk and soma-localized opsin variants to restrict ChR expression to soma membranes5,25 to mitigate Type III cross-talk, but Type II cross-talk remains unresolved. The Type II cross-talk issue is less pronounced in single-photon systems due to GCaMP’s superior light absorption, compared to those of ChRs29,30,31. However, in two-photon systems, cross-talk is exacerbated: ChR2, the most commonly used light gated ion channel protein-exhibits a 10-fold higher two-photon absorption cross-section, which linearly related to the absorption efficiency, than GCaMP32,33, the most widely used GECI. This indicates that ChR2 has tens of times stronger absorption and enables ChR2 to be activated at <10% of the laser power required for GCaMP imaging34.
Efforts to reduce Type II cross-talk via spectral separation by using red-shifted GECIs32,35,36,37,38,39,40 or ChRs6,41,42,43 have achieved substantial achievements. While pairing red GECIs with blue ChRs at 1100 nm can eliminate cross-talk25,44, current red indicators suffer from limited dynamic range and signal-to-noise ratios32,35,36,37,38,39, limiting their broader application. Red-shifted ChRs shift their absorption spectra toward longer wavelengths, reducing activation by imaging light. For example, ChrimsonR exhibits peak two-photon excitation efficiency at 1140 nm, compared to 940 nm for ChR25,45. Combining GCaMP with red-shifted ChRs has been shown to suppress crosstalk2,4,5,7,46. However, imaging conditions must be carefully optimized, as spectral overlap can occur due to broadened absorption profiles of red ChRs towards shorter wavelength. This broadening may result from the complexity of local environment for chromophore, and several distinguish mechanisms in the two-photon excitation such as the enhancement of certain vibronic transitions47,48,49,50,51,52,53. Consequently, Type II cross-talk persists even with the use of current red ChRs.
To address the Type II cross-talk, we developed a dynamic imaging power control strategy that actively and precisely modulates excitation power at each scanning pixel using a fast acousto-optic modulator (AOM). By reducing imaging power at designated pixels, we minimized cross-talk while preserving calcium signal extraction efficiency. Our method utilized ChR expression maps generated by standard fluorescent labeling to create an “Active Pixel Power Control” (APPC) scheme that selectively attenuates or blocks laser intensity at ChR+ pixels during two-photon calcium imaging. Specifically, synchronized with resonant scanning, the AOM dynamically adjusted power in real time, minimizing unintended ChR activation while maintaining calcium signal fidelity through spatial summation of pixel fluorescence intensities.
We validated this approach in the larval zebrafish, a vertebrate model ideal for whole-brain imaging and optogenetics, using a single femtosecond laser to combine two-photon calcium imaging (via GCaMP) with holographic optogenetic stimulation via spatial light modulator (SLM). Calcium signals were deconvolved to estimate spike rates3,54, enabling cross-talk quantification through brain-wide activity patterns. The APPC technology achieved nearly cross-talk-free all-optical interrogation without relying on protein engineering or spectral separation, while enabling precise, high-speed in vivo calcium imaging. By controlling laser power at single-pixel resolution, this approach preserved signal quality while offering a scalable and practical solution for circuit-level studies in behaving animals. This advance opens new avenues for probing causal relationships in brain wide dynamics with higher accuracy and creates opportunities for related research in laboratories equipped with standard single laser two photon microscopes.
Results
All-optical interrogation system with precise pixel-specific excitation power control
To achieve active pixel power control (APPC), which provides dynamic, pixel-specific control of imaging laser power based on ChR expression patterns to minimize cross-talk, we developed a two-photon microscope system with a single excitation source and a custom beam control module (Fig. 1a and Fig. S1). The excitation source is a Ti:Sapphire femtosecond laser (Chameleon Ultra II, Coherent) that can be tuned to wavelengths optimal for imaging both GCaMP and the red fluorescent protein fused to the channelrhodopsin. A polarized beam splitter separates the laser beam into independent imaging and optogenetic stimulation paths. In the imaging path, an electrically tunable lens (ETL) with resonant scanners enables high-speed volumetric imaging, capturing a 10-plane brain volume (400 × 400 × 120 µm³) at 3 Hz.
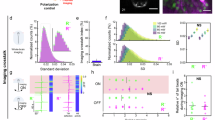
a Simplified schematic representation of the two-photon optical interrogation system. Created using BioRender (https://biorender.com/).(See Fig. S1 for the complete version of the optical system). b AOM driver input voltage vs laser power. The 0th order laser power can be continuously tuned between 5 and 100%, corresponding to a control voltage 5–0 V. c Fluorescent intensity vs laser power. The two-photon excited signal was measured by the fluorescence intensity of fluorescent beads, fitted by a second-order polynomial. d APPC image of a fluorescent plate. Left: original input image. Right: two-photon fluorescence image of the fluorescent plate with the input image loaded on the AOM. Scale bar: 20 µm. e Selective GCaMP6s stimulation of neurons via APPC in zebrafish larvae (n = 5 experiments with consistent results). Left: Calcium image (red) without APPC. Middle: (Green) APPC applied to block excitation of neuron A and B, where the image laser was tuned off in the block. Right: Merged image showing a comparison between conditions without and with APPC. Scale bar: 20 µm. f Fluorescence trace of GCaMP for neurons A and B, with and without APPC.
Next, we focus on the selection of an optical modulator for power control. The AOM modulates light power by applying sound waves to an AOM crystal, causing the light to be diffracted. The bandwidth of the AOM is limited by both the beam diameter and the acoustic velocity in the crystal. To enhance bandwidth, we reduce the beam diameter to 0.5 mm. The AOM exhibits a rise time (\({\tau }_{{on}}\)) of 85 ns, corresponding to an 11.8 MHz bandwidth, which is compatible with a pixel dwell time of 180 ns (5.6 MHz bandwidth). This configuration allows for single-pixel resolution modulation. The Electro-Optic Modulator (EOM) is also widely applied to control laser power by rotating light polarization and responds quickly to electric fields. However, its bandwidth is limited by the control signal, and larger EOMs require high-voltage amplifiers, which typically have limited gain bandwidth ( ≤ 1 MHz). Given this limitation, we have opted for the AOM for rapid control of laser power. The AOM is synchronized with the resonant scanner to dynamically regulates the excitation power, while a pair of dispersion prisms compensates for the wavelength-dependent dispersion induced by the AOM. Using the 0th-order beam (5–100% power range), we calibrated the relationship between AOM driver voltage and output power (Fig. 1b). We then confirm the second-order relationship of excitation power and fluorescence intensity, which has a maximum excitation contrast ratio of ~400:1 (Fig. 1c). After adjusting the time delay (see Methods and Fig. S2), we applied APPC imaging to a uniform fluorescent plate, demonstrating robust continuous control of excitation power at designated pixels (Fig. 1d). Next, we tested the APPC on larval zebrafish and verified its ability to efficiently turn off the two-photon excitation of specific target neurons (Fig. 1e, f). When applied to live, large-volume brain imaging in larval zebrafish, the AOM turned off the scanning laser in the eye region during functional imaging sessions (Fig. S3a), effectively mitigating laser-induced agitation (Fig. S3b-d).
In the optogenetics stimulation path, we programmed the SLM to generate scanning spot arrays targeting neuronal soma. Co-calibration of imaging and stimulation modules using fluorescent plates ensured spatial precision (see Methods).
Cross-talk between imaging and stimulation for optogenetic targeted neurons
We first assessed the cross-talk of ChR2, the most widely used optogenetic tool across various animal models, that has largely overlapped absorption spectrum with GCaMP6s. This stringent condition enabled us to validate that the proposed APPC method can effectively minimize cross-talk. Our strategy was to perform multi-plane volumetric imaging (3 Hz) under standard zebrafish imaging conditions9,55 while applying APPC to adjust laser intensity specifically in ChR2+ pretectal neurons. Next, we analyzed the calcium trace and evaluated pretectal neuron activity at different laser powers for the following APPC pattern updating (Fig. 2a). In this study, we utilized a transgenic fish line: Tg(KalTAu508; UAS-ChR2(H134R)-mCherry; UAS-GCaMP6s)9,56,57, in which the Gal4 variant protein KalTA4 drives expression of both ChR2-mCherry and GCaMP6s in specific neuron populations. Structural imaging confirmed colocalization of green (GCaMP6s) and red (mCherry-tagged ChR2) fluorescence in KalTA4u508+ pretectal neurons (Fig. 2b).
