Abstract
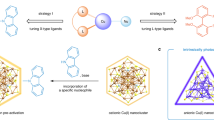
The abundance, low cost, and distinctive properties of copper are driving increased interest in copper nanoclusters. However, modular synthesis of such clusters remains underdeveloped. Here, we report a modular strategy for constructing copper nanoclusters via the synergistic combination of photochemical activation and nanoscale ligand-exchange. Using a readily scalable precursor—[Cu14(TXP35)4(BnSe)12H]+ (Cu14)—we demonstrate its photo-facilitated ligand-exchange with exogenous phosphine ligands, enabling the precise synthesis of 18 copper nanoclusters. This approach also allows functional motifs, such as chiral ligands, to be incorporated into the cluster framework. Mechanistic studies supported by in situ characterization reveal that upon light exposure, Cu14 decomposes into diverse intermediates—including copper atoms/clusters, selenium, benzyl groups, TXP35, and their combinations—which then reassemble in the presence of external ligands to form new nanoclusters. Our modular nanosynthesis establishes a versatile platform for the atomic-level engineering of copper nanoclusters and is expected to advance the tailored design of cluster-based nanomaterials.
Similar content being viewed by others
Introduction
The modular and precise synthesis of nanomaterials with tailored structures and properties represents a central objective and a potentially transformative advance in materials science1,2,3. Achieving such synthetic control is essential for elucidating fundamental mechanisms and establishing clear structure–property relationships4,5,6. Within this framework, the synthesis of structurally well-defined metal nanoclusters marks significant progress7,8,9,10. These nanoclusters comprise assemblies ranging from several to hundreds of metal atoms, stabilized by protecting ligands, and are characterized by atomic-level structural resolution11,12,13,14,15,16,17,18. This precision allows for deliberate fine-tuning of their properties19,20,21. Owing to their diverse structural motifs, high stability, ease of functionalization, and unique physicochemical characteristics, metal nanoclusters have attracted considerable interest for applications in catalysis, energy storage, optics, electronics, and biology22,23,24,25,26,27,28,29,30,31,32. Over the past decades, substantial efforts have been devoted to developing diverse synthetic strategies for structurally precise metal nanoclusters, primarily focusing on Au, Ag, and their bimetallic systems33,34,35,36,37,38,39,40. In contrast, methods for the modular and precise synthesis of Cu nanoclusters remain markedly underdeveloped41, despite the notable advantages of Cu nanoclusters-such as their greater earth-abundance, cost-effectiveness, and functional properties that are comparable to, if not broader than, those of Au and Ag analogs42,43,44,45,46,47. For example, recent pioneering studies have clearly demonstrated the great potential of Cu nanoclusters in catalysis48,49,50,51,52.
The predominant strategy for synthesizing copper nanoclusters involves stimuli-driven bottom-up assembly of Cu atoms in the presence of coordinating ligands42,53. Various—such as chemical agents (e.g., NaBH4 and silanes), thermal energy, and photoexcitation (Fig. 1a)—have been employed to initiate this process44,54,55,56. By carefully controlling synthetic parameters, including the type of stimulus, reaction temperature, solvent, and ligand structure, diverse compositions and architectures of Cu nanoclusters can be achieved57. For instance, reduction with NaBH4 often yields hydride-functionalized copper nanoclusters, where hydride ligands are integrated within the cluster framework, significantly influencing both electronic properties and structural stability58,59,60,61. Ligand exchange has emerged as another powerful strategy for the modular and precise synthesis of Cu nanoclusters62,63. This approach utilizes well-defined precursor clusters and introduces exogenous ligands, enabling the preparation of nanoclusters with tailored compositions, structures, and tunable physicochemical properties (Fig. 1b)64,65. The atomic precision of both precursors and products offers insights into ligand-exchange mechanisms, as well as the correlation between structural transformations and corresponding property modifications66,67. Despite these advances, both strategies face significant challenges. Stimuli-driven bottom-up assembly often suffers from poor size control, limited monodispersity, a narrow ligand scope, and challenges in mechanistic studies. The conventional ligand-exchange approach remains underexplored to copper nanoclusters due to issues such as incomplete exchange, purification and characterization hurdles, and the limited stability of parent clusters68,69.
a The strategy of stimuli-driven bottom-up assembly established in prior studies. b The strategy of ligand-exchange established in prior studies. c The strategy of photosynergetic ligand-exchange developed in this work.
Herein, we present a photosynergetic ligand-exchange strategy for the modular and robust synthesis of copper nanoclusters. This approach combines ligand exchange, which permits precise control over surface chemistry and functionality, with the photochemical activation of precursor clusters, enabling synergistic regulation over both reaction kinetics and thermodynamics. As a result, nanoclusters with tailored compositions and architectures are readily accessible (Fig. 1c). We employed a readily scalable and structurally well-defined [Cu14(TXP35)4(BnSe)12H]+ cluster (denoted as Cu14, where TXP35 = tris(3,5-dimethylphenyl)phosphine and Bn = benzyl), co-stabilized by benzyl selenolate and phosphine ligands, as a molecularly precise precursor. Under light irradiation, Cu14 undergoes simultaneous decomposition into diverse intermediate species while participating in ligand exchange with exogenous ligands, ultimately leading to the formation of 18 distinct atomically precise copper nanoclusters. Below, we delineate the synthetic procedures, underlying mechanisms, molecular structures, and functional exploration of all 19 copper nanoclusters.
Results
Synthesis, molecular structure and photosensitivity of Cu14 precursors
A simple one-pot reduction strategy was employed to synthesize the target copper nanocluster precursors50. As illustrated in Fig. 2a, Cu(MeCN)4BF4 was dissolved in acetonitrile as the copper source, while tris(3,5-dimethylphenyl)phosphine (TXP35) and dibenzyldiselenide (BnSe–SeBn) were co-dissolved in chloroform. A methanolic solution of sodium borohydride was then added dropwise to the mixture. After stirring at room temperature for one hour, the solution color transitioned gradually from yellow to orange-red. The mixture was centrifuged, and the precipitate was collected and redissolved in chloroform (see the Supplementary Information (SI) for detailed experimental procedures). The solution was stored statically in the dark for two weeks, yielding orange crystals with an approximate yield of 36.5% (Fig. S1). The crystalline product was employed in all experiments to characterize the cluster’s properties. It is worth noting that this synthetic procedure can be readily scaled up for bulk preparation, providing a highly pure and easily accessible precursor for subsequent transformations (Fig. S2).
a The large-scale synthetic prototype of Cu14. b Molecular structure of Cu14. c The ESI-MS spectra of Cu14(H/D) in positive mode. Inset: Experimental (blue) and simulated (green) isotopic distributions of [Cu14(TXP35)4(BnSe)12(H/D)]+. In situ UV–vis absorption spectra (d) and ESI-MS spectra (e) of Cu14 in chloroform under illumination.
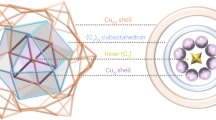
The molecular structure of the precursor cluster was elucidated by single-crystal X-ray diffraction (SCXRD), indicating that Cu14 crystallizes in the cubic space group \(F\)__4\(3c\) (see Figs. S3-S4 and Table S1 in the SI). The cluster adopts a quasi-tetrahedral architecture (Fig. 2b). Structurally, it consists of a Cu8(BnSe)12(TXP35)4 shell encapsulating an octahedral H@Cu6 core. Within the outer Cu8 shell, each copper atom is bridged in a triangular fashion by three adjacent selenide ligands. Moreover, the four copper atoms situated at the tetrahedral vertices are each capped by an auxiliary phosphorus-donor ligand. The Cu–Cu bond distances in the H@Cu6 core are uniform at 2.817 Å. In the outer shell, the Cu–P bond length remains constant at 2.2139 Å, while the Cu–Se bond lengths vary between 2.357 and 2.464 Å. These values align well with those reported for other well-characterized copper nanoclusters40,70.
To unequivocally confirm the presence of a hydride and determine the precise molecular formula of the Cu14 cluster, the compound was synthesized using both NaBH4 and NaBD4 as reducing agents. The resulting hydride and deuteride analogs were characterized by electrospray ionization mass spectrometry (ESI-MS) in positive ion mode. The crossover isotope-labeling experiment revealed a + 1 m/z shift in the mass peak of the deuteride cluster relative to the hydride form, confirming that the hydride originates from the borohydride or borodeuteride reagent and does not exchange with protic solvents (Fig. 2c). Although the dominant peak at m/z = 4317.4 corresponds to the molecular ion [Cu14(TXP35)4(BnSe)12H]+, additional signals attributable to fragment ions or adducts derived from the parent cluster were also observed. The excellent match between experimental and simulated isotopic patterns supports the proposed molecular assignment (Fig. S5). X-ray photoelectron spectroscopy (XPS) further confirmed that all copper atoms are in the +1 oxidation state, as evidenced by the Cu 2p peaks at 952.53 and 932.72 eV and a Cu LMM Auger peak at 916.42 eV (Fig. S6), which is consistent with the assigned molecular formula61. The crystallographic characteristics, elemental composition, and phase purity of Cu14 were further characterized by energy-dispersive X-ray spectroscopy (EDS) mapping (Fig. S7) and nuclear magnetic resonance (NMR) spectroscopy (Figs. S8 and S9), as provided in the SI.
