Abstract
The cGAS-STING pathway is a critical regulator of type I Interferon (IFN) and inflammation upon cytosolic DNA-sensing. cGAS-STING signaling termination is regulated by lysosomal-mediated degradation of STING; however, the mechanisms controlling the inhibitory targeting of STING are incompletely understood. Here, we identify the selective autophagy receptor TAX1BP1 as a negative regulator of the cGAS-STING pathway. TAX1BP1-deficient macrophages activated by cGAS or STING agonists accumulate higher-order STING aggregates, exhibit heightened STING signaling, and increased production of type I IFN and proinflammatory cytokines. Mechanistically, TAX1BP1 promotes STING degradation through microautophagy by facilitating the interaction of STING with the ESCRT-0 protein HGS. Furthermore, STING activation is associated with the swelling and fragmentation of the Golgi apparatus, and TAX1BP1 and p62/SQSTM1 are essential for the autophagic degradation of fragmented Golgi (Golgiphagy). Our findings suggest that STING activation at the Golgi is coupled to its downregulation by Golgiphagy to restrict innate immune responses.
Similar content being viewed by others
Introduction
The cGAS-STING pathway is an ancient DNA-sensing pathway critical for inflammatory responses to bacteria, DNA viruses and retroviruses1. Cyclic GMP-AMP synthase (cGAS) is a DNA sensor that binds double-stranded DNA and catalyzes the cyclic dinucleotide and second messenger 2’3’-cyclic GMP-AMP (2’3’-cGAMP; hereafter referred to as cGAMP), which binds to and activates Stimulator of Interferon Genes (STING) at the endoplasmic reticulum (ER)2,3,4. STING then translocates from the ER to the Golgi via coat protein complex II (COPII) vesicles5. At the trans-Golgi, oligomerized STING activates the kinase TBK1 and transcription factor IRF3, leading to the induction of type I interferon (IFN)6.
The activation of the cGAS-STING pathway is tightly regulated to prevent overexuberant or chronic inflammation. Upon activation at the trans-Golgi, STING is targeted for degradation in lysosomes. The selective autophagy receptor p62/SQSTM1 was previously shown to promote STING degradation7; however, the underlying mechanisms are not well understood. STING also has a primordial function, inducing autophagy independent of TBK1 activation and the induction of type I IFN8,9. STING can interact with LC3 and induce autophagy dependent on ATG5 but independent of ULK1 and Beclin18. STING-induced autophagy plays a role in the clearance of viruses and DNA from the cytoplasm. Recent studies have indicated that microautophagy is the main STING degradation pathway that terminates STING signaling. STING exits the trans-Golgi network (TGN) by adaptor protein complex 1 (AP-1)-mediated sorting into clathrin-coated vesicles and is ubiquitinated by lysine 63 (K63)-linked polyubiquitin (polyUb) chains, which are sensed by the endosomal sorting complexes required for transport (ESCRT) machinery10,11,12,13. STING vesicles are then directly encapsulated into LAMP1-positive lysosomes, resulting in STING degradation11. STING degradation is critical to prevent prolonged activation and induction of type I IFN and proinflammatory cytokines; however, the mechanisms underlying STING downregulation after its activation at the trans-Golgi are incompletely understood.
Tax1 Binding Protein 1 (TAX1BP1) was initially identified as an interacting protein of the human T-cell leukemia virus 1 (HTLV-1) Tax protein, and the NF-κB regulatory proteins A20 (also known as TNFAIP3) and TRAF614,15. TAX1BP1 was shown to inhibit NF-κB signaling by serving as an adaptor for A20 and its substrates targeted for deubiquitination16. In addition to regulating NF-κB, TAX1BP1 also inhibits the RIG-I-like receptor (RLR) pathway and the induction of type I IFN triggered by RNA virus infection17,18. TAX1BP1 also blocks RLR-mediated apoptosis by interacting with MAVS and promoting its degradation19. In the absence of TAX1BP1, MAVS aggregates spontaneously form and accumulate further with RNA virus infection20. In addition, TAX1BP1 suppresses the Toll-like receptor (TLR) 3/4 pathways by targeting the adaptor TRIF for degradation21,22. TAX1BP1 inhibits NF-κB and innate immune signaling pathways to quell inflammation and prevent autoimmunity since TAX1BP1 expression was shown to be downregulated in patients with systemic lupus erythematosus (SLE)23. TAX1BP1 functions as a selective autophagy receptor that contains two LC3 interaction regions (LIR) in the amino (N)-terminus and a ubiquitin binding zinc finger domain 2 (ZnF2) in the carboxy (C)-terminus24,25,26,27. TAX1BP1 detects ubiquitinated cargo via its ZnF2 domain, which is then recruited to developing autophagosomes via its LIR domains. TAX1BP1 also interacts with myosin VI, a cytoskeletal actin-based motor protein regulating vesicular transport, to induce autophagosome maturation26. Therefore, TAX1BP1 exerts multiple roles in autophagy, including cargo selection and autophagosome maturation. Although TAX1BP1 has been shown to inhibit NF-κB, RLR and TLR signaling, a potential role of TAX1BP1 in the regulation of the cGAS-STING pathway has not yet been investigated.
In this study, we identify TAX1BP1 as a negative regulator of the cGAS-STING pathway by targeting STING for degradation through autophagy of the Golgi apparatus (Golgiphagy) and ESCRT-mediated microautophagy. Loss of TAX1BP1 impairs STING degradation, leading to the accumulation of fragmented Golgi and STING aggregates and increased production of type I IFN and proinflammatory cytokines. Mice lacking TAX1BP1 in myeloid cells exhibit enhanced STING-mediated inflammatory responses and are resistant to poxvirus infection. These findings indicate that TAX1BP1 targets STING by Golgiphagy and microautophagy to limit the magnitude and duration of innate immune activation.
Results
TAX1BP1 inhibits cGAS-STING-mediated type I IFN and cytokine activation
We previously generated TAX1BP1-deficient mice using a gene trapping strategy but these mice were embryonic lethal, thus precluding in vivo studies16. To understand the physiological roles of TAX1BP1 we generated “floxed” conditional knockout mice on a C57BL/6 background. Embryonic stem (ES) cells were obtained with exon 4 of Tax1bp1 flanked by LoxP sites to facilitate excision by Cre recombinase (Supplementary Fig. 1). Tax1bp1fl/fl mice were crossed with LysM-Cre mice to delete TAX1BP1 in myeloid cells. We generated bone marrow-derived macrophage (BMDM) cells from control LysM-Cre mice and Tax1bp1fl/flLysM-Cre mice (Tax1bp1–/–). Tax1bp1 mRNA and protein were undetectable in Tax1bp1–/– BMDMs as shown by real-time qRT-PCR and immunoblotting (Supplementary Fig. 2A, B). We also deleted TAX1BP1 in the human monocytic cell line, THP-1, using CRISPR-Cas9, and confirmed knockout by real-time qRT-PCR and immunoblotting (Supplementary Fig. 2C, D). We next sought to determine if the cGAS-STING pathway was dysregulated in TAX1BP1-KO THP-1 cells and Tax1bp1–/– BMDMs.
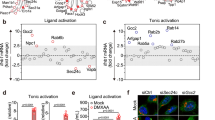
TAX1BP1-KO THP-1 cells were transfected with G3-YSD, a Y-form 26-mer DNA sequence from the HIV-1 genome, and cGAS-dependent agonist. TAX1BP1-KO THP-1 cells transfected with G3-YSD expressed significantly more IFN-β and IL-6 mRNA and protein compared to control THP-1 cells expressing a control nontargeting gRNA as shown by real-time qRT-PCR and ELISA (Fig. 1A-D). To determine if TAX1BP1 inhibits the pathway at or downstream of STING, we next permeabilized control and TAX1BP1-KO THP-1 cells and stimulated with the STING agonist cGAMP. TAX1BP1-KO THP-1 cells expressed significantly more IFN-β and IL-6 mRNA and protein compared to control cells after cGAMP stimulation (Fig. 1E-H). Tax1bp1–/– BMDMs similarly exhibited heightened IFN-β and IL-6 induction with both G3-YSD transfection and cGAMP stimulation (Fig. 1I-L). Therefore, TAX1BP1 is a negative regulator of the cGAS-STING pathway that can inhibit STING induction of downstream target genes.
A Real-time qRT-PCR of Ifnβ mRNA in control and TAX1BP1-KO THP-1 cells transfected with G3-YSD for the indicated times. P values from left to right: P > 0.9999, P = 0.7867, P < 0.0001, P < 0.0001, P < 0.0001, P = 0.0504, P = 0.0365. B IFN-β ELISA using supernatants from control and TAX1BP1-KO THP-1 cells transfected with G3-YSD for the indicated times. P values from left to right: P > 0.9999, P < 0.0001, P < 0.0001, P < 0.0001. C Real-time qRT-PCR of Il6 mRNA in control and TAX1BP1-KO THP-1 cells transfected with G3-YSD for the indicated times. P values from left to right: P > 0.9999, P > 0.9999, P > 0.9999, P < 0.0001, P < 0.0001, P = 0.2985, P = 0.1396. D IL-6 ELISA using supernatants from control and TAX1BP1-KO THP-1 cells transfected with G3-YSD for the indicated times. P values from left to right: P = 0.9991, P < 0.0001, P < 0.0001, P < 0.0001. E Real-time qRT-PCR of Ifnβ mRNA in control and TAX1BP1-KO THP-1 cells treated with cGAMP for the indicated times. P values from left to right: P > 0.9999, P = 0.7159, P = 0.9850, P = 0.0005, P = 0.6471, P = 0.9978, P > 0.9999. F IFN-β ELISA using supernatants from control and TAX1BP1-KO THP-1 cells treated with cGAMP for the indicated times. P = 0.9873, P < 0.0001, P < 0.0001, P = 0.0004. G Real-time qRT-PCR of Il6 mRNA in control and TAX1BP1-KO THP-1 cells treated with cGAMP for the indicated times. P values from left to right: P > 0.9999, P > 0.9999, P = 0.2322, P = 0.0771, P = 0.0005, P < 0.0001, P = 0.0007. H IL-6 ELISA using supernatants from control and TAX1BP1-KO THP-1 cells treated with cGAMP for the indicated times. P values from left to right: P = 0.9933, P < 0.0001, P < 0.0001, P < 0.0001. I Real-time qRT-PCR of Ifnβ mRNA in control and Tax1bp1–/– BMDMs transfected with G3-YSD for the indicated times. P values from left to right: P > 0.9999, P < 0.0001, P = 0.9098. J Real-time qRT-PCR of Il6 mRNA in control and Tax1bp1–/– BMDMs transfected with G3-YSD for the indicated times. P values from left to right: P > 0.9999, P < 0.0001, P = 0.0772. K Real-time qRT-PCR of Ifnβ mRNA in control and Tax1bp1–/– BMDMs treated with cGAMP for the indicated times. P values from left to right: P > 0.9999, P > 0.9999, P < 0.0001, P < 0.0001, P < 0.0001, P = 0.9345, P > 0.9999. L Real-time qRT-PCR of Il6 mRNA in control and Tax1bp1–/– BMDMs treated with cGAMP for the indicated times. P values from left to right: P > 0.9999, P > 0.9999, P = 0.9992, P < 0.0001, P < 0.0001, P = 0.0024, P > 0.9999. The results are expressed as the mean ± SD (n = 3). ****P < 0.0001; ***P < 0.001; **P < 0.01; *P < 0.05; ns = not significant. Two-way ANOVA with Sidak’s multiple comparisons test was used for experiments in (A–L). M Incucyte S3 live-cell imaging of control and Tax1bp1–/– Mx1-GFP BMDMs transfected with G3-YSD. The results are expressed as the mean ± SD (n = 6). N Incucyte S3 live-cell imaging of control and Tax1bp1–/– Mx1-GFP BMDMs treated with diABZI. The results are expressed as the mean ± SD (n = 6). Source data are provided as a Source data file.
