Abstract
Voltage-gated sodium channels are vital for regulating excitability in muscle and nerve cells, and their dysregulation is linked to a range of diseases. However, therapeutic targeting of Nav channels remains challenging due to a limited understanding of their gating mechanisms. Here, we present a cryo-EM structure of human Nav1.5 in an intermediate open state, stabilized by interactions between the N-terminal domain and the S6I segment. This structure reveals a possible Na+ binding site adjacent to the conserved inactivation (IFM) motif. Molecular dynamics simulations demonstrate that monovalent cations stably occupy this site, while electrophysiological recordings demonstrate that ion binding modulates IFM motif docking and fast inactivation kinetics. Our findings reveal that IFM accessibility is dynamically regulated in this intermediate state, refining the canonical door-wedge model of fast inactivation. Collectively, our study provides a revised structural framework for Nav1.5 gating mechanisms, suggesting an alternative pathway for ion accessibility that may inform better mechanistic and therapeutic strategies for treating Nav1.5-related cardiac arrhythmias.
Similar content being viewed by others
Introduction
Voltage-gated sodium channels are fundamental to the initiation and propagation of action potentials. In the heart, the predominant sodium channel isoform Nav1.5 (SCN5A) is indispensable for the rapid depolarization phase of the cardiac action potential1. Upon membrane depolarization, Nav1.5 channels activate with sub-millisecond kinetics, permitting a transient inward sodium current (INa) that drives the rapid upstroke of the action potential2. Channel opening is rapidly followed by fast inactivation, a tightly coupled process that closes the pore and terminates sodium influx within milliseconds3. Proper coordination of activation and inactivation is essential for maintaining normal cardiac electrical activity4. Perturbations in Nav1.5 function, such as mutations that impair fast inactivation, are implicated in a spectrum of inherited and acquired cardiac arrhythmias5,6,7. Thus, the precise regulation of Nav1.5 gating transitions is essential for maintaining normal cardiac excitability and rhythm.
Nav1.5 is composed of a large pore-forming α-subunit organized into four homologous domains (DI to DIV), each comprising six transmembrane segments (S1–S6) arranged in a pseudo-tetrameric configuration8. Within each domain, the S1–S4 helices form the voltage-sensing domain (VSD), whereas the S5–S6 helices and the intervening reentrant loop constitute the pore domain9,10,11. The activation gate is formed by a cluster of hydrophobic residues that line the intracellular ends of the S6 helices10,11. Fast inactivation is governed by the highly conserved isoleucine-phenylalanine-methionine (IFM) motif, also known as the inactivation particle, located in the intracellular DIII–DIV linker9,10,11,12,13. Emerging evidence suggests that the IFM motif does not simply occlude the pore but rather engages in an allosteric mechanism that modulates fenestrations near the intracellular activation gate9,10,11. An alternative model proposes that hydrophobic residues in the activation gate, rather than the IFM motif alone, play a direct role in the fast inactivation process14, suggesting a more complex interplay between the pore domain and the IFM.
Previous functional studies demonstrated that the transition of the IFM motif from an unbound, solvent-exposed state to a bound state within the IFM receptor pocket is essential for activation gate closure during fast inactivation15,16. However, in the open state structure in which the IFM motif is mutated to QQQ, the III-IV linker, including the IFM motif, is unresolved10. In our recent full-length structure of human Nav1.5 in an open conformation, we identified a stabilizing interaction between the DIII–DIV linker and the EF-hand domain of the C-terminal domain (CTD), along with a loosely bound conformation of the IFM motif within the IFM receptor pocket11. These structural insights differ from the conventional model of IFM motif displacement which was primarily established from functional studies utilizing substituted cysteine accessibility method approaches10,12,15,17,18,19.
In this study, we investigate the gating mechanisms of human Nav1.5 (hNav1.5) using structure-function analysis. The cryo-EM structure of full-length hNav1.5 in the intermediate open state is stabilized by interactions between the N-terminal domain (NTD) and the intracellular end of the S6 segment in DI (S6I). This structure conducts Na+ ions during molecular dynamics simulations. Electrophysiological measurements reveal that salt-bridge interactions maintaining the IFM motif shape are critical for fast inactivation. We also identified a second Na+ ion binding site near the IFM motif. We propose that this binding site is also accessible to other positively charged ions, such as Ag+, which is a common probe used as a labeling agent in cysteine accessibility assays15. Together, our findings reveal an alternative pathway for IFM motif accessibility by positive ions during channel opening, even when the IFM is sequestered within its canonical receptor site.
Results
Structure of human Nav1.5
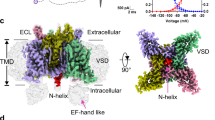
To investigate the gating mechanisms of Nav1.5, we purified full-length protein (Supplementary Figs. 1, 2) for cryo-EM studies. Image processing and the three-dimensional reconstruction resulted in a cryo-EM density map with a global resolution of 3.48 Å (Fig. 1, Supplementary Fig. 2). Subsequent two rounds of local refinement further improved the EM density across most regions (Fig. 1e, Supplementary Fig. 2). The atomic model built from this map includes the transmembrane core, extracellular domains, the III–IV linker, a helical segment of the II-III linker, and the NTD (Fig. 1a, b). Structural comparisons with the previously reported Nav1.5 structures revealed a dilation of the pore domain (Fig. 1c, Supplementary Fig. 3b, c). Notably, we identified density corresponding to a glyco-diosgenin (GDN) molecule at the activation gate region, which stabilizes the dilated conformation (Fig. 1c). This stabilization is accompanied by lateral shifts of all four S6 segments and the outward displacement of the S4–S5 linker helices (Supplementary Fig. 3c). We further observed a counterclockwise rotation of VSDs relative to the pore domain when viewed from the intracellular side (Supplementary Fig. 3b). Superimposition with the partially inactivated state structure Nav1.5-E1784K indicated larger root mean square deviations (RMSDs) for the S6I, S6II, and S6III segments at the activation gate region (Supplementary Fig. 3d), while the regions near the selectivity filter remained largely unchanged. These conformational changes reflect a transition from an α-helical to a π-helical configuration in the S6I and S6III segments. The binding of the GDN molecule stabilizes the π-helix conformation around the flexible ‘Gly-Ser’ (GS) motif of S6I and S6III (Fig. 1f, g), while S6II and S6IV maintain an α-helical conformation (Fig. 1g). The S4 segments of all VSDs are shifted outward with minimal alteration in the side-chain conformations of gating charges, resembling an activated state (Fig. 1d, Supplementary Fig. 3a).
a Cryo-EM map viewed from the membrane plane. Individual domains, inter-domain linkers, and detergent micelles are segmented and color-coded. b Ribbon model of hNav1.5 viewed from the membrane plane, with a glycol-diosgenin (GDN, gray) molecule at the AG. N-acetylglucosamine (NAG, gray) moieties are shown as sticks. All domains and interdomain linkers are segmented and color-coded according to the density map. c Structural superimposition of hNav1.5 with Nav1.5-E1784K (PDB ID: 7DTC, transparent teal) shows a lateral dilation of the overall structure. The arrangement of VSDI-IV is shown. The density of GDN is highlighted. d Comparison of S4 segments and associated gating charges (GCs) residues across VSDI-IV between hNav1.5 and Nav1.5-E1784K by superimposing each VSD over its S2 segment. GC residues are depicted as sticks. An1 and An2 denote anion1 and anion2, respectively. OR denotes the occluding residue. e Cryo-EM densities of SF and IFM motif residues. f Cryo-EM densities of the π-helical regions in segments S6I and S6III. g Ribbon representation illustrating π-bulging in segments S6I and S6III. Segments S6II and S6IV remain in α-helical conformations. Residues involved in π-bulging are labeled.