We further investigated the electronic structure and stability of the cluster. As shown in Fig. S10, the cluster in chloroform appears yellow and displays a characteristic absorption peak at 242 nm with a shoulder peak at 266 nm in the ultraviolet–visible (UV–vis) spectrum. The cluster remains stable in solution in the dark (Fig. S11), a property likely due to its large electrochemical gap of 1.83 eV, as determined by cyclic voltammetry (CV, Fig. S12). Notably, the cluster solution undergoes a rapid transformation upon light exposure, a phenomenon not observed in other copper clusters (Fig. S13)71,72,73. As illustrated in Fig. 2d, the UV–vis profile of Cu14 changed significantly under illumination: after 45 min, the shoulder peak disappeared, and a new sharp peak emerged at 270 nm, indicating a photochemical transformation that led to the formation of new species. To probe the reaction pathway, we employed in situ ESI-MS to analyze the products formed during irradiation (see Fig. S14 for the setup). At time zero, the peaks were grouped into three categories: peaks 1–4 corresponded to fragments where the core kernel had begun to disintegrate, with complete or partial loss of selenium atoms from the core; peaks 5–7, exhibiting the highest intensities, indicated that the residual kernel after the detachment of benzyl groups and phosphine ligands, which subsequently lost partial or complete copper atoms from the outer shell; finally, peaks 8-11 largely retained intact structures (Fig. 2e). During the initial 45 min, the spectral changes followed a monotonic trend: peaks 5-11 decreased to varying degrees, while fragment peaks 1–4 increased significantly. A distinct transition occurred between 45 and 60 min, marked by a sharp rise in peaks 1 and 2, accompanied by a pronounced decline or complete disappearance of peaks 3–7. Notably, molecular ion peak 10 also disappeared within this interval. By 120 min, the transformation of Cu14 was nearly complete, with peaks 1 and 2 becoming dominant. The uniformly spaced peaks observed at this stage were identified primarily as adducts of peak 2 with diethyl ether molecules. These results suggest that Cu14 gradually loses phosphine ligands, benzyl groups, and selenide atoms, yielding both relatively intact kernels and fragmented core structures under light exposure. This indicates that the transformation kinetics and thermodynamics of Cu14 can be modulated through photochemical treatment. It should be noted that further extension of the exposure time to 300 and 600 min results in progressively smaller cluster species, culminating in minute fragments that cannot be distinctly identified by ESI-MS.
Cu14 for photosynergetic ligand-exchange synthesis of 16 copper nanoclusters
As mentioned above, light exposure of Cu14 produced both relatively intact kernel structures along with fragmented cores. Building on these observations, we anticipated that introducing external ligands could promote its in situ transformation into other clusters. In a typical photo-synergized ligand-exchange reaction, one equivalent of Cu14 was mixed with 0.4 equivalents of external phosphine ligands (denoted as P, Fig. 3) in a mixture of chloroform and methanol at room temperature. The reaction mixture was irradiated for 5 hours under aerobic conditions to facilitate complete conversion. The crude product was then collected and purified, affording single crystals suitable for structural analysis. In some cases, additives such as NaSbF6, PPh4Cl, or PPh4BPh4 were introduced as counterions to promote crystallization.
Electron-donating (P1-P4)/withdrawing (P5-P7) triarylphosphines afford Cu32-P1-P7 (orange crystals). The phosphine ligands with functional groups (halides, pyridine, amino, methoxyl; P6-P10) are tolerated, giving Cu32-P6-P10. Alkyl phosphines (P11-P12) yield Cu32-P11-P12. P(pyrr)3 decomposes to PyrrCa, forming Cu38-PyrrCa. Multidentate phosphines (DPPM, BINAP, DPPP) produce Cu3-DPPM, Cu2-BINAP, and Cu8-DPPP, respectively. All products were obtained as high-quality single crystals (right panel).
Importantly, this photosynergetic ligand-exchange strategy applies to a broad range of structurally diverse substrates. As illustrated in Fig. 3, both electron-donating (P1-P4) and electron-withdrawing (P5-P7) triarylphosphines proved effective, consistently affording orange crystals of [Cu32(P1)6(BnSe)18Se7]+ (Cu32-P1), [Cu32(P2)6(BnSe)18Se7]+ (Cu32-P2), [Cu32(P3)6(BnSe)18Se7]+ (Cu32-P3), [Cu32(P4)6(BnSe)18Se7]+ (Cu32-P4), [Cu32(P5)6(BnSe)18Se7]+ (Cu32-P5), [Cu32(P6)6(BnSe)18Se7]+ (Cu32-P6), and [Cu32(P7)6(BnSe)18Se7]+ (Cu32-P7). The transformation tolerated various functional groups, including halides (P6-P7), pyridine (P8), amino (P9), and methoxyl (P10) groups, giving rise to Cu32-P6, Cu32-P7, [Cu32(P8)6(BnSe)18Se7]+ (Cu32-P8), [Cu32(P9)6(BnSe)18Se7]+ (Cu32-P9), and [Cu32(P10)6(BnSe)18Se7]+ (Cu32-P10). Alkyl phosphines, such as dicyclohexylphenylphosphine (P12), also performed to produce [Cu32(P11)6(BnSe)18Se7]+ (Cu32-P11), and [Cu32(P12)6(BnSe)18Se7]+ (Cu32-P12). It is noteworthy that tri(pyrrolidin-1-yl) phosphine (P(pyrr)3) decomposed under the reaction conditions, yielding pyrrolidine-1-carboxylic acid (PyrrCa) that subsequently stabilized the cluster architecture (Cu38(PyrrCa)6(BnSe)24Se7, denoted as Cu38-PyrrCa). In addition to monodentate phosphines, multidentate phosphine ligands (bis(diphenylphosphino)methane, abbreviated as DPPM; 1.1´-binaphthyl-2.2’-diphemyl phosphine, abbreviated as BINAP; 1,3-Bis(diphenylphosphino)propane, abbreviated as DPPP) were also compatible with the transformation of Cu14, leading to clusters with varying nuclearities depending on the functional groups present ([Cu3(DPPM)3(BnSe)2]+; denoted as Cu3-DPPM; Cu2(BINAP)2(BnSe)2, denoted as Cu2-BINAP; and Cu8(DPPP)2(BnSe)8, denoted as Cu8-DPPP). In all cases, the corresponding cluster products were obtained as high-quality single crystals (Fig. 3, right).
The structures of all cluster products were unambiguously established by SCXRD. Their composition, copper oxidation states, purity, and electronic properties were further characterized through a combination of techniques, including XPS, 1H NMR spectroscopy, UV–vis absorption spectroscopy, and CV. Detailed data are provided in the SI (Fig. S15–S105 and Tables S2–S17). As shown in Fig. 4, Cu32 clusters were obtained with most monodentate phosphine ligands (P1-P12), whereas the P(pyrr)3 ligand led to the formation of a Cu38-PyrrCa cluster. Bidentate phosphines yielded Cu2-BINAP, Cu3-DPPM and Cu8-DPPP clusters, respectively, with structural variations depending on the specific ligand architecture. It is noteworthy that further structural diversity in cluster assembly may be achieved by adjusting reaction parameters such as phosphine ligand design, stoichiometry, light intensity, temperature, and solvent choice.
Atomic color: blue, Cu; green, Se; purple, P; yellow, C; dark blue, N; red, O; turquoise F; dark green, Cl; purplish red, Br.
The detailed structural analysis of all clusters is provided below. Given that all Cu32 clusters exhibit similar molecular structures, only Cu32-P1 is anatomized as a representative example. As depicted in Fig. 5a, the Cu32-P1 cluster adopts an overall flattened sandwich-like architecture. The central region comprises four Cu atomic layers arranged with threefold symmetry. The two inner Cu layers are interconnected through six directly bonded Cu2 dimers as the fundamental building units, which are further stabilized by peripheral BnSe⁻ ligands via bridging coordination. A single Se atom is encapsulated at the geometric center between these two inner layers. In contrast, each of the two outer (terminal) Cu layers consists of one Cu3Se3 hexagonal ring and one central Cu atom. Collectively, these four Cu layers and the incorporated Se atoms form a Cu20Se19 sandwich core. Each end of this central motif is capped by an outer structural unit with C3 symmetry, formulated as (Cu9Se9P3). The core of this outer motif features a crown-like Cu6 substructure. Three isolated Cu atoms extend from this Cu6 core through BnSe⁻-mediated bridging and are situated at the vertices of the motif. Each vertex Cu atom is further coordinated by a P atom from the PPh3 ligand, completing the structural stabilization of the outer motif.
a Total structure and structure anatomy of Cu32-P1. It describes the fusion of a Cu20Se19 core and two Cu9Se9P3 motifs. b Total structure and structure anatomy of Cu38-PyrrCa. The structure of Cu38-PyrrCa is derived from Cu32-P1 via P(pyrr)3 decomposition (pyrrolidine-1-carboxylic acid) and Cu–Cu bond cleavage/reconstruction. c Total structure and structure anatomy of Cu2-BINAP. d Total structure and structure anatomy of Cu3-DPPM. e Total structure and structure anatomy of Cu8-DPPP.
Among the transformation reactions involving various monodentate phosphine ligands, an interesting exception was observed: P(pyrr)3 decomposes under the reaction conditions to form a pyrrolidine-1-carboxylic acid species, which subsequently exerts a stabilizing effect on the cluster. This transformation involves the cleavage of two Cu–Cu bonds that connect the terminal copper cores in the original Cu32 structure, accompanied by the formation of new bonds with additional Cu atoms (Fig. 5b). These newly incorporated Cu atoms are further bridged by BnSe- ligands, whose strong Se–Cu interactions promote the cleavage of the original Cu–Cu bonds. The three Cu atoms originally coordinated to P in Cu32 now bind to an O atom from the generated pyrrolidine-1-carboxylic acid species, ultimately leading to the formation of the unique Cu38(PyrrCa)6(BnSe)24Se7 cluster. Notably, the central Cu20Se19 sandwich moiety remains largely consistent between Cu32 and Cu38-PyrrCa structures.