a Experimental strategy for minimizing cross-talk of optical interrogation of ensembles. The ChR+ and ChR- neurons are identified after 3D structure imaging. Then, the APPC image was generated based on the ChR+ neuron pattern for each plane. We conducted volumetric calcium imaging with the imaging power on ChR+ neurons was reduced using APPC. After calcium imaging, the neuronal activities were extracted and deconvoluted. Based on the neuronal activity level, the imaging power on each ChR+ neuron was then further updated on APPC image. b Left: Maximum intensity projection of a two-photon image stack encompassing the midbrain of a 7 dpf triple transgenic Tg(KalTA4u508;UAS:GCaMP6s; UAS:ChR2(H134R)-mCherry). Green: GcaMP6s, Red: ChR2/Chrimson. Right top: enlarged ROI for cross-talk evaluation. Right middle: enlarged ROI for cross-talk evaluation of 7 dpf triple transgenic Tg(KalTA4u508;UAS:GCaMP6s; UAS:ChrimsonR-mKate). Right bottom: ROI of control group, Tg(KalTA4u508;UAS:GCaMP6s). Scale bar: 50 µm (left) and 10 µm (right). c Calcium signal recorded from GCaMP and with and without channelrhodopsin co-expressed neurons with different imaging power. Top group: ChR2+ neurons, middle group: ChrimsonR+ neurons, bottom group (control): ChR- neurons. d Comparison of mean calcium fluctuation (mean±s.d.) of ChR2+ group at different image power (n = 29 neurons, 7 fish, Friedman test with Dunn’s multiple comparisons test, two-sided; 5 mW: 0.27 ± 0.17, 10 mW: 0.33 ± 0.23, 15 mW: 0.59 ± 0.19). e same as (d) but for ChrimsonR+ group (n = 37 neurons, 8 fish, one-way analysis of variance (ANOVA) with Holm-Šídák test, two-sided; 5 mW: 0.21 ± 0.13, 10 mW: 0.32 ± 0.15, 15 mW: 0.40 ± 0.12) and (f) same as (d) but for ChR- group (n = 47 neurons, 10 fish, Friedman test with Dunn’s multiple comparisons test, two-sided; 5 mW: 0.24 ± 0.13, 10 mW: 0.22 ± 0.15, 15 mW: 0.26 ± 0.16). g, Spikes value (mean ± s.d.) of ChR2+ group (n = 29 neurons, 7 fish, Friedman test with Dunn’s multiple comparisons test, two-sided; 5 mW: 5.38 ± 3.42, 10 mW: 6.65 ± 4.46, 15 mW: 12.15 ± 4.03) at different image power. h same as (g) but for ChrimsonR+ group (n = 37 neurons, 8 fish, Friedman test with Dunn’s multiple comparisons test, two-sided; 5 mW: 4.74 ± 3.34, 10 mW: 7.08 ± 3.30, 15 mW: 9.18 ± 3.44) and (i) same as (g) but for ChR- group (n = 47 neurons, 10 fish, Friedman test with Dunn’s multiple comparisons, two-sided; 5 mW: 4.54 ± 2.71, 10 mW: 4.39 ± 2.89, 15 mW: 5.16 ± 3.28). Scale bar in (e) left is 50 μm and in (e) right is 10 μm.
For calcium imaging, the laser power on pretectal ChR2+ neurons was set to 5 mW, 10 mW, and 15 mW by APPC, the commonly used imaging power for zebrafish9. As the imaging power decreases, the signal-to-noise ratio (SNR) of calcium traces become lower while the ΔF/F remains the same (Fig. S4). We determined that 5 mW was the minimum laser power required to maintain an acceptable SNR for measuring calcium traces (ΔF/F) in individual ChR2+ neurons (Fig. S4)54. Next, we performed three trials (100 s each) for every power condition, resulting in nine trials presented in a randomized sequence. Calcium traces (ΔF/F) from each neuron were normalized to their corresponding 15 mW trials (see Methods). Representative traces are shown in Fig. 2c. Our results demonstrated that the normalized mean ΔF/F decreased with reduced imaging power, indicating diminished neuronal activity caused by cross-talk (Fig. 2d). Specifically, the mean ΔF/F value at 15 mW excitation is more than double that at 5 mW. To further quantify neuronal activity, we applied the CNMF algorithm to deconvolve calcium traces and estimate “spike values”, which approximate spiking frequency54 (see Methods and Fig. S5). Compared to 15 mW excitation, neuronal activity (spike values) was significantly reduced at 5 mW and 10 mW excitation (Fig. 2g). On the contrary, in control experiments using ChR2- larvae (Tg(KalTA4u508; UAS-GCaMP6s)), neither mean ΔF/F nor spike values in KalTA4u508+ neurons varied across three excitation power conditions (Fig. 2f, 2i). Notably, at 5 mW excitation, both metrics were statistically indistinguishable between ChR2+ and ChR- neurons (Fig. S6), suggesting the absence of detectable cross-talk at low excitation power. This finding is crucial for the effectiveness of the APPC method, as it selectively attenuates laser intensity in ChR2+ neurons to minimize cross-talk while maintaining the optogenetically stimulated calcium signal from ChR2+ neurons at an acceptable SNR.
ChrimsonR, a widely used red-shifted channelrhodopsin, is favored due to the separation of its single-photon excitation spectrum from that of GCaMP58. To test whether two-photon calcium imaging at 920 nm excitation would induce significant cross-talk in ChrimsonR expressing neurons, we conducted a similar experiment in the Tg(KalTA4u508; UAS-GCaMP6s; UAS-ChrimsonR-mKate2) larvae, using APPC to control the imaging power on ChrimsonR+ neurons58. The results showed that despite spectral separation of single-photon excitation, neurons expressing ChrimsonR exhibited similar behaviors of cross-talk as ChR2 for two-photon imaging: the reduced artefactual neuronal activity was observed under low-power excitation (Fig. 2e). Spike values at 5 mW decreased significantly compared to 15 mW and 10 mW (Fig. 2h), indicating the presence of residual cross-talk at high excitation power, which likely arises from ChrimsonR’s strong two-photon excitation at 920 nm (Fig. S7). Again, both mean ΔF/F and spike values did not differ significantly between ChrimsonR+ and ChR- neurons at 5 mW illumination (Fig. S6). Taken together, these results indicate that localized low-power imaging effectively mitigates cross-talk. It is important to note that ChrimsonR was fused with mKate2, which exhibits higher fluorescence intensity than the mCherry marker fused to ChR251, thus complicating direct comparisons of channel expression levels based solely on fluorescent protein signals. A plausible explanation for the unexpectedly high cross-talk with ChrimsonR may be attributed to the elevated membrane expression of ChrimsonR, combined with sufficient two-photon excitation efficiency at 920 nm, collectively leading to the substantial cross-talk observed in our ChrimsonR-expressing neurons5.
In summary, we determined that 5 mW is a nearly cross-talk-free power for both ChR2 and ChrimsonR under our experimental conditions. The calcium traces of ChR+ neurons could be reliably segmented due to their strong responses to optogenetic stimulation (Fig. S4). We validated our holographic optogenetic stimulation on multiple neurons (Fig. S8) and found that downstream neurons typically exhibit significantly weaker or distinct responses compared to upstream neurons (Fig. S9). Under low signal-to-noise ratio (SNR) conditions, downstream neurons with weak responses cannot be reliably identified or analyzed using the CNMF algorithm54(Fig. S10). Therefore, it is essential to set the laser power in ChR- regions to 15 mW to ensure the quality of calcium signal data, which is crucial for accurately dissecting neural circuits.
Interrogation of neural circuits with minimized cross-talk via active pixel power control
We first investigated how cross-talk at the ChR+ neurons affects their downstream neurons. To identify these downstream neurons, we applied two-photon holographic optogenetic stimulation targeted at pretectal u508+ neurons while simultaneously performing volumetric whole-brain imaging. Concurrently, we utilized APPC on the target u508+ neurons to control laser power for subsequent cross-talk evaluation (Fig. 3a). In these experiments, we systematically reduced the imaging power at the ChR+ pixels from 15 mW to 10 mW, 5 mW, and “0 mW,” while maintaining 15 mW imaging power at other pixels to ensure high-quality calcium signals from downstream neurons. For this, we used larvae pan-neuronally expressing nucleus-localized GCaMP6s, while restricting ChR2 or ChrimsonR expression to the u508+ neurons Tg(elavl3:Hsa.H2B-GCaMP6s;KalTA4u508; UAS:ChR2 (H134R)-mCherry)9,56,59 or Tg(elavl3:Hsa.H2B-GCaMP6s; KalTA4u508; UAS:ChrimsonR-mKate2)58. We demonstrated that 920 nm optogenetic stimulation effectively activated neurons expressing either ChR2 or ChrimsonR (Fig. 3b). However, we observed that the colocalization of GCaMP6s and ChRs was very low in both strains of zebrafish larvae. Consequently, we performed holographic optogenetic stimulation on all pretectal ChR+ neurons. Noticed that we used APPC to block ChR+ neuropil and prevent unintended crosstalk stimulation (Fig. S11a–d). Here, we present the case of activating a single neuron co-expressing ChR and GCaMP6s in Fig. 3c, d and Fig. S11e–g, showing successful activation of the target upstream neuron and identification of its downstream connections. The case of activation of multiple neurons is shown in (Fig. S11h–k). Our study then focused on the calcium responses from downstream neurons. The putative downstream neurons were identified by their highly correlated activity patterns with the targeted ChR+ u508+ neurons60. Since the stimulation light could scatter onto the adjacent retina and potentially introduce visual artifacts, we conducted control experiments to exclude sensory neurons responding to this scattered light. Specifically, we positioned the optogenetic stimulation light on randomly selected ChR- neurons in a neighboring region (15–20 μm) from the ChR+ neurons (Fig. 3c and Fig. S12) as an “off target control”. The “off target control” neurons did not show neuronal activities respond to holographic optogenetic stimulation at ChR+ neurons, indicating that there are no direct or indirect connections between them.
