Abstract
Insect wing development involves tissue patterning, cell fate transitions, and hormone signaling, yet its spatiotemporal logic remains unclear. The silkworm, with large wing discs and defined stages, provides an ideal model for high resolution analysis. Here, we construct a spatiotemporal single-cell atlas of the silkworm wing disc across 10 timepoints, identifying 12 major cell types and their developmental transitions. Wing morphogenesis (Wm) cells act as central progenitors, differentiating into epithelial and cuticle lineages under lineage-specific transcription factors. Time‑resolved snRNA‑seq reveals hierarchical transcriptional reprogramming, with Wm cells functioning as early signaling hubs. Functional modules and signaling pathways were activated in spatiotemporal controlled manner. 20‑hydroxyecdysone treatment rapidly accelerates fate transitions and gene expression, recapitulating natural development within hours. Integration of morphology, hormone levels, and gene expression supports a five-stage Gene Transition Model describing progressive fate resolution. This work reveals wing development in silkworm and provides insights into hormone-driven organogenesis and potential manipulation of insect development in agriculture.
Similar content being viewed by others
Introduction
Insects are the only invertebrates that can fly, a key evolutionary innovation that allows them to travel long distances, escape predators, locate mates, and expand their foraging range. This ability has enabled insects to occupy diverse ecological niches and become the most species-rich group of animals on Earth1. Understanding how insect wings develop is thus fundamental to developmental biology and has practical value for pest control, evolutionary studies, and bio-inspired engineering. In holometabolous insects, their wings develop from specialized larval structures known as imaginal discs, particularly wing discs2. Wing discs are composed of diverse cell types that undergo continuous morphogenetic changes, ultimately forming the fully developed insect wing3,4. Knowing the spatial distribution and functional roles of these cells is important for understanding how wings form. Previous studies have shown that hormones and several signaling pathways play key roles in wing disc development5,6. Among them, 20-hydroxyecdysone (20E) is the main hormone, driving the metamorphosis from larvae to adult stages by guiding cell proliferation, differentiation, and apoptosis5,7. Meanwhile, classic pathways such as Hedgehog (Hh), Wnt, and Notch also play roles in cell fate, tissue organization, and organ formation8,9,10.
Although single-cell analyses have been conducted for the wing disc of Drosophila melanogaster (D. melanogaster) and Bicyclus anynana (B. anynana)11,12, they have mainly focused on cell type composition and gene expression patterns at isolated stages. These studies fail to capture the full continuum of cellular transitions across developmental time and often lack spatial resolution. Consequently, it remains unclear how different cell types interact, how their spatial arrangement evolves, and how they collectively respond to dynamic 20E signaling. A comprehensive, spatiotemporal view of wing disc development is imperative to fill these gaps.
The silkworm Bombyx mori (B. mori), is an important lepidopteran model of economic and developmental significance and serve as an ideal system for investigating developmental processes13,14. Bulk transcriptomic studies have identified temporal gene expression dynamics involving regulators such as BmBlimp-115, BmIGFLP16, BmChiNAG17, and BmSd18. However, these studies often focus on either temporal or spatial dimensions alone. Furthermore, bulk transcriptomic approaches average signals across heterogeneous tissues, lacking the resolution to uncover cell-type-specific dynamics or spatial regulatory cues. This makes it hard to understand how different cells change and interact within the tissue. Recent advances in single-cell RNA sequencing (scRNA-seq) and spatial transcriptomic technologies provide powerful tools to resolve cellular heterogeneity and gene expression dynamics at high resolution19,20,21. By integrating these approaches, it is possible to see how different cell types are arranged, how they develop over time, and how they communicate22,23,24,25,26.
In this work, we combined scRNA-seq, single-nucleus RNA sequencing (snRNA-seq), and spatial transcriptomics (Stereo-seq) to construct a high-resolution spatiotemporal transcriptomic atlas of B. mori wing discs from larvae to pupa. We systematically mapped the dynamics of 12 functional cell populations in wing disc and revealed molecular mechanisms underlying their differentiation trajectories, which are regulated by key transcription factors such as Rfx, Blimp-1, Dll, and Pur-alpha. Performing snRNA-seq on wing disc subjected to time-constrained 20E treatment, we discovered a “time-axis compression” effect. In this process, 20E caused rapid reprogramming of cell fates. Finally, we built a Gene Transition Model (GTM) explaining how cellular signal cascades respond to 20E levels changes to guide wing development. In summary, our work reveals the dynamic processes that build insect wings and provides new insights in studying wing development and lepidopteran pest insect’s control.
Results
Temporal and spatial atlas of silkworm wing disc cells
The silkworm wing disc undergoes progressive morphological transitions, shifting from tissue remodeling during the larval stage (fifth instar) to spinning and ultimately forming mature wings during the pupal stage (Fig. 1a). To systematically resolve developmental trajectories of wing disc cell populations, we exploited the scRNA-seq technology, generating 11 libraries from samples across 10 timepoints spanning 3 developmental stages: larval (L5D1 to L5D7), spinning (W1, W2), and pupal stages (P6, the sixth day of the pupa) (Fig. 1b and Supplementary Data 1). After quality control, a total of 126,041 high-quality single-cell transcriptomes were retained, covering 16,122 genes with a median gene count per cell of 1989 (Supplementary Fig. 1a, b). Using dimension reduction and Uniform Manifold Approximation and Projection (UMAP), the cells were grouped into 12 clusters.
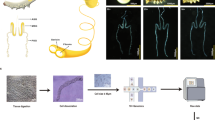
a Morphological changes of the silkworm wing disc were examined across the following stages: the 5th instar larva at days 1–7 (L5D1–L5D7), wandering stage days 1 and 2 (W1–W2), and pupal stage day 6 (Pupa-D6). scRNA-seq was performed on samples from all stages, with additional Stereo-seq analysis conducted specifically on the L5D2, L5D3, and W2 stages. b Overview of experimental workflow for scRNA and Stereo-seq, encompassing tissue dissection, sequencing, and data integration. Stereo-seq spatial resolution: 1 cm × 1 cm. c Dot plot showing cluster-specific marker genes used to define each cell type. d UMAP plot of 126,041 high-quality single cells of the wing disc from 11 samples across 10 timepoints spanning 3 developmental stages: larval (L5D1 to L5D7), spinning (W1, W2), and pupal stages (P6, the sixth day of the pupa), grouped into 12 distinct cell types based on gene expression profiles. e Spatial distribution of annotated cell types in wing discs from three developmental stages (L5D2, L5D3, and W2), illustrated with two biological replicates per stage. f Schematic representation of wing disc anatomy and spatial organization of major cell types based on integrated scRNA- and Stereo-seq data. Cell types: Wm (wing morphogenesis), Im (immune), Ma (matrix), Mp (metabolic process), Ad (axon development), Ci (ciliated), C1–C3 (cuticle 1–3), and Ep1–Ep3 (epithelial 1–3). Source data are provided as a Source Data file.
Based on known marker genes of B. mori22, and other lepidopterans11,12, a Drosophila single-cell RNA-seq dataset27, and enriched Gene Ontology (GO) processes of the top 200 ranked differential expressed genes from each cell cluster (Fig. 1c, Supplementary Fig. 1 and Supplementary Data 2), 12 major cell types were defined (Fig. 1d). Among these, three clusters showed specific expression of cuticle-related genes. These clusters were characterized as: Cuticle 1 (C 1) with high serpin genes (serpin-5 and serpin-1028) expression and its proportion gradually decreased from larval stage (L5D4) to pupal stage (P6), Cuticle 2 (C 2) with high fibroblast growth factor receptor (Btl11) expression, and Cuticle 3 (C 3) with high cuticle protein (CPH28,CPG3 and CPG929) expression and a high proportion of pupal stage (Supplementary Fig. 1c, d). Three epithelial cell types were identified based on markers such as Mf-cpa30 (Supplementary Fig. 1e). Epithelial 1 (Ep 1) showed a stable proportion from L5D1 to W2 and almost disappeared at P6. Epithelial 2 (Ep 2) was prominently abundant at W2 and almost disappeared at P6 too. Epithelial 3 (Ep 3) very rare in whole stages exception (Supplementary Fig. 1f)11,12,22,27. We also identified six other major cell types, including Wing morphogenesis (Wm), Immune (Im), Matrix (Ma), Metabolic process (Mp), Axon development (Ad), and Ciliated (Ci) cells (Supplementary Fig. 1 and Supplementary Data 2), giving a total of 12 major cell types (Fig. 1d). Wm cells, marked with key genes such as Fat31 and NID32, were mainly enriched in pathways related to cell morphogenesis and wing patterning, including tube morphogenesis and imaginal disc-derived wing margin formation (Fig. 1c and Supplementary Fig. 1g). Im cells were enriched for hematopoiesis and immune organ development, suggesting their involvement in immune cell lineage formation. Ma cells were associated with extracellular matrix organization, including basement membrane and respiratory tube development. Mp cells showed enrichment in pathways regulating cuticle development and metabolic processes. Ad cells were linked to axon development, neuron projection, and cell–cell recognition, reflecting roles in neural wiring. Ci cells exhibited significant enrichment in cilium assembly and organization, microtubule-based transport, and cell projection assembly. These enrichment profiles indicate that each cell type contributes specialized developmental or functional roles during wing disc development.
To characterize the spatial distribution of these cell types, we generated Stereo-seq spatial transcriptome data for wing discs at L5D2, L5D3, and W2 timepoints (Supplementary Data 3). Stereo-seq data were annotated through deconvolution mapping of gene signatures derived from 12 cell types, at bin20 (10 µm × 10 µm) resolution (Fig. 1e and Supplementary Fig. 2a). Combing spatial information (Supplementary Fig. 2b) and histological features (Supplementary Fig. 3a–d), we recapitulated the spatial distribution of each cell type at L5D2, including larva cuticle (Cuticle 1–3), epidermis (Epithelial 1–3), wing bud (Wm cells), and hematopoietic organ (Immune and Matrix cells) (Fig. 1f). The spatial distribution pattern of cells shifts during wing disc development. At the L5D2 and L5D3 stages, cuticle cells (Cuticle 1 and Cuticle 2) were mainly distributed at the outermost layer of the wing disc, followed by epithelial cells (Epithelial 1 and Epithelial 2). In contrast, Wm cells were primarily localized in the wing bud region (Fig. 1e and Supplementary Fig. 3e). By the W2 stage, cuticle cells still made up the outermost layer, but the proportion of Wm cells in the central region of the wing disc had significantly decreased, and epithelial cells began to occupy a broader area, which may be related to cellular differentiation and migration (Fig. 1e and Supplementary Fig. 3e).