As illustrated in Fig. 5c, the two copper atoms in Cu2-BINAP are bridged by two selenium atoms in a µ₂ mode, with both metal centers further stabilized by two bidentate BINAP ligands. Structural characterization (Fig. 5d) shows that the Cu3-DPPM cluster core comprises three discrete copper atoms bridged by a central selenium atom in a μ3-mode, with two of the copper atoms additionally connected via an additional selenium atom. The Cu–Se bond distances are measured to be 2.380 Å and 2.438 Å. This core is additionally stabilized by three bidentate DPPM ligands arranged around its periphery.
Figure 5e displays the molecular structure of Cu8-DPPP. At its core lies a Cu4 tetrahedral unit, with two of its vertices each connected to an additional copper atom, thereby forming an overall Cu8 framework. The coordination modes between copper atoms and BnSe ligands are diverse, encompassing three μ2-, four μ3-, and one μ4-bridging selenium atoms. This diversity in coordination is further reflected in the variation of Cu–Se bond lengths, which primarily fall within the range of 1.500–3.000 Å. The cluster framework is encapsulated by two bidentate DPPP ligands, coordinated to copper atoms via their phosphorus donors. Similar to BINAP and DPPM ligands, the characteristic steric bulk and bent coordination geometry of DPPP are crucial for maintaining the structural integrity of the cluster. The steric hindrance imposed by these bidentate phosphine ligands effectively suppresses aggregation of the metal framework, while their distinct coordination geometries contribute to structural diversity in the metal core.
Chiral copper nanoclusters via photosynergetic ligand-exchange
The photosynergetic ligand-exchange strategy enables the introduction of chirality into copper nanoclusters (Fig. S106). As illustrated in Fig. 6a, Cu14 was efficiently converted into [Cu10(2 R,4 R/2S,4S-BDPP)3(BnSe)9]⁺ (denoted as Cu10-R/S) in the presence of the chiral ligand (2 R,4 R/2S,4S-BDPP) (where BDPP = 2,4-bis(diphenylphosphino)pentane) under light irradiation. Figure 6b and Fig. S107 confirm that the resultant product is derived entirely from the parent Cu14 cluster. A prominent peak at m/z 3488.6 in the mass spectrum corresponds to the molecular ion [Cu10(2 R,4 R/2S,4S-BDPP)3(BnSe)9]⁺. The UV–vis spectra of Cu10-R and Cu10-S are identical, both exhibiting a characteristic absorption band at 246 nm (Fig. 6c). As expected, their circular dichroism (CD) spectra display mirror-image symmetry in the 250–700 nm range (Fig. 6d). Specifically, Cu10-R displays positive Cotton effects at 290 and 351 nm, and negative Cotton effects at 268, 319, and 389 nm. The optical activity in the visible region is attributed to ligand-to-metal charge transfer (LMCT), as the intrinsic optical activity of the organic ligands is typically silent in this spectral region (Fig. S108)74,75,76. For additional characterizations of Cu10-R/S, refer to Fig. S109–S114.
a Overall reaction. b The ESI-MS spectra in positive mode. c The UV–vis spectrum in chloroform. d The circular dichroism spectrum. e Anatomical representations of the two enantiomers.
The chiral structures of the Cu10-R/S clusters were characterized by SCXRD (Fig. S115–S118 and Tables S18–S19). Crystallographic analysis reveals that the Cu10-R and Cu10-S crystallize in the chiral space groups P32 and P31, respectively. The space groups are analogous to left- and right-handed quartz crystals: while macroscopically identical, they exhibit opposite helical atomic arrangements, resulting in nonsuperimposable mirror-image structures (Fig. S119)77. The overall structure of Cu10 features a Cu4 core stabilized by a shell composed of Cu6(BDPP)3(BnSe)9. As shown in Fig. 6e, chirality in the clusters is manifested at multiple levels: the metal core geometry, the metal-ligand shell organization, and the ligand spatial arrangement. In both enantiomers, the chirality of the Cu4 core arises from distortions in its tetrahedral geometry, evidenced by variations in Cu–Cu bond lengths and Cu–Cu–Cu bond angles (Fig. S120-S121). Further analysis shows that this structural distortion is primarily induced by the irregular arrangement of surface selenolate ligands (Fig. 6e), whose steric repulsion reduces the point group symmetry of the core from tetrahedral Td to C1, which possesses only the identity symmetry operation. The core is surrounded by three Se–Cu–Se–Cu–Se motifs, which adopt either clockwise or counterclockwise rotational arrangements along the pseudo-C3 axis. Furthermore, the inherent chirality of the 2 R,4R- and 2S,4S-BDPP ligands on the cluster surface imparts additional stereochemical specificity, consistent with the distinct CD signals. Overall, the chirality of Cu10-R/S results from the synergistic interplay of a distorted metal core, chiral ligand arrangement, and intrinsic ligand configuration78,79,80.
Mechanism studies
We further proceeded to investigate the mechanism of the photosynergetic ligand-exchange process, selecting the conversion of Cu14 to Cu10 as a representative model owing to the well-defined ESI-MS spectra of both clusters (Fig. 7a). This study focused on three key questions: (1) Is photoactivation essential for the transformation? How does the photosynergetic ligand-exchange process differ from conventional ligand-exchange? (2) Which step occurs more rapidly: photoexcitation or ligand-exchange? (3) How does the structural framework reorganize from Cu14 to Cu10 during photochemical mediation?
a Overall reaction for the conversion of Cu14 to Cu10. b In situ UV–vis absorption spectroscopy, c In situ EPR spectroscopy. Chloroform as the solvent, tested at 293 K with a frequency of 9.423276 GHz. d In situ mass spectroscopy. e Schematic diagram.
To address these questions, we first monitored the reaction using in situ UV–vis absorption spectroscopy, with and without light irradiation. Upon adding BDPP to Cu14 in chloroform, the UV–vis spectrum of the mixture remained unchanged in the absence of light, indicating negligible reaction progress under dark conditions (Fig. S122). In contrast, under light irradiation, significant spectral changes were observed (Fig. 7b). The initial spectrum at 0 min displayed a distinct peak at 245 nm, accompanied by a shoulder peak at 266 nm. Within the first 45 min of illumination, the peak intensity gradually diminished to its minimum while the position remained largely unchanged. Subsequently, the intensity increased concurrently with a red shift, ultimately stabilizing at 275 nm upon continuous irradiation.
These results clearly demonstrate that light irradiation is essential to drive the reaction. To further elucidate the mechanism, electron paramagnetic resonance (EPR) spectroscopy was performed using 5,5-dimethyl-1-pyrroline N-oxide (DMPO) as a radical trapping agent. As depicted in Fig. 7c, distinct EPR signals were detected for the [Cu14 + DMPO] system under light irradiation, confirming radical generation from Cu14. The spectral parameters (g = 2.0061, AN = 14.3, AH = 20.1) match those of a DMPO-benzyl radical adduct, strongly supporting photodissociation of benzyl groups from the cluster (Fig. S123)81. The formation of benzyl radicals is consistent with their high thermodynamic stability82. Together, these results demonstrate that the transformation proceeds via a light-induced radical mechanism.
In situ mass spectrometry analysis of Cu14 upon the addition of BDPP revealed further insight into the transformation pathway. During the initial 45 min, the spectral pattern aligned with previous observations, dominated by cluster fragments 1–3—indicative of core structural decomposition—as well as relatively intact clusters 4–6. These signals gradually decreased and eventually disappeared during this period, confirming that the photo-driven transformation initiates immediately upon irradiation. Additionally, a peak corresponding to the [Cu14-BDPP complex] was detected from the outset, indicating that ligand exchange occurs promptly after mixing the external ligand with Cu14. These results suggest that photoactivation and ligand-exchange proceed on a comparable timescale.
In good agreement with the observations from in situ UV–vis absorption spectroscopy, characteristic peaks corresponding to the transformed Cu10 species emerged after 60 min of irradiation (Fig. 7d). Peaks 8–10 were attributed to incomplete kernel fragments. Their intensities exhibited oscillatory behavior, reflecting a dynamic equilibrium between the formation and decomposition of these species. By 600 min, the spectrum was dominated by peak 8, which represents the core kernel fragment of Cu10, thus confirming the identity of the photo-transformation product.
Based on the comprehensive experimental evidence, we propose the following reaction mechanism (Fig. 7e): (1) Photoactivation and concurrent ligand-exchange: under light irradiation, partial cleavage of C–Se bonds in the BnSe ligands on Cu14 generates benzyl and selenium-centered radicals. Simultaneously, ligand-exchange takes place between BDPP and the parent Cu14 cluster, yielding BDPP-functionalized Cu14 species. (2) Peripheral-to-core exfoliation: both parent and BDPP-functionalized Cu14 clusters undergo sequential exfoliation from the periphery inward. Outer phosphine ligands and benzyl groups are generally released first, while copper atoms directly coordinated to phosphine ligands may detach upon ligand dissociation. Selenium atoms are released only after full structural exposure. (3) Binding and fragmentation: free BDPP molecules bind to incomplete kernel fragments, but the resulting adducts remain unstable and undergo further fragmentation into molecular species including Cu, Se, benzyl groups, and phosphine ligands. (4) Reassembly into Cu10: these molecular fragments subsequently reassemble to form the kernel of the transformed Cu10 cluster, ultimately leading to the final product.