To examine type I IFN responses, Tax1bp1fl/flLysM-Cre mice were crossed with interferon-stimulated response element (ISRE) reporter mice (Mx1-GFP)28, which have GFP knocked into the Mx1 locus, and thus GFP expression can be used to monitor type I IFN responses at the single-cell level. BMDMs were generated from Tax1bp1fl/flLysM-Cre-Mx1-GFP (Tax1bp1–/–Mx1-GFP) and control LysM-Cre-Mx1-GFP mice and were either transfected with G3-YSD to activate cGAS or stimulated with the STING agonist diABZI (compound 3). Incucyte S3 live-cell imaging was used to detect GFP expression in real-time after stimulation. Significantly more GFP was detected in Tax1bp1–/–Mx1-GFP BMDMs than control BMDMs transfected with G3-YSD or stimulated with diABZI (Fig. 1M, N). We also examined the expression of IFN-α4 and interferon-stimulated genes (ISG) by real-time qRT-PCR in TAX1BP1-KO THP-1 cells and Tax1bp1–/– BMDMs after cGAS-STING activation. TAX1BP1-KO THP-1 cells expressed significantly more Mx1, Isg15, Irf7 and Ifna4 mRNAs compared to control cells after transfection with G3-YSD, herring testes DNA (HT-DNA) or stimulation with cGAMP (Supplementary Fig. 3A-N). Tax1bp1–/– BMDMs similarly expressed more Mx1, Isg15, Irf7 and Ifna4 mRNAs compared to control BMDMs after stimulation with G3-YSD or cGAMP (Supplementary Fig. 3O-V).
TAX1BP1-deficient macrophages have heightened expression of HSV-1-induced type I IFN and ISGs
The cGAS-STING pathway is activated in response to herpes simplex virus type 1 (HSV-1) infection to induce type I IFN, although HSV-1 can antagonize cGAS-STING activation29,30. Control and TAX1BP1-KO THP-1 cells were infected with HSV-1 expressing yellow fluorescent protein (HSV-1-YFP) and subjected to Incucyte S3 live-cell imaging to monitor infection in real-time. TAX1BP1-KO THP-1 cells exhibited significantly decreased yellow fluorescence compared to control cells indicating that TAX1BP1-KO THP-1 cells are more resistant to HSV-1 infection (Fig. 2A). TAX1BP1-KO THP-1 cells infected with HSV-1 expressed more Ifnβ mRNA and protein (Fig. 2B, C) compared to control cells, which is likely mediating the increased resistance. HSV-1-infected TAX1BP1-KO THP-1 cells also expressed more Ifnα4, Mx1, Isg15 and Irf7 mRNAs compared to control HSV-1-infected cells (Fig. 2D-G).
A Incucyte S3 live-cell imaging of control and TAX1BP1-KO THP-1 cells infected with HSV-1 YFP. The results are expressed as the mean ± SD (n = 6). B Real-time qRT-PCR of Ifnβ mRNA in control and TAX1BP1-KO THP-1 cells infected with HSV-1 for the indicated times. P values from left to right: P > 0.9999, P = 0.8342, P < 0.0001, P = 0.0570. C IFN-β ELISA using supernatants from control and TAX1BP1-KO THP-1 cells infected with HSV-1 for the indicated times. P values from left to right: P > 0.9999, P = 0.2968, P = 0.8105, P < 0.0001. D Real-time qRT-PCR of Mx1 mRNA in control and TAX1BP1-KO THP-1 cells infected with HSV-1 for the indicated times. P values from left to right: P > 0.9999, P = 0.0940, P < 0.0001, P = 0.0174. E Real-time qRT-PCR of Isg15 mRNA in control and TAX1BP1-KO THP-1 cells infected with HSV-1 for the indicated times. P values from left to right: P > 0.9999, P = 0.2163, P < 0.0001, P = 0.3577. F Real-time qRT-PCR of Irf7 mRNA in control and TAX1BP1-KO THP-1 cells infected with HSV-1 for the indicated times. P values from left to right: P > 0.9999, P = 0.1352, P = 0.0015, P = 0.2259. G Real-time qRT-PCR of IFNα4 mRNA in control and TAX1BP1-KO THP-1 cells infected with HSV-1 for the indicated times. P values from left to right: P > 0.9999, P = 0.9008, P < 0.0001, P = 0.0521. The results are expressed as the mean ± SD (n = 3). ****P < 0.0001; ***P < 0.001; **P < 0.01; *P < 0.05; ns = not significant. Two-way ANOVA with Sidak’s multiple comparisons test was used for experiments in panels B-G. Source data are provided as a Source data file.
Enhanced STING signaling and impaired STING degradation in TAX1BP1-deficient macrophages
STING activation induces the expression of type I IFN and proinflammatory cytokines via activation of TBK1-IRF3 and IKK-NF-κB pathways31,32,33. Therefore, we next examined the activation of TBK1-IRF3 and IKK-NF-κB downstream of STING by immunoblotting with phospho-specific antibodies after stimulation with diABZI. There was enhanced and prolonged diABZI-induced TBK1 and IRF3 activation in TAX1BP1-KO THP-1 cells compared to control cells (Fig. 3A, C, D). Similarly, there was more robust diABZI-induced IKKα/β phosphorylation in TAX1BP1-KO THP-1 cells compared to control cells (Fig. 3A, G). After binding to cGAMP, STING is phosphorylated by TBK1 on Ser366 to promote activation of IRF3 and induction of type I IFN6. STING phosphorylation was stronger and more persistent in diABZI-stimulated TAX1BP1-KO THP-1 cells compared to control cells (Fig. 3A, E). Interestingly, STING degradation was partially impaired in TAX1BP1-KO THP-1 cells treated with diABZI (Fig. 3A, F). We next examined STING signaling induced by HSV-1 infection of control and TAX1BP1-KO THP-1 cells. There was enhanced HSV-1-induced TBK1, IRF3, STING and IKKα/β phosphorylation in TAX1BP1-KO THP-1 cells compared to control cells (Fig. 3B, H-L). Therefore, TAX1BP1 inhibits cGAS-STING-mediated signaling induced by diABZI stimulation and HSV-1 infection.
A Immunoblotting with the indicated antibodies using lysates from control and TAX1BP1-KO THP-1 cells treated with diABZI for the indicated times. The experiment is representative of three independent experiments with similar results. B Immunoblotting with the indicated antibodies using lysates from control and TAX1BP1-KO THP-1 cells infected with HSV-1 for the indicated times. The experiment is representative of three independent experiments with similar results. C Quantification of p-TBK1/TBK1 in WT and TAX1BP1-KO THP-1 cells treated with diABZI from three independent experiments. P values from left to right: P > 0.9999, P = 0.9421, P = 0.0026, P = 0.0031, P = 0.1855, P = 0.4814. D Quantification of p-IRF3/IRF3 in WT and TAX1BP1-KO THP-1 cells treated with diABZI from three independent experiments. P values from left to right: P > 0.9999, P > 0.9999, P = 0.0380, P = 0.9870, P = 0.9790, P > 0.9999. E Quantification of p-STING/STING in WT and TAX1BP1-KO THP-1 cells treated with diABZI from three independent experiments. P values from left to right: P > 0.9999, P = 0.8248, P = 0.0006, P = 0.0413, P = 0.1932, P > 0.9042. F Quantification of STING/Vinculin in WT and TAX1BP1-KO THP-1 cells treated with diABZI from three independent experiments. P values from left to right: P = 0.4754, P = 0.3446, P = 0.0017, P = 0.0182, P = 0.7086, P = 0.6129. G Quantification of p-IKKα/β/IKKβ in WT and TAX1BP1-KO THP-1 cells treated with diABZI from three independent experiments. P values from left to right: P > 0.9999, P = 0.9998, P > 0.9999, P = 0.0355, P = 0.1342, P > 0.9999. H Quantification of p-TBK1/TBK1 in WT and TAX1BP1-KO THP-1 cells infected with HSV-1 from three independent experiments. P values from left to right: P = 0.7489, P = 0.0007, P = 0.1135, P = 0.1434. I Quantification of p-IRF3/IRF3 in WT and TAX1BP1-KO THP-1 cells infected with HSV-1 from three independent experiments. P values from left to right: P = 0.5191, P = 0.0017, P = 0.0007, P = 0.0433. J Quantification of p-STING/STING in WT and TAX1BP1-KO THP-1 cells infected with HSV-1 from three independent experiments. P values from left to right: P = 0.9951, P = 0.0164, P = 0.0192, P = 0.5873. K Quantification of STING/Vinculin in WT and TAX1BP1-KO THP-1 cells infected with HSV-1 from three independent experiments. P values from left to right: P = 0.4272, P = 0.9871, P = 0.9598, P = 0.6638. L Quantification of p-IKKα/β/IKKβ in WT and TAX1BP1-KO THP-1 cells infected with HSV-1 from three independent experiments. P values from left to right: P > 0.9999, P = 0.4109, P = 0.0021, P = 0.1372. The results are expressed as the mean ± SD (n = 3). ****P < 0.0001; ***P < 0.001; **P < 0.01; *P < 0.05; ns = not significant. Two-way ANOVA with Sidak’s multiple comparisons test was used for experiments in (C–L). M Immunoblotting with the indicated antibodies using lysates from control (LysM-Cre) and Tax1bp1–/– (Tax1bp1fl/flLysM-Cre) BMDMs treated with diABZI for the indicated times. The experiment is representative of three independent experiments with similar results. N Immunoblotting with anti-LC3 using lysates from WT and TAX1BP1-KO THP-1 cells treated with diABZI for the indicated times. The experiment is representative of three independent experiments with similar results. Source data are provided as a Source data file.
STING degradation is a critical mechanism that downregulates its activation to ensure a transient type I IFN response. STING degradation mainly occurs through ESCRT-mediated microautophagy where clathrin-coated STING vesicles egress from the trans-Golgi and become directly encapsulated in lysosomes to degrade STING11,12,13. We next examined STING degradation in Tax1bp1–/– BMDMs treated with diABZI. As expected, in control BMDMs, STING was degraded beginning at 2 h post-stimulation and was almost completely degraded by 6 h (Fig. 3M). However, in Tax1bp1–/– BMDMs treated with diABZI, STING degradation was partially impaired (Fig. 3M). Lipidation of Atg8 family members (i.e., LC3B) is a hallmark of autophagosome biogenesis and lipidated LC3 (LC3-II) migrates faster compared to unlipidated (LC3-I) on SDS-PAGE gels. STING can induce the lipidation of ATG8 family members onto single-membrane vesicles in a process known as conjugation of ATG8 to single membranes (CASM)34. However, STING-induced LC3 lipidation was comparable in WT and Tax1bp1–/– BMDMs (Fig. 3M) and WT and TAX1BP1-KO THP-1 cells treated with diABZI (Fig. 3N). Therefore, loss of TAX1BP1 has no effect on STING-mediated LC3B lipidation.
To further examine the role of TAX1BP1 in STING degradation, we next performed a quantitative STING degradation flow cytometry assay using control THP-1 and TAX1BP1-KO THP-1 cells stably expressing STING-eGFP (Supplementary Fig. 4). Treatment of control THP-1 STING-eGFP cells with diABZI for 0, 2, 4, 6 and 8 h led to a progressive loss of eGFP fluorescence over time due to STING-eGFP degradation (Fig. 4A, B). However, diABZI-induced STING-eGFP degradation was significantly impaired in TAX1BP1-KO THP-1 cells (Fig. 4A, B). To determine the contribution of microautophagy to STING degradation in these cells, we used siRNA to knock down the ESCRT-I protein TSG101, which was previously shown to be important for STING degradation11. As expected, knockdown of TSG101 largely impaired diABZI-induced STING degradation in control THP-1 cells (Fig. 4C, D). STING degradation was also impaired in TAX1BP1-KO THP-1 cells, and TSG101 knockdown completely blocked STING degradation in these cells (Fig. 4C, D). Knockdown of TSG101 and degradation of STING-eGFP were confirmed by immunoblotting (Fig. 4E). These data suggest that the residual degradation of STING in the absence of TAX1BP1 occurs through microautophagy.