Intermediate open state conformation of the activation gate
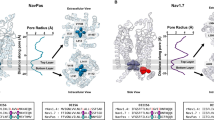
Two hydrophobic rings located at the lower part of the S6 segments define the pore diameter, which varies depending on the kinetic state of Nav channels11. Previous studies have reported that the diameter of an open activation gate is ~10 Å to allow the passage of hydrated Na+ ions (~7.2 Å in diameter)10. In our hNav1.5 structure, the average pore diameters at the upper and lower layers are ~10.2 Å and ~10 Å, respectively (Fig. 2a, Supplementary Fig. 4a). The residues contributing to the upper layer include L409, L935, I1466, and I1768 (Fig. 2a), while those forming the lower layer are A413, L938, I1470, and I1771 (Fig. 2a). Analysis of the pore radius profile relative to other Nav1.5 structures revealed that the pore radius at the selectivity filter region closely matches that of other inactivated state structures (Fig. 2b, Supplementary Fig. 4b). In contrast, the region from the central cavity to the intracellular pore opening, shows a pronounced dilation (Fig. 2b, Supplementary Fig. 4b). An asymmetric dilation is observed at the activation gate compared to partially open structures (Fig. 2b, Supplementary Fig. 3d). This dilation is more pronounced in the S6I, S6II, and S6III segments than in the S6IV segment (Supplementary Fig. 3d).
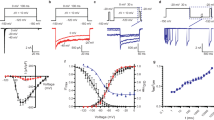
a Intracellular view of the activation gate formed by the S6 helices from D1 to DIV. The average diameters at the top (black dashed lines) and bottom (orange dashed lines) layers of the gate are ~10 Å. Side chains of AG lining residues are shown as sticks. b Comparison of pore radius profiles along the ion conduction pathway for hNav1.5 (orange), Nav1.5-E1784K (black), and rNav1.5c-BTX-B. (PDB ID: 7DTC, green). c Time-averaged water number densities along the pore axis under transmembrane voltages (TMV) of 0 mV (black), 745 mV (red), and 969 mV (blue). At 0 mV, the activation gate region is dehydrated; hydration increases at 745 mV and becomes continuous at 969 mV. The cyan dotted line indicates bulk water density, and shaded regions represent standard error. d Representative frames from an MD simulation showing Na+ (magenta sphere) permeating through the SF, CC, and AG. e Structural alignment of hNav1.5 with rNav1.5c-BTX-B (PDB: 8T6L, color: pale cyan) shows the relative position of NTD and S4-S5I, based on superposition of S1–S3 segments of VSDI. In hNav1.5, the hydrogen bond between S61 (NTD) and S234 (S4–S5 linker) observed in rNav1.5c-BTX-B (S62 and S235) is disrupted, and the NTD is displaced downward. f Cryo-EM densities of residues involved in the NTD-S6I interaction. g Intracellular view of the interface between the NTD and the S6I segment. R433 and K430 of S6I are positioned near D57 of the NTD. Inset i, potential salt bridges between D57 and both R433 and K430. Inset ii, structural alignment with rNav1.5c-BTX-B reveals a 12.5° tilt and a downward shift of S6I toward the NTD. h Time course of inactivation (τ) from sodium current recordings shows that K430D and D57K have a slower inactivation decay. Four independent experiments (N) were performed to obtain the number of cells (n) in each group. The data are presented as mean ± SEM. One-way ANOVA and Post-Hoc Tukey test were used for statistical analysis, *p < 0.05, WT vs K430D * p = 0.008 and WT vs D57K * p = 0.01.
To further characterize Na+ conductance in our intermediate open state structure, we performed MD simulations with harmonic restraints applied to Cα positions to prevent pore collapse. In the absence of transmembrane voltage (TMV), the pore remained largely dehydrated and nonconductive (Supplementary Fig. 4c), with water density significantly lower than that of the bulk solvent. This dehydration is likely attributable to hydrophobic residues lining the activation gate region (Fig. 2c). Application of a TMV of 745 mV enhanced hydration likely by promoting conformational transitions toward a partially hydrated state (Supplementary Fig. 4c). The water density in this region remained lower than bulk solvent levels, suggesting the presence of substantial free energy barriers to Na+ permeation (Fig. 2c). At a higher TMV of 969 mV, the pore became fully hydrated and conductive (Fig. 2c, Supplementary Fig. 4c, and Supplementary Movie 1). We quantified ion conduction by monitoring cumulative permeation events over 500 ns MD trajectories. A conduction event was recorded after the complete passage of a Na+ ion from the extracellular side to the intracellular side (Fig. 2d). During the 500 ns simulation at 969 mV, we observed three spontaneous Na+ conduction events (Supplementary Movie 1). In comparison, our previous open state structure demonstrated three Na+ permeation events at a lower TMV of 713 mV11. The requirement for a higher TMV in our current structure positions it between the canonical open state and the fast-inactivated state, establishing it as a bona fide intermediate open conformation in the Nav1.5 gating continuum.
The S6I segment - NTD interaction regulates activation and inactivation
The NTD of Nav1.5 in our structure is positioned intracellularly at the base of the VSDI, in agreement with other Nav1.5 structures (Figs. 1b, 2e, Supplementary Fig. 4d)11,20. A previous study identified a hydrogen bond between S62 of the NTD and S235 of the S4–S5I linker as a key stabilizing interaction for the NTD conformation (Fig. 2e, Supplementary Data Fig. 4e, f)20. Structural alignment of the VSDI-NTD region between hNav1.5 and rNav1.5c-BTX-B (PDB ID: 8T6L) revealed a downward displacement of the NTD and a concomitant shift of the S4–S5I linker in hNav1.5 (Fig. 2e). This displacement increased the distance between the two serine residues and precluded the formation of a hydrogen bond (Fig. 2e, Supplementary Fig. 4f). In hNav1.5, the stabilization of the NTD is mediated by the intracellular end of S6I, where D57 in the NTD forms salt bridges with K430 and R433 of S6I (Fig. 2f, g). In addition, an overall dilation of the pore and a tilt of the intracellular end of S6I toward the NTD were observed (Supplementary Fig. 4g, h). The helical region encompassing K430 to R433 exhibits a tilt angle of 12.5° relative to rNav1.5c-BTX-B (Fig. 2g). Although the side chain conformation of R433 remains unchanged, K430 undergoes a ~ 180° rotation toward D57 (Fig. 2g). To assess the functional importance of the S6I-NTD interaction on channel gating, site-directed mutagenesis and electrophysiological recordings were performed. Charge-reversal mutations at key interacting residues (D57K, K430D, and R433D) did not significantly alter activation, steady-state inactivation, recovery from inactivation, or late currents (Supplementary Fig. 5a–d, Supplementary Table 2). However, the D57K mutation resulted in a significant reduction in current density (Supplementary Fig. 5c), and western blot analysis confirmed that the protein expression for these mutants was not affected (Supplementary Fig. 5e, f), suggesting that the reduction in current density is due to a change in channel function. The D57K and K430D mutations were associated with a slowed time course of inactivation (Fig. 2h).
