Abstract
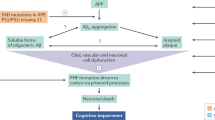
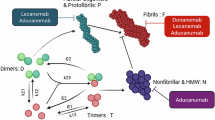
Collinolactone, featuring a 7/10/6 tricyclic core, has been proposed to be biosynthesized via a transannular [6 + 4] cycloaddition reaction. Besides its intriguing architecture, collinolactone holds pharmaceutical promises due to its ability to disrupt amyloid-β (Aβ) and tau aggregation, which are specifically found as disease culprits in the brains of Alzheimer’s disease (AD) patients and are key targets in current drug discovery efforts. However, challenges associated with its acquisition from a natural source and limited pharmacokinetic properties have hampered its further studies. Herein, we report the design, synthesis, and biological evaluation of 3-desoxycollinoketone B, a collinolactone derivative with improved pharmacokinetics for AD treatment. A stereoselective transannular [6 + 4] cycloaddition efficiently constructs the tricyclic core, allowing its scalable synthesis. AI-assisted binding prediction and simulations not only indicate superior binding of 3-desoxycollinoketone B to Aβ and tau aggregates to collinolactone, but also suggest a mechanistic basis for fibril destabilization. In vitro studies confirm its inhibition and dissociation of Aβ and tau fibrils, while in vivo experiments in AD mouse models show substantial amelioration of cognitive functions and Aβ/tau-associated pathology.
Similar content being viewed by others
Data availability
The data that support the findings of this study are available within the paper and its supplementary information files. Source data underlying the figures are provided with this paper. CCDC 2391862 contains the supplementary crystallographic data for compound 8a. These data can be obtained free of charge from The Cambridge Crystallographic Data Centre via [www.ccdc.cam.ac.uk/structures]. Source data are provided with this paper.
Code availability
The source codes for the models used in this study are publicly available at the following repositories: DiffDock-L [https://github.com/gcorso/DiffDock], NeuralPLexer2 [https://github.com/zrqiao/NeuralPLexer], and PIGNet2 [https://github.com/mseok/PIGNet2].
References
Yu, P., Patel, A. & Houk, K. N. Transannular [6 + 4] and ambimodal cycloaddition in the biosynthesis of heronamide A. J. Am. Chem. Soc. 137, 13518–13523 (2015).
Chen, N., Zhang, F., Wu, R. & Hess, B. A. Jr Biosynthesis of spinosyn A: A [4 + 2] or [6 + 4] cycloaddition? ACS Catal. 8, 2353–2358 (2018).
Kanoh, N. et al. Asymmetric total synthesis of heronamides A-C: stereochemical confirmation and impact of long-range stereochemical communication on the biological activity. Chem. Eur. J. 22, 8586–8595 (2016).
Zhang, B. et al. Enzyme-catalysed [6+4] cycloadditions in the biosynthesis of natural products. Nature 568, 122–126 (2019).
Wang, K. B. et al. A [6+4]-cycloaddition adduct is the biosynthetic intermediate in streptoseomycin biosynthesis. Nat. Commun. 12, 2092 (2021).
Schiewe, H. J. & Zeeck, A. Cineromycins, gamma-butyrolactones and ansamycins by analysis of the secondary metabolite pattern created by a single strain of Streptomyces. J. Antibiot. 52, 635–642 (1999).
Bode, H. B., Bethe, B., Höfs, R. & Zeeck, A. Big effects from small changes: possible ways to explore nature’s chemical diversity. Chembiochem 3, 619–627 (2002).
Schmid, J. C. et al. The structure of cyclodecatriene collinolactone, its biosynthesis, and semisynthetic analogues: effects of monoastral phenotype and protection from intracellular oxidative stress. Angew. Chem. Int. Ed. 60, 23212–23216 (2021).
Decker, R. L., Schray, D., Pfeffer, H. I., Grond, S. & Wagner, J. P. Conformations and rearrangements of collinolactone - experiments and theory on a dynamic cyclodecatriene. Chem. Eur. J. 30, e202303435 (2024).
Heo, S. et al. Synthesis of 3-dehydroxy-5-epi-collinoketone and its cognitive benefits by reducing aggregates of amyloid-β and tau. Preprint at Chemrxiv https://doi.org/10.26434/chemrxiv-2025-cqswq (2025)
Reiter, H. et al. Synthesis of Collinoketones via Biomimetic [6 + 4] Cycloaddition. J. Am. Chem. Soc. 147, 43205–43212 (2025).
Hardy, J. Alzheimer’s disease: the amyloid cascade hypothesis: an update and reappraisal. J. Alzheimers Dis. 9, 151–153 (2006).
Yao, W., Yang, H. & Yang, J. Small-molecule drugs development for Alzheimer’s disease. Front. Aging Neurosci. 14, 1019412 (2022).