Discussion
In summary, we have developed a modular strategy for synthesizing diverse copper nanoclusters through the synergistic combination of photochemical activation and nanoscale ligand-exchange. Central to this approach is the use of benzyl-functionalized precursor clusters, which undergo light-induced transformation into various intermediates. The pronounced photosensitivity of benzyl-functionalized Cu14 demonstrated here is expected to inspire further exploration of this cluster family. This strategy proves highly efficient, enabling the precise synthesis of 18 atomically precise copper nanoclusters, along with tailored functionalization—including chiral induction—via rational design of external ligands. From a practical standpoint, this technically straightforward method facilitates accelerated exploration of nanoclusters beyond copper. Mechanistic insights suggest that cluster formation follows principles of modular chemistry, a concept that warrants deeper investigation. Studies expanding this strategy to nanocluster synthesis, functionalization, mechanistic elucidation, and applications in photocatalysis are currently underway in our laboratories.
Methods
Reagents
Cuprous oxide (Cu2O, A.R.) and tetrafluoroboric acid (HBF4, 50 wt.%) were purchased from Macklin (Shanghai, China). Acetonitrile (CH3CN, A.R.) was purchased from Tianjin Xinbote Chemical Co., Ltd. (Tianjin, China). Sodium borohydride (NaBH4, 98%) and methanol (CH3OH, A.R.) were purchased from Tianjin Fengboat Chemical reagent Technology Co., Ltd. (Tianjin, China). Triphenylphosphine (C18H15P, 99%) was purchased from Adamas (Shanghai, China). Dibenzyldiselenide (C14H14Se2, 95.63%), sodium hexafluoroantimonate (NaSbF6, 98%), and other phosphine ligands were purchased from Bide Pharmatech Ltd. (Shanghai, China). (2S,4S)-2,4-Bis (diphenylphosphino)pentane and (2 R,4 R)-2,4-Bis (diphenylphosphino)pentane (C29H30P2, 98%) were purchased from ACMEC (Shanghai, China). Sodium borodeuteride (NaBD4, 95%), 2-pyridyldiphenylphosphine (C15H12N3P, 97%), and bis(diphenylphosphino)methane (C25H22P2, 98%) were purchased from Energy Chemical. (Anhui, China). Tetraphenylphosphonium chloride (PPh4Cl, 98%) was purchased from Damas-beta (Shanghai, China). Tetraphenylphosphine tetraphenylboron (PPh4BPh4, 95%) was purchased from TCI (Shanghai, China). Chloroform (CHCl3, A.R.) and diethyl ether (C4H10O, A.R.) were purchased from Shanghai Titan Scientific Co. Ltd. (Shanghai, China).
Instrument and characterization
Ultraviolet–visible (UV–vis) and circular dichroism (CD) spectroscopy
UV–vis spectra were collected by a JASCO V-780 Spectrophotometer and CD spectra were recorded on a JASCO J-815 CD Spectrometer using a quartz cuvette with a 1 mm path length. The scanning speed is 1000 nm min–1. The spectra were recorded in diluted solutions of chloroform, and the signal of the blank solvent was subtracted. The spectra were recorded at room temperature.
Electrospray ionization mass measurement
Electrospray ionization mass spectra (ESI-MS) were recorded using an Agilent 6224 time-of-flight mass spectrometer. The samples dissolved in dichloromethane were directly infused at a flow rate of 1.2 mL/h by a syringe pump. Typical parameters used for the measurements were as follows: capillary voltage: 4.0 kV; drying gas temp: 150°C; drying gas flow: 4 L/min; nebulizer pressure: 20 psi.
X-ray photoelectron spectroscopy (XPS) measurements
X-ray Photoelectron Spectroscopy (XPS) data were acquired using an ESCALAB XI+ spectrometer at ambient temperature, and the obtained spectra were subsequently calibrated with respect to the C 1 s peak at 284.8 eV.
Electron paramagnetic resonance test
Electron paramagnetic resonance (EPR) spectra were acquired using an EMXplus electron paramagnetic resonance spectrometer.
Dispersive X-ray spectrometry images (EDS)
EDS spectroscopy analysis was recorded on a Bruker XFlash6100 system.
Nuclear magnetic resonance spectroscopy (NMR)
NMR (298 K, CDCl3) spectra of the samples were acquired on a Bruker AV-600 MHz spectrometer and were calibrated using the residual solvent peak as an internal reference (reported in ppm); all data were processed utilizing MestReNova software.
Cyclic voltammetry (CV) experiments
The electrochemical properties of cluster were studied by CV experiments, and the electrochemical measurements were carried out on CHI760E electrochemical workstation (Shanghai Chenhua Co., China) with a typical three-electrode system under N2 atmosphere at room temperature. During all measurements, the Pt wire and silver/silver chloride electrode (Ag/AgCl) were served as the counter and reference electrodes, respectively. The working electrode was a glassy carbon electrode (GCE) with a geometric area of approximately 0.126 cm2 (calculated as π × (0.2 cm)2). Moreover, the solvent/electrolyte system was used in a small glass cell containing DCM/0.1 M TBAPF6. For the CV experiments, the voltage range we used is –2.0 V ~ 2.0 V (vs Ag/AgCl), the scan rate is 0.05 V/s.
X-ray single crystal diffraction
The single crystal diffraction data of the clusters was collected at 100 K using an Agilent Technologies Super Nova System X-ray diffractometer. The data was collected at Cu Kα (λ = 1.54184 Å) and processed using CrysAlisPro after testing. The structure was solved and refined using the full matrix least squares method based on F2 in ShelXT83, and ShelXL84 within Olex285. The geometric shape and displacement parameters of the atoms in the cluster were properly restricted and constrained. The thermal ellipsoid in the ORTEP diagram was completed with a 50% probability. Detailed crystal data and structure refinements for this compound can be found in Table S1. CCDC 2484271-2484275, 2484277-2484289, and 2499652 contain the supplementary crystallographic data for this paper. Further details can be obtained from the cif files deposited at the Cambridge Crystallographic Data Center and are available free of charge upon request.
Synthesis procedure
Synthesis of Cu14
[Cu(MeCN)4]BF4 (50 mg, 0.16 mmol) was dissolved in acetonitrile (1 mL). A chloroform (1 mL) solution of tris(3,5-dimethylphenyl)phosphine (62.5 mg, 0.18 mmol) was sequentially added, followed by a chloroform (0.5 mL) solution of dibenzyl diselenide (37.4 mg, 0.11 mmol) and a methanol (0.5 mL) solution of sodium hexafluoroantimonate (25 mg, 0.10 mmol). Then, a freshly prepared methanol solution (1 mL) of NaBH4 (50 mg, 1.3 mmol) was added dropwise as the reducing agent, yielding a clear red solution. The mixture was stirred for one hour and then centrifuged. The filtrate is subjected to crystallization via ether vapor diffusion. After approximately one-week, orange block crystals with the chemical formula [Cu14(C24H27P)4(C7H7Se)12H]+ (Cu14) were obtained (36.5% yield, based on Cu). 1H NMR (600 MHz, CDCl3) δ 8.10, 8.09, 7.47, 7.26, 7.15, 6.95, 4.69, 4.68, 3.49, 2.32, 2.20, 2.02, 1.54, 1.26, 1.22, 1.21, 1.20, 0.89, 0.88, 0.87. 13C NMR (151 MHz, CDCl3) δ 77.21, 77.00, 76.79, 65.84, 21.22, 15.26, 1.01, -0.02.
Large-scale synthesis of Cu14
[Cu(MeCN)4]BF4 (5 g, 16 mmol) was dissolved in acetonitrile (100 mL). A chloroform (100 mL) solution of tris(3,5-dimethylphenyl)phosphine (6.25 g, 18 mmol) was sequentially added, followed by a chloroform (50 mL) solution of dibenzyl diselenide (3.74 g, 11 mmol) and a methanol (50 mL) solution of sodium hexafluoroantimonate (2.5 g, 10 mmol). Then, a freshly prepared methanol (100 mL) solution of NaBH4 (5 g, 130 mmol) was added dropwise as the reducing agent, yielding a clear red solution. The mixture was stirred for one hour and then centrifuged. The filtrate was subjected to crystallization via ether vapor diffusion. After approximately one-week, yellow crystals were obtained as the final product (2.5948 g).