To dissect the conformational dynamics of the S6I segment, we performed 3D variability analysis (3DVA) to resolve motions along principal components21. Analysis along the leading principal component revealed 14 frames that showed a swing-like motion of the intracellular end of S6I (Fig. 3a, Supplementary Fig. 6a). The S6I density peaked at frame 1, decreased to a minimum at frame 7, and subsequently increased through frame 14 (Supplementary Movie 2). In frame 1, the intracellular end of S6I projects away from the activation gate, and the pore is open (Fig. 3b–d, Supplementary Data Fig. 6b). Frame 14 shows a distinct rotation of the intracellular end of S6I that results in a narrower activation gate (Fig. 3b–d, Supplementary Fig. 6a, b). Structural alignment of these models revealed a shortening of S6I in frame 14 that is accompanied by a ~ 13° tilt towards the center of the pore (Fig. 3d). To resolve side-chain positions, we further partitioned the principal component into two clusters and obtained reconstructions at ~3.8 Å resolution (Supplementary Fig. 7a, b). Comparison of the maps revealed distinct intracellular positions of S6I (Supplementary Fig. 7c), with an improved local resolution for S6I in cluster 1 (Fig. 3e). In cluster 1, R433 forms a salt bridge with D66 of the NTD, while K430 remains distant from the NTD (Fig. 3f, Supplementary Fig. 7d). In cluster 2, both K430 and R433 engage in salt-bridge interactions with D57 of the NTD (Fig. 3f, Supplementary Fig. 7d). These observations signify that R433 makes only a secondary, state-dependent contact with the NTD, whereas the primary latch involves D57-K430. In addition, disrupting D57-K430 (D57K or K430D) slows inactivation, while mutation of R433 alone does not alter the time constant of fast inactivation (Fig. 2h). This indicates that R433 plays an auxiliary role, and its loss does not significantly affect the fast inactivation and pore closure.
a 3D variability analysis of the cryo-EM map after consensus refinement and viewed from the membrane plane. The first frame map (Frame 1) and the last frame map (Frame 14) are aligned. The relative positions of the S6I segments' density are marked. S6I density in Frame 14 is shorter as compared to Frame 1. b Intracellular view of Frame 1 and Frame 14. These maps highlight the shift of the S6I segment toward the AG, which correlates with a transition from an open to a more constricted, inactivated-like conformation. c The S6 segments (S6I-IV) fitted into the densities of Frame 1 and Frame 14 exhibit varying positions of S6I. d Superimposed intracellular views reveal a 13° inward tilt of the S6I segment in Frame 14 relative to Frame 1. e Cryo-EM densities for the S6I segments. The densities are contoured at 5σ for Cluster 1 and Cluster 2. Selected residues are labeled. f NTD and S6I segment interactions in Cluster 1 and Cluster 2. In Cluster 1, R433 of S6I forms a salt bridge with D66 of the NTD (inset ‘i’). In Cluster 2, K430 and R433 of S6I are positioned close to D57 of the NTD, forming a distinct salt bridge network (inset ‘ii’). g Pore radius profiles along the conduction pathway for Cluster 1 (salmon) and Cluster 2 (marine) show a narrower activation gate in Cluster 2. h Ion permeation paths (gray dots) overlaid with S6I and S6III segments. Regions corresponding to the SF, CC, and AG are indicated.
To further analyze the reduction in pore diameter observed across frames 1 to 14 in the principal component analysis, we compared the pore radius profiles between structures from cluster 1 and cluster 2, which showed significant differences. In cluster 2, the pore widens just below the central cavity but then narrows again at the activation gate, whereas in cluster 1, the pore remains uniformly wider (Fig. 3g, h). Residues at the activation gate are well resolved in both structures (Supplementary Fig. 7e). The average pore diameters are ~10 Å in the upper layer and ~9.8 Å in the lower layer in cluster 1 (Supplementary Fig. 7f). Cluster 2 shows reduced averaged pore diameters of ~9.3 Å (upper layer) and ~9.5 Å (lower layer) (Supplementary Fig. 7f). An activation gate diameter <10 Å in cluster 2 suggests that it precludes the passage of hydrated Na+ ions.
IFM conformation and accessibility
The positioning of the IFM motif relative to its receptor is crucial for the fast inactivation mechanisms of mammalian Nav channels. The activation gate adopts either a closed or partially open conformation, while the IFM motif remains sequestered within its hydrophobic receptor in reported Nav1.5 structures8,11,20,22. Previous biochemical and mutagenesis studies suggest that detachment of the IFM motif from its receptor is a prerequisite for channel opening and Na+ permeation15,16. In our Nav1.5 structure, the IFM motif maintains its characteristic ‘U’-shaped conformation and is inserted into its receptor (Fig. 4a). Subtle repositioning of the IFM motif, accompanied by a downward displacement of the adjacent short α-helix in the III–IV linker, is observed (Fig. 4b). Comparative analysis of side chain conformations across available Nav1.5 structures reveals a conserved interaction pattern within this region. Specifically, the backbone oxygen of D1484 forms a highly conserved salt bridge with K1492, located on the short α-helix of the III-IV linker (Fig. 4c–f, Supplementary Fig. 8a). In contrast, the presence of the second salt bridge between K1492 and the backbone oxygen of M1487 within the IFM motif varies with the activation state of the channel: in Nav1.5-E1784K, the interaction distance is shorter (2.7 Å) compared to hNav1.5 (4.4 Å) and rNav1.5c-BTX-B (4.2 Å) (Fig. 4d–f, Supplementary Fig. 8a). The distance between the backbone oxygens of D1484 and M1487 remains consistent (4–5 Å), which is essential for preserving the ‘U’-shaped conformation of the IFM motif (Fig. 4f). To assess the importance of this interaction in the inactivation kinetics of hNav1.5, we used site-directed mutagenesis to substitute K1492 with a negatively charged residue (K1492D). Electrophysiological measurements (Fig. 4g) showed a markedly impaired fast inactivation decay (Fig. 4h, Supplementary Fig. 8d) for K1492D, while other biophysical properties such as steady-state inactivation, conductance, recovery from inactivation and late currents were not significantly affected (Supplementary Fig. 8b, c, e, f, Supplementary Table 2).
a Electrostatic surface of the IFM receptor pocket. The side chains of I1485, F1486, and M1487 are labeled and shown as sticks. b Structural superposition of hNaᵥ1.5 and Nav1.5-E1784K shows conformational changes in IFM residues along with a downward displacement. c Cryo-EM densities of the residues involved in salt bridge interactions at the IFM motif. d Backbone oxygens of D1484 and M1487 form a stabilizing salt bridge network with K1492 near the IFM motif. e Reduced distance between M1487 and K1492 indicates a more stable salt bridge network at the IFM motif in the Naᵥ1.5-E1784K structure. f Variation in the average distances between M1487 to D1484 (M to D is 4.7 Å), M1487 to K1492 (M to K is 3.8 Å), and D1484 to K1492 (D to K is 3.2 Å). These average distances are calculated using cryo-EM structures [PDB ids: 7DTC, 6LQA, 8T6L, 6UZ0, 7K18, and 8F6P; map resolution is ≤3.3 Å]. Error bars represent the standard deviation (SD); n = 7 structures deposited in the PDB. g Representative sodium current traces from patch-clamp recordings of K1492D. h Time constants of inactivation (τ) for WT and K1492D. Results show that K1492D displayed a slowed inactivation lacking voltage dependency. Four independent experiments (N) were performed to obtain the number of cells (n) in each group. The data are presented as mean ± SEM. One-way ANOVA and Post-Hoc Tukey test were used for statistical analysis, *p = 0.0001, WT vs K1492D. i Current-Voltage (I–V) relationship for WT and K1492D. The mutant did not significantly affect current density. Four independent experiments (N) were performed to obtain the number of cells (n) in each group. The data are presented as mean ± SEM. One-way ANOVA and Post-Hoc Tukey test were used for statistical analysis.