Cummings, J., Zhou, Y., Lee, G., Zhong, K., Froseca, J. & Cheng, F. Alzheimer’s disease drug development pipeline: 2024. Alzheimers Dement. 10, e12465 (2024).
Singh, B., Day, C. M., Abdella, S. & Garg, S. Alzheimer’s disease current therapies, novel drug delivery systems and future directions for better disease management. J. Control. Release 367, 402–424 (2024).
Rashad, A. et al. Donanemab for Alzheimer’s disease: a systematic review of clinical trials. Healthcare 11, 32 (2022).
Wojtunik-Kulesza, K., Rudkowska, M. & Orzeł-Sajdłowska, A. Aducanumab—Hope or disappointment for Alzheimer’s disease. Int. J. Mol. Sci. 24, 4367 (2023).
Cummings, J. L. et al. The therapeutic landscape of tauopathies: challenges and prospects. Alzheimers Res. Ther. 15, 168 (2023).
Busche, M. A. & Hyman, B. T. Synergy between amyloid-β and tau in Alzheimer’s disease. Nat. Neurosci. 23, 1183–1193 (2020).
Pickett, E. K. et al. Amyloid beta and tau cooperate to cause reversible behavioral and transcriptional deficits in a model of Alzheimer’s disease. Cell Rep. 29, 3592–3604.e5 (2019).
Pasieka, A. et al. Dual inhibitors of Amyloid-β and Tau aggregation with Amyloid-β disaggregating properties: extended in cellulo, in silico, and kinetic studies of multifunctional anti-Alzheimer’s agents. ACS Chem. Neurosci. 12, 2057–2068 (2021).
Malafaia, D., Albuquerque, H. M. T. & Silva, A. M. S. Amyloid-β and tau aggregation dual-inhibitors: A synthetic and structure-activity relationship focused review. Eur. J. Med. Chem. 214, 113209 (2021).
Gandini, A. et al. Discovery of dual Aβ/Tau inhibitors and evaluation of their therapeutic effect on a Drosophila model of Alzheimer’s disease. ACS Chem. Neurosci. 13, 3314–3329 (2022).
Kwon, Y. et al. Rhizolutin, a novel 7/10/6-tricyclic dilactone, dissociates misfolded protein aggregates and reduces apoptosis/inflammation associated with Alzheimer’s disease. Angew. Chem. Int. Ed. 59, 22994–22998 (2020).
Kim, T., Lee, S. I., Kim, S., Shim, S. Y. & Ryu, D. H. Total synthesis of PGF2α and 6,15-diketo-PGF1α and formal synthesis of 6-keto-PGF1α via three-component coupling. Tetrahedron 75, 130593 (2019).
Roethle, P. A., Chen, I. T. & Trauner, D. Total synthesis of (-)-archazolid B. J. Am. Chem. Soc. 129, 8960–8961 (2007).
Roulland, E. & Ermolenko, M. S. Synthesis of the C12-C19 fragment of (+)-peloruside A through a diastereomer-discriminating RCM reaction. Org. Lett. 7, 2225–2228 (2005).
Surendra, K., Rajendar, G. & Corey, E. J. Useful catalytic enantioselective cationic double annulation reactions initiated at an internal π-bond: method and applications. J. Am. Chem. Soc. 136, 642–645 (2014).
Ding, X.-B., Furkert, D. P. & Brimble, M. A. 2-Nitropyrrole cross-coupling enables a second generation synthesis of the heronapyrrole antibiotic natural product family. Chem. Commun. 52, 12638–12641 (2016).
Schomaker, J. M., Pulgam, V. R. & Borhan, B. Synthesis of diastereomerically and enantiomerically pure 2,3-disubstituted tetrahydrofurans using a sulfoxonium ylide. J. Am. Chem. Soc. 126, 13600–13601 (2004).
Tseng, N.-W. & Lautens, M. Rhodium-catalyzed vinylcyclopropanation/cyclopentenation of strained alkenes via a sequential carborhodation process. J. Org. Chem. 74, 2521–2526 (2009).
Paterson, I., Yeung, K.-S. & Smaill, J.-B. The Horner-Wadsworth-Emmons reaction in natural products synthesis: expedient construction of complex (E)-enones using barium hydroxide. Synlett 10, 774–776 (1993).
Symkenberg, G. & Kalesse, M. Structure elucidation and total synthesis of kulkenon. Angew. Chem. Int. Ed. 53, 1795–1798 (2014).
Balas, L. et al. Linotrins: Omega-3 oxylipins featuring an E,Z,E conjugated triene motif are present in the plant kingdom and alleviate inflammation in LPS-challenged microglial cells. Eur. J. Med. Chem. 231, 114157 (2022).