To exemplify the utility of the annotated dataset, we examined 53 genes known to play important roles during wing disc development (Supplementary Data 4). These genes were mapped onto both our single-cell and spatial transcriptomic datasets and exhibited cell-type and region-specific expression patterns, providing refined insights into how they function at the cellular and spatial distribution levels in wing disc development. For instance, STAT33,34 was highly expressed in Cuticle1 cells located in the larval cuticle, Dpp35,36 showed peak expression in Wm cells within the wing bud at L5D5 (Supplementary Fig. 4), and Fng37,38,39 was enriched in Epithelial 1 cells within the wing bud and epidermis at L5D7 (Supplementary Fig. 4). Overall, by combining Stereo-seq and scRNA-seq-based spatiotemporal transcriptomics, we revealed the heterogeneity and dynamic spatiotemporal changes in cell type composition in the silkworm wing disc, along with their distinct spatial distribution patterns across developmental stages.
Wm cells as a differentiation hub in wing development
Based on the classification of major cell types in the wing disc (Fig. 1d), we analyzed how their proportions changed across developmental stages. From the larval stage (L5) to the pupal stage (P6), several cell types—including Wm, epithelial, cuticle, immune, and matrix cells—showed notable shifts in abundance (Supplementary Fig. 5a). For instance, cuticle cells increased rapidly during the pupal stage, suggesting active morphological transformation (Supplementary Fig. 5b, c). Some cell types, such as Cuticle 3 and Epithelial 2, appeared at later stages and became more prominent during wing formation (Supplementary Fig. 5c). This is consistent with the major structural changes that occur near pupation. Notably, Wm cells gradually decreased from L5 to P6 and were almost absent by the pupal stage (Supplementary Fig. 5d, e).
To investigate how cell fate transitions contribute to wing disc development, we analyzed the differentiation trajectories of various cell types using TOME-tradj40. Most cell types remained in stable states without clear differentiation trajectories (Supplementary Fig. 6). However, Wm cells showed complex and continuous transitions, suggesting a dynamic differentiation process primarily toward epithelial and cuticle lineages (Supplementary Fig. 6g). Additionally, Wm, epithelial, and cuticle cells were found in close proximity in the UMAP embedding space (Fig. 1d), which indicates similarity in gene expression profiles among these cells.
To further investigate the fate of Wm cells, we reconstructed their differentiation trajectories with Monocle2 and identified three distinct cellular states (Fig. 2a). Wm cells were mainly enriched in state 1, while epithelial and cuticle cells were primarily found in states 2 and 3, respectively (Fig. 2b). Based on their temporal distribution, state 1 was identified as the starting point (Fig. 2c). Pseudotime analysis further confirmed a bifurcated trajectory, with sequential emergence of Epithelial 1–3 and Cuticle 1–3 subtypes, showing an intrinsic maturation gradient within each lineage (Fig. 2d and Supplementary Fig. 7a, b). Early subtypes (Epithelial 1, Cuticle 1) appeared in larval stages, while later subtypes (Epithelial 3, Cuticle 3) peaked during the wandering and pupal stages (Fig. 2e). These findings indicate that Wm cells differentiate into two main cell lineages: epithelial and cuticle cells.
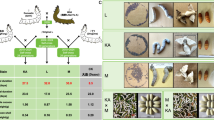
a Pseudotime trajectory analysis of Wm cells showing a bifurcated lineage pattern. b Proportional distribution of annotated cell types across the three trajectory states. c Branching plot showing Wm cell distribution along pseudotime trajectories from L5D1 to P6. d Predicted pseudotime trajectory analysis showing the differentiation trends of Wm cells. e Ridge plots showing the temporal progression of Wm, Epithelial, and Cuticle subtypes along the pseudotime axis. f Heatmap showing the expression dynamics of differentially expressed genes along the pseudotime trajectory, clustered into functional gene modules. Selected enriched pathways are highlighted. The enrichment analysis was performed via Over-Representation Analysis (ORA), with the statistical significance for each pathway evaluated using a two-sided hypergeometric test. P-values were adjusted for multiple comparisons using the Benjamini–Hochberg false discovery rate (FDR) correction. The enrichment results are visualized in a bubble plot. The color gradient of the bubbles represents the adjusted p-value (p.adjust), as indicated in the scale, with more intense color (lower p.adjust) denoting higher statistical significance. The size of the bubbles corresponds to the number of genes (Count) enriched in each pathway. Dynamic changes of Toll/Imd signaling pathway activity (g) and Hippo signaling pathway activity (h) across ten developmental stages. The x-axis represents distinct developmental stages: larval (L5D1 to L5D7), spinning (W-1, W-2), and pupal stages (P6, the sixth day of the pupa). The box plot elements are defined as follows: the center line indicates the median (50th percentile); the lower and upper bounds of the box indicate the 25th and 75th percentiles, respectively, defining the interquartile range (IQR); the whiskers extend to the minimum and maximum data points within 1.5× IQR from the respective box bounds; data points beyond the whiskers are plotted individually as outliers. A dashed horizontal line at y = 0 serves as a reference. i Regulatory network of key transcription factors (TFs) enriched in Wm cells and their predicted downstream target genes. j Quantitative RT-PCR analysis showing significant downregulation of BmRfx expression after RNAi (Data are shown from three biological replicates, with n = 30 independent wing discs used for each replicate; mean ± SD; two-tailed unpaired t test, ***p < 0.001). k Morphological effect of BmRfx gene knockdown on wing disc development. Dashed white boxes indicate the wing bud region. Trans as transverse section. Arrowheads indicate regions of morphological alteration in the wing disc following RNAi treatment. The upper panels show the overall morphology of wing discs, whereas the lower panels display transverse histological sections of the same samples. l Effects of BmRfx gene knockdown on wing morphology at the adult stage. m Quantitative RT-PCR analysis showing significant downregulation of SfRfx expression after RNAi (Data are shown from three biological replicates, with n = 20 independent wing discs used for each replicate; mean ± SD; two-tailed unpaired t test, **p < 0.01). n Morphological effect of SfRfx gene knockdown on wing disc development. Trans as transverse section. Arrowheads indicate regions of morphological alteration in the wing disc following RNAi treatment. o Effects of SfRfx gene knockdown on wing morphology at the adult stage. Cell types: Wm (wing morphogenesis). Source data are provided as a Source Data file.
To gain mechanistic insights into the regulation of these cell fate trajectories, we analyzed differentially expressed genes (DEGs) along pseudotime (Supplementary Data 5). In fate 1, DEGs were mainly enriched in Toll/Imd signaling, longevity-regulating pathways, and endoplasmic reticulum (ER)-related processes (Fig. 2f and Supplementary Fig. 7c). These pathways are associated with tissue remodeling during development. Toll/Imd signaling showed low activity in early larval stages but became more active in the pupal stages, suggesting a role in immune regulation and tissue morphogenesis during later development (Fig. 2g).
In contrast, fate 2 cells, which became dominant during the W1 and W2 period, showed enrichment in ribosome, Hippo, and MAPK pathways (Fig. 2f and Supplementary Fig. 7c). Notably, the Hippo pathway, which is known to regulate somatic growth and cell proliferation41, showed a marked increase in activity from L5D5 to W2, followed by a rapid decline at P6 (Fig. 2h). Consistent with this, Hippo-related genes such as A3 and Lix1 showed distinct expression patterns across the two fate branches (Supplementary Fig. 8a). This pattern suggests that Hippo signaling may regulate Wm cell proliferation and tissue morphogenesis before final differentiation.
To identify upstream regulators of Wm differentiation, we performed SCENIC analysis to identify key transcription factors (TFs) with high activity across developmental time points in Wm cells (Supplementary Fig. 8b). Based on the Regulon Specificity Score (RSS), we selected the top five TFs at two key stages during differentiation: early (Dll, sqz, Pur-alpha, Rfx) and W period (Blimp-1) (Fig. 2i). Enrichment analysis of their downstream target genes indicated that genes regulated by Dll and sqz were mainly associated with wing morphogenesis and displayed elevated RSS values at L5D1 and P6, suggesting their functional role in guiding Wm differentiation toward the cuticle fate (Supplementary Fig. 8c, d). In contrast, target genes of Pur-alpha were primarily associated with epithelial differentiation, whereas those regulated by Blimp-1 showed enriched in Wm-related genes and exhibited high RSS scores from L5D7 to W1, suggesting a potential role in promoting the transition from Wm to Epithelial cells (Supplementary Fig. 8e, f). Additionally, Rfx was identified as a cell-type-specific central regulator in Wm cells with the highest RSS value at L5D1 (Supplementary Fig. 8g).
To functionally validate the role of transcriptional regulators implicated in Wm differentiation, we focused on Rfx, a transcription factor specifically expressed in Wm cells. RNA interference (RNAi)-mediated knockdown of Rfx significantly disrupted its downstream regulatory network, including several key target genes involved in the differentiation of Wm cells (Fig. 2j and Supplementary Fig. 9). Among these, several genes have well-characterized roles in tissue organization and morphogenesis, such as bsh42, dpy43, futsch44 and pnr45. Within 48 hours after RNAi treatment, wing discs showed clear morphological defects, including lobulation in the central region, irregular epithelial invagination, and disorganized tracheal branching (Fig. 2k). In adults, Rfx knockdown caused severe wing deformities, such as shortened veins, malformed marginal bristles, and jagged posterior wing edges (Fig. 2l and Supplementary Data 6). In addition, Rfx is highly conserved across lepidoptera terms of phylogenetic relationship, motif organization, and domain architecture (Supplementary Fig. 10). Knockdown of its ortholog in the fall armyworm Spodoptera frugiperda (SfRfx, XM_050701463.1) resulted in similar morphological defects, indicating that Rfx plays a conserved and essential role in wing disc morphogenesis across lepidoptera (Fig. 2m–o). Furthermore, RNAi-mediated knockdown of other candidate TFs, including Dll, and Pur-alpha, also caused structural abnormalities in the wing disc (Supplementary Fig. 11). Blimp-1, which has previously been implicated in regulating wing disc development, showed consistent functional effects on wing patterning15. Together, these findings support that Rfx, together with Blimp-1, Dll, and Pur-alpha, forms a transcriptional regulatory module that drives the differentiation of Wm cells during wing morphogenesis.
