Abstract
Directed evolution has transformed biomolecular engineering but remains largely untapped for probiotic optimization, despite its immense promise for human health maintenance and disease therapy. Here, we present an in vivo, host-mediated directed-evolution platform that harnesses the gut’s endogenous selective pressures to drive functional enhancement of probiotics. Using Bifidobacterium animalis subsp. lactis as a model, we expose germ-free male mice to stepwise increases in bile-acid stress via a high-fat, high-cholesterol diet. Compared to in vitro evolution, which fails to produce any functional gains, our host-mediated approach yields a variant exhibiting a 77% increase in bile acid metabolism. Multi-omics analysis identifies two critical single-nucleotide variants (SNVs) simultaneously: one in the upstream region of cbh, encoding bile salt hydrolase, and a non-synonymous mutation in mdr, a bile-acid efflux transporter. Functional validation assays confirm that these mutations drive increased corresponding gene expression and enhance substrate binding efficiency. Moreover, to demonstrate its translational potential, we administer the parental and adapted strains daily in a male diet-induced mouse model of non-alcoholic fatty liver disease (NAFLD). We find that the adapted strain significantly improves bile-acid homeostasis, reduces hepatic steatosis, lowers inflammatory and lipid biomarkers, and enhances histological liver health compared to the parental strain. Our work establishes the host gut as a living evolutionary bioreactor for precision engineering of probiotics, enabling targeted phenotypic enhancement in vivo through natural selection. This framework paves the way for personalized, functionally tailored microbiome therapeutics and sets a foundation for next-generation live biotherapeutic agents.
Similar content being viewed by others
Introduction
Directed evolution is an iterative process of mutagenesis and phenotypic screening that enables efficient optimization of biomolecular functions for specific traits. It harnesses evolutionary principles by introducing genetic diversity under selective pressure, driving the emergence of elite mutants through progressive screening cycles1. Described by the Nobel Committee as harnessing evolution for the greatest benefit to humankind, this methodology has found wide transformative applications in enzyme engineering, antibody development, and metabolic pathway refinement2,3,4. Currently, the most widely used strategies for generating targeted microbial adaptation include two major strategies: (1) induction-based methods, such as adaptive laboratory evolution (ALE)5,6,7; and (2) gene-editing-based methods, such as CRISPR-Cas9-based genome editing7,8. For example, short-term ALE using phenolic acids (such as ferulic acid, p-hydroxybenzoic acid, and vanillic acid) as model inhibitors yielded Saccharomyces cerevisiae strains with 2- to 3-fold increased growth rates and enhanced tolerance to lignin-derived phenolics9. Similarly, as an example of gene editing, CRISPR-Cas9-mediated modification of repetitive elements in the phytosterol biosynthesis pathway of Solanaceae plants enabled a fivefold increase in 7-dehydrocholesterol (7-DHC), a vitamin D₃ precursor, in leaves and immature fruits without impairing plant development10.
Probiotics have garnered considerable attention in recent years due to their multifaceted roles in maintaining gut homeostasis11, regulating metabolic functions12, and reinforcing immunological barriers13. Nevertheless, most studies to date concentrate on screening native strains14 and characterizing their phenotypic functions15 or probiotic properties16, while systematic optimization and directed evolution of their functional potential remain largely unexplored. A few reports have applied ALE in vitro, exposing strains to prolonged passaging under stressors such as acidity17, osmotic pressure18, or antibiotics19 to select for mutants with improved survival or functional traits. Likewise, gene-editing approaches (e.g., CRISPR-Cas) have been used to reconfigure metabolic pathways to enhance gut colonization20, target specific functions21, or boost metabolite production22. However, these in vitro strategies rely on low-throughput screening pipelines that limit scalability in probiotic engineering23. Moreover, strains evolved under laboratory conditions frequently fail to achieve ecological adaptation and functional remodeling when reintroduced into authentic host-gut environments.