Secondary Na+ binding site
During our structural analysis, we observed prominent density within a hydrophilic pocket adjacent to the IFM motif (Fig. 5a). This density was deduced as a Na+ binding site for the following reasons: First, Na+ ions are the only cations present throughout the purification process. Second, the density is linked to the negatively charged side chain of D1484 (Fig. 5a). Third, a similar density for Na+ ion has been reported for D361 at the selectivity filter region in the human Nav1.7-β1–β2 complex23. We propose that D1484 and surrounding residues act as a binding site for Na+ or other monovalent cations in hNav1.5. Additionally, the IFM motif is surrounded on one side by the hydrophobic IFM receptor pocket and on the other side by the hydrophilic Na+ ion binding pocket (Fig. 5b). We compared the positions of key residues forming the hydrophilic Na⁺ binding pocket in our intermediate open state hNav1.5 structure and in the partially inactivated Nav1.5-E1784K structure. In our structure, we observed that D1484, E1773, and Y1495 are shifted outward, forming an electronegative pocket capable of accommodating Na+ (Fig. 5c). Additionally, there is a subtle repositioning of the IFM residues. The volume of this pocket measures 282 Å3 and exhibits two distinct regions of electrostatic surfaces (Fig. 5d). The outer region of this pocket is created by the side chains of Q1483, N1496, and K1499 (Supplementary Data Fig. 8g). The inner region is surrounded by D1484, F1486, M1487, Y1495, S1653, L1772, E1773, and S1776 (Supplementary Fig. 8h). As a consequence, the side chains of F1486 and M1487 in the IFM motif are positioned closer to the bound Na+ ion (4.7–5.1 Å), making them more accessible than I1485, which is positioned farther away at 7.2 Å (Supplementary Fig. 8i). Comparative analysis showed the presence of a significantly smaller pocket with a volume of 86 Å3 in Nav1.5-E1784K. This pocket is situated on the intracellular side of the channel, directly beneath the IFM motif (Fig. 5e). The conformation of the side chain of D1484 prevents Na+ coordination and binding.
a Cryo-EM density around the IFM region of hNaᵥ1.5: a spherical density (blue) corresponds to a Na+ ion coordinated by the carboxylate moiety of D1484. b Electrostatic surface view of the intracellular vestibule highlighting a negatively charged Na+ pocket (green outline) adjacent to the IFM pocket (yellow) and the AG. c Superposition of hNaᵥ1.5 and Naᵥ1.5-E1784K structures shows a Na+-D1484 interaction (3.2 Å) and a concerted displacement of E1773, Y1495 and the III-IV linker helix. d Secondary binding site analysis of hNaᵥ1.5 reveals a large (282 Å3) electronegative Na+ binding pocket near the IFM residues. The blue and magenta colors of the pocket region indicate positive and negative electrostatic potential, respectively. e Secondary binding site analysis of Naᵥ1.5-E1784K shows a markedly reduced binding pocket volume (86 Å3). This structure lacks the negatively charged pocket seen in WT hNaᵥ1.5. f Steady-state inactivation relationship showed that E1773Q presents a left shift, whereas D1484K is shifted to depolarized potentials (see Supplementary Table 2). Four independent experiments (N) were performed to obtain the number of cells (n) in each group. The data are presented as mean ± SEM. One-way ANOVA and Post-Hoc Tukey test were used for statistical analysis. g I-V relationship represents electrophysiological recordings from D1484K, Y1495F, and E1773Q. Four independent experiments (N) were performed to obtain the number of cells (n) in each group. The data are presented as mean ± SEM. One-way ANOVA and Post-Hoc Tukey test were used for statistical analysis. h D1484K displayed a left shift in the conductance curve in comparison to Y1495F and E1773Q (see Supplementary Table 2). Four independent experiments (N) were performed to obtain the number of cells (n) in each group. The data are presented as mean ± SEM. One-way ANOVA and Post-Hoc Tukey test were used for statistical analysis. i Time course inactivation relationship showed that D1484K and Y1495F resulted in slower inactivation, whereas E1773Q was not affected. Four independent experiments (N) were performed to obtain the number of cells (n) in each group. The data are presented as mean ± SEM. One-way ANOVA and Post-Hoc Tukey test were used for statistical analysis. *p < 0.05, WT vs 1484D p = 0.001 and WT vs Y1495F p = 0.04.
To assess the accessibility of the Na+ binding pocket to silver ions (Ag+), which can serve as labeling agents for cysteine-substituted IFM residues15,24, we conducted an all-atom MD simulation under isothermal-isobaric (NPT) conditions. The system was solvated in 150 mM AgCl and subjected to a transmembrane potential of 360 mV. It is important to note that Ag+ is not commonly parameterized in standard biomolecular force fields. In this simulation, Ag+ was used to mimic NaCl conditions, rather than to specifically model Ag+-protein interactions. Despite the modest applied voltage, Ag+ permeated the Na+ binding site within the first 16.5 ns of simulation (Supplementary Movie 3). The bound Ag+ remained coordinated in the pocket for the remainder of the 110 ns trajectory (Supplementary Fig 9a–c, Supplementary Movie 3). Structural analysis revealed that Ag+ is coordinated by the carboxylate side chains of D1484 and E1773 (Supplementary Fig 9d). In addition, the ion is localized near the IFM motif, suggesting a possible interaction (Supplementary Fig 9a–c). Therefore, cysteine-substituted IFM motif residues are likely to form stable thiolate-Ag+ coordination complexes and possibly alter fast inactivation kinetics.
To understand the importance of Na+ coordination inside this pocket in the intermediate open state structure of hNav1.5, multiple charge reversal and altered electrostatic mutations were generated for electrophysiological measurements (Supplementary Fig. 10a). Supporting the important role of D1484 in channel function, charge reversal of D1484 to a lysine (D1484K) affected several biophysical properties of the channel (Fig. 5f–i, Supplementary Table 2). Specifically, the steady-state inactivation was shifted toward more depolarized potentials (Fig. 5f), activation was shifted toward hyperpolarized voltages (Fig. 5h), and inactivation decay was significantly slowed (Fig. 5i). However, recovery from inactivation was not affected (Supplementary Fig. 10b). Mutation of E1773 to Q1773 in the inner region of the pocket led to the stabilization of steady-state inactivation as seen by a significant hyperpolarizing shift (Fig. 5f, Supplementary Table 2) without affecting other biophysical properties of this mutant channel (Fig. 5g–i, Supplementary Fig. 10, and Supplementary Table 2). Mutation of residue Y1495 to F1495 in this same inner region of the pocket slightly slowed the inactivation decay (Fig. 5i) without affecting any other biophysical properties of the channel (Fig. 5f–h, Supplementary Fig. 10 and Supplementary Table 2). None of the mutations had a significant impact on current density (Fig. 5g) or the late sodium current (Supplementary Fig. 10c). These functional data are consistent with the structural environment of the pocket. E1773Q disrupts the negative charge and enables Q1773 to form a hydrogen bond with D1484 or Y1495, thereby clamping the Na+ pocket, reducing ion occupancy, and stabilizing IFM docking (hyperpolarizing shift). Conversely, D1484K replaces a dynamic, anionic coordination site with a permanent cationic side chain, repelling Na+ ions and disrupting the tunable microenvironment. This weakens IFM docking and requires stronger depolarization for inactivation, consistent with the depolarizing shift observed in steady-state inactivation (Fig. 5f). Y1495 stabilizes the IFM motif via a π-sulfur interaction with M1487. The Y1495F mutation preserves this aromatic π-sulfur interaction but removes the hydroxyl group, resulting in a modest slowing of inactivation without substantial gating changes (Fig. 5f–i).