20E regulates wing disc developmental dynamics
20E is a central hormonal regulator in insect development, known to control cell differentiation and tissue morphogenesis2,46. To explore how 20E influences silkworm wing disc development, we first examined the expression patterns of 20E-related genes in naturally developing tissues. These genes displayed strong spatiotemporal specificity across cell types (Supplementary Data 7). For example, 20E receptor genes such as USP47 were predominantly expressed in Wm cells, whereas cuticle cells highly expressed downstream signaling cascades like E7448 (Supplementary Fig. 12a). Integration with spatial transcriptomics further confirmed cell-type-specific expression of 20E-responsive genes (Supplementary Fig. 12b, c), suggesting that 20E plays a key role in guiding cell fate transitions within the wing disc.
To capture the rapid and transient transcriptional responses induced by 20E—beyond the resolution of natural development—we established an in vitro 20E stimulation system and performed time-resolved snRNA-seq across nine time points, ranging from 0 to 360 minutes (mins) after treatment (Fig. 3a and Supplementary Data 8). Wing discs were dissected at the L5D2 stage, which is characterized by low endogenous 20E levels49, providing a clean baseline for hormonal response. A 20E concentration of 2 µg/mL was selected based on dose–response optimization, as it robustly induced canonical target genes such as EcR and USP (Supplementary Fig. 13a). After quality control, a total of 125,717 high-quality cells were retained with a median gene number of 1304. Using scRNA-seq data from naturally developing wing discs as a reference, we identified six major cell types in the 20E-treated samples: Wm, Cuticle 1, Cuticle 2, Epithelial 1, Immune, and Matrix cells (Fig. 3b and Supplementary Fig. 13b, c). Gene expression exhibited a clear cascade response following 20E exposure. Within the first 20 min, genes involved in larval cuticle development were significantly upregulated. At 40 min, genes related to lipid response became activated. Between 50 and 60 min, genes involved in epithelial cell and hemocyte differentiation were highly expressed. By 4–6 hours (h), genes related to cytoskeleton organization were strongly upregulated (Fig. 3c and Supplementary Data 9). All cell types responded to 20E at different levels during this period. In the early phase (0–30 min), Wm, Cuticle, and Epithelial cells showed the strongest transcriptional responses. At later stages, the response shifted toward Ma and Im cells. This pattern indicates a time-dependent regulatory sequence (Supplementary Fig. 13d).
a Schematic diagram of the experimental workflow. Wing discs were dissected from L5D2 larvae and treated in vitro with 20E. Samples were collected at nine time points post-treatment (0, 10, 20, 30, 40, 50, 60, 240, and 360 min) for snRNA-seq analysis. b UMAP plot showing six major cell types identified across all time points based on snRNA-seq data (10 libraries from 10 samples across 9 timepoints post-treatment (0, 10, 20, 30, 40, 50, 60, 240, and 360 min, Supplementary Data 8). c Heatmap showing temporal changes in gene expression and pathway enrichment across time points from 0 to 360 min following 20E treatment. d Relative information flow of signaling pathways across time points. Relative information flow represents the normalized strength and direction of ligand–receptor–mediated signaling interactions among cell types, inferred from snRNA‑seq expression profiles and prior knowledge of ligand–receptor pairs. e Cell type-specific expression and distribution of gene modules M1 and M2 across different populations. f Temporal expression patterns of gene modules M1 and M2 across nine time points after 20E treatment. g Activity levels of transcription factors (TFs) at specific timepoints post-20E treatment. h Radar plots and UMAP overlays showing transcriptomic similarity between 20E-treated samples and natural developmental stages, indicating that 20E induces stage-specific gene programs in a time-compressed manner. Cell types: Wm (wing morphogenesis).
To assess how 20E influences intercellular communication, we analyzed the dynamics of signaling pathway activity among cell types. Notably, the TENASCIN pathway showed a sharp increase in signaling strength at 40 min. In contrast, the BMP, COLLAGEN, Notch, PARs, FGF, and SEMA4 pathways became strongly activated at later stages, especially at 4 and 6 h after treatment (Fig. 3d). Among these, the FGF pathway was activated earliest, with strong activity observed in Wm cells. BMP and Notch signaling peaked around 30 min and stayed active, although their levels gradually diminish over time. By 6 h, most pathways reached their highest activity levels. These results suggest that 20E triggers a stepwise and coordinated activation of multiple signaling pathways (Supplementary Fig. 14a). Before 40 min, Wm cells were the main source of outgoing signals, particularly through the BMP, Notch, PARs, COLLAGEN, and FGF pathways. After 40 min, Im cells became the dominant signaling source. This indicates a shift in intercellular communication patterns between different cell types during the 20E response (Supplementary Fig. 14b).
To further investigate the gene expression programs regulated by 20E, we identified two gene expression modules: Module 1 (M1) and Module 2 (M2) (Supplementary Fig. 15a). Genes in M1 was highly expressed in Wm, epithelial, and cuticle cells during the early phase (0–30 min). It was enriched in genes related to morphogenesis and cuticle development. This indicates that Wm differentiation begins before 30 min and is driven by morphogenetic signaling pathways (Fig. 3e, f and Supplementary Fig. 15b). In contrast, genes in M2 were mainly expressed in Ma and Im cells during the later stages (40–360 min). (Fig. 3e, f and Supplementary Fig. 15c). This shift in module activity occurred alongside a reduction in morphogenetic cell types and an increase in Ma and Im cell (Supplementary Fig. 13c). These results indicate that 20E controls a stage-specific switch in gene expression programs.
To identify key regulatory factors, we used SCENIC to analyzed TFs whose activity changed over time in response to 20E. Based on RSS values, we selected the top six key TFs, which show specific enrichment in different cell types (Supplementary Fig. 16a). In the early stage (before 30 min), Hr38 (LOC693064), Nubbin (LOC101744796), and Six2 (LOC101735990) showed strongly activity. These TFs regulated genes involved in Wm, Epithelial 1, and Cuticle 1 cells (Fig. 3g and Supplementary Fig. 16b–d). After 30 min, the activity of CREB (LOC692871), FoxN3 (LOC101739133), and GATA-A (LOC10174013) were involved in Immune and Matrix cells. These factors targeted genes related to cell migration and immune responses (Fig. 3g and Supplementary Fig. 16e–g). These results reveal a time-dependent shift in transcriptional regulation controlled by 20E.
Compared with gene expression profiles of wing discs under natural development (scRNA-seq dataset), 20E-treated wing discs (snRNA-seq dataset) exhibited a dual transcriptional signature, showing both L5D2-like baseline features and characteristics of later developmental timepoints (Fig. 3h). With increasing durations of 20E exposure, the induced transcriptional profile progressively transitioned from L5D4 (before 30 min) to L5D5 (30 min), and further to W1 stage (50 min to 6 h), suggesting that the 30-min time point may represent a critical tipping point for 20E-induced transcriptional reprogramming (Fig. 3h). Pseudotime trajectory analysis revealed that Wm cells from the treated group closely followed a trajectory like natural differentiation (Supplementary Fig. 17a, b), gradually transitioned into epithelial and cuticle lineages. Moreover, the key Wm regulators showed a distinct shift in expression before and after 30 min in both datasets (Supplementary Fig. 17c). Together, these findings suggest that 20E induces natural developmental-like transcriptional programs but in a faster, time-compressed manner.
Although the major cell types remained the same in both conditions, their relative abundances and activation timings were different. For instance, the proportions of Wm, Cuticle, and Epithelial lineages gradually changed from L5D1 to W1 in natural development. In contrast, 20E treatment caused a rapid and short-term expansion of these lineages within the first 30 min. After 40 min, Matrix and Immune cells became dominant. Matrix cells showed a continuous increase over time in both groups (Supplementary Fig. 18a). GO and KEGG enrichment analyses revealed that genes enriched in endocytosis, mTOR signaling pathway, structural constituent of cuticle, and DNA-binding transcription factor binding were specifically activated during natural development, reflecting sustained biosynthetic and developmental activity during natural wing disc maturation. In contrast, genes enriched in ribosome binding, mRNA binding, and oxidative phosphorylation were activated upon 20E stimulation, indicating a stronger emphasis on protein synthesis, energy metabolism, and transcriptional reprogramming (Supplementary Fig. 18b, c). A set of 273 shared upregulated genes between 20E-treated and naturally developing wing disc were identified as candidate direct targets of 20E stimulation (Supplementary Fig. 19a and Supplementary Data 10). GO and KEGG enrichment analysis of these genes revealed significant involvement in developmental processes such as cellular and organ morphogenesis, and growth, as well as in stress and immune responses. Notably, several pathways associated with protein homeostasis and signal transduction—such as autophagy, MAPK signaling, and ER protein processing—were also enriched, suggesting that 20E may coordinate both structural remodeling and cellular stress responses during wing disc development (Supplementary Fig. 19b). We further analyzed the dynamic expression patterns of these genes across natural development and 20E-treated (Supplementary Fig. 19c). These genes showed a progressive, stage-specific activation from L5D1 to P6 under natural development, transitioning from clusters C1 to C10. Upon 20E treatment, genes in clusters C3–C6 responded rapidly. In contrast, a subset of genes in clusters C8–C10 was activated within the first 30 min, while the remaining genes in these clusters showed gradual activation thereafter. Together, these results provide a high-resolution view of 20E-induced transcriptional reprogramming and underscore its role in accelerating developmental transitions in the wing disc.