a Experimental strategy for evaluating cross-talk spreading in the downstream neurons. We first conduct holographic optogenetic stimulation on single or multiple ChR+ neurons while simultaneously performing volumetric calcium imaging to identify the putative downstream neurons. Next, the imaging power on ChR+ neurons is reduced using APPC, and we then conduct volumetric calcium imaging to exam the cross-talk by measuring the neuronal activity of putative downstream neurons. b 920 nm two-photon optogenetics stimulation effectively activates both ChR2 (Blue) and ChrimsonR (Red). The somatic GCaMP signals (mean ± s.d.) averaged across ten trails under the same optogenetic, with onset indicated by the red bar for 1 s. c Field of view of GCaMP6s fluorescent signal of zebrafish brain depth projection across 80–180 µm. Red patch: optogenetic stimulation target. Orange patch: off-target control stimulation position. Green: putative downstream neurons. Scale bar: 50 µm . d Heatmap displaying the activities of the optogenetic target neuron and 13 downstream neurons. The optogenetic target neuron was imaged at 5 mW, 10 mW and 15 mW, while the downstream neurons are imaged at 15 mW. The downstream neurons are determined based on the correlation between the regressor related to optogenetics stimulation (stimulation onset at red lines). e Comparison of mean calcium fluctuation (mean±s.d.) of ChR2+ group downstream neurons at different image power on upstream neuron (n = 42 neurons, 8 fish, Friedman test with Dunn’s multiple comparisons test, two-sided; no optogenetic stimulation: 0 mW: 0.53 ± 0.41, 5 mW: 0.59 ± 0.51, 10 mW: 0.60 ± 0.47, 15 mW: 0.98 ± 0.74) (f) same as (e) but for ChrimsonR+ group (n = 30 neurons, 7 fish, Friedman test with Dunn’s multiple comparisons test, two-sided; no optogenetic stimulation: 0 mW: 0.34 ± 0.34, 5 mW: 0.35 ± 0.29, 10 mW: 0.60 ± 0.50, 15 mW: 0.66 ± 0.48). g Spikes value (mean±s.d.) of ChR2+ group downstream neurons at different image power on upstream neuron (n = 42 neurons, 8 fish, Friedman test with Dunn’s multiple comparisons test, two-sided; no optogenetic stimulation: 0 mW: 13.40 ± 13.47, 5 mW: 13.18 ± 11.43, 10 mW: 17.21 ± 17.86, 15 mW: 24.13 ± 21.93). h Same as (g) but for ChrimsonR+ group (n = 30 neurons, 7 fish, Friedman test with Dunn’s multiple comparisons test, two-sided; no optogenetic stimulation: 0 mW: 14.31 ± 9.40, 5 mW: 14.00 ± 11.59, 10 mW: 18.89 ± 13.95, 15 mW: 28.67 ± 20.86).
Only neurons that responded specifically to activation of “target” pretectal ChR+ neuron, and not generally to light (Fig. S13), were selected as downstream neurons (Fig. 3d). These neurons exhibited strong correlations with the optogenetic stimulation applied to the targeted ChR+ neuron across various imaging conditions (Fig. 3d). In zebrafish expressing either ChR2 or ChrimsonR in the u508+ neurons, analysis of the downstream neurons revealed that their artefactual activity diminished as the imaging power applied to the upstream neurons was reduced (Fig. 3e–h). Importantly, these results demonstrate that applying 15 mW imaging power to ChR+ regions caused strong cross-talk, which affected downstream neurons and perturbed the neuronal circuit. Furthermore, neuronal activity levels were statistically indistinguishable when the upstream neurons were scanned at 5 mW versus “0 mW” (Fig. 3e–h). This led to the conclusion that 5 mW is the “near cross-talk free” power for upstream neurons, effectively minimizing cross-talk and providing a solution for negligible interference within the neuronal circuit. This means that in a strain of zebrafish expressing calcium indicators and ChRs in upstream neurons, alongside calcium indicators in downstream neurons, “near cross-talk free” optical interrogation can be achieved using the APPC approach.
To extend our method to different imaging scenarios, we investigated how to optimize imaging conditions to minimize cross-talk. A particularly important consideration is the extent to which ChRs can be activated by out-of-focus scanning light in 3D volumetric imaging. First, we modeled the activation of ChR2 molecules during imaging (see Methods) and assessed the extent of ChR2 molecule activation under various scanning conditions. Our simulation indicates that the ChR2 molecule can be effectively activated at a typical low power density of ~0.2 mW/µm2 with a stimulation duration of 1 ms, consistent with numerous in vivo studies5,7,9,34 (Fig. 4a and Fig. S14). The excitation probability is proportional to laser power density and stimulation time. However, during calcium imaging, the power density at the focal point can be 50 times higher than the typical activation power density, significantly reducing the stimulation time required for ChR activation3,9,46. At an excitation power of 15 mW, a channelrhodopsin-2 (ChR2) molecule has an activation probability of ~8.1% under each scan during imaging. This probability decreases to ~0.9% at an excitation power of 5 mW, given a pixel dwell time of 68 ns under our imaging conditions (Fig. 4a). The trend is consistent with our previous in vivo experiment results. In addition, a recent study also demonstrated that rapid raster scanning can lead to power-dependent cross-talk45. Therefore, by controlling imaging power at the pixel level (e.g., 5 mW), we can effectively mitigate cross-talk. Suitable imaging power can be further adjusted according to the change of FOV. A previous study has shown that setting stationary out-of-focus light with a high numerical aperture (NA) can activate more ChRs across the entire membrane compared to focusing on the soma equator33. While dendrites and axons can contribute to photoactivation, their distribution and relationship with the neuronal membrane potential are complex. Therefore, to effectively simulate the scanning scenario, we have simplified our model to concentrate primarily on the main source of cross-talk, which is the ChR located at the soma membrane. For this simulation, we used a neuronal soma diameter of 10 µm, which is consistent with the sizes of neurons we observed in larvae zebrafish. The imaging light was scanned above or below the soma equator at varying distances and power levels (Fig. 4b). Our simulation indicated that the total photocurrent decreases as the distance between the focal plane and the equator increases (Fig. 4c, Fig. S15, and Fig. S16). Given the symmetry of the photocurrent generated by the out-of-focus plane, we focused only on the scenarios where the out-of-focus plane was positioned above the equator. To validate our simulation results in vivo, we performed quasi two plane imaging to validate out of focus activation on ChR2+ neurons in larva zebrafish (Tg(KalTAu508; UAS-ChR2(H134R)-mCherry; UAS-GCaMP6s))59. One imaging plane was set at the equatorial (central) plane of the neuron, while the other was placed at an offset latitudinal plane located at distances varied from 0 µm to 14 µm using the ETL, exceeding the typical sampling gap9,55. And the two planes were scanned sequentially (offset plane first) under ETL control, ensuring that the equatorial plane was not simultaneously excited with the offset plane.