A Quantitative flow cytometry assay to examine STING-eGFP degradation. Control and TAX1BP1-KO THP-1 cells stably expressing eGFP were treated with diABZI for the indicated times and subjected to flow cytometry for eGFP fluorescence. The experiment is representative of three independent experiments with similar results. B Quantification of eGFP fluorescence from (A). P values from left to right: P = 0.9982, P = 0.0067, P < 0.0001, P < 0.0001, P < 0.0001. C Control and TAX1BP1-KO THP-1 cells stably expressing eGFP were transfected with control or TSG101 siRNAs for 48 h and then treated with diABZI for the indicated times and subjected to flow cytometry for eGFP fluorescence. The experiment is representative of three independent experiments with similar results. D Quantification of eGFP fluorescence from (C). P values from left to right: P > 0.9999, P > 0.9999, P > 0.9999, P < 0.0001, P < 0.0001, P < 0.9944, P < 0.0001, P < 0.0001, P < 0.0002. The results are expressed as the mean ± SD (n = 3). ****P < 0.0001; ***P < 0.001; **P < 0.01; *P < 0.05; ns = not significant. Two-way ANOVA with Sidak’s multiple comparisons test was used for experiments in (B, D). E Immunoblotting with the indicated antibodies using protein lysates from (C). The experiment is representative of three independent experiments with similar results. F Immunofluorescence confocal microscopy with anti-HGS-Alexa Fluor 594 (pseudo magenta) in control and TAX1BP1-KO HeLa cells stably expressing STING-eGFP and treated with diABZI for 4 h. Scale bar: 10 μM. G Quantification of STING and HGS colocalization by Manders’ colocalization coefficient. P values from left to right: P = 0.6727, P < 0.0001, P < 0.0001, P < 0.0001. The results are expressed as the mean ± SD (n = 30 cells). ****P < 0.0001; ns = not significant. Two-way ANOVA with Tukey’s multiple comparisons test was used. H Co-IP assay with either control IgG or anti-STING and immunoblotting with HGS using lysates from THP-1 cells treated with diABZI for 4 h. The experiment is representative of three independent experiments with similar results. Source data are provided as a Source data file.
We next sought to determine if TAX1BP1 played a role in STING regulation by microautophagy. Given that p62/SQSTM1 was previously shown to colocalize with STING and the ESCRT-0 protein HGS12, we hypothesized that TAX1BP1, STING and HGS were in the same complex and TAX1BP1 linked ubiquitinated STING with the ESCRT machinery. Control HeLa and HeLa TAX1BP1-KO cells stably expressing STING-eGFP were generated for imaging studies (Supplementary Fig. 5C). We performed immunofluorescence microscopy using control HeLa-STING-eGFP and HeLa TAX1BP1-KO STING-eGFP cells stained with anti-HGS. There was no colocalization of STING and HGS in unstimulated cells, but the STING-HGS colocalization was strongly induced by diABZI stimulation (Fig. 4F, G). However, in TAX1BP1-KO HeLa cells the STING-HGS colocalization was partially impaired (Fig. 4F, G). Furthermore, a diABZI-induced STING-HGS interaction was detected by a co-IP assay, and the interaction was decreased in TAX1BP1-KO THP-1 cells (Fig. 4H). These data suggest that TAX1BP1 promotes an interaction of STING with the ESCRT machinery.
TAX1BP1 interacts with STING
We next examined whether endogenous TAX1BP1 could interact with STING. A co-immunoprecipitation (co-IP) assay was performed using WT THP-1 cells treated with diABZI in a time-course experiment. We immunoprecipitated with anti-TAX1BP1 and immunoblotted for STING. TAX1BP1 interacted with STING under basal conditions and the interaction further increased upon diABZI stimulation (Fig. 5A). The reciprocal IP was also performed (IP: anti-STING; IB: anti-TAX1BP1) and TAX1BP1 was detected in STING immunoprecipitates which was further enhanced with diABZI treatment (Fig. 5A). A proximity ligation assay (PLA) also confirmed basal TAX1BP1-STING interaction that was significantly increased by diABZI stimulation (Fig. 5B, C). Immunofluorescence and confocal microscopy were next performed to determine the cellular location of TAX1BP1-STING interaction. Since STING-induced TBK1 activation occurs at the trans-Golgi we stained cells with a marker of the trans-Golgi, TGN46. TAX1BP1 colocalized with STING, predominantly at the trans-Golgi, after diABZI stimulation (Fig. 5D, E). Together, these results suggest that TAX1BP1 interacts with STING basally, and this interaction is enhanced upon STING activation.
A Co-IP assay with either control IgG, anti-TAX1BP1 or anti-STING using lysates from WT THP-1 cells treated with diABZI for the indicated times. The experiment is representative of three independent experiments with similar results. B Proximity ligation assay with anti-STING and anti-TAX1BP1 in WT THP-1 cells either untreated or treated with diABZI for 2 h. Scale bar: 10 μM. C Quantification of number of PLA puncta from panel B (n = 10). P < 0.0001. The results are expressed as the mean ± SD (n = 10 cells). ****P < 0.0001; ns = not significant. Unpaired Student’s t test was used. D Immunofluorescence confocal microscopy with anti-TAX1BP1-Alexa Fluor 647 (pseudo magenta) and anti-TGN46-CL594 (pseudo yellow) in WT STING-eGFP HeLa cells treated with diABZI for 2 h. Scale bar: 10 μM. E Quantification of TAX1BP1-Alexa Fluor 647 (pseudo magenta) and STING (eGFP) colocalization by Manders’ colocalization coefficient. P < 0.0001. The results are expressed as the mean ± SD (n = 30 cells). ****P < 0.0001; ns = not significant. Unpaired t test was used. F Schematic of Lg-BiT and Sm-BiT TAX1BP1 and STING fusions used for the NanoBiT assay. G NanoBiT assay with the indicated LgBiT and Sm-BiT fusion proteins. P < 0.0001. One-way ANOVA, ****p < 0.0001, ns = not significant. The results are expressed as the mean ± SD (n = 6 from two independent experiments). Immunoblotting with the indicated antibodies using protein lysates from 293 T cells transfected with NanoBiT plasmids. The experiment is representative of two independent experiments. Source data are provided as a Source data file.
To determine if the interaction between TAX1BP1 and STING was direct we next performed a NanoBiT complementation assay. In this assay, the luminescent protein NanoLuc is split into two subunits, Small BiT (SmB) and Large BiT (LgB) which interact very weakly. The interaction of target proteins tagged with the subunits brings SmB and LgB into an active luminescent complex35. Fusion of TAX1BP1 with LgB and STING with SmBiT resulted in luciferase activity (Fig. 5F, G), suggesting that TAX1BP1 and STING can directly interact.
TAX1BP1 promotes STING localization to lysosomes
Because STING degradation was partially impaired in both TAX1BP1-KO THP-1 cells and Tax1bp1–/– BMDMs treated with diABZI (Figs. 3, 4), we hypothesized that TAX1BP1 regulated STING targeting to lysosomes. Immunofluorescence and confocal microscopy experiments were next performed using control HeLa STING-eGFP and HeLa TAX1BP1-KO STING-eGFP cells to examine STING colocalization with the trans-Golgi marker Golgin-97 and the lysosomal marker LAMP2. STING trafficking proceeded as expected in control HeLa cells as STING colocalization with Golgin-97 increased 30 min post-diABZI treatment followed by a subsequent increase in colocalization with the lysosome (LAMP2) at later times (Fig. 6A, C, D, Supplementary Fig. 5A). However, in HeLa TAX1BP1-KO STING-eGFP cells, STING colocalization with Golgin-97 was enhanced at 1 and 2 h post-stimulation with diABZI, but STING colocalization with LAMP2 was diminished in TAX1BP1-KO HeLa cells treated with diABZI (Fig. 6A, C, D, Supplementary Fig. 5A).
A Immunofluorescence confocal microscopy with anti-Golgin-97-Alexa Fluor 594 (pseudo magenta) and anti-LAMP2-Alexa Fluor 647 (pseudo yellow) in WT STING-eGFP HeLa and TAX1BP1-KO STING-eGFP HeLa cells treated with diABZI and leupeptin for the indicated times. Scale bar: 10 μM. Uncropped single-color images are shown in Supplementary Fig. 5. B Immunofluorescence confocal microscopy with anti-TfnR-Alexa Fluor 594 (pseudo magenta) and anti-LAMP2-Alexa Fluor 647 (pseudo yellow) in WT STING-eGFP HeLa and TAX1BP1-KO STING-eGFP HeLa cells treated with diABZI and leupeptin for the indicated times. Scale bar: 10 μM. Uncropped single-color images are shown in Supplementary Fig. 5. C Quantification of the colocalization of STING (eGFP) with Golgin-97-Alexa Fluor 594 (pseudo magenta) using Manders’ colocalization coefficients. P values from left to right: P = 0.9994, P > 0.9431, P > 0.0035, P < 0.0001. D Quantification of the colocalization of STING (eGFP) with LAMP2-Alexa Fluor 647 (pseudo yellow) using Manders’ colocalization coefficients. P values from left to right: P = 0.9999, P > 0.9995, P > 0.0133, P < 0.0002. E Quantification of the colocalization of STING (eGFP) with TfnR-Alexa Fluor 594 (pseudo magenta) using Manders’ colocalization coefficients. P values from left to right: P = 0.9024, P > 0.9999, P = 0.0257, P < 0.0002. The results are expressed as the mean ± SD (n = 25 cells). ****P < 0.0001; ***P < 0.001; **P < 0.01; *P < 0.05; ns = not significant. Two-way ANOVA with Sidak’s multiple comparisons test was used for experiments in (C–E). Source data are provided as a Source data file.
Upon activation at the Golgi, STING is sorted into clathrin-coated vesicles derived from the recycling endosome (RE)10,11. To further define the block in STING trafficking in TAX1BP1-KO cells, we next examined STING trafficking to REs using transferrin receptor (TfnR) as a marker11. As expected, in control HeLa STING-eGFP cells STING colocalization with TfnR increased with 1 and 2 h diABZI treatment (Fig. 6B, E, Supplementary Fig. 5B). However, in HeLa TAX1BP1-KO STING-eGFP cells, STING colocalization with TfnR after 1 and 2 h diABZI stimulation was significantly decreased (Fig. 6B, E). Taken together, TAX1BP1 regulates the targeting of STING from the trans-Golgi to REs and lysosomes, and in the absence of TAX1BP1, STING accumulates at the trans-Golgi.