Gene transition model defines wing disc developmental stages
To systematically characterize the transcriptional dynamics and developmental driven regulated by 20E in the wing disc, by combing gene expression patterns similarity (Fig. 4a), local 20E concentrations (Fig. 4b, Supplementary Fig. 20a and Supplementary Data 11), and morphological changes (Supplementary Fig. 20b–e), we propose a five-stage gene transition model of wing disc development: Stage 1 (L5D1–L5D2), Stage 2 (L5D3–L5D6), Stage 3 (L5D7), Stage 4 (W1–W2), and Stage 5 (P6). DEGs analysis showed distinct gene expression peaks in each stage (Fig. 4c, Supplementary Fig. 21a–c, and Supplementary Data 12). This model reflected the molecular shifts underlying each developmental phase.
a Temporal correlation map of wing disc development across sequential time points. Data were analyzed using the permutation test, with significance at *, 0.01 ≤ p < 0.05, **, p < 0.01. b Concentration dynamics of 20E in wing discs across developmental stages. c Temporal expression profiles of stage-specific gene sets across five developmental stages. d Proportional changes in the number of different cell types during wing disc development across five stages. e Spatial expression patterns of stage-specific differentially expressed genes. Dashed-line boxes highlight corresponding anatomical structures within the wing disc. f Stage-specific expression patterns of 20E-associated genes (Stages 1–5) visualized through a heatmap. g–k Temporal dynamics of key signaling pathway activities across five developmental stages. l Dynamic changes in cell–cell interaction networks across five developmental stages. m–q Dot heatmap showing statistically significant ligand–receptor interactions across five developmental stages (Stages 1–5), (m) Stage 1, (n) Stage 2, (o) Stage 3, (p) Stage 4, (q) Stage 5. The x-axis indicates the direction of communication between cell types; the y-axis lists the ligand-receptor pairs analyzed. The size of the dots denotes the statistical significance of the interaction: small open circles (p > 0.05), medium filled circles (0.01 <p < 0.05), large filled circles (p < 0.01). The color of the dots represents the communication probability (Commun. Prob.), as indicated by the color bar, ranging from max (red) to min (blue). The statistical significance of each predicted interaction (represented by the colored dots) was determined by performing a permutation test. The resulting p-values were adjusted for multiple comparisons across all ligand-receptor pairs and cell type comparisons within each panel using the Benjamini–Hochberg false discovery rate (FDR) correction method. Cell types: Wm (wing morphogenesis), Im (immune), Ma (matrix), Mp (metabolic process), Ad (axon development), Ci (ciliated), C1–C3 (cuticle 1–3), and Ep1–Ep3 (epithelial 1–3). Source data are provided as a Source Data file.
Specifically, in Stage 1, 20E concentrations were low (Supplementary Fig. 20a), and wing disc morphology remained unexpanded (Supplementary Fig. 20b). L5D1 and L5D2 exhibited strong transcriptional similarity, indicating closely coordinated gene expression in early stage. This period featured co-activation of the Toll/Imd and Hippo pathways, initiating early morphogenesis, alongside chromatin remodeling that reflected enhanced transcriptional plasticity (Supplementary Fig. 21d). As 20E levels began to rise in Stage 2, wing disc cells became more structure, and the tissue displayed increasing organization, with visible formation of distinct layers and regions (Supplementary Fig. 20a, c). Transcriptional similarity remained high from L5D3 to L5D6, suggesting sustained gene activity. Pathways involved in homologous recombination and DNA replication were strongly enriched (Supplementary Fig. 21e), likely to maintain genome stability during rapid cell proliferation. Stage 3 represented a major transition point. L5D7 displayed strong transcriptional separation, coinciding with peak 20E levels and wing disc entered a remodeling phase (Supplementary Fig. 20a, d). Morphogenesis and MAPK signaling pathways were likely involved to tissue boundary reorganization (Supplementary Fig. 21f). In Stage 4, W1 and W2 formed separate transcriptional clusters, showing stage-specific transcriptome shifts. 20E levels remained elevated, and pathways such as oxidative phosphorylation and ribosome biogenesis were activated, providing the energy and protein synthesis required for tissue maturation (Supplementary Fig. 21g). Morphologically, the wing disc showed defined regional structure (Supplementary Fig. 20e). By Stage 5, P6 displayed strong transcriptional separation, suggesting a major gene expression shift. 20E levels were highest at this stage, and coordinated activation of Wnt and MAPK signaling helped refine tissue architecture, while longevity-associated pathways contributed to maintaining tissue homeostasis (Supplementary Fig. 21h). Cuticle cells became the dominant population (Fig. 4d), consistent with high expression of cuticle-related genes such as CPH32, CPH42, CPG46, and CPG39 (Fig. 1c). These genes likely support the final formation of the adult wing cuticle. The wing disc morphology reached its final form, with all major structures and cell types fully differentiated and functionally mature. GO and KEGG enrichment analyses of stage-specific genes revealed a cascade of signaling pathways that were sequentially activated during developmental transitions.
Spatial mapping of stage-specific DEGs revealed their enrichment in distinct regions, indicating functional compartmentalization (Fig. 4e). For example, gene expression was concentrated in the wing bud at Stage 1, shifted to the epidermis at Stage 2, and returned to the wing bud at Stage 4. These spatial dynamics of gene expression closely mirrored tissue morphology, highlighting the tight coordination between gene activity and structural development (Fig. 4c, e). Meanwhile, the expression of 20E-responsive genes, such as USP and E75, also showed dynamic and stage-specific expression peaks (Fig. 4f). Notably, the 20E receptor gene USP was substantially upregulated in Stage 3, when 20E levels reached their peak (Fig. 4b), suggesting its key role in triggering 20E-driven metamorphic remodeling.
To dissect how cell–cell interactions (CCIs) regulate wing disc development, we performed CCI analysis across stages. Overall, the interaction network grew more complex from Stage 1 to 4 and then simplified at Stage 5, suggesting a shift from active remodeling to stabilization (Fig. 4l). Seven signaling pathways—Wnt, Notch, Hh, BMP, FGF, PARs, and SEMA4—showed stage-specific activation (Fig. 4g–k and Supplementary Fig. 22). During Stage 1, intercellular signaling was mainly concentrated in Wm, Im, and Ma cells. Notch signaling was prominently initiated by Ma cells and received by Wm, Ep1, and C1 cells, with key ligand–receptor pairs, including DLK1–NOTCH3 (Ma-secreted) and JAG1–NOTCH3 (Wm-secreted) (Fig. 4g, m and Supplementary Fig. 22a). These interactions indicate that Wm cells served as primary recipients of morphogenetic cues during early patterning. The preferential Notch reception by structural cell types (Wm, Ep1, C1) highlighted Notch’s central role in establishing spatial organization at this stage.
In Stage 2, Im cells emerged as major signaling hubs by integrating multiple pathways such as BMP, Hh, Notch, SEMA4, and FGF. Meanwhile, PAR signaling was predominantly initiated by Wm, Ep1, and Im cells, with Ma cells as principal targets. Notably, ligand–receptor interactions like PRSS1–PARD3 (from Wm or Im to Ma) suggested a conserved signaling axis potentially involved in immune–metabolic coordination (Fig. 4h, n and Supplementary Fig. 22b). In Stage 3, Notch signaling became more diverse, with Wm, Ep1/Ep2, C1/2, and Im cells acting as sources. Wm cells retaining dual roles as both primary senders and receivers. Wnt signals mainly targeted C1, Wm, and Ep1, while FGF ligands from Ma, Im, and Wm cells acted on Ep1 and C1/2. A key FGF1–FGFR2 axis (Wm to Ep1) was identified, potentially driving epithelial morphogenesis at this stage (Fig. 4i, o and Supplementary Fig. 22c).
At Stage 4, Wm and Ep1/Ep2 cells dominated the reception and transmission of BMP, Notch, SEMA4, and Wnt signals. Notably, Ep2 cells stood out by sending WNT2 and SEMA4G ligands to Wm and Ep1 through WNT2–(FZD2 + LRP5) and SEMA4G–PLXNB2 pairs. These interactions likely promote epithelial layering and boundary formation. (Fig. 4j, p and Supplementary Fig. 22d). By Stage 5, C1 cells mainly received Wnt signaling, while Ma cells received Hh signals. BMP signals from Ad cells targeted C2, and C3 showed strong intra-type BMP activity via GDF5–(BMPR1B + ACVR2B). PAR signaling was largely reduced at this stage. (Fig. 4k, q and Supplementary Fig. 22e). Notably, Ma cells continued to receive PAR signals across all stages. Both Ma and Im cells remained active participants in Hh signaling and had extensive contacts with other cells. These findings indicate that Ma and Im cells serve as persistent signaling hubs throughout development (Supplementary Fig. 22). Together, these findings reveal a dynamic and lineage-dependent signaling framework that ensures stage-specific coordination of silkworm wing disc morphogenesis.
To summarize these coordinated changes, we propose the Gene Transition Model (GTM), which divides wing disc development into five stages: (1) Developmental blueprint (L5D1–L5D2), characterized by early morphogenetic signaling and high transcriptional plasticity; (2) Cellular foundation (L5D3–L5D6), marked by cell growth and genomic stabilization; (3) Remodeling and sculpting (L5D7), defined by peak 20E and structural boundary reorganization; (4) Structural formation (W1–W2), characterized by tissue layering and increased metabolism; and (5) Maturation and stability (P6), where final differentiation and homeostasis shape the adult wing (Fig. 5). This model outlines a hormone-gradient-driven temporal axis, spatial decoding via region-specific activation of 20E-responsive genes, and modular coordination through dynamic signaling networks such as Wnt, Notch, Hh, and MAPK. Together, these findings provide a unified framework linking hormonal regulation, gene expression, and tissue morphogenesis during insect organ development.
Stage 1 represents the developmental blueprint (L5D1), followed by Stage 2 cellular foundation (L5D2–L5D6), Stage 3 structural formation (L5D7), Stage 4 remodeling and sculpting during spinning (W1–W2), and Stage 5 maturation and stability in the pupa–moth transition (P6). The upper panel illustrates morphological transitions of the wing disc across stages. The middle panel summarizes dynamic changes in major cell types, including Wm, Epithelial 1–3, Cuticle 1–3, immune, and matrix, as well as 20E levels in wing discs. The lower panel depicts coordinated cell state transitions and highlights key signaling pathways (Notch, PARs, FGF, Hippo, BMP, Hh, SEMA4, and Wnt) that regulate wing morphogenesis from larval wing disc to mature adult wing. Timepoint:the 5th instar larva at days 1–7 (L5D1–L5D7), wandering stage days 1 and 2 (W1–W2), and pupal stage day 6 (Pupa-D6). Wm:wing morphogenesis.