a The activation probability of a ChR2 molecule under stimulation by two-photon raster scanning at different powers of a defocused laser (Simulated N.A. = 0.9). The dashed lines correspond to the stimulation times for 100 μm, 200 μm, and 400 μm FOVs using a resonant scanner with an 8 kHz resonance frequency. The results account for the attenuation caused by the sample tissue (see “Methods”). For simulation results that do not consider attenuation for in vitro experiments as reference, see Fig. S14. b Illustration of raster scanning through the entire focal plane at a distance z from the cell equator plane (orange). Green circle with red outline: soma. c Simulated photocurrents stimulated by raster scanning at various distances Z related to the equator. d The spikes value (mean ± s.d.) of target neuron with an extra scanning plane outside of cell equator (n = 24 neurons, 15 fish, Friedman test with Dunn’s multiple comparisons test, two-sided; 0 µm: 7.68 ± 4.57, 2 µm: 6.75 ± 2.97, 5 µm: 6.54 ± 3.16, 10 µm: 5.33 ± 2.44, 14 µm: 4.79 ± 2.48, OFF: 4.35 ± 1.85). Black dot and black line: simulated results in (c) and fitted to spike value. e Illustration of the volumetric calcium imaging (10 planes) experiment with plane gap of 2–14 µm. The equatorial plane of target neuron located at the 6th plane. f The spikes value (mean ± s.d.) of target neuron with an extra scanning plane outside of cell equator (n = 30 neurons, 10 fish, Friedman test with Dunn’s multiple comparisons test, two-sided; 2 µm: 6.79 ± 4.46, 5 µm: 5.81 ± 4.51, 8 µm: 5.08 ± 3.18, 10 µm: 4.76 ± 2.95, 14 µm: 4.61 ± 2.57, OFF: 4.21 ± 2.42). g Simulated photocurrents evoked by raster scanning at various distances Z from the equator under different imaging conditions. h The spikes value (mean±s.d.) of target neuron at 5 mw whole cell imaging, 15 mw sub-soma imaging, and 15 mw whole cell imaging. (n = 12 neurons, 5 fish, one-way ANOVA with Tukey’s multiple comparisons test, two-sided; 5 mW whole cell: 4.146 ± 2.09, 15 mW sub-soma: 3.91 ± 1.67, 15 mW sub-soma: 8.47 ± 4.46).
With APPC, we applied 5 mW of imaging power to the equatorial plane and 15 mW to the offset plane. Since our previous results (Figs. 2 and 3) suggested that 5 mW imaging power does not induce detectable cross-talk, the calcium signal recorded at the equatorial plane alone was used as the near cross-talk–free reference. To evaluate out-of-focus activation, we measured calcium signals at the equatorial plane while the offset imaging plane was excited at a higher power (15 mW). These signals were then compared with the reference condition, in which the offset plane was not excited, to quantify cross-talk originating from out-of-focus activation. We found that out-of-focus light at 10 µm or farther away from the equatorial plane leads to a negligible cross-talk (Fig. 4d). This experiment mimics the case of the neuronal equator located in the gap between imaging planes and demonstrated the in vivo experiment results are consistent with our simulation model.
Next, we evaluate the impact of 10-plane volumetric calcium imaging on cross-talk as more imaging planes may induce more severe cross-talk. In our experimental setup, the imaging planes are positioned at distances of 2, 5, 8, 10, and 14 µm, with the equatorial plane of monitored neuron situated at the 6th plane from the top. All the offset imaging planes are at 15 mW while applying 5 mW imaging at target neuron at the 6th plane using APPC to avoid high cross-talk baseline. In the control group, the imaging power for the other 9 planes was turned off (single-plane imaging). Consistent with previous results, cross-talk from out-of-focus excitation becomes non-negligible when the distance between planes is smaller than 8 µm, as shown in Fig. 4f and Fig. S17, indicating the further imaging plane provide minimum contribution. Taking together, these results provide critical guidance for determining optimal depth intervals in 3D volumetric imaging protocols.
To further investigate the performance of APPC, we examined its potential to spatially prevent excitation of ChR at the cell membrane and in the axons and dendrites near the soma. Given that ChR is localized at the cell membrane while GCaMP is situated within the cytoplasm or nucleus, we propose that restricting the imaging laser to the sub-soma region may significantly reduce cross-talk. We conducted numerical simulations with varying sizes of restrictions on the soma, as illustrated in Fig. S16. When scanning at the equatorial plane, most ChR activation occurs in the equator. Consequently, confining excitation inside this region markedly diminishes activation across the entire neuron, as depicted in Fig. 4g. We then performed in vivo experiments to validate our simulation results, where we scanned the whole neuron by 5 mW and 15 mW laser, and 15 mW laser confined to sub-soma region with a diameter of 4 µm due to neuron’s irregular shape. The results presented in Fig. 4h and Fig. S18a demonstrate that laser confinement effectively reduces cross-talk caused by the activation of ChR at the equator and adjacent axons and dendrites. Although this approach results in a decrease in the calcium signal collected of 73%, it still reflects overall cellular activity (Fig. S18b).
Discussion
In summary, we present the APPC approach, an innovative all-optical methodology that enables simultaneous monitoring and manipulation of neuronal activity while markedly reducing Type II cross-talk interference. Leveraging the rapid tuning capacity of AOM, our method achieves continuous modulation of scanning light intensity with single-pixel resolution. Guided by ChR expression patterns, we implemented dynamic control of imaging power within defined regions of interest to prevent inadvertent activation of ChRs by the imaging beam. In larval zebrafish, this strategy enabled comprehensive whole-brain functional imaging while mitigating laser-induced behavioral artifacts through selective suppression of scanning in the ocular region. Collectively, these findings highlight the potential of APPC to preserve the functional integrity of diverse light-sensitive domains within the imaging volume.
In particular, we evaluated the efficiency of APPC in two experimental contexts. First, we combined ChR2 with GCaMP6s, the widely used light-sensitive ion channel and genetically encoded calcium indicator, respectively. This pairing represents a stringent test for two-photon optical interrogation with minimal cross-talk, given their overlapping excitation spectra. Second, we examined ChrimsonR, the far red-shifted ChR (excitation peak at 1130 nm), in combination with GCaMP6s45. Our findings revealed that both ChR2 and ChrimsonR exhibit significant cross-talk under calcium imaging excitation at 920 nm, thereby potentially altering downstream neuronal dynamics. These results indicate that the calcium imaging laser not only directly perturbs ChR+ neurons but also disrupts broader circuit function by activating ChR+ populations. Importantly, we successfully suppress the cross-talk across the neuronal circuit using APPC and demonstrate cross-talk minimized, accurate two-photon optical interrogation can be achieved using the single-laser systems commonly available in most laboratories worldwide. APPC aims to suppress Type II cross-talk; however, single-laser excitation is not optimal due to the presence of Type I cross-talk and the lower excitation efficiency for red ChR activation. These issues can be mitigated by extending APPC to dual-laser configurations, where one laser with a longer wavelength optimally stimulates red-shifted ChRs, such as ChrimsonR, to avoid Type I cross-talk, while APPC modulates the other calcium imaging laser to minimize Type II cross-talk. Taken together, our study establishes APPC as both a cost-effective solution for laboratories relying on single-laser two-photon systems and a performance-enhancing strategy for dual-laser platforms by suppressing the cross-talk between imaging and manipulation.
We observed significant cross-talk on ChrimsonR during imaging at 920 nm, attributable to the broadened and blue-shifted two-photon absorption spectrum of ChRs. Several factors are likely to contribute to this effect. First, the complex chromophore environment surrounding the protonated Schiff base (PSB) introduces uncertainty under two-photon excitation, as interactions between the PSB and adjacent counterions may generate new electronic states49,53. Second, the seven-transmembrane scaffold constrains the retinal chromophore, reducing centrosymmetry and relaxing parity restrictions, thereby permitting transitions that would otherwise be forbidden47,48,52. Third, vibronic transitions can be selectively enhanced by two-photon excitation51. Finally, resonance enhancement allows access to higher excited singlet states that are less accessible under single-photon excitation50. As a result, some ChRs, including ChrimsonR, exhibit excitation peaks at shorter effective wavelengths5. The magnitude of cross-talk varies with ChR expression levels and intrinsic properties, including excitation spectrum, kinetics, and photocurrent3,13,26. Therefore, the broadening of the two-photon excitation spectra of ChRs presents significant challenges for achieving optical interrogation with minimal cross-talk. However, we encourage the application of a combination of spectral separation between GECI and ChR, as this approach inherently offers a lower potential for cross-talk. In principle, pairing red GECIs with blue ChRs that possess non-overlapping excitation spectra offers an optimal strategy for eliminating cross-talk25,44. However, despite notable progress in red GECI development, current variants still have limitations. Many red GECIs perform well under single-photon excitation32,36,37,38,39,40, yet the brightest indicators are still dimmer than GCaMPs or lack validation under two-photon excitation32,36,38, and most exhibit relatively low ΔF/F ratios under high-activity conditions due to weaker coupling between calmodulin (CaM) and the red fluorescent protein (RFP) chromophore, restricting their ability to capture dynamic neuronal responses32,36,37,38,39,40.
In this study, we present a comprehensive framework for optically quantifying cross-talk and adaptively optimizing imaging conditions. We demonstrate that low imaging power hinders the reliable identification of downstream neurons, underscoring the need to maintain normal laser power in ChR- regions to preserve calcium signal quality and ensure accurate circuit dissection. Our approach dynamically adjusts imaging power on a per-neuron basis according to depth, ChR expression levels, and intrinsic neuronal properties, thereby minimizing cross-talk while preserving high-fidelity calcium readouts. To support this framework, we developed a computational simulation model that predicts viable imaging parameters and guides experimental design. By integrating optical activity estimation with dynamic excitation power control, we identified near cross-talk-free imaging conditions that minimize interference in both upstream and downstream pathways, and we validated these conditions through computational simulations and in vivo volumetric imaging.