TAX1BP1 clears STING aggregates
TAX1BP1 functions as an aggrephagy receptor and clears protein aggregates together with p62/SQSTM136. To examine protein aggregates in TAX1BP1-deficient macrophages, we used PROTEOSTAT dye which emits red fluorescence upon binding to protein aggregates37. Control and TAX1BP1-KO THP-1 cells were incubated with PROTEOSTAT and subjected to flow cytometry to quantify fluorescence. As a positive control, cells were treated with the proteasome inhibitor MG132 to promote the accumulation of protein aggregates. Interestingly, there was more PROTEOSTAT staining in TAX1BP1-KO THP-1 cells after MG132 treatment suggesting more protein aggregates in the absence of TAX1BP1 (Fig. 7A, B, Supplementary Fig. 6). There were also more protein aggregates in TAX1BP1-KO THP-1 cells infected with HSV-1 or stimulated with cGAMP (Fig. 7A, B). We next examined STING aggregates using an SDD-AGE assay, an agarose gel-based electrophoresis assay that enables detection of large molecular weight protein complexes. SDD-AGE has previously been used to examine STING aggregates38,39. Indeed, STING aggregates were increased in TAX1BP1-KO THP-1 cells infected with HSV-1 compared to control cells (Fig. 7C). Control and TAX1BP1-KO THP-1 cells were also treated with diABZI, and STING aggregates were examined by SDD-AGE. STING aggregates were also increased in TAX1BP1-KO THP-1 cells upon diABZI stimulation (Fig. 7D). Together, these data suggest that TAX1BP1 functions as an aggrephagy receptor in the cGAS-STING pathway, and STING aggregates accumulate in the absence of TAX1BP1.
A PROTEOSTAT detection of protein aggregates by flow cytometry in control and TAX1BP1-KO THP-1 cells infected with HSV-1 for the indicated times. MG132 was added as a positive control. B PROTEOSTAT detection of protein aggregates by flow cytometry in control and TAX1BP1-KO THP-1 cells treated with cGAMP for the indicated times. MG132 was added as a positive control. C Immunoblotting with the indicated antibodies following SDD-AGE and SDS-PAGE using lysates from control and TAX1BP1-KO THP-1 cells infected with HSV-1 for the indicated times. D Immunoblotting with the indicated antibodies following SDD-AGE and SDS-PAGE using lysates from control and TAX1BP1-KO THP-1 cells stimulated with diABZI for the indicated times. The experiment is representative of three independent experiments with similar results in panels A-D. Source data are provided as a Source data file.
TAX1BP1 requires LC3, FIP200/TBK1 and Ub binding to inhibit STING
TAX1BP1 contains two N-terminal LIRs, three central coiled-coil domains and two C-terminal ZnF domains, of which ZnF2 binds to polyUb chains (Supplementary Fig. 7A). A recent study demonstrated that a TAX1BP1 point mutant (A114Q) was impaired in the interaction with the ULK1 scaffold protein FIP200 as well as TBK140. Point mutations were introduced in TAX1BP1 expressed in a lentiviral vector to disrupt the two LIRs (W49A, V143S), FIP200/TBK1 binding (A114Q) and polyUb binding (C760A). Silent mutations were also made in the cDNA to render TAX1BP1 resistant to the gRNA expressed in the CRISPR knockout THP-1 cells. Lentiviruses expressing WT TAX1BP1 or the indicated mutants were used to transduce TAX1BP1-KO THP-1 cells. Cells were then treated with diABZI and the expression of IFN-β and IL-6 was examined by real-time qRT-PCR. WT TAX1BP1 fully rescued the inhibition of STING-induced IFN-β and IL-6, which was comparable to control THP-1 cells (Supplementary Fig. 7B-D). The TAX1BP1 FIP200/TBK1 (A114Q), LIR (W49A, V143S) and ubiquitin binding (C760A) mutants were all impaired to varying degrees for the inhibition of STING-induced IFN-β and IL-6 (Supplementary Fig. 7B-D). Thus, TAX1BP1 interaction with LC3, Ub, FIP200 and/or TBK1 contributes to the inhibition of STING.
STING induces TAX1BP1-dependent Golgiphagy
We noticed that the Golgi appeared more fragmented in TAX1BP1-KO cells compared to control cells treated with diABZI (Fig. 6); therefore, we hypothesized that STING activation triggered Golgi stress and fragmentation which then elicited TAX1BP1-dependent autophagy of the Golgi apparatus (Golgiphagy). To test this hypothesis, we performed immunoblotting using trans-Golgi (Golgin-97) and cis-Golgi (GM130) markers. In control THP-1 cells treated with diABZI, there was degradation of both Golgin-97 and GM130; however, this degradation was not evident in TAX1BP1-KO THP-1 cells (Fig. 8A). Similar results were obtained in WT and Tax1bp1–/– BMDMs treated with diABZI (Fig. 8B). However, Golgiphagy induction by golgicide A, an inhibitor of cis-Golgi ADP ribosylation factor guanine nucleotide exchange factor (ARFGEF) GBF1 and Golgi dispersal agent, promoted the degradation of Golgi proteins independently of TAX1BP1 (Fig. 8A, B). These results indicate that TAX1BP1 is specific for STING-induced Golgiphagy and is not required for Golgiphagy induced by golgicide A.
A Immunoblotting with the indicated antibodies using lysates from control and TAX1BP1-KO THP-1 cells treated with diABZI alone or diABZI+BafA1 for the indicated times. The experiment is representative of three independent experiments with similar results. B Immunoblotting with the indicated antibodies using lysates from control (LysM-Cre) and Tax1bp1–/– (Tax1bp1fl/flLysM-Cre) BMDMs treated with diABZI alone or diABZI+BafA1 for the indicated times. The experiment is representative of three independent experiments with similar results. C Immunofluorescence confocal microscopy with control and TAX1BP1-KO HeLa cells stably expressing a Golgiphagy reporter (pseudo magenta for mCherry) either untreated or treated with diABZI for 4 h. D Quantification of the number of mCherry+ eGFP– puncta in (C) (n = 25). P < 0.0001. The results are expressed as the mean ± SD (n = 25 cells). ****P < 0.0001; ns = significant. Two-way ANOVA with Sidak’s multiple comparisons test was used. E Immunoblotting with the indicated antibodies using lysates from control and p62-KO THP-1 cells. F Immunoblotting with the indicated antibodies using lysates from control and p62-KO THP-1 cells stimulated with diABZI for the indicated times. G Immunoblotting with the indicated antibodies using lysates from control and STING-KO THP-1 cells stimulated with diABZI for the indicated times. The experiment is representative of three independent experiments with similar results. Source data are provided as a Source data file.
To determine if diABZI-induced degradation of Golgi proteins was mediated by LC3-dependent autophagy, we performed immunofluorescence and confocal microscopy experiments using control HeLa STING-eGFP and HeLa TAX1BP1-KO STING-eGFP cells. Cells were treated with diABZI and bafilomycin A1 (BafA1) to block autophagic flux and then stained with anti-Golgin-97 and anti-LC3 to visualize Golgi and LC3+ autophagosomes, respectively. As expected, STING colocalized with Golgin-97 in control cells treated with diABZI, and this colocalization was enhanced in TAX1BP1-KO cells (Supplementary Fig. 8A). However, LC3 did not colocalize with Golgin-97 in control or TAX1BP1-KO cells, and there was only partial colocalization of LC3 with STING in control cells that was comparable to TAX1BP1-KO cells (Supplementary Fig. 8A-C). Therefore, TAX1BP1 does not appear to target Golgi or STING to LC3+ autophagosomes.
We next generated a Golgiphagy tandem fluorescence reporter which expresses mCherry and EGFP targeted to the Golgi by a minimal Golgi targeting sequence and alternative open reading frame (altORF) within the mRNA of the centromere protein CENP-R (Supplementary Fig. 8D). Under steady state conditions, Golgi will fluoresce both red and green; however, during late stages of Golgiphagy, fragmented Golgi are delivered to lysosomes where mCherry fluorescence is stable, but GFP fluorescence becomes quenched by the acidic microenvironment. Therefore, cells undergoing Golgiphagy will exhibit mCherry+eGFP– foci. Control and TAX1BP1-KO HeLa cells were generated that stably express the Golgiphagy reporter. First, we confirmed that mCherry and GFP fluorescence were localized to the Golgi by staining with anti-Golgin-97. As expected, mCherry and GFP puncta were perinuclear and colocalized with Golgin-97 (Supplementary Fig. 8E). We next utilized the Golgiphagy reporter to determine if STING activation induced Golgiphagy. Immunofluorescence and confocal microscopy experiments revealed that Golgiphagy occurred in control HeLa cells treated with diABZI as revealed by an increase of mCherry+eGFP– foci; however, mCherry+eGFP– foci were significantly decreased in TAX1BP1-KO HeLa cells (Fig. 8C, D). These data suggest that TAX1BP1 is critical for STING-induced Golgiphagy.
TAX1BP1 clears protein aggregates together with p62/SQSTM141, which has also been implicated in STING degradation7. To determine if p62/SQSTM1 was involved in STING-induced Golgiphagy we generated p62/SQSTM1 knockout (p62-KO) THP-1 cells using CRISPR/Cas9. Knockout of p62/SQSTM1 was validated by immunoblotting (Fig. 8E). We treated control and p62-KO THP-1 cells with diABZI and examined the expression of Golgin-97, GM130 and STING. Interestingly, diABZI-mediated degradation of Golgi proteins (Golgin-97 and GM130) and STING (Fig. 8F) was significantly impaired in p62-KO THP-1 cells. Reconstitution of Flag-p62 into p62-KO THP-1 cells using a lentiviral vector expressing Flag-p62 restored diABZI-induced Golgin-97 and STING degradation (Supplementary Fig. 9A). Therefore, p62/SQSTM1 and TAX1BP1 are both required for STING-induced Golgiphagy.
We next examined a potential role of macroautophagy in STING-induced Golgiphagy by generating ATG3 knockout (ATG3-KO) THP-1 cells using CRISPR-Cas9. ATG3 functions as an E2-like enzyme for LC3 lipidation that is essential for macroautophagy42. We also generated FIP200 (also known as RB1CC1) knockout (FIP200-KO) THP-1 cells by CRISPR-Cas9. FIP200 is a scaffold protein for the ULK1 complex required for the initial stages of autophagosome formation43. FIP200 has also been implicated in TAX1BP1-mediated noncanonical autophagy, whereby TAX1BP1 clusters FIP200 around cargo to form autophagosomes independent of LC3 lipidation40. ATG3-KO and FIP200-KO THP-1 cells were validated by immunoblotting (Supplementary Fig. 9B, D). Control, ATG3-KO and FIP200-KO THP-1 cells were treated with diABZI and immunoblotting was performed to examine the degradation of Golgin-97 and STING. Golgin-97 degradation seemed to be enhanced in diABZI-stimulated ATG3-KO and FIP200-KO THP-1 cells compared to control cells, and there was no significant difference in STING degradation in ATG3-KO and FIP200-KO THP-1 cells compared to control cells (Supplementary Fig. 9C, E). These data suggest that macroautophagy and FIP200-mediated noncanonical autophagy are not involved in STING degradation and STING-induced Golgiphagy.
To determine if microautophagy played a role in STING-induced Golgiphagy we used siRNA to knockdown the ESCRT-0 protein TSG101. The knockdown of TSG101 in THP-1 cells was confirmed by immunoblotting (Supplementary Fig. 9F). However, knockdown of TSG101 had no effect on STING-induced Golgin-97 degradation compared to cells transfected with nontargeting control siRNA (Supplementary Fig. 9F). VPS4A and VPS4B function as AAA ATPases that are essential for the function of the ESCRT machinery and are required for STING degradation by microautophagy11. Knockdown of VPS4A/B had no effect on Golgin-97 degradation induced by diABZI stimulation (Supplementary Fig. 9G) providing further evidence that this Golgi degradation pathway is independent of microautophagy.
To determine if STING was required for diABZI-induced Golgi degradation, we treated control THP-1 and STING-KO THP-1 cells with diABZI and examined the expression of Golgin-97 by immunoblotting. Interestingly, basal levels of Golgin-97 were elevated in STING-KO cells (Fig. 8G) suggesting that tonic activation of STING may promote Golgiphagy. Furthermore, diABZI-induced Golgin-97 and GM130 degradation was completely impaired in the absence of STING (Fig. 8G).