Discussion
Our cryo-EM structure of wild-type hNav1.5 reveals an intermediate open conformation of the activation gate, despite VSDIV adopting a fully activated state and VSDs from DI–DIII appearing in an inactivated conformation. Our structure provides important structural insights into activation-gate dynamics and alternative IFM accessibility.
Notably, the open state structure rNav1.5c/QQQ lacks the canonical IFM motif, an essential component of fast inactivation10. In the inactivated structures, the IFM motif is docked into its hydrophobic receptor site, stabilizing the closed or partially closed state of the activation gate8,22,25. The prevailing model posits that the release of the IFM motif from this receptor is required for gate opening, a concept derived primarily from substituted cysteine accessibility assays15. However, structural evidence delineating conformational transitions of the IFM motif during gating remains limited and thus impedes a full understanding of the structural rearrangements required for Na+ conduction. Our recent wild-type hNav1.5 open state structures revealed a repositioning of the IFM motif, which is stabilized through interactions with the CTD11. In the present study, we demonstrate the regulatory influence of NTD-S6I interactions on activation gate opening and propose an alternative model of IFM motif accessibility in the open state, wherein a secondary pocket near the IFM region accommodates positively charged ions.
Our cryo-EM structure of hNav1.5 represents a critical transition state between fully open and inactivated states. This conformation is functionally relevant, as it allows Na+ permeation during MD simulations under depolarizing voltage. Unlike fully open structures that require strong depolarization for conduction, our structure reveals a partially conductive pore stabilized without applied voltage, likely enabled by a GDN molecule trapped at the activation gate. This GDN molecule occupies a conserved hydrophobic site also seen in other voltage-gated sodium channel structures with wider activation gates10,23 and likely plays a key role in mechanically holding the gate open by separating the hydrophobic residues that otherwise collapse during inactivation. The stabilization by GDN is accompanied by a substantial dilation of the pore domain and coordinated rearrangements across all four S6 segments. Notably, this includes α-to-π helical transitions in S6I and S6III, consistent with emerging evidence that such transitions are conserved in voltage-gated ion channels20,23,26,27. Together, these findings position our structure as a functional intermediate between closed and open states, providing critical structural insights into the gating transitions that control sodium channel activation and fast inactivation.
Unlike most cryo-EM structures of Nav1.5, where the NTD and the intracellular end of S6I are poorly resolved, our structure captures a well-ordered interaction interface between these two regions. We observed distinct salt bridges between residues in the NTD and S6I, suggesting that this interdomain contact plays a stabilizing role in maintaining the open conformation. To further explore this coupling, we performed 3D variability analysis (3DVA), which revealed changes in S6I density as the activation gate transitions from an open to a partially closed state (Supplementary Fig. 6a, b). This resembles observations from inactivated state Nav structures, where the intracellular S6I is often unresolved13,28,29. 3DVA analysis also uncovered two conformational clusters in which S6I adopts alternate interactions with the NTD: R433 and K430 of S6I interact with D66 in cluster 1 and with D57 in cluster 2, correlating with changes in pore diameter (Fig. 3f). These findings provide structural evidence for a dynamic coupling between S6I and the NTD, implicating these salt bridges as modulators of gating transitions. While charge-reversal mutations at these sites did not significantly alter inactivation kinetics, the reduced current density observed in the D57K mutant supports a functional role for this interaction. Together, our data establish the NTD-S6I interface as a critical regulatory element in activation gate stability, conformational tuning, and consistent with prior DI-S6I mutations that abolish inactivation even when the IFM remains docked30,31,32.
We observed that activation gate opening is coupled with subtle remodeling of the IFM motif and its binding pocket, while the motif retains a stable U-shaped conformation within its hydrophobic receptor (Fig. 4a, b). Salt bridge interactions between K1492, located on a short α-helix near the IFM motif, and the backbone oxygens of D1484 and M1487 are crucial for stabilizing this conformation. Notably, the K1492-M1487 interaction varies depending on the conformational state of the activation gate. The charge-reversal mutation K1492D significantly impaired fast inactivation kinetics (Fig. 4h) without altering other gating parameters (Supplementary Fig. 8b, c, e, f, Supplementary Table 2). This suggests that K1492D decouples VSDIV activation from IFM docking and underscores the role of the salt bridge network in electromechanical coupling. These findings further establish that precise repositioning of the IFM motif, but not its full exposure to solvent, is critical for gate opening.
Our structure also uncovered an electronegative Na+ binding pocket directly adjacent to the docked IFM motif. This pocket, formed in part by D1484 and lined by F1486 and M1487, becomes accessible only when the activation gate is open and collapses in the inactivated state (Fig. 5d, e), rendering D1484 inaccessible to cations. This finding fundamentally challenges the prevailing interpretation of the substituted cysteine accessibility method (SCAM) that is based on the notion that IFM labeling by positively charged reagents indicates full solvent exposure following IFM displacement15,24. Instead, our results suggest that Ag+ and MTS reagents can access substituted cysteine residues through this pocket even when the IFM remains docked, providing an alternate mechanistic explanation for prior labeling results15,19,24. Furthermore, the kinetic phenotype of the D1484K mutant mirrors the effects of Ag+ labeling at F1486C and M1487C, reinforcing the functional importance of this pocket15,19,24. Collectively, these structural and functional observations indicate that this pocket acts as an active modulator of inactivation rather than a passive cavity. By coordinating permeant cations, the pocket directly tunes IFM docking in a state-dependent manner. When the pore is permissive, Na+ binding destabilizes the docked IFM, whereas loss of occupancy favors IFM engagement. This mechanism provides a structural explanation for classical observations that permeant Na⁺ alters inactivation kinetics33.
Together, these findings refine the current understanding of IFM accessibility, uncover a functionally coupled and structurally tunable Na+ pocket, and identify an allosteric site with therapeutic potential for modulating inactivation in Nav1.5 disease-causing mutations.
Methods
Cloning of the full-length human Nav1.5 expression vectors
Codon-optimized full-length human Nav1.5 (hNav1.5) cDNA was synthesized (GenScript, NJ) and cloned into a modified pcDNA3.1 vector containing an N-terminal twin-Strep tag followed by a TEV protease site and a C-terminal HRV 3 C site and FLAG tag. Point mutations D57K, K430D, R433D, K1492D, D1484K, Y1495F, and E1773Q were introduced by site-directed mutagenesis using the QuickChange II mutagenesis kit (Agilent). All constructs were verified by Sanger sequencing. All primers used in this study are included in Source Data file 1.