Photosynergetic ligand-exchange transformation to synthesize Cu32-P1
A total of 25 mg (0.006 mmol) of the Cu14 cluster product was dissolved in chloroform. It was mixed with a chloroform solution of PPh3 (4.5 mg, 0.017 mmol). The mixture was exposed to irradiation using either a PLS-SXE300+ Xenon lamp or an incandescent lamp for a specific duration. The resulting mixture was filtered, and the filtrate was subjected to ether vapor diffusion for crystallization. The transformed cluster crystals Cu32-P1 (6.4 mg, 33.8%) were obtained within approximately one week. 1H NMR (600 MHz, CDCl3) δ 8.08, 7.26, 3.49, 3.49, 3.48, 3.47, 3.47, 1.56, 1.26, 1.22, 1.22, 1.21, 1.21, 1.20, 1.19. 13C NMR (151 MHz, CDCl3) δ 133.82, 133.70, 132.14, 128.72, 128.47, 128.43, 77.22, 77.01, 76.80, 65.84, 50.63, 15.25, 1.00, -0.03. Other Cu32 clusters were synthesized following the same procedure but with different phosphine ligands. Cu32-P2 (4.6 mg, 23.4%) 1H NMR (600 MHz, CDCl3) δ 7.27, 7.26, 7.24, 7.23, 7.22, 7.08, 7.07, 7.06, 6.73, 6.72, 6.71, 3.50, 3.48, 3.47, 3.46, 2.39, 1.61, 1.26, 1.22, 1.21, 1.20, 0.07, 0.00. 13C NMR (151 MHz, CDCl3) δ 142.82, 142.64, 134.51, 134.44, 133.07, 130.08, 130.04, 128.66, 126.14, 77.23, 77.02, 76.81, 65.85, 50.85, 21.24, 21.10, 15.27, 1.02, -0.01; Cu32-P3 (9.4 mg, 47.9%) 1H NMR (600 MHz, CDCl3) δ 7.27, 7.26, 7.24, 7.23, 7.22, 7.21, 7.08, 7.07, 7.06, 6.73, 6.72, 6.71, 3.50, 3.49, 3.47, 3.46, 2.39, 1.61, 1.26, 1.22, 1.21, 1.20, 1.03, 1.03, 0.07, 0.00. 13C NMR (151 MHz, CDCl3) δ 142.82, 142.65, 134.51, 134.44, 133.07, 130.08, 130.04, 128.66, 126.14, 77.23, 77.02, 76.81, 65.86, 50.86, 29.71, 21.25, 21.10, 15.27, 1.02, -0.01; Cu32-P4 (8.6 mg, 44.8%) 1H NMR (600 MHz, CDCl3) δ 7.34, 7.28, 7.26, 7.24, 7.21, 6.77, 6.72, 3.50, 3.48, 3.47, 3.46, 2.39, 2.02, 2.01, 2.00, 1.61, 1.26, 1.22, 1.21, 1.21, 1.20, 0.89, 0.88, 0.87, 0.08, 0.07, 0.00. 13C NMR (151 MHz, CDCl3) δ 142.82, 142.29, 142.13, 134.09, 133.96, 133.06, 132.74, 130.10, 129.90, 128.78, 128.66, 128.59, 128.54, 126.15, 126.00, 77.23, 77.02, 76.81, 65.86, 21.32, 21.25, 21.19, 21.11, 15.28, 1.02, -0.01; Cu32-P5 (6.7 mg, 33.8%) 1H NMR (600 MHz, CDCl3) δ 7.50, 7.49, 7.36, 7.35, 7.34, 7.32, 7.31, 7.26, 7.26, 7.20, 7.19, 7.18, 7.11, 7.10, 7.08, 7.08, 7.07, 7.06, 7.05, 7.05, 6.95, 6.94, 6.92, 3.49, 3.48, 3.47, 3.46, 3.07, 2.23, 2.21, 2.20, 2.03, 2.01, 2.00, 1.99, 1.60, 1.26, 1.22, 1.20, 1.19, 1.02, 1.01, 0.89, 0.88, 0.87, 0.84, 0.07, 0.07, -0.00, -0.00. 13C NMR (151 MHz, CDCl3) δ 163.75, 163.70, 162.10, 162.05, 138.67, 130.42, 130.36, 130.31, 129.93, 129.50, 129.48, 129.37, 129.34, 129.05, 129.00, 128.79, 128.30, 120.20, 120.08, 120.06, 119.93, 116.48, 116.34, 77.23, 77.02, 76.81, 65.86, 50.87, 29.78, 29.71, 29.53, 29.48, 29.33, 29.25, 27.22, 25.54, 15.27, 1.02, -0.01; Cu32-P6 (8.2 mg, 39.9%) 1H NMR (600 MHz, CDCl3) δ 7.32, 7.31, 7.26, 7.20, 7.19, 7.18, 3.49, 3.48, 3.47, 3.46, 1.61, 1.25, 1.22, 1.20, 1.19, 1.04, 1.03, 1.02, 1.01, 0.89, 0.88, 0.87, 0.07. 13C NMR (151 MHz, CDCl3) δ 135.66, 134.91, 134.78, 134.64, 134.58, 129.20, 129.04, 128.99, 77.23, 77.02, 76.81, 65.86, 50.86, 29.78, 29.71, 29.62, 29.53, 29.48, 29.33, 29.25, 15.27, 14.12, 1.02, -0.01; Cu32-P7 (12.0 mg, 59.4%) 1H NMR (600 MHz, CDCl3) δ 7.58, 7.37, 7.36, 7.34, 7.30, 7.26, 7.26, 7.21, 6.77, 3.49, 3.48, 3.47, 3.46, 2.39, 2.31, 1.63, 1.25, 1.22, 1.21, 1.19, 0.89, 0.88, 0.87, 0.84, 0.83, 0.07, 0.07, -0.00, -0.00. 13C NMR (151 MHz, CDCl3) δ 134.52, 134.13, 134.00, 133.07, 130.38, 129.23, 128.71, 128.67, 127.43, 77.23, 77.02, 76.81, 65.86, 50.86, 29.33, 15.27, 1.02, -0.01. Cu32-P8 (7.4 mg, 39.0%) 1H NMR (600 MHz, CDCl3) δ 8.70, 8.70, 8.69, 7.55, 7.54, 7.53, 7.50, 7.49, 7.40, 7.39, 7.36, 7.35, 7.34, 7.26, 7.17, 7.16, 7.15, 7.11, 7.10, 5.35, 5.34, 5.34, 3.49, 3.49, 3.48, 3.48, 3.47, 3.46, 3.07, 2.23, 2.21, 2.20, 2.03, 2.01, 2.00, 1.99, 1.59, 1.22, 1.20, 1.19, 0.99, 0.98, 0.97, 0.96, 0.89, 0.88, 0.87, 0.84, 0.07, -0.00. 13C NMR (151 MHz, CDCl3) δ 150.38, 150.29, 135.72, 135.70, 134.26, 134.14, 132.14, 129.12, 128.79, 128.63, 128.58, 128.31, 128.07, 122.26, 77.23, 77.02, 76.81, 65.86, 15.28, 1.02, -0.00; Cu32-P9 (11.6 mg, 62.9%) 1H NMR (600 MHz, CDCl3) δ 7.30, 7.30, 7.29, 7.28, 7.26, 7.24, 7.23, 6.65, 6.64, 3.55, 3.51, 3.49, 3.48, 3.47, 3.46, 3.01, 2.95, 1.25, 1.22, 1.20, 1.19, 0.07, 0.07, -0.00. 13C NMR (151 MHz, CDCl3) δ 151.04, 135.69, 135.66, 133.43, 133.31, 128.46, 128.38, 128.33, 128.31, 128.28, 128.25, 112.23, 112.17, 77.24, 77.03, 76.82, 65.86, 40.14, 39.96, 29.79, 29.71, 29.61, 29.56, 29.53, 29.33, 29.26, 15.28, 1.02, -0.00; Cu32-P10 (7.9 mg, 38.8%) 1H NMR (600 MHz, CDCl3) δ 7.34, 7.33, 7.26, 7.24, 6.90, 6.89, 6.88, 6.87, 3.80, 3.72, 3.50, 3.49, 3.48, 3.47, 3.46, 2.39, 1.58, 1.37, 1.29, 1.26, 1.22, 1.21, 1.20, 0.89, 0.88, 0.87, 0.84, 0.07, –0.00. 13C NMR (151 MHz, CDCl3) δ 159.57, 159.51, 134.09, 133.96, 129.55, 129.49, 128.78, 128.58, 128.54, 126.13, 126.01, 119.09, 118.95, 114.67, 77.23, 77.02, 76.81, 65.86, 55.45, 55.18, 15.28, 1.02, -0.00; Cu32-P11 (11.0 mg, 63.1%) 1H NMR (600 MHz, CDCl3) δ 7.71, 7.70, 7.50, 7.48, 7.47, 7.32, 7.31, 7.26, 3.49, 3.48, 3.47, 3.46, 2.31, 2.02, 2.00, 1.99, 1.98, 1.91, 1.89, 1.88, 1.87, 1.76, 1.74, 1.22, 1.20, 1.19, 1.15, 1.13, 1.12, 1.10, 0.99, 0.97, 0.96, 0.95, 0.94, 0.07, –0.01. 13C NMR (151 MHz, CDCl3) δ 132.14, 131.94, 131.83, 131.52, 130.60, 130.54, 129.02, 128.67, 128.65, 128.57, 128.44, 128.28, 128.23, 77.23, 77.02, 76.81, 65.86, 50.83, 22.53, 22.07, 18.64, 15.27, 8.91, 5.58, 5.55, 1.02, -0.01; Cu32-P12 (7.8 mg, 40.7%) 1H NMR (600 MHz, CDCl3) δ 7.59, 7.57, 7.51, 7.50, 7.48, 7.47, 7.39, 7.37, 7.36, 7.32, 7.31, 7.30, 7.26, 3.49, 3.48, 3.47, 3.46, 2.03, 2.01, 1.74, 1.66, 1.63, 1.60, 1.52, 1.50, 1.21, 1.20, 1.19, 0.88, 0.87, 0.86, 0.83, 0.06, –0.01. 13C NMR (151 MHz, CDCl3) δ 134.74, 131.52, 131.47, 131.21, 129.02, 128.44, 128.26, 128.20, 128.07, 77.23, 77.02, 76.81, 65.86, 50.83, 29.14, 28.27, 26.99, 26.81, 26.52, 26.41, 26.33, 26.12, 26.02, 25.87, 25.53, 25.28, 24.66, 24.63, 15.27, 8.89, 1.02, –0.01.