Fragmented Golgi accumulate in TAX1BP1-deficient macrophages upon STING activation
To examine Golgi with higher resolution, we next performed transmission electron microscopy (TEM) using WT and Tax1bp1–/– BMDMs treated with diABZI (Fig. 9, Supplementary Fig. 10). In WT BMDMs, intact Golgi stacks and cisternae were readily detected; however, after diABZI treatment the Golgi became swollen and fragmented consistent with a recent study44. Interestingly, in WT BMDMs we could detect phagophore formation around fragmented Golgi, and fragmented Golgi localized in autophagosomes and autolysosomes (Fig. 9A-D). In Tax1bp1–/– BMDMs, fragmented Golgi accumulated in diABZI-treated Tax1bp1–/– BMDMs and these were not found in autophagosomes or autolysosomes (Fig. 9A-H, Supplementary Fig. 10). Tax1bp1–/– BMDMs treated with diABZI exhibited a significant decrease in the number of autophagic vesicles (Fig. 9I). Furthermore, small electron dense structures in Golgi that may represent protein aggregates (potentially STING) were observed in WT BMDMs treated with diABZI, and these were increased in diABZI-treated Tax1bp1–/– BMDMs (Fig. 9A-H). Therefore, STING activation induces Golgi stress and fragmentation that triggers Golgiphagy mediated by TAX1BP1.
A Transmission electron microscopy showing ultrastructure of Golgi in control (LysM-Cre) BMDMs indicating intact Golgi stacks and cisternae. G- Golgi, M- Mitochondria, N-Nucleus. B Ultrastructure of Golgi in control BMDMs treated with diABZI for 4 h reveals swollen and fragmented Golgi with phagophores. Black arrow-swollen and fragmented Golgi, double-headed arrow- phagophores, black arrowhead- autolysosomes. C Control BMDMs treated with diABZI for 4 h. Black arrow-swollen and fragmented Golgi, double-headed arrow-phagophores, black arrowhead- autolysosomes. D Ultrastructure of Golgi in control BMDMs treated with diABZI for 4 h reveals autolysosomes with fragmented Golgi undergoing degradation. Black arrow-swollen and fragmented Golgi, black arrowhead- autolysosomes. E Ultrastructure of Golgi in Tax1bp1–/– (Tax1bp1fl/flLysM-Cre) BMDMs showing Golgi stacks and cisternae. G- Golgi, M- Mitochondria, N-Nucleus. F Tax1bp1–/– (Tax1bp1fl/flLysM-Cre) BMDMs treated with diABZI for 4 h reveals swollen and fragmented Golgi. Black arrow-swollen and fragmented Golgi, yellow arrowhead- dark patches of protein aggregates. G Ultrastructure of Golgi in Tax1bp1–/– (Tax1bp1fl/flLysM-Cre) BMDMs treated with diABZI for 4 h. Black arrow-swollen and fragmented Golgi, yellow arrowhead- dark patches of protein aggregates. H Ultrastructure of Golgi in Tax1bp1–/– BMDMs treated with diABZI for 4 h. Black arrow-swollen and fragmented Golgi. I Quantification of autophagic vacuoles/vesicles (AV) per cell. P values from left to right: P = 0.8734, P < 0.0001, P < 0.0001, P = 0.0023. The results are expressed as the mean ± SD (n = 10 micrographs per condition). ****P < 0.0001; **P < 0.01; *P < 0.05; ns = significant. Two-way ANOVA with Sidak’s multiple comparisons test was used. J Immunoblotting with the indicated antibodies using lysates from control and TAX1BP1-KO THP-1 cells treated with diABZI and/or C53 for 6 h. The experiment is representative of three independent experiments with similar results. Source data are provided as a Source data file.
Recent studies have demonstrated that STING exerts proton channel activity upon trafficking to the Golgi44,45. The STING proton channel activity increases the pH in the Golgi, leading to LC3B lipidation and inflammasome activation, functions distinct from IRF3 activation and type I IFN induction. We hypothesized that STING proton channel activity promoted Golgi stress and fragmentation which then triggered Golgiphagy. To assess the importance of STING proton channel function in STING-induced Golgiphagy, we used the STING agonist C53, which was identified as an inhibitor of its proton channel function and downstream LC3 lipidation45. We treated control and TAX1BP1-KO THP-1 cells with diABZI for 30 min to allow for STING to traffic to the Golgi and then added compound C53 to inhibit STING proton channel function. Treatment of control and TAX1BP1-KO THP1 cells with C53 did not block the degradation of Golgin-97 induced by diABZI, although C53 suppressed diABZI-induced LC3 lipidation as expected (Fig. 9J). Therefore, STING-mediated Golgiphagy does not appear to depend on its proton channel activity.
Tax1bp1 fl/flLysM-Cre mice exhibit heightened STING activation
To understand the physiological relevance of TAX1BP1 inhibition of cGAS-STING, Tax1bp1fl/flLysM-Cre and LysM-Cre mice were injected intraperitoneally (i.p.) with diABZI and serum levels of type I IFN and inflammatory cytokines were quantified by the LEGENDplexTM mouse inflammation panel using flow cytometry. Serum levels of IFN-β, IL-6 and TNF were significantly increased in Tax1bp1fl/flLysM-Cre mice (Fig. 10A-C). The other cytokines in the inflammation panel were not significantly changed in Tax1bp1fl/flLysM-Cre mice injected with diABZI (Supplementary Fig. 11). Tax1bp1fl/flLysM-Cre and LysM-Cre mice were next infected with Ectromelia virus (ECTV), a mouse DNA virus and orthopoxvirus that causes mousepox and to which the innate immune response is dependent upon STING46 and cGAS47. Mice were infected in the footpad and viral loads in the draining lymph node and liver quantified by plaque assays. ECTV viral loads were significantly diminished in the draining lymph node of Tax1bp1fl/flLysM-Cre mice (Fig. 10D); there was also a trend of lower viral load in the liver of Tax1bp1fl/flLysM-Cre mice (although not statistically significant) (Fig. 10E). Therefore, deletion of TAX1BP1 in myeloid cells promotes resistance to ECTV replication and/or spread in infected mice.
A LysM-Cre and Tax1bp1fl/flLysM-Cre mice (n = 10 per group) were injected i.p. with diABZI (2.5 mg/kg) and serum was collected after 3 h to examine IFN-β expression using the LEGENDplexTM mouse inflammation panel by flow cytometry. P = 0.0016. B LysM-Cre and Tax1bp1fl/flLysM-Cre mice (n = 10 per group) were injected i.p. with diABZI (2.5 mg/kg) and serum was collected after 3 h to examine IL-6 expression using the LEGENDplexTM mouse inflammation panel by flow cytometry. P = 0.0137. C LysM-Cre and Tax1bp1fl/flLysM-Cre mice (n = 10 per group) were injected i.p. with diABZI (2.5 mg/kg) and serum was collected after 3 h to examine TNF expression using the LEGENDplexTM mouse inflammation panel by flow cytometry. P = 0.0241. The results are expressed as the mean ± SD. **P < 0.01; *P < 0.05; ns = not significant. Unpaired Student’s t test was used for experiments in (A–C). D LysM-Cre and Tax1bp1fl/flLysM-Cre mice were infected in the footpad with ECTV and viral loads were quantified by plaque assays in the draining lymph node (n = 10) at day 7 post-infection. P = 0.0030. E LysM-Cre and Tax1bp1fl/flLysM-Cre mice were infected in the footpad with ECTV and viral loads were quantified by plaque assays in the liver (n = 7) at day 7 post-infection. P = 0.0529. The results are expressed as the mean ± SD. **P < 0.01; ns = not significant. Unpaired Student’s t test with Welch’s correction was used for experiments in (D, E). F Model depicting STING-induced Golgiphagy. After binding to cGAMP, STING translocates to the Golgi, forms higher order aggregates and activates TBK1 and IKK. As a byproduct of STING activation, the Golgi becomes swollen and fragmented which triggers Golgiphagy. TAX1BP1 is required for STING-induced Golgiphagy that degrades a pool of activated STING molecules and thus downregulates STING in a negative feedback loop. In addition to STING-induced Golgiphagy, TAX1BP1 also promotes ESCRT-mediated STING degradation. There is a partial impairment in STING degradation in TAX1BP1-deficient cells that may be due to redundancy with p62/SQSTM1. The schematic was created with BioRender.com (Suklabaidya, S., 2025) https://BioRender.com/1c4aq0r). Source data are provided as a Source data file.
Discussion
The cGAS-STING pathway plays critical roles in host immunity to pathogens and anti-tumor responses1. Activation of the cGAS-STING pathway leads to potent induction of inflammatory responses; therefore, cGAS and STING are tightly regulated by numerous negative regulatory mechanisms, including post-translational modifications (i.e., phosphorylation, ubiquitination) that prevent or downregulate their activities48,49,50. STING degradation is a critical mechanism to resolve STING signaling and this mainly occurs by ESCRT-mediated microautophagy11,12,13. Negative regulation of cGAS-STING is essential for immune homeostasis since dysregulated or aberrant activation of the cGAS-STING pathway by self-DNA has been linked to autoimmune, autoinflammatory and neurodegenerative diseases, and aging-related chronic inflammation1,51,52. Our findings indicate that STING-induced Golgiphagy serves as a negative feedback mechanism that couples STING activation at the trans-Golgi with its degradation. STING activation gives rise to Golgi swelling and fragmentation that elicits the recruitment of TAX1BP1 to STING anchored in the trans-Golgi membrane to induce Golgiphagy (see model in Fig. 10F). STING-induced Golgiphagy appears to be a unique STING degradation pathway distinct from the direct encapsulation of STING vesicles by lysosomes.
STING accumulates at the TGN in the absence of TAX1BP1 and its trafficking to REs and lysosomes is impaired (Fig. 6). However, STING-induced LC3-lipidation was not impaired in Tax1bp1–/– BMDMs or TAX1BP1-KO THP-1 cells treated with diABZI (Fig. 3M, N), and activated STING anchored in the trans-Golgi did not colocalize with LC3 (Supplementary Fig. 8A), suggesting that STING is primarily degraded in an LC3-independent manner. Since diABZI-induced STING degradation was not fully impaired in TAX1BP1-deficient macrophages (Figs. 3, 4), and TSG101 knockdown completely blocked STING degradation in TAX1BP1-KO THP-1 cells (Fig. 4C-E), residual microautophagy is likely responsible for the partial degradation of STING in the absence of TAX1BP1. Thus, while TAX1BP1 facilitates STING degradation by promoting STING-HGS binding to induce microautophagy, there may be redundancy with p62/SQSTM1. Our results therefore suggest that ESCRT-mediated STING degradation is a selective form of microautophagy driven by autophagy receptors p62/SQSTM1 and TAX1BP1. It remains unclear if p62/SQSTM1 and/or TAX1BP1 regulate microautophagy in other contexts, such as nutrient starvation.
Our results are in agreement with a recent study that showed that TAX1BP1 targeted STING for degradation53. It was shown that TAX1BP1 promotes the degradation of STING K63-ubiquitinated at lysine 224 (K224) to impair anti-tumor immunity. However, the pathway of TAX1BP1-mediated STING degradation was not addressed in the study. We found that TAX1BP1 induces STING degradation by two independent mechanisms: microautophagy by facilitating an interaction between STING and the ESCRT machinery as well as STING-induced Golgiphagy (Fig. 10F).