Purification of the full-length full length hNav1.5
Full-length human Nav1.5 was expressed in Expi293F GnTI- suspension cells (ThermoFisher) grown in Expi293 expression medium (ThermoFisher) at 37 °C with 8% CO₂. Cells were transiently transfected with pCDNA3.1-hNav1.5 using ExpiFectamine 293 (ThermoFisher) and harvested ~48 h post-transfection. Pellets were washed with PBS. The cell pellets were then suspended in lysis buffer containing 25 mM HEPES (pH 7.4), 150 mM NaCl, 0.1 mM EGTA, and 10% glycerol (buffer A), including protease inhibitors. Following homogenization, the membrane was isolated through ultracentrifugation and carefully mixed in buffer A supplemented with protease inhibitors, 1% (w/v) n-dodecyl-β-D maltopyranoside (DDM, Anatrace), and 0.1% (w/v) cholesteryl hemisuccinate (CHS, Anatrace) at 4 °C for 2 h. Following ultracentrifugation, the supernatant was incubated for 2 h with 5 mL of anti-Flag M2 affinity gel in buffer B (buffer A supplemented with 0.06% (w/v) glycol-diosgenin (GDN, Anatrace) and protease inhibitor cocktail). The column was washed with buffer B, followed by the elution of the protein using buffer B supplemented with 200 μg/ml of FLAG peptide. The eluted protein was subjected to affinity purification using Strep-Tactin XT 4Flow (IBA). Following a wash with buffer B, the bound protein was eluted using buffer B supplemented with 50 mM biotin (IBA). The eluent was concentrated and subsequently purified using a Superose 6 increase 10/300 gl column (Cytiva) in buffer C (25 mM HEPES at pH 7.4, 150 mM NaCl, 0.1 mM EGTA, and 0.06% GDN). The peak fractions of the purified protein were pooled and concentrated to around 8 µM for structural studies.
Cell culture and expression of hNav1.5 for electrophysiological recordings and western blots
HEK293 cells, from ATCC, were cultivated in DMEM high glucose media (Gibco) supplemented with 10% fetal bovine serum (Gibco) and 1% penicillin-streptomycin (Sigma) at 37 °C in 5% CO2. The Nav1.5-WT and various mutation plasmids were transfected into the HEK293 cells by electroporation using ATx from MaxCyte (Gaithersburg, MD) for maximal transfection efficiency according to the manufacturer’s instructions. Cells were dissociated at about 70% confluency and mixed with the target plasmid (200 ng/µl) and transfection buffer (MaxCyte). The electroporated cells were incubated for 20 min at 37 °C in 5% CO2 and then transferred to the maintenance DMEM media for about 48 h until electrophysiological recordings or protein isolation for western blots. Western blot experiments using 40 μg of proteins per sample were performed to reveal protein expression levels using the following primary antibodies: antibody against Nav1.5 (kind gift from Dr. Thomas Hund) and GAPDH antibody (Sigma) was used for loading control. The secondary antibody used was IRDye® 800 CW goat anti-rabbit (Li-COR). Uncropped western blot gels and corresponding raw data are provided in the Source Data file 2.
Whole-cell electrophysiology
Electrophysiological experiments were performed using a high-throughput automated patch-clamp with the SyncroPatch 394i (Nanion, Munich, Germany). Briefly, single-hole, low-resistance recording chips from the same manufacturer were used to record sodium currents. The extracellular solution contained 140 mM NaCl, 4 mM KCl, 2 mM CaCl2, 1 mM MgCl2, 5 mM D-glucose, 10 mM HEPES. The intracellular solution contained: 10 mM HEPES, 10 mM EGTA, 110 mM CsF, 10 mM NaCl, and 10 mM CsCl. Voltage protocols generation, data collection, and data analysis were performed using PatchController384 V.1.3.2 and DataController384 V1.10.1 (Nanion, Munich, Germany).
The current-voltage curves of the sodium currents were elicited by holding the cells to −120mV and stepping from −80mV to +60 mV in 5 mV intervals (each step held for 30 ms). For activation, the G/V curve was obtained by fitting the linear part before the peak of the current-voltage curve with a Boltzmann function:
The time course of inactivation was obtained by fitting the current traces obtained from the current-voltage relationship with a single exponential fitting of each current trace from peak current to the end of the pulse (30 ms) using the following equation:
where A is the amplitude, τ is the time constant, I is the current, and t is the time.
The recovery from inactivation was recorded with a two-pulse protocol. The pre-pulse and the test-pulse duration are 30 ms, stepping from −120 mV to −30 mV. The interval between the two pulses ranges from 1 ms to 250 ms. Currents from the recovery from inactivation were fitted to the following equation:
The steady-state inactivation was studied with a 500 ms pre-pulse ranging from −140 mV to −30 mV, followed by a 30 ms test pulse stepping from −120 mV to −30 mV. The currents for the steady-state inactivation were fitted to a Boltzmann distribution using the following equation:
The late Na current was recorded with a 300 ms pulse ranging from −120 mV to −30 mV. Late current was calculated from the percentage of the current measured at 250 ms to the peak current.
The plots for steady-state inactivation and recovery from inactivation analysis were generated with Origin 10.1.1 software (OriginLab Corporation, Northampton, MA). The leak subtraction protocol was used.
Statistical analysis
Statistical analyses for electrophysiology data were performed using the standard statistical package in Origin 10.1.1 (OriginLab Corporation, Northampton, MA). The student’s t-test was performed at a significance level of p < 0.05 for a single comparison after a normality test with the Shapiro-Wilk method for sample sizes 7–50. Two-sided p-values less than 0.05 were considered statistically significant. Multiple comparisons with the different mutants were performed using one-way ANOVA, and p-values less than 0.05 were considered statistically significant. Results were presented as mean ± SEM. A summary of the electrophysiological parameters for WT and mutant Nav1.5 channels, including activation, steady-state inactivation, and recovery from inactivation, is provided in Supplementary Table 2.
Sample preparation and cryo-EM data collection
A 3.5 µL of full-length hNav1.5 protein was applied to glow-discharged UltrAuFoil 300 mesh, R1.2/1.3 grids (Quantifoil) at 4 °C and 100% humidity. Grids were blotted for 3.5 s and then plunge-frozen in liquid ethane using Vitrobot Mark IV (Thermo Fisher). After initial screening using Glacios (Thermo Fisher Scientific), optimal grids were transferred to a Titan Krios G3i microscope (Thermo Fisher Scientific) at 300 kV equipped with a K3 direct electron detector, a BioQuantum energy filter, and a Cs image corrector. A total of 4967 movies were collected at 81,000x magnification in super-resolution mode (pixel size: 0.4495 Å). The defocus range during collecting was kept from −0.5 to −2.5 μm. The total electron dosage per movie was 60 e−/Å2. For data processing, the summed and dose-weighted micrographs were binned to 0.899 Å/pixel. Data collection statistics are shown in Supplementary Table 1.
Image processing and 3D reconstruction
All raw movies were aligned, drift-corrected and dose-weighted in cryoSPARC v4.4.1 using Patch Motion Correction34, and per-micrograph defocus, and CTF parameters were estimated with Patch CTF. An initial blob picking step generated a set of templates for template picking from a small subset of micrographs. The picked particles were subjected to reference-free 2D classification to yield high-quality templates. After two rounds of iterative 2D classification, a clean set of ~1000 particles was used as a reference for Topaz training and a further two rounds of 2D classification. The extracted particles were cleaned by three rounds of iterative 2D classification. A single ab initio 3D reconstruction was generated from 281,571 particles extracted with a box size of 256 pixels. After one round of homogeneous refinement and non-uniform refinement, a 3D classification was performed to split the particles into four classes. Class 1 (75,907 particles) and Class 2 (65,347 particles) showed features for the transmembrane region as well as density for the EF-hand domain of the CTD. Using these two classes with CTD, we published the role of CTD in the Nav1.5 function11. Class-4 particles are discarded because of junk particles with empty micelles.