An essential step in our framework is the evaluation of the “near cross-talk-free” imaging power. While two-photon excitation of ChR theoretically scales quadratically with excitation power, the resultant ion influx does not always yield reliable action potential generation due to multiple nonlinear factors. As imaging power increases, out of focus excitation of ChR is enhanced, thereby enlarging the effective stimulation volume relative to lower power. Moreover, the thresholding nature of neuronal spiking and membrane potential saturation under strong excitation further shape the nonlinear outcomes7,45,46. Cross-talk behavior also varies with neuronal properties and opsin-specific features such as sensitivity and kinetics4,7,45,46. Consequently, the “near cross-talk free” imaging power must be empirically determined for each opsin-expressing neuron within its experimental context. For neurons expressing only opsins, the imaging laser should be fully blanked at their pixels using APPC to eliminate cross-talk. For neurons co-expressing ChRs and GECIs, imaging power should be gradually increased from zero until the calcium signal achieves the minimal acceptable SNR (e.g. ~3 for reliable analysis54); if further increases do not yield statistically significant activity changes, the minimal-SNR power can be considered “near cross-talk-free”. Conversely, if neuronal activity scales with power, an optimal value must be selected by balancing cross-talk minimization with reliable calcium signal extraction.
It should be noted that we demonstrate APPC’s efficiency on KalTA4u508 neurons, which are sparse labeled. Based on our results, we expect the complexity of applying APPC on several scenarios. In the first scenario, ChR+ neurons are densely packed and located within the inter-plane gaps. Acquiring 3D information about ChR+ neurons is essential for accurately determining the imaging laser power distribution. Specifically, once the position of the imaging plane is established, the pixel power is reduced if a ChR+ neuron is located on the image plane or just a few microns away from it. This approach, however, may reduce calcium signal quality when a large number of ChR+ neurons are closely packed. In the second scenario, typically related to Type III cross-talk, ChR expression extends beyond the soma to axons and dendrites, which are often too small to be clearly visualized. In this case, we take advantage of the distinct localization patterns of GCaMP and ChR: GCaMP is primarily expressed in the cytoplasm or nucleus, whereas ChR is predominantly localized to the cell membrane. With APPC, we could restrict the calcium imaging laser exclusively to sub-soma region, the intracellular space of the soma excluding membrane, while completely turning off illumination outside the soma of GCaMP expressing neurons. Both our simulation and experiment results indicate that by confining excitation to the center of the soma, the activation of ChR molecules on the neuronal membrane can be significantly reduced. Another promising approach is to engineer protein tags that restrict ChR expression to the soma membrane, for example, by fusing a soma-targeting sequence from the voltage-gated K⁺ channel Kv2.1 to CoChR. The resulting stCoChR variant would markedly reduce expression in distal dendrites and axons, further enhancing the precision of optical interrogation44.
Since APPC relies on one-to-one spatiotemporal mapping, it is possible to implement it in other systems, such as light-beads microscopy, which also adheres to this mapping relationship. In this technique, light beads are generated by splitting the beam and introducing varying time delays to each “bead”. In such cases, an AOM/EOM could be integrated into the individual optical path of each bead, although this would increase the system’s complexity. However, APPC cannot be applied to microscopes that utilize more complex spatiotemporal mappings, such as light-sheet and Bessel beam systems. Alternatively, by temporarily increasing the imaging power to an appropriate level for targeting ChR+ neurons, we can, theoretically, achieve optogenetic stimulation using only the imaging light, as demonstrated in previous studies45. However, it may induce higher latencies and higher jitters compared with holographic area stimulation4. In addition, this method cannot simultaneously activate multiple neurons located at different positions (depths) in a 3D volume.
Due to the limited SNR of ChR+ neurons under APPC, weak activity may not be reliably extracted. Therefore, it may be necessary to use optogenetic stimulation to elicit stronger activity in order to examine network connections. In certain experiments that utilize the virus to express GCaMP, increasing the virus concentration can help enhancing the SNR. In addition, we conducted off-line 3D registration as the zebrafish larvae was well embedded in agarose. With APPC on eye region, we observed negligible tissue drifts. However, real time registration will be important to maintain the accuracy of APPC on target region during in vivo experiments, particularly under conditions where tissue drift occurs, such as in awake mice.
We demonstrate holographic optogenetic stimulation using 920 nm laser, which strongly excites GCaMP6s and induces a substantial baseline fluorescence signal, leading to Type I cross-talk. To mitigate this issue, we implemented a baseline subtraction method, where the baseline is estimated from the signals collected during the scanning flyback period at the edge of each line, when the imaging laser power is turned off by the APPC. Subtracting this baseline from the GCaMP signal results in an undistorted calcium trace. This approach allowed us to accurately extract calcium responses without identifying inactive regions. Our results indicate that we can activate up to 25 neurons without PMT saturation, which is adequate for most in vivo optogenetics studies3,23,24. However, the background fluorescence signal can saturate PMT when a large group of neurons are activated, potentially causing damage to PMT. We recommend incorporating a mechanical shutter for PMT to temporarily block strong fluorescence or dynamically adjust the PMT gain by controlling the driving voltage. If the duration of optogenetic stimulation increases to second level, recovering the calcium trace becomes more challenging. In such cases, it is necessary to employ a dual-laser configuration to reduce Type I cross-talk.
Methods
Animals
All procedures conformed to the guidelines by the animal ethics committee of HKUST. Animals were kept under a standard 14:10 light cycle at 28 °C. Zebrafish larvae carrying mutations in the mitfa allele (nacre)61 were used for all experiments. Larvae were kept in Danieau’s solution, and fed with paramecia (Paramecium multimicronucleatum, Carolina Biological Supply Company) on 5 to 7 days post fertilization (dpf).
The following transgenic lines were used for this study: Tg(KalTA4u508;UAS:GCaMP6s; UAS:ChR2(H134R)-mCherry)9,57, Tg(KalTA4u508;UAS:GCaMP6s)56, Tg(KalTA4u508; UAS:GCaMP6s; UAS:ChrimsonR-mKate2)58, Tg(elavl3:Hsa.H2B-GCaMP6s; KalTA4u508; UAS:ChR2(H134R)-mCherry)59, and Tg(elavl3:Hsa.H2B-GCaMP6s;UAS:ChrimsonR-mKate2)59.
Zebrafish preparation
6-8 dpf larvae were embedded in 2% low melt agarose (Invitrogen) on 90 mm round (Thermo Scientific (cat#101VR20)) plastic petri dish. The agarose surrounding the tail and the eyes was removed by surgical blade (Paragon p303) to allow for tail and eye movements. Fish were allowed to recover overnight prior to the experiment.
Two-photon imaging microscope
A home-built two-photon light scanning microscope controlled by a C# based custom software is modified from previous setup. The laser source is a Ti:Sapphire femtosecond laser (Chameleon Ultra II, Coherent), repetition rate: 80 Mhz, and can be tuned between 680 nm and 1080 nm. The laser beam first passes through the half-wave plate to adjust polarization direction. Then, a polarized beam-splitter separates the laser beam into two paths, one for imaging and the other for optogenetic stimulation.
At imaging path, the laser beam first pass through a 4 f system of scale factor x0.467 (AC254-075-AB-ML F = 75 mm, Thorlabs and AC254-035-AB-ML F = 35 mm, Thorlabs) in order to not overfill the AOM (CAOM-080-010-TEC-740-1020-AF-A17, Castech) aperture (1.0 mm). After passing through the AOM, the laser beam is expanded by a factor of ×3 through a 4 f system (AC254-050-AB-ML F = 50 mm, Thorlabs and AC254-150-AB-ML F = 150 mm, Thorlabs). Two dispersion compensating prism (SF10 25.4 ×25.4 mm Ultrafast Prism, Edmund optics) is set and optimized the dispersion compensation at 920 nm. The first prism is fixed, and the second prism is located at a linear translation stage with actuator (CONEX-LTA-HL, Newport). The working direction of the linear translation stage is perpendicular to the prism base plane. For each wavelength used, the position of the prism is adjusted by maximizing the fluorescent intensity from the fluorescent plate. The total system group dispersion was estimated from the optical components used to calculate the distance between two prisms when the laser worked at 920 nm. The pulse duration after the objective is 118 fs, measured by pulseCheck autocorrelators (NX 150, A.P.E.) to verify the dispersion compensation.