Photosynergetic ligand-exchange transformation to synthesize Cu38-PyrrCa
A total of 25 mg (0.006 mmol) of the Cu14 cluster product was dissolved in chloroform. It was mixed with a chloroform solution of P(C4H8N)3 (4.1 mg, 0.017 mmol) and a methanolic solution containing PPh4Cl (12.5 mg, 0.05 mmol). The mixture was exposed to irradiation using either a PLS-SXE300+ Xenon lamp or an incandescent lamp for a specific duration. The resulting mixture was filtered, and the filtrate was subjected to ether vapor diffusion for crystallization. The transformed cluster crystals Cu38-PyrrCa (10.1 mg, 63.5%) were obtained within approximately one week. 1H NMR (600 MHz, CDCl3) δ 8.10, 8.09, 7.26, 5.35, 4.69, 4.68, 3.50, 3.49, 3.48, 3.48, 3.47, 3.46, 2.22, 2.02, 2.01, 1.64, 1.53, 1.26, 1.22, 1.22, 1.21, 1.21, 1.20, 1.20, 0.89, 0.88, 0.87, 0.84. 13C NMR (151 MHz, CDCl3) δ 128.32, 77.21, 77.00, 76.79, 65.81, 29.30, 27.20, 21.20, 15.23.
Photosynergetic ligand-exchange transformation to synthesize Cu3-DPPM
A total of 25 mg (0.006 mmol) of the Cu14 cluster product was dissolved in chloroform. It was mixed with a chloroform solution of DPPM (3.3 mg, 0.008 mmol) and a methanolic solution containing NaSbF6 (12.5 mg, 0.05 mmol). The mixture was exposed to irradiation using either a PLS-SXE300+ Xenon lamp or an incandescent lamp for a specific duration. The resulting mixture was filtered, and the filtrate was subjected to ether vapor diffusion for crystallization. The transformed cluster crystals Cu3-DPPM (9.0 mg, 19.1%) were obtained within approximately one week. 1H NMR (600 MHz, CDCl3) δ 8.10, 8.09, 7.26, 7.15, 7.04, 7.02, 5.35, 4.69, 4.68, 3.47, 2.31, 2.29, 1.53, 1.22, 1.21, 1.20, 0.88, 0.87. 13C NMR (151 MHz, CDCl3) δ 138.79, 133.79, 133.66, 132.89, 132.82, 132.75, 128.65, 128.43, 128.34, 77.22, 77.01, 76.80, 65.82, 50.54, 27.20, 15.22, 0.98.
Photosynergetic ligand-exchange transformation to synthesize Cu2-BINAP
A total of 25 mg (0.006 mmol) of the Cu14 cluster product was dissolved in chloroform. It was mixed with a chloroform solution of BINAP (5.3 mg, 0.008 mmol). The mixture was exposed to irradiation using either a PLS-SXE300+ Xenon lamp or an incandescent lamp for a specific duration. The resulting mixture was filtered, and the filtrate was subjected to ether vapor diffusion for crystallization. The transformed cluster crystals Cu2-BINAP (10.9 mg, 15.2%) were obtained within approximately one week. 1H NMR (600 MHz, CDCl3) δ 8.10, 8.09, 7.47, 7.26, 7.15, 7.14, 6.85, 6.84, 4.69, 4.68, 3.48, 2.23, 2.22, 2.21, 2.01, 1.64, 1.54, 1.26, 1.22, 1.21, 1.20, 0.89, 0.88, 0.87. 13C NMR (151 MHz, CDCl3) δ 77.22, 77.01, 76.80, 65.83, 50.76, 15.24, 0.99.
Photosynergetic ligand-exchange transformation to synthesize Cu8-DPPP
A total of 25 mg (0.006 mmol) of the Cu14 cluster product was dissolved in chloroform. It was mixed with a chloroform solution of DPPP (3.5 mg, 0.017 mmol) and a methanolic solution containing PPh4Cl (12.5 mg, 0.05 mmol). The mixture was exposed to irradiation using either a PLS-SXE300+ Xenon lamp or an incandescent lamp for a specific duration. The resulting mixture was filtered, and the filtrate was subjected to ether vapor diffusion for crystallization. The transformed cluster crystals Cu8-DPPP (13.4 mg, 47.4%) were obtained within approximately one week. 1H NMR (600 MHz, CDCl3) δ 8.10, 8.08, 7.64, 7.26, 7.13, 7.07, 7.04, 6.98, 3.48, 1.57, 1.22, 1.21, 1.20, 0.84. 13C NMR (151 MHz, CDCl3) δ 132.77, 132.66, 132.44, 128.88, 128.72, 128.32, 77.23, 77.02, 76.81, 65.81, 50.44, 15.21.
Photosynergetic ligand-exchange transformation to synthesize Cu10-R/S
A total of 25 mg (0.006 mmol) of the Cu14 cluster product was dissolved in chloroform. It was mixed with a chloroform solution of R/S-BDPP (3.8 mg, 0.017 mmol). The mixture was exposed to irradiation using either a PLS-SXE300+ Xenon lamp or an incandescent lamp for a specific duration. The resulting mixture was filtered, and the filtrate was subjected to ether vapor diffusion for crystallization. The transformed cluster crystals Cu10-R/S (16.0 mg, 54.6%) were obtained within approximately one week. The NMR spectrum of the Cu10-S cluster: 1H NMR (600 MHz, CDCl3) δ 7.26, 7.20, 7.18, 7.17, 3.49, 3.48, 3.47, 3.46, 2.94, 1.27, 1.26, 1.25, 1.22, 1.20, 1.19, 1.06, 0.89, 0.88, 0.87, 0.84, 0.84, 0.83, 0.07, -0.00. 13C NMR (151 MHz, CDCl3) δ 135.10, 134.98, 129.92, 129.02, 128.71, 128.44, 128.10, 127.92, 77.23, 77.02, 76.81, 65.86, 50.70, 35.85, 31.91, 29.78, 29.71, 29.56, 29.53, 29.33, 29.26, 27.22, 25.52, 22.69, 15.28, 14.12, 1.02, -0.01. The NMR spectrum of the Cu10-R cluster: 1H NMR (600 MHz, CDCl3) δ 8.10, 8.08, 7.69, 7.68, 7.49, 7.48, 7.44, 7.43, 7.26, 7.14, 7.08, 7.06, 7.02, 7.00, 6.96, 6.94, 6.93, 6.86, 3.48, 2.35, 2.33, 2.23, 2.21, 2.20, 1.58, 1.22, 1.20, 1.19. 13C NMR (151 MHz, CDCl3) δ 129.73, 128.23, 127.89, 77.22, 77.01, 76.80, 65.83, 50.78, 27.13, 15.25, 0.99.
Synthesis of Cu61(StBu)26S6Cl6H14 cluster
The cluster was synthesized as previously reported71. A sample (315 mg) of [Cu(CH3CN)4]BF4 was dissolved in acetonitrile (15 mL). Addition of tert-butyl thiol (180 µL) and triethylamine (1 mL) yielded a light yellow solution, which was stirred for 3 h. After solvent removal, the residue was dissolved in chloroform. A 3 mL aliquot was treated with borane tert-butylamine complex (25 mg), causing a color change to dark red over 4 h. Ethanol was added, and the solution was slowly evaporated in the dark. Dark red crystals formed after one week.
Synthesis of Cu14(BEN)6(AdmS)8
Cu14(BEN)6(AdmS)8 was prepared by a modified procedure reported in literature72. Typically, a mixture of Cu(C6H5COO)2 and AdmSH in a 1:1 molar ratio was dissolved in a mixed solvent of ethylene glycol and toluene. The reaction mixture was heated at 80 °C for 48 h, yielding a yellow solution. After careful filtration, the solution was concentrated to dryness. The resulting solid was then treated with dichloromethane to give a yellow solution. Yellow block crystals were obtained by vapor diffusion of diethyl ether into this solution.
Synthesis of the [Cu4(PPh2py)4Cl2]+
The cluster was synthesized as previously reported73. Typically, 200 mg of HAuCl4·3H2O and 100 mg of Cu(CH3COO)2·H2O were added to a mixed solvent of methanol (15 mL) and toluene (15 mL) under continuous stirring. After 10 min, 50 μL of 1,3-dimercaptopropane and 100 mg of PPh2py were introduced. Following an additional 30 min of stirring, an ice‑cold aqueous solution of borane tert-butylamine complex was added. Sodium tetraphenylborate was also introduced as a counter‑ion to assist crystallization. Yellow crystals of the Cu4 nanocluster were grown by layering n‑hexane over a CH2Cl2 solution of the product.
Data availability
Data supporting the findings of this work are available within the article and its Supplementary Information. All data that support the findings of this study are available from the corresponding author upon request. Crystallographic data for the structures reported in this Article have been deposited at the Cambridge Crystallographic Data Center, under deposition numbers CCDC 2484271 (Cu10-R), 2484272 (Cu10-S), 2484273 (Cu14), 2484274 (Cu2-BINAP), 2484275 (Cu3-DPPM), 2484277 (Cu32-P4), 2484278 (Cu32-P10), 2484279 (Cu32-P6), 2484280 (Cu32-P2), 2484281 (Cu32-P1), 2484282 (Cu32-P5), 2484283 (Cu32-P11), 2484284 (Cu32-P8), 2484285 (Cu32-P12), 2484286 (Cu32-P7), 2484287 (Cu32-P3), 2484288 (Cu32-P9), 2484289 (Cu38-PyrrCa), and 2499652 (Cu8-DPPP). Copies of the data can be obtained free of charge via https://www.ccdc.cam.ac.uk/structures/.