Oka, N., Yamada, T., Sajiki, H., Akai, S. & Ikawa, T. Aryl boronic esters are stable on silica gel and reactive under Suzuki-Miyaura coupling conditions. Org. Lett. 24, 3510–3514 (2022).
Gade, N. R. & Iqbal, J. Stereoselective formal synthesis of macrolidecore of migrastatin using late stage C–H oxidation. Tetrahedron Lett. 54, 4225–4227 (2013).
Kim, H. J. et al. Chemoenzymatic synthesis of spinosyn A. Angew. Chem. Int. Ed. 53, 13553–13557 (2014).
Pracht, P., Bohle, F. & Grimme, S. Automated exploration of the low-energy chemical space with fast quantum chemical methods. Phys. Chem. Chem. Phys. 22, 7169–7192 (2020).
Lee, K., Lee, J., Park, S. & Kim, W. Y. Facilitating Transition State Search with Minimal Conformational Sampling Using Reaction Graph. J. Chem. Theory Comput. 21, 2487–2500 (2025).
Sperling, R. A. et al. Toward defining the preclinical stages of Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement. 7, 280–292 (2011).
Burley, S. K. et al. Updated resources for exploring experimentally-determined PDB structures and Computed Structure Models at the RCSB Protein Data Bank. Nucleic Acids Res. 53, D564–D574 (2025).
Lührs, T. et al. 3D structure of Alzheimer’s amyloid-β(1-42) fibrils. Proc. Natl. Acad. Sci. USA 102, 17342–17347 (2005).
Fitzpatrick, A. W. P. et al. Cryo-EM structures of tau filaments from Alzheimer’s disease. Nature 547, 185–190 (2017).
Corso, G., Stärk, H., Jing, B., Barzilay, R. & Jaakkola, T. DiffDock: diffusion steps, twists, and turns for molecular docking. In Proc. International Conference on Learning Representations (ICLR, 2023).
Corso, G. et al. Deep confident steps to new pockets: strategies for docking generalization. In Proc. International Conference on Learning Representations (ICLR, 2024).
Chen, S., Zheng, J. & Li, J. The impact of sample size after sampling on the accuracy of machine learning models. in Proc. 2024 International Conference on Computers, Information Processing and Advanced Education (CIPAE) 61–66 (IEEE, 2024).
Koes, D. R., Baumgartner, M. P. & Camacho, C. J. Lessons learned in empirical scoring with smina from the CSAR 2011 benchmarking exercise. J. Chem. Inf. Model. 53, 1893–1904 (2013).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Salentin, S., Schreiber, S., Haupt, V. J., Adasme, M. F. & Schroeder, M. PLIP: fully automated protein-ligand interaction profiler. Nucleic Acids Res. 43, W443–W447 (2015).
Schake, P., Bolz, S. N., Linnemann, K. & Schroeder, M. PLIP 2025: introducing protein-protein interactions to the protein-ligand interaction profiler. Nucleic Acids Res. 53, W463–W465 (2025).
Qiao, Z., Nie, W., Vahdat, A., Miller, T. F. III & Anandkumar, A. State-specific protein–ligand complex structure prediction with a multiscale deep generative model. Nat. Mach. Intell. 6, 195–208 (2024).
Bouysset, C. & Fiorucci, S. ProLIF: a library to encode molecular interactions as fingerprints. J. Cheminform. 13, 72 (2021).
Gremer, L. et al. Fibril structure of amyloid-β(1-42) by cryo-electron microscopy. Science 358, 116–119 (2017).
Im, D. et al. Decoding the roles of amyloid-β (1-42)’s key oligomerization domains toward designing epitope-specific aggregation inhibitors. JACS Au 3, 1065–1075 (2023).
Kunach, P. et al. Cryo-EM structure of Alzheimer’s disease tau filaments with PET ligand MK-6240. Nat. Commun. 15, 8497 (2024).
LeVine, H. 3rd. Quantification of beta-sheet amyloid fibril structures with thioflavin T. Methods Enzymol. 309, 274–284 (1999).
Ferreira, S. T. & Klein, W. L. The Aβ oligomer hypothesis for synapse failure and memory loss in Alzheimer’s disease. Neurobiol. Learn. Mem. 96, 529–543 (2011).
Cho, I. et al. Immobilized amyloid hexamer fragments to map active sites of amyloid-targeting chemicals. ACS Chem. Neurosci. 14, 9–18 (2023).
von Bergen, M., Barghorn, S., Biernat, J., Mandelkow, E.-M. & Mandelkow, E. Tau aggregation is driven by a transition from random coil to beta sheet structure. Biochim. Biophys. Acta 1739, 158–166 (2005).