Discussion
Here, we constructed a high-resolution spatiotemporal atlas of the silkworm wing disc by integrating scRNA-seq, snRNA-seq, and Stereo-seq data, defining 12 cell types and capturing their developmental trajectories. Importantly, we proposed a five-stage GTM that connects hormone-driven gene expression, spatial organization, and cell fate transitions and regulatory. This model reveals how 20E signal guide region-specific morphogenesis. Our findings provide a detailed view of the dynamic changes that occur during wing disc development and highlight the stage-specific regulatory roles of 20E and related gene networks. Together, these results advance our understanding of insect developmental biology and offer a framework for studying similar processes in other species.
Among the identified cell types, Wm cells emerged as central progenitors that differentiate into two major structural cell lineages: epithelial (Epithelial 1–3) and cuticle (Cuticle 1–3) cells. Pseudotime trajectory analysis revealed that Wm cells express genes related to proliferation and morphogenesis in early state, while activated regulatory networks associated with tissue remodeling and organ formation in later state. SCENIC analysis further revealed a set of TFs potentially controlling Wm cell fate and guide lineage differentiation, such as Rfx, sqz, Blimp-1, Dll, and Pur-alpha. Several TF–target relationships further support their important roles during development. For instance, Dll regulates Fng, which is essential for wing morphogenesis37,38,39. Sqz are target genes in the Hippo pathway, particularly kibra, a gene associated with wing size regulation in B. mori18. Blimp-1 targets Hh, a key gene involved in wing disc development and larval tissue remodeling50. Importantly, Rfx emerged as a central regulator of Wm cell identity and downstream structural programs, targeting genes including bsh, dpy, futsch and pnr, which contribute to neuronal compartmentalization42, maintaining epidermal integrity43, cytoskeletal organization44, and dorsoventral patterning45, respectively. Notably, functional assay in B. mori and S. frugiperda confirmed Rfx as a conserved regulator essential for wing structural remodeling across lepidoptera. Together, these findings suggest that Rfx, Blimp-1, and Dll form a temporally and spatially coordinated transcriptional network directing Wm bifurcation and terminal differentiation. Given its conserved role in wing morphogenesis, Rfx represents a promising genetic target for pest management. More broadly, understanding how these TFs interface with signaling pathways such as Hippo, Toll/Imd, and MAPK will help uncover the regulatory logic underlying epithelial and cuticle lineage specification in lepidopteran wing development.
Beyond tracking natural developmental trajectories, our study reveals how 20E acts as both a temporal accelerator and a spatially specific regulator during wing disc morphogenesis in B. mori. Using in vitro 20E stimulation combined with time-resolved snRNA-seq, we identified a critical response window of ~30 min. During this period, transcriptional reprogramming was rapidly initiated in Wm, Cuticle, and Epithelial cells, followed by delayed responses in Immune and Matrix cells. These observations indicate that 20E induces a temporal compression of developmental events—triggering transcriptional and signaling cascades that normally require days within just hours. Importantly, this 20E-driven acceleration is cell-type-specific. Wm cells respond earliest, functioning as intermediate progenitors that give rise to epithelial and cuticle lineages, consistent with patterns observed during natural development. In contrast, Ma and Im cells show delayed activation, reflecting a hierarchical temporal order in how different cell types decode endocrine signals. While the systemic role of 20E in coordinating metamorphosis has been well established in insects, previous studies have primarily focused on its organism-wide endocrine actions2,5,7. Our data refine this understanding by mapping early 20E responses to spatially distinct clusters—particularly Wm cells—highlighting a highly localized and cell type-specific interpretation of systemic hormonal cues. These findings expand the current paradigm of hormonal regulation by providing a fine-resolution view of how 20E modulates morphogenetic processes at both spatial and temporal levels. This spatial decoding of endocrine signals offers a framework to dissect how global hormonal cues achieve precise tissue patterning during insect development.
To dissect the stage-specific developmental logic of the silkworm wing disc, we proposed a five-stage Gene Transition Model (GTM) that integrates gene expression dynamics, hormone concentrations, morphological progression, and spatial patterning during wing disc development. This model captures key transitions from early transcriptional plasticity to terminal differentiation, offering a stage-based scaffold to interpret how global endocrine cues are decoded into localized developmental outcomes. The GTM also reflects modular shifts in signaling pathway activity (e.g., Notch, Wnt, MAPK) and cell–cell interactions, indicating that tissue morphogenesis is coordinated through temporally layered and spatially compartmentalized regulatory programs. By linking 20E gradients to discrete developmental modules, this model advances our understanding of how systemic hormonal signals are translated into stage-specific gene regulatory cascades and cell identity decisions. This model provides a conceptual framework for dissecting organogenesis in holometabolous insects and may inform broader applications in developmental biology and evolutionary morphogenesis.
In addition, our work provides precise cellular contexts for many previously reported genes involved in wing disc development. By combining single-cell and spatial transcriptomic data, we identified the specific cell types and anatomical subregions where these genes are active at different stages. This fine resolution reveals that the expression of these classic “wing disc genes” is not uniform, but rather confined to distinct spatiotemporal niches. Such mapping allows a more refined understanding of how these genes integrate into broader developmental networks. It also offers a clear guide for designing targeted functional assays. Beyond B. mori, this framework may support future investigations in other lepidopteran insects or related model species, where spatial and temporal precision is essential for dissecting developmental programs.
Comparing our findings in B. mori with that of D. melanogaster and B. anynana revealed both conserved regulatory modules and lineage-specific developmental strategies11 (Supplementary Fig. 23a, b). Core pathways such as Wnt, Notch, and MAPK were consistently involved in cell fate determination and tissue patterning across species. However, our findings highlight several specific adaptations in B. mori. Specifically, the Wm cell in B. mori followed a bifurcated developmental trajectory toward epithelial and cuticle fates. This distinct progenitor identity has not been clearly reported in D. melanogaster, suggesting a silkworm-specific mechanism for lineage allocation. In B. anynana, a more diversified set of scale precursor cells supports elaborate wing coloration and patterning, highlighting another lineage-specific adaptation12 (Supplementary Fig. 23c). Furthermore, although 20E functions as a conserved hormone regulating metamorphosis, our data point to a silkworm-specific mode of spatial and temporal hormone responsiveness. In B. mori, Wm cells responded rapidly and locally to 20E, initiating early morphogenetic programs. This contrasts with the more systemic and less spatially resolved 20E response observed in Drosophila. Taken together, these comparisons demonstrate that while fundamental signaling frameworks are preserved, each lineage has evolved tailored regulatory strategies—shaped by ecological pressures and morphogenetic needs. Our single-cell and spatial atlas of B. mori not only confirms these shared principles but also provides a valuable reference for exploring how evolutionary flexibility in hormone integration and progenitor dynamics contributes to wing diversity in insects.
Our results highlight the value of combining single-cell and spatial transcriptomics to dissect the spatial architecture and 20E-mediated regulatory dynamics in wing disc development. However, several limitations remain. Although we validated key TFs such as Rfx using RNA interference, more precise approaches like conditional CRISPR/Cas9 (cell type and time-specific) would help dissect gene function with single-cell resolution and verify downstream protein interactions. In addition, the current five-stage GTM, while informative, may be further refined by extend temporal sampling to later metamorphic stages and increase resolution during critical transition windows. Finally, although we identified 12 major cell types, finer subpopulations might emerge with deeper sequencing or enhanced clustering resolution. Despite these challenges, our study demonstrates the potential of combining single-cell and spatial transcriptomics to capture the dynamic architecture of insect development. This framework may serve as a roadmap for future high-resolution tissue atlases and enable more targeted investigations into the cellular logic of morphogenesis. As spatial omics technologies continue to evolve, they will likely deepen our understanding of developmental processes and accelerate the discovery of lineage-specific regulatory mechanisms across species.
Overall, our study provides a high-resolution, integrative view of B. mori wing disc development, revealing how progenitor cell transitions, signaling pathway activity, and hormonal regulation are tightly coordinated in both time and space. This dataset provides a valuable foundation for exploring the molecular logic of cell fate determination and morphogenetic patterning in holometabolous insects. Together, our findings not only deepen the mechanistic understanding of wing disc development in B. mori but also offer a broadly applicable resource for comparative developmental and evolutionary studies across insect taxa. The spatiotemporal gene expression atlas and the proposed GTM model set the stage for future functional analyses and cross-species investigations into the principles of organogenesis.
Methods
Experimental animal preparation and sample collection
The B. mori strain Dazao was reared under controlled conditions at 27 ± 1 °C with 60% relative humidity under a natural photoperiod. Larvae were fed fresh mulberry leaves until the spinning stage. Wing discs were collected from both larval and pupal stages. Larvae and pupae of both sexes were randomly selected for all experiments. S. frugiperda larvae were reared in plastic containers at 26 ± 2 °C, 66 ± 5% relative humidity under a 16-h light/8-h dark cycle. They were fed an artificial diet until pupation. After pupation, individuals were moved to separate containers and maintained until adult emergence. All silkworm and S. frugiperda were provided by the Biological Science Research Center at Southwest University, China.
For tissue collection, gentle pressure was applied to the region below the spiracles of the second and third thoracic segments to make the wing disc area slightly protrude. Then, the epidermis was pierced using the tip of forceps (Dumont, 0209-5-PO, Switzerland), and the wing disc was carefully extruded. Each disc was immediately transferred into phosphate-buffered saline (PBS) at 4 °C. Under a stereo microscope (Leica, dm2700 p, Germany), visible fat body, tracheae, and other unwanted tissues were carefully removed. The entire procedure was completed within 5 min to preserve tissue integrity.
Microscopic observation of wing disc
Wing discs were observed using a stereo microscope (Olympus, Bx53, Japan). Each disc was carefully mounted on a glass slide to maintain stability and ensure clear imaging. High-resolution images were captured and analyzed using Photoshop (v22.4.3, Adobe, USA). The major (long) and minor (short) axes of each disc were measured to assess size and morphology.