Class 3 map showed no CTD, and we focused on this class for further analysis. All subsequent data processing steps for Class 3 were performed in cryoSPARC v4.6.2. Non-uniform refinement, along with separate global and local CTF refinements, was applied to the Class 3 map after 3D classification, yielding a global resolution of 3.7 Å from 100,809 particles. To further improve the map, three decoy classes were generated from Class 4 junk particles, specifically with empty micelles. These decoy classes were used alongside Class 3 in a round of heterogeneous refinement. Particles with clear Nav1.5 features were then re-extracted with a box size of 320 pixels and subjected to another round of non-uniform refinement, improving the global resolution to 3.57 Å with 90,155 particles.
Next, the micelle subtraction was performed by masking the core regions, followed by local refinement using the subtracted particles, resulting in a final global resolution of 3.48 Å. The final 3D map was sharpened in cryoSPARC and followed by DeepEMhancer35. A long α-helical density corresponding to the S6 segment of domain I was observed; however, the local resolution at the intracellular end of the S6 segment was approximately 6 Å. To improve the quality of this region, along with the N-terminal domain, we prepared a local mask for this region by using the Gaussian filtered volume of the local-refined map of Class 3. After local refinement, the density of this region improved substantially. Data collection statistics and image processing summary are shown in Supplementary Table 1, Supplementary Fig. 2.
Model building, refinement, and validation
The model of hNav1.5 was built using the sharpened map, and the structure of our Class 1 map with PDB ID: 8VYJ was used as a template [https://doi.org/10.2210/pdb8VYJ/pdb]. The initial structure was docked into the Class 3 map using the Phenix Dock in Map tool. Subsequently, the CTD and most of the unresolved intracellular regions of the docked model were removed in Coot. Iterative model building was performed using real-space refinement in Phenix and Coot to remove the outliers and improve the model refinement statistics. The final structure, refined using the Class 3 map, has a total of 1258 residues. Additionally, 8 NAG, 1 GDN, and 1 sodium ion were fitted into the density. Figures were prepared with PyMOL (Schrödinger, LLC) and ChimeraX. Final figures were assembled but not created in BioRender (www.BioRender.com). Statistics for cryo-EM model refinement are summarized in Supplementary Table 1.
3D variability analysis (3DVA) of the EM map of hNav1.5
The EM map after local refinement was subjected to 3D variability analysis (3DVA) in cryoSPARC v4.6.234. A local mask was used, which included all four S6 segments and the NTD. A single principal component was used for the analyses. A movie for the component was prepared with UCSF Chimera using frames generated by the 3DVA display program in a simple mode. Followed by another 3DVA display program in cluster mode to separate the frames into two clusters. The initial structure from the consensus refinement of the Class 3 model was docked into the Cluster 1 and Cluster 2 maps using the Phenix Dock in Map tool. All the ligands were removed, and iterative model building was performed using real-space refinement in Phenix and Coot.
MD simulations
Since pore hydration is a prerequisite for conduction in sodium channels and provides36 more information about the functionally open state of the pore compared to the geometric radius alone, we examined the hydration profiles and conductance of Nav1.5 channel pore using classical molecular dynamics (MD) simulations in GROMACS37 at varying transmembrane voltage (TMV). In our MD simulations, higher TMVs were applied solely to accelerate rare permeation events within the limited simulation timescale. We do not interpret absolute conductance from these trajectories. Instead, our analysis is restricted to qualitative features, such as changes in hydration at the activation gate and the relative state-dependence of permeation.
System preparation for MD simulations
Using the Membrane Builder38 from CHARMM-GUI, the structure was embedded in a homogenous lipid bilayer of 1-palmitoyl-2-oleoyl-glycero-3-phosphocholine (POPC) molecules. Water was added to both sides of the membrane, and the system was charge neutralized along with the addition of 150 mM NaCl. The periodic cell dimensions were 13.3 × 13.3 × 14.9 nm and comprised ~250,000 atoms. The Charmm36m all-atom force field39 was used to describe the interactions involving protein, lipid, and ions, while the TIP3P model40 was used for water. Electrostatic interactions were calculated using the particle-mesh Ewald (PME) algorithm41 and the bonds were constrained using the LINCS algorithm42.
Equilibration MD simulations
The energy was minimized using 5000 steps of steepest descent, which was followed by canonical (NVT) equilibration at 300 K using a 2 fs time integration step with harmonic position restraints on the protein and lipid. Then, isothermal-isobaric (NPT) simulations were run using a timestep of 3 fs, Parrinello-Rahman pressure coupling43 at 1 bar, and temperature coupling at 300 K using velocity rescaling with a stochastic term44 for a total MD time duration of 50 ns, such that the protein’s backbone atoms were restrained (5000 kJ mol−1 nm−2) to their initial positions. The equilibrated structure so obtained was further subjected to production MD simulations at varying TMV, as described below, to calculate the hydration profile of the pore.
Production MD simulations
Varying transmembrane voltage (TMV = EzLz) in the form of an electric field (Ez) was applied along the membrane normal (z-direction) such that Lz represents the simulation box length along the electric field direction. To prevent the pore from collapsing to a nonconducting conformation, the Cα positions of the protein were restrained by imposing harmonic potentials with force constants of 1000 kJ mol−1 nm−2. Electric fields of strength: Ez = 0, 0.025, 0.05, and 0.065 V/nm were applied, which correspond to the TMV of around 0, 373, 745, and 969 mV, respectively, in order to translocate the sodium ions from the selectivity filter to the activation gate. Each production MD simulation was run for 500 ns using a timestep of 3 fs. Channel annotation package (CHAP)45 was employed for the hydration profile calculations, such that the initial 100 ns data was discarded, and the averages were computed over the frames extracted every 1.5 ns. The applied voltages are used solely for probing qualitative features and are not representative of physiological conditions.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The cryo-EM maps generated in this study have been deposited in the Electron Microscopy Data Bank (EMDB) under accession code EMD-71158. The corresponding atomic coordinates have been deposited in the Protein Data Bank (PDB) under accession code 9P24. MD simulation files are uploaded to the figshare repository under https://figshare.com/s/0d0cde04c69d289e14cd. All other data supporting the findings of this study are available within the article and its Supplementary Information. Primers sequences are submitted as a separate Source Data file. All the electrophysiology data used to generate the main and Supplementary Figs. are submitted as a separate Source Data file. Source Data is provided as a Source Data file. Source data are provided with this paper.
References
Fozzard, H. A. & Hanck, D. A. Structure and function of voltage-dependent sodium channels: comparison of brain II and cardiac isoforms. Physiol. Rev. 76, 887–926 (1996).
Gamal El-Din, T. M. When the gates swing open only: arrhythmia mutations that target the fast inactivation gate of Na(v)1.5. Cells 11, 3714 (2022).
Goldin, A. L. Mechanisms of sodium channel inactivation. Curr. Opin. Neurobiol. 13, 284–290 (2003).
Ulbricht, W. Sodium channel inactivation: molecular determinants and modulation. Physiol. Rev. 85, 1271–1301 (2005).