Jakes, R., Novak, M., Davison, M. & Wischik, C. M. Identification of 3- and 4-repeat tau isoforms within the PHF in Alzheimer’s disease. EMBO J. 10, 2725–2729 (1991).
Ambadipudi, S., Biernat, J., Riedel, D., Mandelkow, E. & Zweckstetter, M. Liquid-liquid phase separation of the microtubule-binding repeats of the Alzheimer-related protein Tau. Nat. Commun. 8, 275 (2017).
Vergara, C. et al. Amyloid-β pathology enhances pathological fibrillary tau seeding induced by Alzheimer PHF in vivo. Acta Neuropathol. 137, 397–412 (2019).
He, Z. et al. Amyloid-β plaques enhance Alzheimer’s brain tau-seeded pathologies by facilitating neuritic plaque tau aggregation. Nat. Med. 24, 29–38 (2018).
Nelson, P. T., Braak, H. & Markesbery, W. R. Neuropathology and cognitive impairment in Alzheimer disease: a complex but coherent relationship. J. Neuropathol. Exp. Neurol. 68, 1–14 (2009).
Luna-Viramontes, N. I. et al. PHF-core tau as the potential initiating event for tau pathology in Alzheimer’s disease. Front. Cell. Neurosci. 14, 247 (2020).
Shao, C. Y., Mirra, S. S., Sait, H. B. R., Sacktor, T. C. & Sigurdsson, E. M. Postsynaptic degeneration as revealed by PSD-95 reduction occurs after advanced Aβ and tau pathology in transgenic mouse models of Alzheimer’s disease. Acta Neuropathol. 122, 285–292 (2011).
Liu, J. et al. Amyloid-β induces caspase-dependent loss of PSD-95 and synaptophysin through NMDA receptors. J. Alzheimers Dis. 22, 541–556 (2010).
Yoon, S. et al. Drug discovery and screening tool development for tauopathies by focusing on pathogenic tau repeat 3 oligomers. Angew. Chem. Int. Ed. 63, e202411942 (2024).
von Bergen, M. et al. Assembly of tau protein into Alzheimer paired helical filaments depends on a local sequence motif ((306)VQIVYK(311)) forming beta structure. Proc. Natl. Acad. Sci. USA 97, 5129–5134 (2000).
Lasagna-Reeves, C. A. et al. Tau oligomers impair memory and induce synaptic and mitochondrial dysfunction in wild-type mice. Mol. Neurodegener. 6, 39 (2011).
Yang, S. H. et al. Nec-1 alleviates cognitive impairment with reduction of Aβ and tau abnormalities in APP/PS 1 mice. EMBO Mol. Med. 9, 61–77 (2017).
Yang, S. H. et al. A small molecule Nec-1 directly induces amyloid clearance in the brains of aged APP/PS1 mice. Sci. Rep. 9, 4183 (2019).
Acknowledgements
We thank Dr. Dongwook Kim for analyzing the crystal structure of 8a, and Dr. Kyunghoon Lee for his assistance with the transition state calculations for the [6 + 4] cycloaddition. This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korean government (MSIT) (NRF-2021R1A2C2011203, NRF-2018R1A5A1025208, NRF-2021R1A2C2093916, Y.K., RS-2023-00257479) and by the BK21 FOUR Project. This research was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2019R1A6A1A10073887). We acknowledge support by Korea Dementia Research Project through the Korea Dementia Research Center (KDRC) (RS-2024-00349158, Y.K.). This research was also supported by KAIST Cross-Generation Collaborative Lab Project.
Author information
Authors and Affiliations
Contributions
Seongrok H. synthesized and characterized chemicals. M.C. conducted the full series of pharmacological evaluations from in vitro to in vivo studies. W.Z. performed the molecular simulation analyses. Seongrok H., M.C., Y.K., and Sunkyu H. wrote the original draft. J.K. participated in 5XFAD long-term drug administration. S.K. assisted in large-scale compound synthesis. J.K. and H.S. were involved in mouse breeding. I.C. and M.P. synthesized fluorescent Aβ peptides. Soljee Y. and W.S. synthesized the R3 peptide. W.S. and Suhyun Y. synthesized Aβ42 peptide. J.H. and H.H. carried out the ADME analyses. Seongrok H., M.C., W.Z., W.Y.K., Y.K., and Sunkyu H. analyzed and interpreted the data and wrote the paper. W.Y.K., Y.K., and Sunkyu H. supervised the entire work.
Corresponding authors
Ethics declarations
Competing interests
Y.K. is an employee of Amyloid Solution Inc. and received equity or equity options. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Heo, S., Cha, M., Zhung, W. et al. Synthesis of 3-desoxycollinoketone B and its ability to reduce Alzheimer-associated misfolded proteins. Nat Commun (2026). https://doi.org/10.1038/s41467-026-69662-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-69662-z