Wing disc sample collection and scRNA-seq sequencing
Wing disc samples were collected from B. mori across developmental timepoints from L5D1 to L5D7, as well as W1, W2, and P6, to ensure comprehensive data for scRNA-seq analysis. Due to the smaller size of the wing discs, 200, 100, and 50 independent wing discs were collected for the L5D1–L5D7, W-1/W-2, and P6 stages, respectively. The wing disc samples collected from each timepoint were disassociated into single-cell suspensions using PBS (BBI, Cat# E607008-0500, China). Before sequencing, cell counting and viability assessment were performed, ensuring a cell viability rate of ≥80%. The prepared cell suspensions were processed using a DNBelab C Series High-Throughput Single Cell RNA Library Prep Kit V2.0 (MGI, Cat# 940-000519-00) according. Briefly, magnetic beads containing Oligonucleotide sequences were encapsulated together with wing disc cells in droplets, forming single-cell gel beads in emulsions. Within the droplets, cells underwent lysis to release mRNA, which was subsequently captured by the cell barcode sequences on the beads. After droplet breaking, reverse transcription was performed to synthesize cDNA. Sequencing librairies were constructed for the barcoded cDNA molecules and sequencing platform (MGI, MGISEQ 2000, China).
scRNA-seq data pre-processing
The raw FASTQ sequencing data were aligned to the NCBI B. mori reference genome (GCF_014905235.1_Bmori_2016v1.0) using the DNB-C4 pipeline (https://github.com/MGI-tech-bioinformatics/DNBelab_C_Series_HT_scRNA-analysis-software, version3.1.5). The alignment and quality control steps resulted in the generation of a gene expression matrix. Subsequent filtering was conducted for each sample using the Seurat package (v4.3.0)51 based on the following criteria:
1) Potential doublets identified using the DoubletFinder (version2.0.3)52 algorithm were removed; 2) cells with reasonable UMI counts (600 ≤ nUMI < max nUMI × 0.95), expressed genes (nGene ≥ 500 were retained. 3) Cells with log10(genes) per log10(nUMI) ≤ 0.8 were excluded. In addition, mitochondrial genes were excluded from downtream analysis. Dimensionality reduction and clustering were performed using the Scanpy package (v1.9.3)53. Highly variable genes (HVGs) were identified, retaining the top 2500 genes. Batch effects were corrected using the Harmony54 algorithm. Principal component analysis (PCA) was applied, retaining the top 30 PCs, followed by neighbor graph construction with n_neighbors set to 30. Clustering was conducted using the Leiden algorithm with a resolution of 0.4. DEGs were identified using the t-test approach, and the top 200 DEGs were selected for GO enrichment analysis. Cell type annotation was performed based on the functional properties of top markers and DEGs.
Wing disc tissue collection and Stereo-seq sequencing
Spatial transcriptomic profiling of wing disc tissue was performed using Stereo-seq (Spatial Enhanced Resolution Omics Sequencing) technology55. Briefly, multiple wing disc tissues from B. mori at different developmental stages (L5D2–L5D3 and W2) were embedded in OCT compound (Sakura, Cat# 4583, Japan), sectioned, and subjected to morphological quality control and RNA integrity assessment. The sections were then mounted onto spatiotemporal FC chips for permeabilization. Fluorescent imaging at various time intervals was performed to determine the optimal permeabilization time.
After identifying the optimal permeabilization time, sections with larger exposed areas of the wing disc were selected and mounted onto spatial SC chips. The mounted sections were fixed and stained with Qubit™ ssDNA Assay Kit (Thermo Fisher, Cat# Q10212, USA) for fluorescence imaging, which recorded tissue morphology and position. The tissues were permeabilized in an incubator at 37 °C for the previously determined optimal time to release RNA, followed by reverse transcription at 42 °C for 3 h to generate cDNA. After tissue removal and RNA release, the samples underwent magnetic bead purification and cDNA amplification. Next, the amplified cDNA products were fragmented and ligated with PCR adapters for further amplification and size selection. Subsequently, DNA nanoballs were prepared and sequenced using the DNBSEQ-T7RS platform. The spatial chips are equipped with capture probes that bind to mRNA molecules released from tissue cells. The captured gene information was spatially resolved based on the probe’s unique CID (Capture ID), which corresponds to the original location within the wing disc.
Stereo-seq data pre-processing
Raw Stereo-seq FASTQ data were aligned to the NCBI B. mori reference genome (GCF_014905235.1_Bmori_2016v1.0) using the SAW pipeline56 (https://github.com/STOmics/SAW version 6.0.0) for quality control. Downstream analysis was performed at a bin20 resolution. Quality control of the data was conducted using the Scanpy package (v1.9.3)53. Cells were filtered using the criteria of min_cells = 20 and min_genes = max (genes) × 0.05. Cell types annotated in the scRNA-seq dataset were used as references. Spatial annotation was achieved using the Cell2location (v0.1.3)57 algorithm.
20-hydroxyecdysone geneset
To identify the 20E hormone-related genes in silkworm wing discs (WDs), we conducted a comprehensive review of relevant literature available on the Web of Science. Based on their functions and roles within signaling pathways, these genes were classified into several categories: 20E Receptor, 20E Signaling Cascade Proteins, Signal Cascade-Related Enzymes, Immune-Related Factors, Functional Proteins, Protein Receptors, Transcription Factors (TFs), and Cuticular Proteins. A detailed list of these genes is provided in Supplementary Data 4. This classification offers a systematic framework for understanding the roles of these genes in WD development and their involvement in the 20E signaling network.
Trajectory analysis and pseudotime analyses
Cellular trajectories were constructed using the Tome pipeline (https://github.com/ChengxiangQiu/tome_code)40. Pseudotime analysis was performed in Monocle2 (v2.28.0)58 using the DDRTree method. To focus on specific cell populations, expression matrixes for epithelial, cuticle, and Wm-related cells were extracted. DEGs were identified using the FindAllMarkers function, with significant genes filtered using an adjusted p-value threshold of p_val_adj <0.05. Pseudotime visualization was conducted using CellRank2 (v2.0.0)59.
Pathway enrichment analysis of gene set
Using the R package “ClusterProfiler” (v4.6.2, Bioconductor, China)60 to perform GO enrichment and KEGG pathway enrichment analyses, with a p-value cutoff set at 0.05. Data visualization was primarily conducted using the ggplot2 (v3.5.1) package.
RNA interference experiment
Double-stranded RNA (dsRNA) targeting specific genes was synthesized using the Ribomax T7 RNA Production System (Promega, Cat# P1300, USA) according to established protocols61,62 and stored at –80 °C for subsequent experiments. Double-stranded primer sequences were provided in Supplementary Data 6. Size-matched fifth-instar B. mori larvae, at the early stage of day 2, were randomly divided into experimental and control groups (three biological replicates, with 30 independent wing disc per replicate). Each larva in the experimental group was injected with 10 μg dsRNA into the third abdominal segment using a fine capillary needle, while control larvae were injected with an equivalent amount of dsRNA targeting EGFP. After 48 h, five larvae from each group were dissected to assess wing disc morphology and evaluate RNAi efficiency via RT-qPCR. The remaining larvae were reared to the pupal stage, during which wing development was monitored. The incidence of wing disc segmentation defects and adult wing abnormalities was statistically compared between groups. The same procedure was applied to assess the role of Rfx in S. frugiperda. Fifth-instar larvae with uniform growth conditions (three biological replicates, with 20 independent wing disc per replicate) were injected with dsRNA targeting SfRfx. Wing discs were dissected 48 h post-injection to examine morphological changes and determine knockdown efficiency using RT-qPCR. Surviving larvae were allowed to pupate, and adult wing morphology was analyzed and compared statistically.
Rfx conservation analysis
The gene and protein sequences of Rfx were obtained from SilkDB3.0 (https://silkdb.bioinfotoolkits.net) and NCBI. Phylogenetic trees were built in MEGA 8.0. Conserved domain analysis was performed using the CDD website (https://www.ncbi.nlm.nih.gov/Structure/bwrpsb/bwrpsb.cgi), and motif analysis was conducted online using the MEME website (https://meme-suite.org/meme/tools/meme).
In vitro culture of wing discs and 20E treatment
To examine the time-dependent transcriptional response of wing discs to 20E, we established an in vitro culture system and applied 20E at different time points. Wing discs were dissected using forceps (Dumont, 0209-5-PO, Switzerland) and washed in PBS to remove surrounding tissues. Cleaned wing discs were transferred into Grace’s medium (Gibco, Cat# 11300-043, USA) supplemented with 10% fetal bovine serum (FBS) (Gibco, Cat# A5669701, USA) and 20E (Yuanye, Cat# S25531, China).
To determine the optimal 20E concentration, we first performed a dose–response test (see Supplementary Fig. S12). Wing discs were cultured in medium containing 0, 0.1, 0.2, 0.5, 1, 2, 5, or 10 μg/mL of 20E, each with 10% FBS. We measured the expression levels of known 20E-responsive genes, including EcR and USP. The results showed that 2 μg/mL of 20E-induced significantly increased the expression of these genes. Previous studies also reported that this concentration effectively activated 20E responses in cultured wing discs.
Silkworms have an open circulatory system, so endogenous 20E in hemolymph can influence multiple tissues. However, the efficiency of 20E uptake into isolated wing discs remains uncertain. To ensure strong and consistent stimulation, we chose 2 μg/mL of 20E to induce a robust transcriptional response. This allowed us to capture the activation timing of 20E target genes and compare it with natural development. Therefore, we used this concentration in the formal experiment.
In the formal experiment, wing discs were cultured in complete medium containing 2 μg/mL of 20E for 10, 20, 30, 40, 50, or 60 min, and for 4 or 6 h. A blank control group (immediately frozen without culture) was also included. After incubation, we transferred each sample into a sterile microcentrifuge tube. We removed the medium, washed the discs twice with PBS, blotted off excess liquid, and immediately froze the samples in liquid nitrogen. These samples were used for snRNA sequencing.