The laser beam with 4 mm in diameter (propagate a large distance and thus expanded) is then expanded by a factor of ×4 through a 4 f system (AC254-050-AB-ML F = 50 mm, Thorlabs and AC254-200-AB-ML F = 200 mm, Thorlabs) to full fill the ETL (EL-16-40-TC-NIR, Optotune AG), which is used for multi-plane functional imaging. Then a 4 f system (AC254-200-AB-ML F = 200 mm, Thorlabs, and AC254-050-AB-ML F = 50 mm, Thorlabs) reduces the beam diameter to 4 mm to avoid overfilling the scanner mirror. A resonant scanner (CRS 8 kHz, Cambridge technology) for high-speed scanning is conjugated with a set of XY galvo scanners (6215H, Cambridge technology) by a 1:1 relay (AC254-100-AB-ML F = 100 mm, Thorlabs). Finally, two lenses (Scan lens SL50-CLS2 F = 50 mm, Thorlabs, and Tube lens TTL200MP F = 200 mm, Thorlabs) are used to correct the aberration and expand the laser beam to fulfill the objective back pupil (objective lens (20x, N.A. 1.0, XLUMPLFLN20XW, Olympus Corporation)). The ETL, resonant scanner, XY galvo scanners, and objective pupil plane are all conjugated. The control voltage of ETL was calibrated by comparing the 3D stack of samples obtained by moving the objective with an actuator and shifting the focal plane using the ETL. We fulfill the ETL aperture to minimize the change in focal power required for shifting the focal plane by the same distance along the Z-axis. Only the upper one tenth of the image plane is slightly tilted, but each plane remains parallel to one another at the same distance.
In the fluorescence collection path, a dichroic mirror (FF705-Di01-25*36, Semrock) is set between the tube lens and objective to reflect the fluorescence light, which then passes through two relay lenses (AC254-150-AB-ML F = 150 mm, Thorlabs and AC254-040-AB-ML F = 40 mm, Thorlabs) into the PMT (H11461-03, Hamamatsu). Two filters (FF03-525/50-25, Semrock, and FF01-600/52-25, Semrock) are installed on a flipper which can switch the filter to collect different fluorescence signals. Voltage signals are converted from PMT output current using a high-speed current amplifier (DHPCA-100, Femto) and digitized using an oscilloscope device (PCIe-9105H, ART technology).
At stimulation path, the laser beam passes through a set of galvo scanners (6215H, Cambridge Technology). Then the light beam is expended by two lenes (AC254-50-AB-ML F = 50 mm, Thorlabs, and AC254-250-AB-ML F = 250 mm, Thorlabs) to fill the spatial light modulator (HSP1K-500-1200-PC8, Meadowlark Optics). Then the SLM is conjugated to the back focal plane of objective by a demagnify relay (AC508-250-AB-ML F = 250 mm, Thorlabs, and AC254-150-AB-ML F = 150 mm, Thorlabs). The photon stimulation light combines with imaging light by a polarized beam splitter (CCM1-PBS252/M; Thorlabs) between tube lens and dichroic mirror.
The LCR projector and camera were employed to provide visual stimuli and record animal behavior, similar to our previous work60. We used the spatial visual stimulator module (DLP® Light Crafter™ 4500 modified by EKB Technologies Ltd) to display stimuli, with 385 nm light source. The Light Crafter (LCr) projected images onto a Teflon fabric screen which is positioned inside the petri dish, underwater, and 1 cm in front of the fish head, with light intensity of ~400 nW/cm2. The prey-like stimuli consisted of UV dots measuring 4° × 2°, moving in an elliptical path of 8° × 4° at a speed of 2 Hz for 2 s against a dark background.
Pixelwise adjustment of laser intensity
The orientation of the AOM is initially adjusted to achieve maximum diffraction efficiency. Here we adopt the 0th order light and measured the intensity modulation dynamic range, which spans from 5 to 100%.
To enable dynamic adjustment of laser intensity on a pixel-by-pixel basis, we developed a GUI that allows for the custom generation of figures using fluorescence images of ChR+ neurons as the pattern for APPC. These figures are encoded into voltage signals and transmitted to the AOM driver (CARD-FE2-080-24D04C-1DA5H-AF, Castech). To synchronize the AOM with the imaging scanning beam, the clock signal generated by the resonant scanner serves as the reference clock signal for the DAQ card (PCIe9105_H, ART technology). The AO port is configured to generate a series of output signals upon receiving a trigger from the resonant scanner clock, thereby eliminating time delay accumulation (Fig. S3). Finally, we captured the fluorescence image from a fluorescence plate with the loaded figure to finely tune the delay by aligning the fluorescence pattern between two adjacent lines.
In vivo calcium imaging and holographic optogenetics stimulation
Two-photon multiple-plane imaging was performed using resonant-galvanometer raster scanning (30 Hz frame rate, 512 × 512 pixel per frame, 400 µm field-of-view, 12-14 µm between imaging plane, 10 planes, 3 Hz volume rate) typical for whole brain imaging in zebrafish60. An objective lens (20x, N.A. 1.0, XLUMPLFLN20XW, Olympus Corporation) was used for all experiments. The femtosecond laser (Chameleon Ultra II, Coherent) was operated at 920 nm wavelength for calcium imaging and 1020 nm for mCherry and mKate imaging. In all the in vivo experiments, the imaging power applied to the sample ranged from 0.5 to 15 mW. At the beginning of the imaging, we first imaging at the very low power, 0.5 mW, to outline the eye contours from the fluorescence signal of the pigments on retina. Then we manually selected the eye region and included it in the APPC pattern, where the laser power was set to be the minimum. For pixel level power control of ChR expression region, the APPC pattern was generated based on the ChR expression pattern verified by fused fluorescent protein. We set the pixel intensity threshold to adjust the pattern region flexibility. We then manually refined the pattern for each plane. The imaging power at these regions were determined based on the experiment requirement.
For optogenetics stimulation, the holograms were calculated by compressive sensing weighted Gerchberg–Saxton algorithm62 and loaded on SLM which controlled by the custom software in MATLAB. A large beam diameter is applied to achieve an effective excitation NA ( ~ 0.5) for axial profile compression. A spot array is then created on each target neuron, consisting of spots that are 10–12 µm in diameter with a 1 µm gap between them, and an average power density of 0.15–0.3 mW/µm². The spot array is scanned over a range of 2 µm to ensure that the entire soma area is stimulated. To mimic natural neuronal response, we first provided visual stimuli and extract the calcium signal from the anterior pretectal u508 neurons. Then we adjust the optogenetic stimulation laser power to trigger similar calcium response. Each optogenetic stimulation lasted for 1 second, with a total of 5 stimulations per session, separated by 30-s intervals. In the experiments that aimed at identifying the downstream neurons, we selected approximately 5–10 anterior pretectal u508 neurons on either the left or right side.
Before each optogenetic experiment, we calibrated the alignment between the imaging light and the photo-stimulation light. We bleach two dark spots on a fluorescent microscope slide (FSK3, Thorlabs) using both imaging light and photo-stimulation light. After measuring the distance between the spots, we included a coordination shift in the hologram calculation.
Calcium imaging data processing
Image analysis was performed using custom-written scripts in Matlab (Mathworks). Non-rigid motion correction for each imaging plane was performed using the NoRMCorre algorithm63. For neuron-based analyses, the locations of regions of interest (ROIs) and their fluorescent signal trace were extracted using the CNMF algorithm from the CaImAn package54. Image sections with axial shift >5 µm were discarded because they were not able to keep precise information. ROIs with the SNR, as computed by CaImAn, below 1.5 were excluded from further analysis. Specifically, the deconvolveCa function returned the baseline, denoised calcium trace, noise level estimated from the spectral density and deconvolved signal (spikes train). We calculated ΔF/F based on the inferred baseline and calcium trace. The mean ΔF/F is calculated as the average ΔF/F value across all frames during the entire video recording. Then we computed the inferred spike value as the summation of the spike value normalized by the baseline. Spike values below one standard deviation of the noise level were discarded.
Data presentation and statistical analyses
All values are presented as mean ± standard deviation (s.d.), unless otherwise specified. Statistical analyses were conducted using appropriate tests, as indicated in the figure legends. These statistical tests were performed using Graph Pad Prism, version 9.5.1, with a p value of <0.05 considered statistically significant. Data normality was first checked using the Shapiro–Wilk normality test. Normally distributed data were analyzed using paired, unpaired one-way ANOVA test. Non-normally distributed data was analyzed using paired, unpaired non-parametric test (Friedman test with Dunn’s multiple comparison test). In the statistical results presented in Figs. 2d–i and 3e–h the neurons depicted were selected based on their distribution, as they exhibited strong cross-talk. Specifically, there was no statistically significant difference in spike values below laser power of 5 mW between the ChR+ and ChR- groups, leading us to consider cross-talk negligible at this power level. Neurons with mean ΔF/F or spike values at 15 mW imaging power exceeding the mean plus one standard deviation at 5 mW were classified as exhibiting strong cross-talk and included in Figs. 2d–i and 3e–h.