Remme, C. A. & Bezzina, C. R. Sodium channel (dys)function and cardiac arrhythmias. Cardiovasc Ther. 28, 287–294 (2010).
Balser, J. R. The cardiac sodium channel: gating function and molecular pharmacology. J. Mol. Cell Cardiol. 33, 599–613 (2001).
Shah, M., Akar, F. G. & Tomaselli, G. F. Molecular basis of arrhythmias. Circulation 112, 2517–2529 (2005).
Jiang, D. et al. Structure of the cardiac sodium channel. Cell 180, 122–134 e10 (2020).
Wisedchaisri, G. et al. Resting-state structure and gating mechanism of a voltage-gated sodium channel. Cell 178, 993–1003 e12 (2019).
Jiang, D. et al. Open-state structure and pore gating mechanism of the cardiac sodium channel. Cell 184, 5151–5162 e11 (2021).
Biswas, R. et al. Structural basis of human Na(v)1.5 gating mechanisms. Proc. Natl. Acad. Sci. USA. 122, e2416181122 (2025).
West, J. W. et al. A cluster of hydrophobic amino acid residues required for fast Na(+)-channel inactivation. Proc. Natl. Acad. Sci. USA. 89, 10910–10914 (1992).
Pan, X. et al. Structure of the human voltage-gated sodium channel Na(v)1.4 in complex with beta1. Science 362, eaau2486 (2018).
Liu, Y., Bassetto, C. A. Z. Jr., Pinto, B. I. & Bezanilla, F. A mechanistic reinterpretation of fast inactivation in voltage-gated Na(+) channels. Nat. Commun. 14, 5072 (2023).
Kellenberger, S., Scheuer, T. & Catterall, W. A. Movement of the Na+ channel inactivation gate during inactivation. J. Biol. Chem. 271, 30971–30979 (1996).
Kellenberger, S., West, J. W., Catterall, W. A. & Scheuer, T. Molecular analysis of potential hinge residues in the inactivation gate of brain type IIA Na+ channels. J. Gen. Physiol. 109, 607–617 (1997).
Karlin, A. & Akabas, M. H. Substituted-cysteine accessibility method. Methods Enzymol. 293, 123–145 (1998).
Tomaselli, G. F. Cysteine mutagenesis in the voltage-dependent sodium channel structural insights and implications. Trends Cardiovasc. Med. 7, 211–218 (1997).
Kellenberger, S., West, J. W., Scheuer, T. & Catterall, W. A. Molecular analysis of the putative inactivation particle in the inactivation gate of brain type IIA Na+ channels. J. Gen. Physiol. 109, 589–605 (1997).
Tonggu, L. et al. Dual receptor-sites reveal the structural basis for hyperactivation of sodium channels by poison-dart toxin batrachotoxin. Nat. Commun. 15, 2306 (2024).
Punjani, A. & Fleet, D. J. 3D variability analysis: resolving continuous flexibility and discrete heterogeneity from single particle cryo-EM. J. Struct. Biol. 213, 107702 (2021).
Li, Z. et al. Structure of human Na(v)1.5 reveals the fast inactivation-related segments as a mutational hotspot for the long QT syndrome. Proc. Natl. Acad. Sci. USA. 118, e2100069118 (2021).
Huang, G. et al. High-resolution structures of human Na(v)1.7 reveal gating modulation through alpha-pi helical transition of S6(IV). Cell Rep. 39, 110735 (2022).
Deschenes, I., Trottier, E. & Chahine, M. Cysteine scanning analysis of the IFM cluster in the inactivation gate of a human heart sodium channel. Cardiovasc Res. 42, 521–529 (1999).
Li, Z. et al. Structural basis for pore blockade of the human cardiac sodium channel Na(v) 1.5 by the antiarrhythmic drug quinidine*. Angew. Chem. Int Ed. Engl. 60, 11474–11480 (2021).
Su, Q. et al. Cryo-EM structure of the polycystic kidney disease-like channel PKD2L1. Nat. Commun. 9, 1192 (2018).
Zhao, Y. et al. Molecular basis for ligand modulation of a mammalian voltage-gated Ca(2+) channel. Cell 177, 1495–1506 e12 (2019).
Shen, H., Liu, D., Wu, K., Lei, J. & Yan, N. Structures of human Na(v)1.7 channel in complex with auxiliary subunits and animal toxins. Science 363, 1303–1308 (2019).
Wu, Q. et al. Structural mapping of Na(v)1.7 antagonists. Nat. Commun. 14, 3224 (2023).
Liu, Y. & Bezanilla, F. A sodium channel mutant removes fast inactivation with the inactivation particle bound. J. Gen. Physiol. 157, e202413667 (2025).
Yarov-Yarovoy, V. et al. Role of amino acid residues in transmembrane segments IS6 and IIS6 of the Na+ channel alpha subunit in voltage-dependent gating and drug block. J. Biol. Chem. 277, 35393–35401 (2002).
Wang, S. Y., Bonner, K., Russell, C. & Wang, G. K. Tryptophan scanning of D1S6 and D4S6 C-termini in voltage-gated sodium channels. Biophys. J. 85, 911–920 (2003).
Kuo, C. C. & Liao, S. Y. Facilitation of recovery from inactivation by external Na+ and location of the activation gate in neuronal Na+ channels. J. Neurosci. 20, 5639–5646 (2000).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput Chem. 29, 1859–1865 (2008).
Bauer, P., Hess, B. & Lindahl, E. GROMACS 2022 manual. https://doi.org/10.5281/zenodo.6103568 (2022).
Lee, J. et al. CHARMM-GUI membrane builder for complex biological membrane simulations with glycolipids and lipoglycans. J. Chem. Theory Comput 15, 775–786 (2019).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: A linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Klesse, G., Rao, S., Sansom, M. S. P. & Tucker, S. J. CHAP: a versatile tool for the structural and functional annotation of ion channel pores. J. Mol. Biol. 431, 3353–3365 (2019).
Acknowledgments
Electron microscopy data were acquired at the Center for Electron Microscopy and Analysis at The Ohio State University. We thank the Ohio Supercomputer Center for high-performance computing resources. This work was supported by National Institutes of Health grant R01HL094450 (I.D.) and the Frick Center for Heart Failure via a Synergy Award from the Dorothy M. Davis Heart and Lung Research Institute at The Ohio State University Wexner Medical Center (I.D.) and start-up funds from The Ohio State University College of Medicine (S.M.H., I.D., and K.C.).
Author information
Authors and Affiliations
Contributions
Conceptualization: K.C.; Supervision: I.D., and K.C.; Resources/analytic tools: X.C., S.M.H., I.D., and K.C.; Investigation: R.B., A.L.L.-S., A.P., A.R.-N., and H.-L.H.; Data Curation: R.B., A.L.L.-S., A.P., X.C., I.D., and K.C.; Formal Analysis: R.B., A.L.L.-S., and A.P.; Writing: Original draft R.B., A.P., I.D., and K.C.; Writing: Review & Editing: R.B., A.L.L.-S., A.P., X.C., S.M.H., I.D., and K.C.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Daohua Jiang and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Biswas, R., López-Serrano, A.L., Purohit, A. et al. Structural and functional mechanisms underlying activation gate dynamics and IFM motif accessibility in human Nav1.5. Nat Commun 17, 2820 (2026). https://doi.org/10.1038/s41467-026-69672-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69672-x