Wing disc tissue collection and snRNA-seq sequencing
The fresh-frozen wing disc tissues were ground and lysed to prepare single-nucleus suspensions. Nuclei were extracted and isolated from fresh-frozen wing disc tissue using PBS (BBI, Cat# E607008-0500, China) and resuspended. Nuclei suspensions were subjected to quality control, only those with cell viability >85% and cell count >100 thousand were retained for downstream experiments. Nuclei were processed using the DNBelab C Series Single-Cell Library Prep Set (MGI, 1000021082, China)63. Single-nucleus RNA libraries were constructed using the DNBSEQ-T7RS High-Throughput Sequencing Kit V3.0 (MGI, 940-000269-00, China) and sequenced on the MGI DNBSEQ-T7RS sequencing platform.
snRNA-seq data pre-processing
The raw sequencing data were aligned to the NCBI B. mori reference genome (GCF_014905235.1_Bmori_2016v1.0) using the DNB-C4 pipeline (https://github.com/MGI-tech-bioinformatics/DNBelab_C_Series_HT_scRNA-analysis-software) for quality control and alignment, resulting in the generation of a gene expression matrix. The gene expression matrix was filtered using the Seurat package (v4.3.0) using the aforementioned criteria. Dimensionality reduction and clustering were performed using Scanpy (v1.9.3) HVGs were identified, retaining the top 3000 genes. Batch effects were corrected using the Harmony algorithm, and PCA was performed, retaining 30 PCs. A neighbor graph was constructed with neighbors set to 10, and clustering was performed using the Leiden algorithm with a resolution of 0.4. Cell type annotation was conducted using SingleR64, with scRNA-seq annotated cell types serving as references.
Inferring cell–cell communication
Using blastp to perform homology alignment between human (GCF_000001405.40) and B. mori to obtain homologous gene correspondences. B. mori genes were mapped to their human homologs to enable downstream analysis. CCIs were inferred using the CellChat package (v1.6.1)65. For each developmental stage, a separate CellChat object was created, and CCI networks were independently calculated. These stage-specific CellChat objects were subsequently merged to facilitate comparative analysis across stages.
Transcription factor analysis
Using blastp to perform homology alignment between D. melanogaster (GCF_000001215.4) and bombyx to obtain homologous gene correspondences. B. mori genes were mapped to their D. melanogaster homologs to enable TF regulon analysis. The SCENIC package (v1.3.1)66 with default settings used to infer statistically active TFs and their targets.
Used with the D. melanogaster TF database to identify TF regulons and their associated downstream target genes. Top TFs were filtered based on an RSS > 0.01 and a zThreshold of 2. The resulting TF–target regulatory networks were visualized using Cytoscape (v3.9.1)67.
Gene network construction
Single-cell weighted gene co-expression network analysis (WGCNA) was performed using the hdWGCNA package (v0.2.26)68. Stage-specific hub gene co-expression networks were identified and visualized using Cytoscape (v3.9.1) to explore the dynamic regulatory relationships across developmental stages.
Similarity analysis
The Harmony algorithm was applied to integrate scRNA- and snRNA-seq datasets. To evaluate the correlation between cells from different time points in the scRNA- and snRNA-seq datasets, the Lorentzian distance metric from the R package “philentropy” (v0.8.0)69 was used to calculate pairwise distances between cells.
Quantification of 20E content in each wing disc
To accurately measure 20E levels during wing disc development, we directly quantified the hormone concentration in the wing disc tissue instead of the hemolymph. This tissue-specific method avoided the dilution effect caused by the silkworm’s open circulatory system. Wing discs were collected at different timepoints. Three biological replicates were used for each timepoint, with 30, 10, and 5 independent wing discs per replicate for the L5D1–L5D7, W-1/W-2, and P1/P3/P5/P7/M stages, respectively. The concentration of 20E was measured using an ELISA kit (Boshen Biotechnology, Cat# BS-E19221O2, China). For each sample, wing discs were homogenized in 100 μL of PBS. The homogenate was centrifuged at 2000 × g for 10 min, and the supernatant was collected for analysis. The ELISA plate was equilibrated at 25 °C for 20 min before use. Standards were prepared, and 50 μL of each standard solution was added to the corresponding wells. For sample wells, 10 μL of the sample supernatant and 40 μL of sample diluent were added. Blank wells were left empty. Then, 100 μL of HRP-conjugated antibody was added to each well, except the blanks. The plate was incubated at 37 °C for 60 min. After incubation, the wells were washed five times. Next, 50 μL each of substrate solutions A and B were added to all wells. The plate was incubated in the dark at 37 °C for 15 min. The reaction was stopped by adding 50 μL of stop solution to each well. Optical density (OD) values were measured within 15 min at 450 nm with a reference wavelength of 630 nm. A standard curve was generated in Excel (Microsoft Corporation, Redmond, WA, USA) by plotting OD values against the standard concentration. The curve followed the equation: Y = −0.001086×2 + 0.1162X + 0.07753, was used to calculate the 20E concentration in each sample. The total 20E content per wing disc was calculated by dividing the total 20E content by the number of discs in each sample.
Drosophila melanogaster wing disc single-cell RNA sequencing (scRNA-seq) data analysis
Sequencing data and aligned matrixes were downloaded from the GEO database (accession code GSE155543). The standard analysis pipeline was the same as that used in the article methods11. Clusters were split into AMP and epithelial cells. Finally, we obtained 12,872 cells and 6552 homologous features with the silkworm. Correlations were calculated using Spearman’s method.
Statistics and reproducibility
Each statistical test used to analyze data is indicated in the figure legends where p values were calculated. The animal experiment data in the figure are presented as mean ± SD. Two-tailed unpaired Student’s t test was used for statistical analysis of the differences between the two groups. Data in Fig. 2g, h were analyzed by one-way ANOVA. ***p < 0.001, **p < 0.01, and *p < 0.05 was accepted as statistically significant. All analysis assumed both populations were normally distributed when parametric tests were used. A confidence interval of 95% was used. Specific quantification methods and p values are indicated in the figure’s legends.
To identify differentially expressed genes (DEGs) between cell types within each developmental stage, we performed cluster-based differential expression analysis. Specifically, for the subset of cells from each developmental stage, and we employed the FindAllMarkers function from the Seurat package (v4.3.0). This function performs pairwise comparisons between each cell cluster and all other cells within that stage to identify its specifically upregulated gene markers. The key parameters used in the analysis were as follows: test.use = “MAST”, only.pos = TRUE. This setting retains only genes that are significantly upregulated in the target cluster. min.pct = 0.01 and min.cells.feature = 3. A gene was included in the analysis only if it was detected in at least 1% of cells within the cluster of interest and in at least 3 cells of that cluster. logfc.threshold = 0.1. Only genes with an average log fold change (base 2) greater than 0.1 were retained. return.thresh = 0.01. Only genes with an adjusted p-value (p_val_adj) less than 0.01 were considered statistically significant DEGs. The function automatically corrects for multiple hypothesis testing using the Bonferroni method, adjusting all raw p-values (p_val) based on the total number of genes tested across the entire dataset, yielding adjusted p-values (p_val_adj). For each developmental stage, the analysis generated a data frame listing the positive marker genes for each cell cluster or each timepoint, along with metrics such as the average log expression, the percentage of cells expressing the gene, p-values, and adjusted p-values.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The scRNA-seq, snRNA-seq and Stereo spatial raw sequeneing data have been deposited in Genome Sequence Archive (GSA) under bioproject PRJCA053684. scRNA-seq and snRNA-seq transcriptomics sequencing data and gene expression matrix files have been deposited in the China National GeneBank Sequence Archive (CNGB) under project CNP0005964. Stereo spatial transcriptomics sequencing data and gene expression matrix files have also been deposited in STOmicsDB under project STT0000176. Previously published Drosophila melanogaster wing disc of scRNA-seq data is available in the NCBI GEO database (GSE155543 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE155543]). All other data are available in the article or Supplementary files. Source data are provided with this paper. The processed data for all figures are available in the source data file associated with this manuscript. Source data are provided with this paper.
Code availability
The R and Python analysis code, encompassing all steps from preprocessing to visualization, is available on GitHub (https://github.com/meimeihe/Bombyx-mori-wing-disc.git) and archived on Zenodo (https://doi.org/10.5281/zenodo.17973196) with an assigned DOI. A detailed README.md file provides execution instructions.
References
Zhang, C. X., Brisson, J. A. & Xu, H. J. Molecular mechanisms of wing polymorphism in insects. Annu. Rev. Entomol. 64, 297–314 (2019).
Baehrecke, E. H. Ecdysone signaling cascade and regulation of Drosophila metamorphosis. Arch. Insect Biochem. 33, 231–244 (1996).
Angelini, D. R. & Kaufman, T. C. Comparative developmental genetics and the evolution of arthropod body plans. Annu. Rev. Genet. 39, 95–119 (2005).
Kango-Singh, M., Singh, A. & Gopinathan, K. P. The wings of develop from larval discs exhibiting an early differentiated state: a preliminary report. J. Biosci. 26, 167–177 (2001).
Mansilla, A., Martin, F. A., Martin, D. & Ferrus, A. Ligand-independent requirements of steroid receptors EcR and USP for cell survival. Cell Death Differ. 23, 405–416 (2016).
Roy, S., Hsiung, F. & Kornberg, T. B. Specificity of Drosophila cytonemes for distinct signaling pathways. Science 332, 354–358 (2011).
Lin, L., Li, H., Zheng, Q., Hu, J. & Wu, W. Research progress on the regulation of autophagy andapoptosis in insects by sterol hormone 20-hydroxyecdysone. Insects 14, 871 (2023).
Bischoff, M. et al. Cytonemes are required for the establishment of a normal Hedgehog morphogen gradient in Drosophila epithelia. Nat. Cell Biol. 15, 1269–1281 (2013).
Baonza, A. & Garcia-Bellido, A. Notch signaling directly controls cell proliferation in the Drosophila wing disc. Proc. Natl. Acad. Sci. USA 97, 2609–2614 (2000).
Ewen-Campen, B. & Perrimon, N. Wnt signaling modulates the response to DNA damage in the Drosophila wing imaginal disc by regulating the EGFR pathway. Plos Biol. 22, e3002547 (2024).
Everetts, N. J., Worley, M. I., Yasutomi, R., Yosef, N. & Hariharan, I. K. Single-cell transcriptomics of the Drosophila wing disc reveals instructive epithelium-to-myoblast interactions. Elife 10, e61276 (2021).
Prakash, A., Dion, E., Banerjee, T. D. & Monteiro, A. The molecular basis of scale development highlighted by a single-cell atlas of Bicyclus anynana butterfly pupal forewings. Cell Rep. 43, 114147 (2024).
Xia, Q. Y., Li, S. & Feng, Q. L. Advances in silkworm studies accelerated by the genome sequencing of Bombyx mori. Annu. Rev. Entomol. 59, 513–536 (2014).