A regressor for single optogenetic stimulation was generated by averaging the calcium trace corresponding to optogenetic stimulation. The downstream neurons depicted in Fig. 3c–h were selected based on the criterion that the correlation between the calcium trace and the target optogenetic regressor exceeded 0.39, while the correlation with the control optogenetic regressor remained below 0.05.
ChRs modeling
A ChR2 molecule has an absorption rate of \(\left(\frac{{\sigma }_{2}}{2}\right)I{\left(t\right)}^{2}\), and the ground state lifetime is given by \({\tau }_{2}=\frac{2}{{\sigma }_{2}{I}^{2}}\), where \({\sigma }_{2}\) is the two-photon absorption cross-section (260GM)33. The laser intensity \(I\) which can be expressed by ref. 64:
where \({g}^{\left(2\right)}\) is the factor of 0.59 for a hyperbolic-secant-squared pulse shape65, \({P}_{{avg}}\) is the total time-average laser power, \({f}_{p}\) is the pulse train repetition rate, \({\tau }_{p}\) is the pulse duration. In our calculations, we used \({f}_{p}=80{MHz}\) and \({\tau }_{p}=120{fs}\). \(h\) is the Planck’s constant, \(c\) is the light speed, \(\lambda\) is the wavelength.
The laser is attenuated in the sample due to scattering and absorption. We determined the attenuation length (AL) from the brain z-stack images, where the average fluorescence signal of the brightest 0.1% of pixels attenuates by \(\frac{1}{{e}^{2}}\) 66 (Fig. S19a). The calculated AL for 7–8 days post-fertilization (dpf) zebrafish larvae is approximately 201 µm (Fig. S19b). At a depth of 140 µm, the laser power reaching the pretectal u508 neurons was estimated to be 50% of the initial power, based on the calculated AL, and this value was used for subsequent photocurrent calculations.
The probability of activating a ChR2 molecule under stationary illumination of time \(\Delta t\) is:
Next, we consider the activation of a ChR2 molecule by a moving focused beam, which has a Gaussian distribution in the lateral direction. Thus, the laser intensity can be expressed as33:
where \(\omega\) is the beam waist. At the focal plane, \({\omega=\omega }_{0}\) and \({\omega }_{0}\) can be determined by measuring the two-photon excitation point-spread function from image 0.2 µm microspheres. The two-photon excitation waist \({\omega }_{{TPE}}\) was obtained by fitting the two-photon excitation point-spread function by the Gaussian distribution. The \(\omega\) can then be calculated from \({\omega }_{0}={\omega }_{{TPE}}\sqrt{2}\). Or \({\omega }_{0}\) can be estimated by \({\omega }_{0}\approx \frac{\lambda }{2{NA}}\). In the perifocal region, the \(\omega\) can be expressed by ref. 34:
To simulate the activation of ChR2 molecules on the neuron, we modeled the neuronal soma as a sphere with a diameter of 10 µm, generating 25,000 points representing ChR2 molecules randomly distributed on the surface, corresponding to approximately 200 proteins proteins/µm². We simulated the focused beam scanning and calculated the photocurrent of each ChR2 molecules by combining equ. (1)-(4) and integrating over time. The kinetic information can refer to Rickgauer’s work33. For ChR2, the photocycle can be completed before the next frame even for a 30 Hz frame rate. Thus, we only considered the ChR2 activation during single frame scanning. The maximum photocurrent for each ChR2 molecule was normalized to 1. The total photocurrent for the entire neuron was then obtained by summing the photocurrent contributions from each individual ChR2 molecule.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The main data supporting the findings of this study are available within the paper and Supplementary Information files. The source data files for all data presented within the figures can be found at: https://github.com/QuLab1234/APPC.
Code availability
The custom codes for image processing are available online at: https://github.com/QuLab1234/APPC.
Change history
26 February 2026
We sincerely apologize for an oversight on our part: an outdated version of the Supplementary Information file was uploaded earlier in the process, resulting in an incorrect figure sequence (Figure S11, S12, S16 and S17) and mismatched in-text citations to supplementary figures in the manuscript. To resolve this, we have attached the corrected and up-to-date Supplementary Information file, in which the figure order has been properly updated, and we kindly request that the current Supplementary Information file be replaced with this attached version.
References
Rickgauer, J. P., Deisseroth, K. & Tank, D. W. Simultaneous cellular-resolution optical perturbation and imaging of place cell firing fields. Nat. Neurosci. 17, 1816–1824 (2014).
Packer, A. M., Russell, L. E., Dalgleish, H. W. P. & Häusser, M. Simultaneous all-optical manipulation and recording of neural circuit activity with cellular resolution in vivo. Nat. Methods 12, 140–146 (2015).
Zhang, Z., Russell, L. E., Packer, A. M., Gauld, O. M. & Häusser, M. Closed-loop all-optical interrogation of neural circuits in vivo. Nat. Methods 15, 1037–1040 (2018).
Yang, W., Carrillo-Reid, L., Bando, Y., Peterka, D. S. & Yuste, R. Simultaneous two-photon imaging and two-photon optogenetics of cortical circuits in three dimensions. Elife 7, 1–21 (2018).
Mardinly, A. R. et al. Precise multimodal optical control of neural ensemble activity. Nat. Neurosci. 21, 881–893 (2018).
Marshel, J. H. et al. Cortical layer-specific critical dynamics triggering perception. Science 365, eaaw5202 (2019).
Chen, I. W. et al. In Vivo submillisecond two-photon optogenetics with temporally focused patterned light. J. Neurosci. 39, 3484–3497 (2019).
Förster, D., Dal Maschio, M., Laurell, E. & Baier, H. An optogenetic toolbox for unbiased discovery of functionally connected cells in neural circuits. Nat Commun 8, 116 (2017).
dal Maschio, M., Donovan, J. C., Helmbrecht, T. O. & Baier, H. Linking neurons to network function and behavior by two-photon holographic optogenetics and volumetric imaging. Neuron 94, 774–789 (2017).
Hampel, S., Franconville, R., Simpson, J. H. & Seeds, A. M. A neural command circuit for grooming movement control. Elife 4, 1–26 (2015).
Shipley, F. B., Clark, C. M., Alkema, M. J. & Leifer, A. M. Simultaneous optogenetic manipulation and calcium imaging in freely moving C. elegans. Front Neural Circuits 8, 1–8 (2014).
Zhang, F. et al. Multimodal fast optical interrogation of neural circuitry. Nature 446, 633–639 (2007).
Emiliani, V., Cohen, A. E., Deisseroth, K. & Häusser, M. All-optical interrogation of neural circuits. J. Neurosci. 35, 13917–13926 (2015).
Grosenick, L., Marshel, J. H. & Deisseroth, K. Closed-loop and activity-guided optogenetic control. Neuron 86, 106–139 (2015).
Jackson, A. Closed-loop optogenetic control of the dynamics of neural activity in non-human primates. https://doi.org/10.1038/s41551-022-00945-8 (2022).
Iaccarino, H. F. et al. Gamma frequency entrainment attenuates amyloid load and modifies microglia. Nature 540, 230–235 (2016).
Etter, G. et al. Optogenetic gamma stimulation rescues memory impairments in an Alzheimer’s disease mouse model. Nat. Commun. 10, 1–11 (2019).
Fougère, M. et al. Optogenetic stimulation of glutamatergic neurons in the cuneiform nucleus controls locomotion in a mouse model of Parkinson’s disease. Proc. Natl. Acad. Sci. USA 118, e2110934118 (2021).
Valverde, S. et al. Deep brain stimulation-guided optogenetic rescue of parkinsonian symptoms. Nat. Commun. 11, 2388 (2020).
Carrillo-Reid, L., Han, S., Yang, W., Akrouh, A. & Yuste, R. Controlling visually guided behavior by holographic recalling of cortical ensembles. Cell 178, 447–457.e5 (2019).
Chettih, S. N. & Harvey, C. D. Single-neuron perturbations reveal feature-specific competition in V1. Nature 567, 334–340 (2019).
Gill, J. V. et al. Precise holographic manipulation of olfactory circuits reveals coding features determining perceptual detection. Neuron 108, 382–393.e5 (2020).
Dalgleish, H. W. P. et al. How many neurons are sufficient for perception of cortical activity? Elife 9, 1–99 (2020).
Vinograd, A., Nair, A., Kim, J., Linderman, S. W. & Anderson, D. J. Causal evidence of a line attractor encoding an affective state. Nature 634, 910–918 (2024).
Forli, A. et al. Two-photon bidirectional control and imaging of neuronal excitability with high spatial resolution in vivo. Cell Rep. 22, 3087–3098 (2018).