References
Zhang, F., Shi, W., Liu, Q. & Wang, X. Modular assembly of polyoxometalate clusters at the sub-1 nm scale. Nat. Protoc. 21, 347–372 (2025).
Jones, M. R., Seeman, N. C. & Mirkin, C. A. Programmable materials and the nature of the DNA bond. Science 347, 1260901 (2015).
Divine, R. et al. Designed proteins assemble antibodies into modular nanocages. Science 372, eabd9994 (2021).
Yao, Y. et al. High-entropy nanoparticles: Synthesis-structure-property relationships and data-driven discovery. Science 376, eabn3103 (2022).
Malola, S. et al. A method for structure prediction of metal-ligand interfaces of hybrid nanoparticles. Nat. Commun. 10, 3973 (2019).
Lim, K. R. G. et al. Nanoscale wetting controls reactive Pd ensembles in synthesis of dilute PdAu alloy catalysts. Nat. Commun. 16, 6293 (2025).
Chakraborty, I. & Pradeep, T. Atomically precise clusters of noble metals: Emerging link between atoms and nanoparticles. Chem. Rev. 117, 8208–8271 (2017).
Jin, R., Zeng, C., Zhou, M. & Chen, Y. Atomically precise colloidal metal nanoclusters and nanoparticles: Fundamentals and opportunities. Chem. Rev. 116, 10346–10413 (2016).
Yao, Q., Chen, T., Yuan, X. & Xie, J. Toward total synthesis of thiolate-protected metal nanoclusters. Acc. Chem. Res. 51, 1338–1348 (2018).
Zhao, H. et al. Assembly of air-stable copper(I) alkynide nanoclusters assisted by tripodal polydentate phosphoramide ligands. Nat. Synth. 3, 517–526 (2024).
Kenzler, S., Schrenk, C. & Schnepf, A. Au108S24(PPh3)16: A highly symmetric nanoscale gold cluster confirms the general concept of metalloid clusters. Angew. Chem. Int. Ed. 56, 393–396 (2016).
Qin, Z. et al. A homoleptic alkynyl-ligated [Au13Ag16L24]3− cluster as a catalytically active eight-electron superatom. Angew. Chem. Int. Ed. 60, 970–975 (2021).
Tang, L. et al. Total structure of bimetallic core–shell [Au42Cd40(SR)52]2− nanocluster and its implications. Angew. Chem. Int. Ed. 60, 17969–17973 (2021).
Takano, S. & Tsukuda, T. Chemically modified gold/silver superatoms as artificial elements at nanoscale: Design principles and synthesis challenges. J. Am. Chem. Soc. 143, 1683–1698 (2021).
Wan, X., Tang, Q., Yuan, S., Jiang, D. & Wang, Q. Au19 nanocluster featuring a V-shaped alkynyl–gold motif. J. Am. Chem. Soc. 137, 652–655 (2015).
Zhu, M., Aikens, C. M., Hollander, F. J., Schatz, G. C. & Jin, R. Correlating the crystal structure of A thiol-protected Au25 cluster and optical properties. J. Am. Chem. Soc. 130, 5883–5885 (2008).
Desireddy, A. et al. Ultrastable silver nanoparticles. Nature 501, 399–402 (2013).
Jadzinsky, P. D., Calero, G., Ackerson, C. J., Bushnell, D. A. & Kornberg, R. D. Structure of a thiol monolayer-protected gold nanoparticle at 1.1 Å resolution. Science 318, 430–433 (2007).
Li, S. et al. Stepwise achievement of circularly polarized luminescence on atomically precise silver clusters. Adv. Sci. 7, 2000738 (2020).
Yonesato, K., Yamazoe, S., Yokogawa, D., Yamaguchi, K. & Suzuki, K. A molecular hybrid of an atomically precise silver nanocluster and polyoxometalates for H2 cleavage into protons and electrons. Angew. Chem. Int. Ed. 60, 16994–16998 (2021).
Zhang, Y. et al. Preorganized nitrogen sites for Au11 amidation: A generalizable strategy toward precision functionalization of metal nanoclusters. J. Am. Chem. Soc. 145, 12164–12172 (2023).
Isozaki, K. et al. Gold nanocluster functionalized with peptide dendron thiolates: Acceleration of the photocatalytic oxidation of an amino alcohol in a supramolecular reaction field. ACS Catal. 11, 13180–13187 (2021).
Yoskamtorn, T. et al. Thiolate-mediated selectivity control in aerobic alcohol oxidation by porous carbon-supported Au25 clusters. ACS Catal. 4, 3696–3700 (2014).
Lim, D. C. et al. Emissive nanoclusters based on subnanometer-sized Au38 cores for boosting the performance of inverted organic photovoltaic cells. Adv. Energy Mater. 5, 1500393 (2015).
Kim, H. et al. Noble metal-based multimetallic nanoparticles for electrocatalytic applications. Adv. Sci. 9, 2104054 (2022).
You, Q. et al. Pd8 nanocluster with nonmetal-to-metal- ring coordination and promising photothermal conversion efficiency. Angew. Chem. Int. Ed. 63, e202313491 (2024).
Zhu, Y. et al. Room-temperature spin transport in metal nanocluster-based spin valves. Angew. Chem. Int. Ed. 62, e202213208 (2022).
Shen, H. et al. Regioselective hydrogenation of alkenes over atomically dispersed Pd sites on NHC-stabilized bimetallic nanoclusters. Chem 8, 2380–2392 (2022).
Chen, S. et al. Near infrared electrochemiluminescence of rod-shape 25-atom auag nanoclusters that is hundreds-fold stronger than that of Ru(bpy)3 standard. J. Am. Chem. Soc. 141, 9603–9609 (2019).
Huang, R. et al. Hypersensitive dual-function luminescence switching of a silver-chalcogenolate cluster-based metal–organic framework. Nat. Chem. 9, 689–697 (2017).
Chen, J. et al. Atomically precise photothermal nanomachines. Nat. Mater. 23, 271–280 (2024).
Wang, X. et al. Ligand-protected metal nanoclusters as low-loss, highly polarized emitters for optical waveguides. Science 381, 784–790 (2024).
Shen, H. et al. Guiding the high-yield synthesis of NHC-ligated gold nanoclusters by 19F NMR spectroscopy. ACS Nanosci. Au 2, 520–526 (2022).
He, L. Z. & Dong, T. G. Progress in controlling the synthesis of atomically precise silver nanoclusters. CrystEngComm 23, 7369–7379 (2021).
Qu, M. et al. Bidentate phosphine-assisted synthesis of an all-alkynyl-protected Ag74 nanocluster. J. Am. Chem. Soc. 139, 12346–12349 (2017).
Tian, F. & Chen, R. Pd-mediated synthesis of Ag33 chiral nanocluster with core–shell structure in T point group. J. Am. Chem. Soc. 141, 7107–7114 (2019).
Tsukamoto, T., Kambe, T., Nakao, A., Imaoka, T. & Yamamoto, K. Atom-hybridization for synthesis of polymetallic clusters. Nat. Commun. 9, 3873 (2018).
Peng, J. et al. Exploration of formation and size-evolution pathways of thiolate-gold nanoclusters in the CO-directed [Au25(SR)18]− synthesis. Small 17, 2000627 (2020).
Shen, H. et al. Simple and selective synthesis of copper-containing metal nanoclusters using (PPh3)2CuBH4 as reducing agent. Small Methods 5, 2000603 (2021).
Ma, F. Y., Abboud, K. A. & Zeng, C. J. Precision synthesis of a CdSe semiconductor nanocluster via cation exchange. Nat. Synth. 2, 949–959 (2023).
Liu, X. & Astruc, D. Atomically precise copper nanoclusters and their applications. Coord. Chem. Rev. 359, 112–126 (2018).
Sharma, S., Chakrahari, K. K., Saillard, J. & Liu, C. W. Structurally precise dichalcogenolate-protected copper and silver superatomic nanoclusters and their alloys. Acc. Chem. Res. 51, 2475–2483 (2018).
Tang, Q. et al. Lattice-hydride mechanism in electrocatalytic CO2 reduction by structurally precise copper-hydride nanoclusters. J. Am. Chem. Soc. 139, 9728–9736 (2017).
Xu, C. et al. A high-nuclearity copper sulfide nanocluster [S-Cu50] featuring a double-shell structure configuration with Cu(II)/Cu(I) valences. J. Am. Chem. Soc. 145, 25673–25685 (2023).
Dong, C. et al. Cu36H10(PET)24(PPh3)6Cl2] reveals surface vacancy defects in ligand-stabilized metal nanoclusters. J. Am. Chem. Soc. 143, 11026–11035 (2021).
Han, B. et al. Polymorphism in atomically precise Cu23 nanocluster incorporating tetrahedral [Cu4]0 kernel. J. Am. Chem. Soc. 142, 5834–5841 (2020).
Nguyen, T. D. et al. A Cu25 nanocluster with partial Cu(0) character. J. Am. Chem. Soc. 137, 13319–13324 (2015).
Wang, Y. et al. An atomically precise pyrazolate-protected copper nanocluster exhibiting exceptional stability and catalytic activity. Angew. Chem. Int. Ed. 62, e202218369 (2023).