Mitter, C., Davis, D. R. & Cummings, M. P. Phylogeny and evolution of lepidoptera. Annu. Rev. Entomol. 62, 265–283 (2017).
Wu, S. Y. et al. BmBlimp-1 gene encoding a C2H2 zinc finger protein is required for wing development in the silkworm Bombyx mori. Int. J. Biol. Sci. 15, 2664–2675 (2019).
Zhang, X. J., Li, D. D., Xu, G. F., Chen, Y. Q. & Zheng, S. C. Signal transducer and activator of transcription is involved in the expression regulation of ecdysteroid-induced insulin-like growth factor-like peptide in the pupal wing disc of silkworm. Insect Sci. 27, 1186–1197 (2020).
Zhang, B. et al. The role of chitooligosaccharidolytic beta-N-acetylglucosamindase in the molting and wing development of the silkworm Bombyx mori. Int. J. Mol. Sci. 23, 3850 (2022).
Yin, J. et al. BmSd gene regulates the silkworm wing size by affecting the Hippo pathway. Insect Sci. 27, 655–664 (2020).
Liu, L. et al. Spatiotemporal omics for biology and medicine. Cell 187, 4488–4519 (2024).
Sun, F. et al. Single-cell omics: experimental workflow, data analyses and applications. Sci. China Life Sci. 68, 5–102 (2024).
Wang, J. et al. Advances and applications in single-cell and spatial genomics. Sci. China Life Sci. 68, 1226–1282 (2024).
Ma, Y. et al. The architecture of silk-secreting organs during the final larval stage of silkworms revealed by single-nucleus and spatial transcriptomics. Cell Rep. 43, 114460 (2024).
Ma, Y. et al. A single-cell transcriptomic atlas characterizes the silk-producing organ in the silkworm. Nat. Commun. 13, 3316 (2022).
Liu, X. et al. Single-cell transcriptomes provide insights into expansion of glial cells in Bombyx mori. Insect Sci. 31, 1041–1054 (2024).
Feng, M. et al. Identification of silkworm hemocyte subsets and analysis of their response to baculovirus infection based on single-cell RNA sequencing. Front. Immunol. 12, 645359 (2021).
Xia, J. et al. Single-nucleus sequencing of silkworm larval midgut reveals the immune escape strategy of BmNPV in the midgut during the late stage of infection. Insect Biochem. Mol. Biol. 164, 104043 (2024).
Li, H. et al. Fly cell atlas: A single-nucleus transcriptomic atlas of the adult fruit fly. Science 375, eabk2432 (2022).
Jiang, L. Insights into the antiviral pathways of the silkworm. Front. Immunol. 12, 639092 (2021).
Fang, C. Y. et al. The Hox gene Antennapedia is essential for wing development in insects. Development 149, dev199841 (2022).
Ote, M. et al. Identification of molting fluid carboxypeptidase A (MF-CPA) in Bombyx mori. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 141, 314–322 (2005).
Medina, E. et al. Structure of the planar cell polarity cadherins Fat4 and Dachsous1. Nat. Commun. 14, 891 (2023).
Bettenhausen, B., Hrabe de Angelis, M., Simon, D., Guenet, J. L. & Gossler, A. Transient and restricted expression during mouse embryogenesis of Dll1, a murine gene closely related to Drosophila Delta. Development 121, 2407–2418 (1995).
Ren, C. et al. Two transcriptional cascades orchestrate cockroach leg regeneration. Cell Rep. 43, 113889 (2024).
Wang, Y. P. et al. Cytokine receptor DOME controls wing disc development in Bombyx mori. Insect Biochem. Mol. Biol. 148, 103828 (2022).
Romanova-Michaelides, M. et al. Morphogen gradient scaling by recycling of intracellular Dpp. Nature 602, 287–293 (2022).
Qian, W. et al. Decapentaplegic retards lipolysis during metamorphosis in Bombyx mori and Drosophila melanogaster. Insect Biochem. Mol. Biol. 155, 103928 (2023).
Ling, L. et al. MiR-2 family targets and to regulate wing morphogenesis in Bombyx mori. Rna Biol. 12, 742–748 (2015).
Rauskolb, C., Correia, T. & Irvine, K. D. Fringe-dependent separation of dorsal and ventral cells in the Drosophila wing. Nature 401, 476–480 (1999).
LeBon, L., Lee, T. V., Sprinzak, D., Jafar-Nejad, H. & Elowitz, M. B. Fringe proteins modulate Notch-ligand cis and trans interactions to specify signaling states. Elife 3, e02950 (2014).
Qiu, C. X. et al. Systematic reconstruction of cellular trajectories across mouse embryogenesis. Nat. Genet. 54, 328–341 (2022).
Li, N. N. et al. Hippo pathway regulates somatic development and cell proliferation of silkworm. Genomics 111, 391–397 (2019).
Xu, C., Ramos, T. B., Marshall, O. J. & Doe, C. Q. Notch signaling and Bsh homeodomain activity are integrated to diversify Drosophila lamina neuron types. Elife 12, RP90136 (2024).
Wilkin, M. B. et al. Dumpy is a gigantic extracellular protein required to maintain tension at epidermal-cuticle attachment sites. Curr. Biol. 10, 559–567 (2000).
Roos, J., Hummel, T., Ng, N., Klämbt, C. & Davis, G. W. Futsch regulates synaptic microtubule organization and is necessary for synaptic growth. Neuron 26, 371–382 (2000).
Calleja, M. et al. Generation of medial and lateral dorsal body domains by pannier gene of Drosophila. Development 127, 3971–3980 (2000).
Yamanaka, N., Rewitz, K. F. & O’Connor, M. B. Ecdysone control of developmental transitions: lessons from Drosophila research. Annu. Rev. Entomol. 58, 497–516 (2013).
Hill, R. J., Billas, I. M., Bonneton, F., Graham, L. D. & Lawrence, M. C. Ecdysone receptors: from the Ashburner model to structural biology. Annu. Rev. Entomol. 58, 251–271 (2013).
Wang, H. B., Iwanaga, M. & Kawasaki, H. Stage-specific activation of the E74B promoter by low ecdysone concentrations in the wing disc of Bombyx mori. Gene 537, 322–327 (2014).
Saito, M., Fujimoto, S. & Kawasaki, H. Ecdysone and gene expressions for chromatin remodeling, histone modification, and Broad Complex in relation to pupal commitment in Bombyx mori. Arch. Insect Biochem. 115, e22076 (2024).
Chen, K. et al. The morphogen Hedgehog is essential for proper adult morphogenesis in Bombyx mori. Insect Biochem. Mol. Biol. 153, 103906 (2023).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Gong, C. et al. SAW: an efficient and accurate data analysis workflow for Stereo-seq spatial transcriptomics. GigaByte 2024, 111 (2024).
Kleshchevnikov, V. et al. Cell2location maps fine-grained cell types in spatial transcriptomics. Nat. Biotechnol. 40, 661–671 (2022).
Qiu, X. et al. Single-cell mRNA quantification and differential analysis with Census. Nat. Methods 14, 309–315 (2017).
Weiler, P., Lange, M., Klein, M., Pe’er, D. & Theis, F. CellRank 2: unified fate mapping in multiview single-cell data. Nat. Methods 21, 1196–1205 (2024).
Wu, T. Z. et al. ClusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innov. Amst. 2, 100141 (2021).
Zhu, K. Y. & Palli, S. R. Mechanisms, applications, and challenges of insect RNA interference. Annu. Rev. Entomol. 65, 293–311 (2020).
Yi, Y. & Wu, G. RNA-seq analysis of LPS-induced immune priming in silkworms (Bombyx mori) and the role of cytochrome P450 detoxification system in the process. Int. J. Biol. Macromolecules 283, 136551 (2024).
Zhu, L. et al. Single-cell sequencing of peripheral mononuclear cells reveals distinct immune response landscapes of COVID-19 and influenza patients. Immunity 53, 685–696 (2020).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Jin, S., Plikus, M. V. & Nie, Q. CellChat for systematic analysis of cell-cell communication from single-cell transcriptomics. Nat. Protoc. 20, 180–219 (2025).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Morabito, S., Reese, F., Rahimzadeh, N., Miyoshi, E. & Swarup, V. hdWGCNA identifies co-expression networks in high-dimensional transcriptomics data. Cell Rep. Methods 3, 100498 (2023).
Drost, H. G. Philentropy: information theory and distance quantification with R. J. Open Source Softw. 3, 765 (2018).
Acknowledgements
We thank Yun He, Xiaoman Zhang, Jianxin Song, and Yu Zhang for assistance with tissue collection. This work was supported by the National Natural Science Foundation of China (Grant No. 32030103 to Q.X.) and the National Key Research and Development Program of China (2022YFD1201600 to Q.X. and 2022YFC3400400 to X.X.). Thanks for the support of the academy for advanced interdisciplinary studies device sharing service platform (equipment No.14A00784), Southwest University. We also would like to thank DCS Cloud (https://cloud.stomics.tech/) for providing the computational resources and software support necessary for this study.
Author information
Authors and Affiliations
Contributions
Q.X., M.X., A.C., P.Z. and Q.L. conceived the idea and designed the project. Q.L, H.C. and Y.F.Z. supervised sample preparation and sequencing data generation. Y.L. and L.Y.W. contributed silkworm materials. Q.L., H.C., X.W., H.N.L., Z.T., L.Z., G.L.L., X.C., X.L. and L.J. contributed to the selection of materials. M.H. and L.X.W., X.Z., H.Y.L, D.R., Y.W., X.J.Z, J.H., and G.C.L performed data processing and analysis. Q.L., M.H., H.C., W.L., P.Z, A.C., M.X. and Q.X. interpreted results and synthesized the scientific contents. H.C. performed RNAi experiment and slice staining. Q.L., M.H., H.C, P.Z., A.C., M.X. and Q.X. wrote the first draft of the manuscript. F.X., G.B.L. and X.X. revised the manuscript. Q.L., M.H., H.C, L.X.W., P.Z., A.C., M.X. and Q.X. wrote the final manuscript. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
W.L., G.C.L., and J.H. are employees at MGI Tech. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, Q., He, M., Chen, H. et al. Single-cell and spatial transcriptomics define 20E-driven developmental reprogramming in silkworm wing disc. Nat Commun 17, 3064 (2026). https://doi.org/10.1038/s41467-026-69518-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69518-6