Chen, I. W., Papagiakoumou, E. & Emiliani, V. Towards circuit optogenetics. Curr. Opin. Neurobiol. 50, 179–189 (2018).
Soor, N. S. et al. All-optical crosstalk-free manipulation and readout of Chronos-expressing neurons. J. Phys. D Appl. Phys. 52, 104002 (2019).
Liu, C., Hao, Y., Lei, B., Zhong, Y. & Kong, L. Removing crosstalk signals in neuron activity by time multiplexed excitations in a two-photon all-optical physiology system. Biomed. Opt. Express 15, 2708 (2024).
Ernst, O. P. et al. Photoactivation of channelrhodopsin. J. Biol. Chem. 283, 1637–1643 (2008).
Guo, Z. V., Hart, A. C. & Ramanathan, S. Optical interrogation of neural circuits in Caenorhabditis elegans. Nat. Methods 6, 891–896 (2009).
Stoltzfus, C. R. et al. Two-photon directed evolution of green fluorescent proteins. Sci. Rep 1–9 https://doi.org/10.1038/srep11968 (2015).
Dana, H. et al. Sensitive red protein calcium indicators for imaging neural activity. Elife 5, 1–24 (2016).
Rickgauer, J. P. & Tank, D. W. Two-photon excitation of channelrhodopsin-2 at saturation. Proc. Natl. Acad. Sci. USA 106, 15025–15030 (2009).
Papagiakoumou, E., Ronzitti, E. & Emiliani, V. Scanless two-photon excitation with temporal focusing. Nat. Methods 17, 571–581 (2020).
Akerboom, J. et al. Genetically encoded calcium indicators for multi-color neural activity imaging and combination with optogenetics. Front Mol. Neurosci. 6, 1–29 (2013).
Shen, Y. et al. A genetically encoded Ca2+ indicator based on circularly permutated sea anemone red fluorescent protein eqFP578. BMC Biol. 16, 1–16 (2018).
Inoue, M. et al. Rational engineering of XCaMPs, a multicolor GECI suite for in vivo imaging of complex brain circuit dynamics. Cell 177, 1346–1360.e24 (2019).
Yokoyama, T. et al. A multicolor suite for deciphering population coding of calcium and cAMP in vivo. Nat. Methods 21, 897–907 (2024).
Dalangin, R. et al. Far-red fluorescent genetically encoded calcium ion indicators. Nat. Commun. 16, 1–14 (2025).
Zhou, S. et al. A sensitive soma-localized red fluorescent calcium indicator for in vivo imaging of neuronal populations at single-cell resolution. PLoS Biol. 23, e3003048 (2025).
Packer, A. M. et al. Two-photon optogenetics of dendritic spines and neural circuits. Nat. Methods 9, 1202–1205 (2012).
Lin, J. Y., Knutsen, P. M., Muller, A., Kleinfeld, D. & Tsien, R. Y. ReaChR: A red-shifted variant of channelrhodopsin enables deep transcranial optogenetic excitation. Nat. Neurosci. 16, 1499–1508 (2013).
Klapoetke, N. C. et al. Independent optical excitation of distinct neural populations. Nat. Methods 11, 338–346 (2014).
Forli, A., Pisoni, M., Printz, Y., Yizhar, O. & Fellin, T. Optogenetic strategies for high-efficiency all-optical interrogation using blue-light-sensitive opsins. Elife 10, 1–25 (2021).
Sridharan, S. et al. High-performance microbial opsins for spatially and temporally precise perturbations of large neuronal networks. Neuron 110, 1139–1155.e6 (2022).
Ronzitti, E. et al. Submillisecond optogenetic control of neuronal firing with two-photon holographic photoactivation of chronos. J. Neurosci. 37, 10679–10689 (2017).
Tajkhorshid, E., Baudry, J., Schulten, K. & Suhai, S. Molecular dynamics study of the nature and origin of retinal’s twisted structure in bacteriorhodopsin. Biophys. J. 78, 683–693 (2000).
Logunov, S. L., Volkov, V. V., Braun, M. & El-Sayed, M. A. The relaxation dynamics of the excited electronic states of retinal in bacteriorhodopsin by two-pump-probe femtosecond studies. Proc. Natl. Acad. Sci. USA 98, 8475–8479 (2001).
Cembran, A., Bernardi, F., Olivucci, M. & Garavelli, M. The retinal chromophore/chloride ion pair: Structure of the photoisomerization path and interplay of charge transfer and covalent states. Proc. Natl. Acad. Sci. USA 102, 6255–6260 (2005).
Drobizhev, M., Makarov, N. S., Hughes, T. & Rebane, A. Resonance enhancement of two-photon absorption in fluorescent proteins. J. Phys. Chem. B 111, 14051–14054 (2007).
Drobizhev, M., Makarov, N. S., Tillo, S. E., Hughes, T. E. & Rebane, A. Two-photon absorption properties of fluorescent proteins. Nat. Methods 8, 393–399 (2011).
Oda, K. et al. Crystal structure of the red light-activated channelrhodopsin Chrimson. Nat. Commun. 9, 1–11 (2018).
Malakar, P. et al. Retinal photoisomerization versus counterion protonation in light and dark-adapted bacteriorhodopsin and its primary photoproduct. Nat. Commun. 15, 1–14 (2024).
Pnevmatikakis, E. A. et al. Simultaneous Denoising, Deconvolution, and Demixing of Calcium Imaging Data. Neuron 89, 285 (2016).
Ahrens, M. B. et al. Brain-wide neuronal dynamics during motor adaptation in zebrafish. Nature 485, 471–477 (2012).
Antinucci, P., Folgueira, M. & Bianco, I. H. Pretectal neurons control hunting behaviour. Elife 8, 1–34 (2019).
Thiele, T. R., Donovan, J. C. & Baier, H. Descending Control of Swim Posture by a Midbrain Nucleus in Zebrafish. Neuron 83, 679–691 (2014).
Shang, C. F. et al. Real-time analysis of large-scale neuronal imaging enables closed-loop investigation of neural dynamics. Nat. Neurosci. 27, 1014–1018 (2024).
Freeman, J. et al. Mapping brain activity at scale with cluster computing. Nat. Methods 11, 941–950 (2014).
Tian, A. G., Ka, T., Lam, C., Yan, G. & He, Y. Binocular integration of prey stimuli in the zebrafish visual system. Current Biology 35, 3228–3240.e5 (2024).
Lister, J.A., Robertson, C.P., Lepage, T., Johnson, S.L. & Raible, D.W. Nacre encodes a zebrafish microphthalmia-related protein that regulates neural-crest-derived pigment cell fate. Development 126, 3757–3767 (1999).
Pozzi, P. et al. Fast calculation of computer generated holograms for 3D photostimulation through compressive-sensing Gerchberg-Saxton algorithm. Methods Protoc. 2, 1–11 (2018).
Pnevmatikakis, E. A. & Giovannucci, A. NoRMCorre: An online algorithm for piecewise rigid motion correction of calcium imaging data. J. Neurosci. Methods 291, 83–94 (2017).
Diaspro, A., Chirico, G. & Collini, M. Two-photon fluorescence excitation and related techniques in biological microscopy. Q Rev. Biophys. 38, 97–166 (2005).
Svelto, O. Principles of lasers. Princ. Lasers https://doi.org/10.1007/978-1-4757-6266-2 (1998).
Horton, N. G. et al. In vivo three-photon microscopy of subcortical structures within an intact mouse brain. Nat. Photonics 7, 205–209 (2013).
Acknowledgements
This study was supported in part by the Research Grants Council of Hong Kong to J.Y.Q. (16102825, 16102123, 16102122, 16102421, 16102920, 16102518); Collaborative Research Fund (C6001-19E); the Innovation and Technology Commission (ITCPD/17-9); the Theme-Based Research Scheme (T12-611/25-N) and the Hong Kong University of Science & Technology through grant 30 for 30 Research Initiative Scheme, and Research Grants Council of Hong Kong to J.L.S. (16101221, 16103224, 16103522, and 16103625).
Author information
Authors and Affiliations
Contributions
G.Y., G.T., and J.Y.Q. conceived the research idea and designed the experiments. G.Y. built the imaging systems and Y.F. and Z.S. provided technical support; G.T. and K.Y.C. prepared animals; G.Y. performed imaging experiments under the supervision of J.Y.Q. and J.L.S.; G.Y. and G.T. analyzed the data with assistance from Y.H.; G.Y., G.T., Y.H., J.L.S., and J.Y.Q. wrote the paper with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks anonymous reviewers for their contribution to the peer review of this work. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yan, G., Tian, G., Fu, Y. et al. Active pixel power control for crosstalk-free all-optical neural interrogation. Nat Commun 17, 3195 (2026). https://doi.org/10.1038/s41467-026-69419-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69419-8