The mechanisms and regulation of Golgiphagy are not well understood but several autophagy receptors for autophagic degradation of Golgi have been identified. The selective autophagy receptor and TAX1BP1 paralog, CALCOCO1, was identified as a Golgiphagy receptor that contributes to Golgi turnover during nutrient stress54. Interestingly, CALCOCO1 requires a short motif termed zDABM to interact with the Golgi-resident protein ZDHHC17 and induce Golgiphagy. TAX1BP1 also harbors a putative zDABM motif at 673-VVCSQP-67854; however, it is unclear if this motif in TAX1BP1 is required for STING-induced Golgiphagy. Recent studies have also identified Yip domain family members 3 and 4 (YIPF3 and YIPF4) as Golgiphagy receptors that function in response to nutrient stress55. YIPF3 and YIPF4 are Golgi membrane proteins that form a complex and recruit ATG8 family members via an LIR motif in YIPF356. Our data indicate that STING-induced Golgiphagy requires TAX1BP1; however, induction of Golgiphagy by the Golgi dispersal agent golgicide A did not require TAX1BP1. Therefore, TAX1BP1 is unlikely to function as a general regulator of Golgiphagy but rather exerts specificity as a Golgiphagy receptor for STING-induced fragmented Golgi, most likely due to TAX1BP1 targeting of STING aggregate formation and/or STING polyubiquitination. STING-induced Golgiphagy is independent of ATG3, FIP200 and TSG101 (Supplementary Fig. 8) suggesting that this pathway does not depend on macroautophagy or microautophagy. However, STING-induced Golgiphagy requires STING, TAX1BP1 and p62/SQSTM1 (Fig. 8). Additional studies are needed to further define the pathway and factors required for STING-induced Golgiphagy.
STING forms higher-order aggregates upon activation and a recent study showed that STING forms clusters at the trans-Golgi that are dependent on STING palmitoylation and cholesterol57. Our data indicate that STING aggregates accumulate in TAX1BP1-deficient macrophages treated with diABZI or infected with HSV-1 (Fig. 7). TAX1BP1 has been shown to function as an aggrephagy receptor together with p62/SQSTM136,58. It is thought that p62/SQSTM1 initially detects ubiquitinated protein aggregates and drives condensate formation, subsequently recruiting NBR1, TAX1BP1, and FIP200 to ubiquitin condensates to clear aggregates via autophagosomes41,59. Given that p62/SQSTM1 has previously been shown to inhibit the cGAS-STING pathway by promoting STING degradation7, it is likely that p62/SQSTM1, NBR1 and TAX1BP1 function in a sequential manner to clear STING aggregates from fragmented Golgi by Golgiphagy as well as targeting post-Golgi STING vesicles for ESCRT-mediated degradation. TAX1BP1 interaction with FIP200 and TBK1 seems to be important for the downregulation of STING activation (Supplementary Fig. 7), and this interaction was shown to mediate the noncanonical LC3-independent autophagy function of TAX1BP140. However, STING-induced Golgiphagy can occur in the absence of FIP200 suggesting a potential FIP200-independent but TBK1-dependent form of noncanonical autophagy. Overall, our findings suggest that STING activation is associated with Golgi fragmentation and TAX1BP1-dependent Golgiphagy, which serves as a critical negative feedback mechanism to downregulate its activation.
Methods
Tax1bp1 knockout mice
An embryonic stem cell (ES) clone (clone ID: HEPD0680_3_H08) with exon 4 of the Tax1bp1 gene flanked by LoxP sites was obtained from the European Conditional Mouse Mutagenesis Program (EUCOMM). The LacZ/Neo cassettes were flanked by FRT sites to facilitate excision by flippase (Flp) recombinase prior to deletion of exon 4 by Cre recombinase. A C57BL/6 embryonic stem cell (ES) clone with a correctly targeted Tax1bp1 allele was microinjected into mouse blastocysts to create chimeric mice. Male chimeras were crossed with female C57BL/6 mice and pups were screened for germline transmission of the mutant allele by PCR genotyping. Germline transmission of the LoxP flanked Tax1bp1 allele was confirmed by PCR genotyping. Heterozygous mice were then crossed with transgenic mice expressing Flp under the control of the universal Pgk1 promoter and deletion of the LacZ/Neo cassettes was confirmed by PCR. Tax1bp1fl/fl mice were crossed with LysM-Cre mice (Jackson Laboratory; strain# 004781) to delete TAX1BP1 in myeloid cells. Tax1bp1fl/flLysM Cre mice were also crossed with Mx1-GFP mice28 (a gift from Adolfo García-Sastre). Animals were housed in a specific pathogen-free (SPF) barrier facility kept at 22 °C with 40–60% humidity and maintained on a 12 h light/12 h dark cycle with ad libitum access to food and water. Experimental animals were bred separately and age- and sex- matched mice were used for each experiment. Animals were euthanized by CO2 inhalation. All animal experiments were approved and performed in accordance with the Penn State College of Medicine Institutional Animal Care and Use Guidelines (Protocol Number 1854).
Cell culture, plasmids, and reagents
THP-1 cells were purchased from ATCC (TIB-202) and cultured in RPMI media supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin (P/S). THP-1 Dual KO-STING cells were purchased from InvivoGen (thpd-kostg). BMDMs were generated by culturing bone marrow cells isolated from LysM-Cre and Tax1bp1fl/flLysM-Cre mouse femurs in DMEM medium supplemented with 30% L929 conditioned medium, 20% FBS and 1% P/S for 7 days as described previously60. TAX1BP1-KO HeLa cells were provided by Richard Youle61. Lenti-X 293T cells were purchased from Takara (632180) and cultured in DMEM medium supplemented with 10% FBS and 1% P/S. Plasmid transfection was performed using GenJet version II (SignaGen Laboratories; SL100489), and G3-YSD transfection was performed using Lipofectamine 3000 reagent (ThermoFisher Scientific; L3000008) following the manufacturer’s instructions. LentiCRISPRv2 was a gift from Feng Zhang (Addgene plasmid# 52691; 422 http://n2t.net/addgene:52691; RRID:Addgene_52691). pUltra-Hot was a gift from Malcolm Moore (Addgene plasmid# 24130; http://n2t.net/addgene:24130; RRID:Addgene_24130). psPAX2 was a gift from Didier Trono (Addgene plasmid#12260; http://n2t.net/addgene:12260; RRID:Addgene_12260). pHR-SFFV-FLAG-SQSTM1 (p62) was a gift from Benjamin Wolozin (Addgene plasmid# 223715; http://n2t.net/addgene:223715; RRID:Addgene_223715). The STING-eGFP plasmid (pMRX-hSTING-eGFP) was a gift from Nan Yan62. PMA (12-011) was purchased from Fisher Scientific. G3-YSD (tlrl-ydna), cGAMP (tlrl-nacga23) and diABZI compound 3 (tlrl-diabzi) were purchased from InvivoGen. HT-DNA (D6898) was purchased from Millipore-Sigma.
Antibodies used for immunoblotting
p-TBK1 (5483S), TBK1 (3504S), p-IRF3 (4947S), IRF3 (4302S), p-STING (50907S), p-IKKα/β (2697S), IKKβ (8943S), TSG101 (72312S) and HGS (15087S) antibodies were purchased from Cell Signaling Technology. TAX1BP1 antibodies were purchased from Abcam (ab176572) and Bethyl (A303-792A). STING antibody was purchased from Cell Signaling Technology (13647S). GFP (B-2) (sc-9996), vinculin (sc-73614) and β-actin (sc-47778) antibodies were purchased from Santa Cruz Biotechnology. Golgin-97 polyclonal antibody (12640-1-AP), TGN46 monoclonal antibody (66477-1-1g), VPS4A polyclonal antibody (14272-1-AP), VPS4B monoclonal antibody (68443-1-Ig), FIP200 (RB1CC1) polyclonal antibody (17250-1-AP), ATG3 polyclonal antibody (11262-2-AP) and p62/SQSTM1 polyclonal antibody (18420-1-AP) were purchased from Proteintech. GM130 polyclonal antibody (PA5-95727) and p-IKKα/β (37515) antibody were purchased from ThermoFisher Scientific. Flag M2 monoclonal antibody (F3165) was purchased from Millipore-Sigma.
Antibodies used for immunofluorescence assays
TAX1BP1 (Ab176572) and Golgin-97 (Ab169287) antibodies were purchased from Abcam. Golgin-97 (12640-1-AP) and CoraLite 594-conjugated TGN46 antibodies were purchased from Proteintech (CL594-13573). LAMP2 antibody (A15464) and Alexa Fluor 647-conjugated donkey anti-mouse IgG (A-31573) were purchased from ThermoFisher Scientific. Transferrin receptor (TfnR)/CD71 antibody (46222) was purchased from Cell Signaling Technology.
Nucleic acid manipulation
pUltra-Hot-TAX1BP1 was generated by subcloning TAX1BP1 PCR product digested with Nhe1-HF (New England Biolabs; R3131S) and BamH1-HF (New England Biolabs; R3136S) into the pUltra-Hot vector. Silent mutations were introduced into the TAX1BP1 cDNA by PCR site-directed mutagenesis to render TAX1BP1 resistant to the TAX1BP1 gRNA. Point mutations for TAX1BP1 LIR double mutant (W49A and V143S), ZnF2 mutant (C760A) and FIP200/TBK1 mutant (A114Q) were introduced by PCR site-directed mutagenesis in the pUltra-Hot-TAX1BP1 plasmid. Site-directed mutagenesis was performed with the QuikChange II mutagenesis kit (Agilent; 200523) using the primers listed in Supplementary Table 1. The Golgiphagy reporter was generated by adding a minimal Golgi targeting sequence (FLWRIFCFRK)56 to the C-terminus of the tandem fusion of fluorescence proteins, mCherry and EGFP, as described previously63. All constructs generated in this study were verified by sequencing.
CRISPR-Cas9-mediated generation of knockout cells
CRISPR-Cas9-mediated knockout of TAX1BP1, ATG3, FIP200 and p62/SQSTM1 was performed as described previously19,64 by cloning control nontargeting gRNA and gRNAs for Tax1bp1, Atg3, Fip200 and p62/SQSTM120,61 (Supplementary Table 2) in the pLentiCRISPRv2 plasmid. THP-1 cells were transduced with lentiviruses expressing Cas9 and control nontargeting gRNA or Tax1bp1, Atg3, Fip200 or p62/SQSTM1 gRNAs to generate control and knockout cell lines. Bulk population, puromycin-resistant THP-1 control-gRNA and knockout cells were validated by immunoblotting and qRT-PCR (for TAX1BP1-KO).
Knockdown of gene expression with siRNAs
THP-1 cells were transfected with control nontargeting (D-001810-01-05) or On Target Plus human TSG101 (L-003549-00-0005), VPS4A (L-013092-00-0005) or VPS4B (L-013119-00-0005) SMARTpool siRNAs (Horizon Discovery) using Lipofectamine RNAiMAX Transfection Reagent (ThermoFisher Scientific; 13778075). Cells were harvested after 48 h and knockdowns were confirmed by immunoblotting.
Virus infections
HSV-1-YFP was a gift from Drs. John Wills and Lynn Enquist65. Cells were starved for 1 h in serum-free RPMI and inoculated with HSV-1-YFP for 1 h at a multiplicity of infection=1 (MOI = 1) in serum-free RPMI and further incubated in complete DRPMI for the indicated times.
Live-cell imaging
For HSV-1 replication, 3 × 105 control or TAX1BP1-KO THP-1 cells were differentiated with PMA in 24-well plates and then infected with HSV-1-YFP (MOI = 1). For Mx1-GFP expression, 3×105 BMDMs prepared from LysM-Cre-Mx1-GFP and Tax1bp1fl/flLysM-Cre-Mx1-GFP mice were seeded in 24-well plates and transfected with either 1 μg G3-YSD or stimulated with 1 μM diABZI. Live-cell imaging was performed with an IncuCyte S3 imaging system (Sartorius). Images were acquired every 2 h using phase contrast and green channels with a 10× objective in triplicate. GFP fluorescence was quantified as green object count per image normalized to phase area confluence.