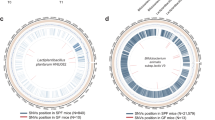
The host intestinal environment serves as a natural evolutionary arena that drives the accumulation of adaptive mutations in gut bacteria24,25. Studies demonstrate that gut microbiota acquire single-nucleotide variants (SNVs) at rates orders of magnitude higher in vivo than in vitro (reaching billions in scale) due to intestinal selection pressures26. For instance, genomic analysis of Lactiplantibacillus plantarum HNU082 isolates after 28-day intestinal colonization in model organisms (human, zebrafish, and mouse) revealed a significant enrichment of SNPs associated with acid resistance, membrane protein synthesis, and carbon source metabolism. In contrast, serially passaged in vitro counterparts exhibited no detectable mutations25. This divergence underscores the unique evolutionary pressures exerted by host intestinal microenvironments. Likewise, a parallel comparison showed that Lactiplantibacillus plantarum P‑8 passaged in vitro (glucose‑limited or standard media) exhibited no genetic changes, while isolates recovered from human feces (in vivo) carried multiple SNPs, including mutations in mucus‑adhesion protein and pyruvate oxidase genes27. Although these findings demonstrate that host-imposed intestinal selection pressures can drive functional optimization of probiotics, most studies remain observational and lack phenotype-driven directed-evolution strategies. We therefore seek to investigate whether the host gut itself could serve as a bioreactor for the directed evolution of probiotic strains.
To validate the efficacy of leveraging the host intestinal microenvironment as a bioreactor for directed evolution, we first select Bifidobacterium animalis subsp. lactis H22B656, a probiotic strain with intrinsic bile acid-metabolizing capacity, as our model. We then implement two induction strategies: (i) an in vivo approach, in which germ‑free mice are mono‑colonized with H22B656 and maintained on a high‑fat, high‑cholesterol diet (studies have shown that dietary fat intake correlates positively with bile acid secretion and expands the bile acid pool, thus imposing increased bile acid-metabolic pressure on the gut28,29); and (ii) an in vitro approach, consisting of serial passaging in conventional liquid medium supplemented with bile acids. Although the in vitro strategy fails to produce any beneficial phenotypic shift, the in vivo, host‑mediated directed evolution yields a strain with a 77% increase in bile acid-metabolizing capacity. Subsequently, we use whole‑genome sequencing (WGS) and transcriptomic profiling to reconstruct the evolutionary trajectory of the adapted strain, pinpointing SNVs underlying enhanced bile acid metabolism. We functionally validate two key SNVs (rs1846941 and rs402499) to demonstrate their causal role in modulating bile acid-metabolizing capacity, thereby elucidating the molecular mechanisms of in vivo adaptation. Finally, to evaluate translational potential, we employ a diet‑induced nonalcoholic fatty liver disease (NAFLD) mouse model and administer the adapted strain as a dietary intervention, demonstrating its metabolic benefits and affirming its biosafety in vivo. Our study not only introduces an in vivo, host‑mediated directed evolution strategy for probiotic optimization but also establishes an approach for targeted interventions against metabolic diseases such as NAFLD, laying a critical foundation for the future clinical translation of disease‑tailored probiotic therapies.
Results
Screening of native strains with bile acid-metabolizing capacity
To evaluate the efficacy of the in vitro and in vivo induction strategies for directed probiotic adaptation, we first sought to identify a strain with bile acid-metabolizing capacity as a model. A panel of 12 B. animalis subsp. lactis strains were randomly selected from our library and systematically assessed across four dimensions: (1) a calcium precipitation assay, (2) specific enzyme activity quantification, (3) stress tolerance testing, and (4) growth curve analysis (Fig. 1a; “Methods”). Specifically, to assess bile salt hydrolase (BSH) activity, we plated strains on bile acid-supplemented agar and measured the diameter of the calcium precipitation zone after 48 h. Strains H22B484, H22B525, and H22B656 produced the largest zones (averaging approximately 2.7 cm), indicating robust enzymatic activity (Fig. 1b and Supplementary Fig. 1a). Next, we quantified BSH-specific activity using a ninhydrin-based colorimetric assay. H22B656 exhibited the highest activity (0.4561 U/mg), followed by H22B484 (0.3570 U/mg) and H22B493 (0.3392 U/mg) (Fig. 1c). For stress tolerance assessment, candidate strains were evaluated under both acidic and bile acid conditions to reflect gut survival potential. Under acidic conditions (pH 2.0 for 2 h), seven strains (H22B748, H22B938, H22B801, H22B525, H22B391, H22B648, and H22B656) maintained viability above 25%. When exposed to bile acids (0.3% w/v for 4 h), H22B656 again exhibited the highest survival rate (83.49%), followed by H22B723 (69.50%) and H22B525 (64.67%) (Fig. 1d). Finally, we measured growth kinetics by tracking OD₆₀₀ every 4 h over 36 h. H22B391 demonstrated the highest biomass accumulation (OD₆₀₀=0.5521 at 24 h), followed by H22B656 (0.5031) and H22B748 (0.4778) (Fig. 1e).