Wu, Q. et al. Atomically precise copper nanoclusters for highly efficient electroreduction of CO2 towards hydrocarbons via breaking the coordination symmetry of Cu site. Angew. Chem. Int. Ed. 62, e202306822 (2023).
Li, S. et al. Anchoring frustrated lewis pair active sites on copper nanoclusters for regioselective hydrogenation. J. Am. Chem. Soc. 146, 27852–27860 (2024).
Alamer, B. et al. Planar core and macrocyclic shell stabilized atomically precise copper nanocluster catalyst for efficient hydroboration of C–C multiple bond. J. Am. Chem. Soc. 146, 16295–16305 (2024).
Bodiuzzaman, M. et al. Modulating decarboxylative oxidation photocatalysis by ligand engineering of atomically precise copper nanoclusters. J. Am. Chem. Soc. 146, 26994–27005 (2024).
Baghdasaryan, A. & Bürgi, T. Copper nanoclusters: Designed synthesis, structural diversity, and multiplatform applications. Nanoscale 13, 6283–6340 (2021).
Sun, C. et al. Atomically precise, thiolated copper–hydride nanoclusters as single-site hydrogenation catalysts for ketones in mild conditions. ACS Nano 13, 5975–5986 (2019).
Yan, B. et al. Flat-shaped copper nanoclusters with near-infrared absorption for enhanced photothermal conversion. JACS Au 5, 1884–1893 (2025).
Ferlazzo, A. et al. Photochemical eco-friendly synthesis of photothermal and emissive copper nanoclusters in water: towards sustainable nanomaterials. Mater. Adv. 5, 8034–8041 (2024).
Zhang, L. L. & Wong, W. Atomically precise copper nanoclusters as ultrasmall molecular aggregates: Appealing compositions, structures, properties, and applications. Aggregate 4, e266 (2023).
Dhayal, R. S. et al. [Cu32(H)20{S2P(OiPr)2}12]: The largest number of hydrides recorded in a molecular nanocluster by neutron diffraction. Chem. Eur. J. 21, 8369–8374 (2015).
Sun, J. et al. Eight-electron copper-hydride nanoclusters: Synthesis, structure, alloying chemistry and photoluminescence. Chem. Sci. 16, 6392–6401 (2025).
Chiu, T. H. et al. Hydride doping effects on the structure and properties of eight-electron Rh/Ag superatoms: The [RhHx@Ag21–x{S2P(OnPr)2}12] (x = 0–2) series. J. Am. Chem. Soc. 145, 16739–16747 (2023).
He, A. et al. Eight-electron Pt/Cu superatom encapsulating three “electron-donating” hydrides. Sci. Adv. 11, eads4488 (2025).
Nguyen, T. D. et al. Ligand-exchange-induced growth of an atomically precise Cu29 nanocluster from a smaller cluster. Chem. Mater. 28, 8385–8390 (2016).
Dhayal, R. S. et al. Diselenophosphate-induced conversion of an achiral [Cu20H11{S2P(OiPr)2}9] into a chiral [Cu20H11{Se2P(OiPr)2}9] polyhydrido nanocluster. Angew. Chem. Int. Ed. 54, 13604–13608 (2015).
Kang, X. & Zhu, M. Transformation of atomically precise nanoclusters by ligand-exchange. Chem. Mater. 31, 9939–9969 (2019).
Shichibu, Y., Negishi, Y., Tsukuda, T. & Teranishi, T. Large-scale synthesis of thiolated Au25 clusters via ligand exchange reactions of phosphine-stabilized Au11 clusters. J. Am. Chem. Soc. 127, 13464–13465 (2005).
Niihori, Y., Kikuchi, Y., Kato, A., Matsuzaki, M. & Negishi, Y. Understanding ligand-exchange reactions on thiolate-protected gold clusters by probing isomer distributions using reversed-phase high-performance liquid chromatography. ACS Nano 9, 9347–9356 (2015).
Wang, Y. & Bürgi, T. Ligand exchange reactions on thiolate-protected gold nanoclusters. Nanoscale Adv. 3, 2710–2727 (2021).
Chen, M. et al. Atomically precise Cu nanoclusters: Recent advances, challenges, and perspectives in synthesis and catalytic applications. Nano-Micro Lett. 17, 83 (2024).
Qin, H. et al. Thiacalix[4]arene etching of an anisotropic Cu70H22 intermediate for accessing robust modularly assembled copper nanoclusters. J. Am. Chem. Soc. 146, 3545–3552 (2024).
Huo, R. et al. Atomically precise Cu32 nanoclusters with selenide doping: Synthesis, bonding, and catalysis. Aggregate 6, e679 (2025).
Ghosh, A. et al. Cu61(StBu)26S6Cl6H14]+: A core–shell superatom nanocluster with a quasi-J36 Cu19 Core and an “18-Crown-6” metal-sulfide-like stabilizing belt. ACS Mater. Lett. 1, 297–302 (2019).
Sun, J. et al. Carboxylate engineering for manipulating the optical and assembly properties of copper clusters. Inorg. Chem. Front. 10, 2618–2625 (2023).
Tian, Y. et al. Reductant-selected formation of copper nanoclusters with crystallization-induced emission enhancement performance. Nanoscale 17, 26123–26128 (2025).
Truttmann, V. et al. Directing intrinsic chirality in gold nanoclusters: Preferential formation of stable enantiopure clusters in high yield and experimentally unveiling the “Super” chirality of Au144. ACS Nano 17, 20376–20386 (2023).
Liu, C. et al. Asymmetric transformation of achiral gold nanoclusters with negative nonlinear dependence between chiroptical activity and enantiomeric excess. Nat. Commun. 14, 3730 (2023).
Dolamic, I., Varnholt, B. & Bürgi, T. Chirality transfer from gold nanocluster to adsorbate evidenced by vibrational circular dichroism. Nat. Commun. 6, 7117 (2015).
Lange, G. F., Pottecher, J. D. F., Robey, C., Monserrat, B. & Peng, B. Negative refraction of weyl phonons at twin quartz interfaces. ACS Mater. Lett. 6, 847–855 (2024).
Zhu, Y. et al. Enantioseparation of Au20(PP3)4Cl4 clusters with intrinsically chiral cores. Angew. Chem. Int. Ed. 57, 9059–9063 (2018).
Lopez-Acevedo, O., Tsunoyama, H., Tsukuda, T., Ha¨kkinen, H. & Aikens, C. M. Chirality and electronic structure of the thiolate-protected Au38 nanocluster. J. Am. Chem. Soc. 132, 8210–8218 (2010).
Zhu, Y., Guo, J., Qiu, X., Zhao, S. & Tang, Z. Optical activity of chiral metal nanoclusters. Acc. Mater. Res. 2, 21–35 (2021).
Wang, D. et al. Direct electrochemical oxidation of alcohols with hydrogen evolution in continuous-flow reactor. Nat. Commun. 10, 2796 (2019).
Spasiano, D. et al. Kinetic modeling of partial oxidation of benzyl alcohol in water by means of Fe(III)/O2/UV–solar simulated process. Chem. Eur. J. 249, 130–142 (2014).
Sheldrick, G. M. SHELXT-integrated space-group and crystal-structure determination. Acta Cryst. A 71, 3–8 (2015).
Hubschle, C. B., Sheldrick, G. M. & Dittrich, B. ShelXle: A Qt graphical user interface for SHELXL. J. Appl. Cryst. 44, 1281–1284 (2011).
Dolomanov, O. V., Bourhis, L. J., Gildea, R. J., Howard, J. A. K. & Puschmann, H. OLEX2: A complete structure solution, refinement and analysis program. J. Appl. Cryst. 42, 339–341 (2009).
Acknowledgements
H. S. acknowledges the financial support from the National Natural Science Foundation of China (22301149 and 22571172), National Key R&D Program of China (2023YFB3507100), Natural Science Foundation of Inner Mongolia (2025JQ026), Program for Young Talents of Science and Technology in Universities of Inner Mongolia Autonomous Region (NJYT23035) and start-up funding of Inner Mongolia University (10000-23112101/043 and 23600-5233710). S. L. acknowledges the financial support from the National Natural Science Foundation of China (22565021), Natural Science Foundation of Inner Mongolia (2025QN02068), and Start-Up Funding of Inner Mongolia University (10000-A24202027). N.F. Z. acknowledges the financial support from the National Natural Science Foundation of China (grant no. 92261207, and NSFC Center for Single-Atom Catalysis under grant no. 22388102) and the New Cornerstone Science Foundation. Y. Gao acknowledges the financial support from National Natural Science Foundation of China (52461037). We thank Hongwei Li in the NMR facility of National Center for Protein Sciences at Peking University for assistance with the NMR measurement.
Author information
Authors and Affiliations
Contributions
M.Y. and H.S. designed the research. M.Y. did most of the experiments. M.Y., Q.L., H.G., R.H., X.G., and Z.G. performed the sample characterization. M.Y., Q.L., C.X., A.H., and H.S. wrote the paper. M.Y., Q.L., Z.X., and S.L. analyzed the data. Y.G. and H.S. secured the funding required for the project. J.P., N.Z., and H.S. supervised the overall study. N.Z. and H.S. reviewed the manuscript. All authors participated in the discussions regarding the research progress and results. M.Y., Q.L., and Z.X. contributed equally to this work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, M., Li, Q., Xie, Z. et al. Photosynergetic ligand-exchange for modular synthesis of copper nanoclusters. Nat Commun 17, 2596 (2026). https://doi.org/10.1038/s41467-026-69444-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69444-7