Real-time quantitative reverse transcription PCR
2 × 106 BMDMs or PMA-differentiated THP-1 cells in 6-well plates were transfected with either 1 μg G3-YSD or 1 μg HT-DNA, infected with HSV-1 (MOI = 1) or stimulated with either 50 nM cGAMP or 1 μM diABZI. Cells were harvested by gentle scraping and processed for RNA isolation using the RNA Spin II Kit (Macherey-Nagel; 740955) using the manufacturer’s instructions. cDNA was synthesized using M-MLV Reverse Transcriptase (ThermoFisher Scientific; 28025-013) and Oligo (dT) (ThermoFisher Scientific; 18418-012), and qRT–PCR (ThermoFisher Scientific; QuantStudio 3) was performed using Power SYBR green PCR master mix (ThermoFisher Scientific; A25742) according to the manufacturer’s protocol. β-Actin was used for normalization of the assay, and fold change in expression was calculated by the 2−ΔΔCT method. Primers used for real-time qRT-PCR are listed in Supplementary Table 3.
Enzyme-linked immunosorbent assay
3 × 106 PMA-differentiated THP-1 cells in six-well plates were transfected with 1 μg G3-YSD, infected with HSV-1 (MOI = 1) or stimulated with 50 nM cGAMP and the culture supernatants were collected at the indicated time points. The concentration of IFN-β and IL-6 in the cell culture supernatants was measured using the Human IFN-β ELISA kit (PBL Assay Science; 41410) and Human IL-6 Quantikine ELISA kit (R&D; D6050) according to the manufacturers’ instructions.
Immunoblotting and co-immunoprecipitation assays
Whole cell lysates were generated by lysing cells in RIPA buffer (50 mM Tris-Cl [pH 7.4], 150 mM NaCl, 1% NP-40, 0.25% sodium deoxycholate, Pierce Protease and Phosphatase Inhibitor) on ice, followed by centrifugation. Lysates were separated using SDS–polyacrylamide gels, transferred onto a nitrocellulose membrane and blocked for 1 h in 5% skim milk followed by incubation with the indicated primary antibodies overnight at 4 °C followed by incubation with HRP-conjugated secondary antibodies for 1 h at room temperature. Immunoreactive bands were visualized using SuperSignal West Pico Chemiluminescent Substrate (Bio-Rad; 34580) and the Bio-Rad ChemiDoc Imaging System. Western blot images were processed using Image Lab software (Bio-Rad Laboratories). The Dynabeads Protein G Immunoprecipitation Kit (ThermoFisher Scientific; 10007D) was used for co-IP assays according to the manufacturer’s recommendations.
Protein aggregation detection assay
The PROTEOSTAT Aggresome detection kit (Enzo Life Sciences; ENZ-51035-0025) was used to detect total protein aggregates according to the manufacturer’s instructions. 1 × 106 control or TAX1BP1-KO THP-1 cells were differentiated with PMA in 6-well plates and either infected with HSV-1-YFP (MOI = 0.1) or stimulated with 50 nM cGAMP. As a positive control, cells were treated with MG132 for 16 h. Cells were then harvested by gentle scraping and centrifuged at 500 g for 5 min followed by fixation with 4% formaldehyde for 30 min at RT, permeabilized with 0.5% Triton X-100 for 30 min on ice, and stained with the PROTEOSTAT dye for 30 min at RT. Samples were then acquired on a BD LSRFortessa flow cytometer and data analysis was performed using FlowJo v10 software. The gating strategy for these experiments is shown in Supplementary Fig. 6.
Semi-denaturing detergent agarose gel electrophoresis
SDD-AGE was performed as previously described35. Briefly, whole cell lysates were resuspended in 2× sample buffer (0.5x Tris-borate-EDTA [TAE], 10% glycerol, 2% SDS, and 0.0025% bromophenol blue) followed by incubation at room temperature for 15 min and loaded onto a 1.5% agarose gel. Electrophoresis was performed in the running buffer (1× TAE and 0.1% SDS) for 35 min with a constant voltage of 100 V at 4 °C, proteins were transferred to a nitrocellulose membrane for immunoblotting.
Immunofluorescence microscopy
Control HeLa and TAX1BP1-KO HeLa cells stably expressing STING-eGFP plasmid were generated with a recombinant retrovirus expressing STING-eGFP. Cells were treated with 10 mM leupeptin and 1 μM diABZI for the indicated time points. Immunofluorescence/confocal microscopy was performed as previously described64. Images were acquired with using a Leica SP8 confocal microscope equipped with a 63× oil objective. Images were processed, analyzed and Manders’ coefficient analysis was performed using Fiji image analysis software (NCBI).
STING quantitative flow cytometry degradation assay
Control THP-1 and TAX1BP1-KO THP-1 cells stably expressing STING-eGFP plasmid were generated with a recombinant retrovirus expressing STING-eGFP. Cells were sorted for high GFP expression. In certain experiments, cells were first transfected with control nontargeting or On Target Plus human TSG101 SMARTpool siRNAs (Horizon Discovery) using Lipofectamine RNAiMAX Transfection Reagent (ThermoFisher Scientific). Cells were treated with 1 μM diABZI and then harvested by gentle scraping and centrifuged at 500 g for 5 min followed by staining with SYTOX Blue. Samples were then acquired on a BD LSRFortessa flow cytometer. Data analysis was performed using FlowJo v10 software. The gating strategy for these experiments is shown in Supplementary Fig. 4.
Transmission electron microscopy
WT and Tax1bp1–/– BMDMs (3 × 106) were seeded in 60 mm cell culture dishes (174888; Nalgene Nunc) and treated with 1 μM diABZI for 4 h. Cells were then gently washed with PBS followed by fixation with 2.5% glutaraldehyde and 2% paraformaldehyde in 0.1 M phosphate buffer (pH 7.4) and further fixed in 1% osmium tetroxide in 0.1 M phosphate buffer (pH 7.4) for 1 h. Samples were dehydrated in a graduated ethanol series, acetone, and embedded in LX-112 (Ladd Research, Williston, VT). Thin sections (65 nm) were stained with uranyl acetate and lead citrate and viewed in a JEOL JEM1400 Transmission Electron Microscope (JEOL USA Inc., Peabody, MA, USA) located at the Penn State College of Medicine TEM Facility (RRID Number: SCR_021200).
Proximity ligation assay
PLA was performed using the DuoLink In Situ Red Starter Kit Mouse/Rabbit (Millipore-Sigma; DUO92101) as recommended by the manufacturer. 5 × 104 PMA-differentiated THP-1 cells were seeded in 8-well chamber slides (Corning; 354118) and stimulated with 1 μM diABZI for the indicated time points. Cells were then fixed, permeabilized, and incubated with primary antibodies anti-rabbit TAX1BP1 (Abcam; ab176572) and anti-mouse STING (Proteintech; 66680-1-Ig). The slides were incubated with Duolink PLA probes, ligated, amplified, and washed. Images were acquired using a Leica SP8 confocal microscope equipped with a 63× oil objective. Images were processed, analyzed, and quantified using Fiji image analysis software (NCBI).
Protein fragment complementation NanoBiT assay
293 T cells were transfected with the NanoBiT binary plasmids (Promega; N2014) expressing TAX1BP1 and STING for 24 h, harvested and resuspended at 5 × 105 cells/ml in Opti-MEM I medium. Next, 50 µl of cell suspension was transferred to a 96-well opaque plate in triplicate and treated with 25 µl of Opti-MEM I medium supplemented with 1% furimazine (Promega; N1110) for 5 min as described previously35. Luminescence intensities were measured using a GloMax 96 microplate luminometer (Promega).
In vivo diABZI treatment
8–10-week-old Tax1bp1fl/flLysM-Cre and LysM-Cre mice (n = 10; male n = 5 and female n = 5), were intraperitoneally administered with 2.5 mg/kg diABZI compound-3 (InvivoGen; tlrl-diabzi) for 3 h. Serum cytokines were measured by the LEGENDplexTM mouse inflammation panel (BioLegend; 740446) and acquired on a BD LSRFortessa flow cytometer according to the manufacturer’s protocol. Data analysis was performed using LEGENDplexTM data analysis software.
Ectromelia virus infection and titration
All experiments used WT ECTV (Moscow strain)66 propagated in L929 cells. 8–10-week-old Tax1bp1fl/flLysM-Cre and LysM-Cre mice (n = 10; male n = 5 and female n = 5) were injected with 3 × 103 plaque-forming units (PFU) of ECTV into one hind footpad or mock-infected with HBSS/0.1% BSA. Mice were monitored daily for morbidity and mortality. Infected footpad, draining lymph node, spleen, and liver were harvested at 7 d.p.i. and frozen in HBSS/0.1% BSA at −80 °C. Tissues were homogenized using a TissueLyser II, and titers were determined by standard plaque assay under carboxymethyl cellulose overlay on WT-19 cells at 3 d.p.i. For weighed tissues, PFU per gram tissue was calculated.
Statistical analysis
Data are presented as mean ± standard deviation from a representative experiment with triplicate samples. Statistical analysis was performed in GraphPad Prism 10.1.2 and indicated in the Figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this Article. The raw numbers for charts and graphs are available in the Source Data file whenever possible. Source data are provided with this paper.
References
Zhang, X., Bai, X. C. & Chen, Z. J. Structures and mechanisms in the cGAS-STING innate immunity pathway. Immunity 53, 43–53 (2020).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Ishikawa, H. & Barber, G. N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455, 674–678 (2008).
Ishikawa, H., Ma, Z. & Barber, G. N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461, 788–792 (2009).
Jeltema, D., Abbott, K. & Yan, N. STING trafficking as a new dimension of immune signaling. J Exp. Med. 220 (2023).
Liu, S. et al. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science 347, aaa2630 (2015).
Prabakaran, T. et al. Attenuation of cGAS-STING signaling is mediated by a p62/SQSTM1-dependent autophagy pathway activated by TBK1. EMBO J. 37 e97858 (2018).
Gui, X. et al. Autophagy induction via STING trafficking is a primordial function of the cGAS pathway. Nature 567, 262–266 (2019).
Liu, D. et al. STING directly activates autophagy to tune the innate immune response. Cell Death Differ. 26, 1735–1749 (2019).
Liu, Y. et al. Clathrin-associated AP-1 controls termination of STING signalling. Nature 610, 761–767 (2022).
Kuchitsu, Y. et al. STING signalling is terminated through ESCRT-dependent microautophagy of vesicles originating from recycling endosomes. Nat. Cell Biol. 25, 453–466 (2023).
Gentili, M. et al. ESCRT-dependent STING degradation inhibits steady-state and cGAMP-induced signalling. Nat. Commun. 14, 611 (2023).
Balka, K. R. et al. Termination of STING responses is mediated via ESCRT-dependent degradation. EMBO J. 42, e112712 (2023).
De Valck, D. et al. The zinc finger protein A20 interacts with a novel anti-apoptotic protein which is cleaved by specific caspases. Oncogene 18, 4182–4190 (1999).
Ling, L. & Goeddel, D. V. T6BP, a TRAF6-interacting protein involved in IL-1 signaling. Proc. Natl. Acad. Sci. USA 97, 9567–9572 (2000).
Shembade, N., Harhaj, N. S., Liebl, D. J. & Harhaj, E. W. Essential role for TAX1BP1 in the termination of TNF-alpha-, IL-1- and LPS-mediated NF-kappaB and JNK signaling. EMBO J. 26, 3910–3922 (2007).
Parvatiyar, K., Barber, G. N. & Harhaj, E. W. TAX1BP1 and A20 inhibit antiviral signaling by targeting TBK1-IKKi kinases. J. Biol. Chem. 285, 14999–15009 (2010).
White, J., Suklabaidya, S., Vo, M. T., Choi, Y. B. & Harhaj, E. W. Multifaceted roles of TAX1BP1 in autophagy. Autophagy 19, 44–53 (2022).
Choi, Y. B., Shembade, N., Parvatiyar, K., Balachandran, S. & Harhaj, E. W. TAX1BP1 restrains virus-induced apoptosis by facilitating itch-mediated degradation of the mitochondrial adaptor MAVS. Mol. Cell Biol. 37, e00422-16 (2017).
White, J. et al. Phosphorylation of the selective autophagy receptor TAX1BP1 by TBK1 and IKBKE/IKKi promotes ATG8-family protein-dependent clearance of MAVS aggregates. Autophagy 21, 160–77 (2024).
Yang, Q. et al. TRIM32-TAX1BP1-dependent selective autophagic degradation of TRIF negatively regulates TLR3/4-mediated innate immune responses. PLoS Pathog. 13, e1006600 (2017).
Samie, M. et al. Selective autophagy of the adaptor TRIF regulates innate inflammatory signaling. Nat. Immunol. 19, 246–254 (2018).
Qian, T. et al. Decreased TAX1BP1 participates in systemic lupus erythematosus by regulating monocyte/macrophage function. Int. Immunol. 35, 483–495 (2023).
Newman, A. C. et al. TBK1 kinase addiction in lung cancer cells is mediated via autophagy of Tax1bp1/Ndp52 and non-canonical NF-kappaB signalling. PLoS ONE 7, e50672 (2012).
Tumbarello, D. A. et al. Autophagy receptors link myosin VI to autophagosomes to mediate Tom1-dependent autophagosome maturation and fusion with the lysosome. Nat. Cell Biol. 14, 1024–1035 (2012).
Tumbarello, D. A. et al. The autophagy receptor TAX1BP1 and the molecular motor myosin VI are required for clearance of Salmonella typhimurium by autophagy. PLoS Pathog. 11, e1005174 (2015).
Iha, H. et al. Inflammatory cardiac valvulitis in TAX1BP1-deficient mice through selective NF-kappaB activation. EMBO J. 27, 629–641 (2008).
Uccellini, M. B. & Garcia-Sastre, A. ISRE-reporter mouse reveals high basal and induced type I IFN responses in inflammatory monocytes. Cell Rep. 25, 2784–2796.e2783 (2018).
Reinert, L. S. et al. Sensing of HSV-1 by the cGAS-STING pathway in microglia orchestrates antiviral defence in the CNS. Nat. Commun. 7, 13348 (2016).
Christensen, M. H. et al. HSV-1 ICP27 targets the TBK1-activated STING signalsome to inhibit virus-induced type I IFN expression. EMBO J. 35, 1385–1399 (2016).
Yum, S., Li, M., Fang, Y. & Chen, Z. J. TBK1 recruitment to STING activates both IRF3 and NF-kappaB that mediate immune defense against tumors and viral infections. Proc. Natl. Acad. Sci. USA 118, e2100225118 (2021).
Abe, T. & Barber, G. N. Cytosolic-DNA-mediated, STING-dependent proinflammatory gene induction necessitates canonical NF-kappaB activation through TBK1. J. Virol. 88, 5328–5341 (2014).
Balka, K. R. et al. TBK1 and IKKepsilon act redundantly to mediate STING-induced NF-kappaB responses in myeloid cells. Cell Rep. 31, 107492 (2020).
Fischer, T. D., Wang, C., Padman, B. S., Lazarou, M. & Youle, R. J. STING induces LC3B lipidation onto single-membrane vesicles via the V-ATPase and ATG16L1-WD40 domain. J. Cell Biol. 219, e202009128 (2020).
Vo, M. T., Smith, B. J., Nicholas, J. & Choi, Y. B. Activation of NIX-mediated mitophagy by an interferon regulatory factor homologue of human herpesvirus. Nat. Commun. 10, 3203 (2019).
Sarraf, S. A. et al. Loss of TAX1BP1-directed autophagy results in protein aggregate accumulation in the brain. Mol. Cell 80, 779–795.e710 (2020).
Shen, D. et al. Novel cell- and tissue-based assays for detecting misfolded and aggregated protein accumulation within aggresomes and inclusion bodies. Cell Biochem. Biophys. 60, 173–185 (2011).
Li, Z. et al. PPM1A regulates antiviral signaling by antagonizing TBK1-mediated STING phosphorylation and aggregation. PLoS Pathog. 11, e1004783 (2015).
Qin, F. et al. Listerin promotes cGAS protein degradation through the ESCRT pathway to negatively regulate cGAS-mediated immune response. Proc. Natl. Acad. Sci. USA 120, e2308853120 (2023).
Ohnstad, A. E. et al. Receptor-mediated clustering of FIP200 bypasses the role of LC3 lipidation in autophagy. EMBO J. 39, e104948 (2020).
Bauer, B., Idinger, J., Schuschnig, M., Ferrari, L. & Martens, S. Recruitment of autophagy initiator TAX1BP1 advances aggrephagy from cargo collection to sequestration. EMBO J. 43, 5910–5940 (2024).
Fang, D., Xie, H., Hu, T., Shan, H. & Li, M. Binding features and functions of ATG3. Front. Cell Dev. Biol. 9, 685625 (2021).
Hara, T. et al. FIP200, a ULK-interacting protein, is required for autophagosome formation in mammalian cells. J. Cell Biol. 181, 497–510 (2008).
Xun, J. et al. A conserved ion channel function of STING mediates noncanonical autophagy and cell death. EMBO Rep. 25, 544–69 (2024).
Liu, B. et al. Human STING is a proton channel. Science 381, 508–514 (2023).
Xu, R. H. et al. Sequential activation of two pathogen-sensing pathways required for type I interferon expression and resistance to an acute DNA virus infection. Immunity 43, 1148–1159 (2015).
Wong, E. B., Montoya, B., Ferez, M., Stotesbury, C. & Sigal, L. J. Resistance to ectromelia virus infection requires cGAS in bone marrow-derived cells which can be bypassed with cGAMP therapy. PLoS Pathog. 15, e1008239 (2019).
Konno, H., Konno, K. & Barber, G. N. Cyclic dinucleotides trigger ULK1 (ATG1) phosphorylation of STING to prevent sustained innate immune signaling. Cell 155, 688–698 (2013).
Li, T. et al. Phosphorylation and chromatin tethering prevent cGAS activation during mitosis. Science 371 (2021).
Xu, P. et al. The CRL5-SPSB3 ubiquitin ligase targets nuclear cGAS for degradation. Nature 627, 873–879 (2024).
Decout, A., Katz, J. D., Venkatraman, S. & Ablasser, A. The cGAS-STING pathway as a therapeutic target in inflammatory diseases. Nat. Rev. Immunol. 21, 548–569 (2021).
Gulen, M. F. et al. cGAS-STING drives ageing-related inflammation and neurodegeneration. Nature 620, 374–380 (2023).
Zhang, R. et al. TAX1BP1-dependent autophagic degradation of STING1 impairs anti-tumor immunity. Autophagy 21, 1802–1823 (2025).
Nthiga, T. M. et al. Regulation of Golgi turnover by CALCOCO1-mediated selective autophagy. J. Cell Biol. 220, e202006128 (2021).
Hickey, K. L. et al. Proteome census upon nutrient stress reveals Golgiphagy membrane receptors. Nature 623, 167–174 (2023).
Kitta, S. et al. YIPF3 and YIPF4 regulate autophagic turnover of the Golgi apparatus. EMBO J. 43, 2954–78 (2024).
Kemmoku, H. et al. Single-molecule localization microscopy reveals STING clustering at the trans-Golgi network through palmitoylation-dependent accumulation of cholesterol. Nat. Commun. 15, 220 (2024).
Pankiv, S. et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J. Biol. Chem. 282, 24131–24145 (2007).
Turco, E. et al. Reconstitution defines the roles of p62, NBR1 and TAX1BP1 in ubiquitin condensate formation and autophagy initiation. Nat. Commun. 12, 5212 (2021).
Shembade, N., Ma, A. & Harhaj, E. W. Inhibition of NF-kappaB signaling by A20 through disruption of ubiquitin enzyme complexes. Science 327, 1135–1139 (2010).
Lazarou, M. et al. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 524, 309–314 (2015).
Gonugunta, V. K. et al. Trafficking-mediated STING degradation requires sorting to acidified endolysosomes and can be targeted to enhance anti-tumor response. Cell Rep. 21, 3234–3242 (2017).
Choi, C. Y., Vo, M. T., Nicholas, J. & Choi, Y. B. Autophagy-competent mitochondrial translation elongation factor TUFM inhibits caspase-8-mediated apoptosis. Cell Death Differ. 29, 451–464 (2022).
Mohanty, S. et al. The E3/E4 ubiquitin conjugation factor UBE4B interacts with and ubiquitinates the HTLV-1 tax oncoprotein to promote NF-kappaB activation. PLoS Pathog. 16, e1008504 (2020).
Taylor, M. P., Kobiler, O. & Enquist, L. W. Alphaherpesvirus axon-to-cell spread involves limited virion transmission. Proc. Natl. Acad. Sci. USA 109, 17046–17051 (2012).
Sei, J. J. et al. Peptide-MHC-I from endogenous antigen outnumber those from exogenous antigen, irrespective of APC phenotype or activation. PLoS Pathog. 11, e1004941 (2015).
Acknowledgements
We thank Dr. John Wills (Penn State College of Medicine) and Lynn Enquist (Princeton University) for HSV-1 YFP, Dr. Richard Youle (National Institutes of Health) for TAX1BP1-KO HeLa cells, Dr. Nan Yan (UT Southwestern) for STING-eGFP plasmid and Dr. Adolfo García-Sastre (Icahn School of Medicine at Mount Sinai) for Mx-1 GFP mice. We thank John Tawil for the critical reading of the manuscript. We also acknowledge Dr. Loic Dragin for initial contributions to this project. We thank Dr. Han Chen for assistance with TEM studies. The experiments in this manuscript used the Penn State College of Medicine Advanced Light Microscopy Core, TEM Core and the Flow Cytometry Core. The Advanced Light Microscopy Core (RRID: SCR_022526), the TEM Core (RRID:SCR_021200) and Flow Cytometry Core (RRID:SCR_021134), services and instruments used in this project were funded, in part, by the Pennsylvania State University College of Medicine via the Office of the Vice Dean of Research and Graduate Students and the Pennsylvania Department of Health using Tobacco Settlement Funds (CURE). The content is solely the responsibility of the authors and does not necessarily represent the official views of the University or the College of Medicine. The Pennsylvania Department of Health specifically disclaims responsibility for any analyses, interpretations, or conclusions. This work was supported by NIH grant R01 AI162815 (to E.W.H.).
Author information
Authors and Affiliations
Contributions
S.S., S.M., C.C.N. and E.W.H. designed the experiments. S.S. and S.M. performed most of the experiments. I.E.R. and C.C.N. performed the Ectromelia infections in mice. S.S., S.M., J.W., C.C.N. and E.W.H. analyzed the data. Y.B.C. and S.M.M. performed initial experiments and Y.B.C. also performed the NanoBiT assays and generated the Golgiphagy reporter. N.S. contributed to the generation of Tax1bp1fl/fl mice. D.C. performed bioinformatics analysis. E.W.H. and S.S. wrote the manuscript. S.S., S.M., J.W., Y.B.C. and C.C.N. edited the manuscript. E.W.H. conceived and supervised the project and acquired funding for it. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Zhi Jian Chen who co-reviewed with Jing Zhang; Terje Johansen, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Suklabaidya, S., Mohanty, S., Reider, I.E. et al. Negative feedback regulation of STING signaling by TAX1BP1-directed Golgiphagy. Nat Commun 17, 2762 (2026). https://doi.org/10.1038/s41467-026-69422-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69422-z