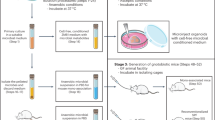
a The experimental design comprises three successive phases: In vitro Screening: Twelve infant-derived strains were initially evaluated based on four criteria: dissociation ability, enzyme activity, tolerance, and growth kinetics. This process identified Bifidobacterium animalis subsp. lactis H22B656 as the original candidate strain. Host-mediated In vivo Adaptation: GF mice served as an evolution bioreactor. Following colonization with H22B656, mice were subjected to increasing dietary fat content (30% to 60%) over 7 weeks. Sixty colonies were isolated from feces at different time points. Based on the specific activity ratio, the adapted strain B. animalis subsp. lactis W5S9 was selected. NAFLD Efficacy Validation: The therapeutic potential was evaluated in SPF mice under a high-fat diet (HFAD). Mice were divided into four groups (Control, HFAD, HFAD + H22B656, and HFAD + W5S9; n = 6 mice per group) for an 8-week intervention. Efficacy was assessed via biochemistry, histology, targeted bile acid (BA) metabolomics, and metagenomics to compare the metabolic regulation capabilities of the original and adapted strains. b Calcium precipitation assay. Twelve candidate strains were cultured anaerobically on MRS-cys agar containing 0.3% (w/v) conjugated bile acids for 48 h. The bar plot shows the mean diameter of the calcium-precipitation halo for each strain. c Specific BSH activity measured by a ninhydrin-based colorimetric assay using glycine-conjugated bile acids as substrate. d Heatmaps of survival rates under acid (pH 2.0, 2 h) and bile acid (0.3% (w/v), 4 h) stress, calculated as OD₆₀₀₍treatment₎/OD₆₀₀₍control₎. Data are from n = 3 biologically independent experiments. e Growth kinetics. Strains were inoculated at 2% (v/v) into MRS-cys broth and incubated anaerobically at 37 °C for 36 h, with OD₆₀₀ recorded every 4 h. For all in vitro experiments (b–e), data are derived from n = 3 biologically independent bacterial cultures. Data in (b), (c), and (e) are presented as mean ± SD. Panels (a–e) were created with BioRender (Zhang, J. (2026) https://BioRender.com/3qyx3m3). Source data are provided as a Source Data file.
To comprehensively evaluate probiotic potential in bile acid metabolism, we integrated these five key indicators from four dimensions and conducted a percentile-based ranking using a multidimensional linear summation score for each strain. The resulting composite scores were visualized as radar plots (Supplementary Fig. 1b). Among the candidates, H22B656 achieved the highest composite score as evidenced by the largest radar plot area (425), followed by H22B525 (375) and H22B391 (366.6667) (Supplementary Fig. 1c). Based on this integrative assessment, H22B656 was selected for subsequent adaptive evolution experiments.
The in vivo induction markedly enhanced the bile acid-metabolizing capacity
To enhance the bile acid-metabolizing capacity of B. animalis subsp. lactis H22B656, we employed both in vitro and in vivo induction strategies and systematically evaluated their efficacy in driving targeted adaptive evolution. Using an in vitro induction protocol, H22B656 was serially passaged under 0.3% (w/v) conjugated bile acid pressure for ten days, with transfers every 24 h into fresh, modified MRS‑cys medium containing the same bile acid concentration (“Methods”). On day 10, we isolated 60 clones from the induced population and measured their BSH-specific activities (Fig. 2a). Under 0.3% bile acid stress, none of the induced clones exhibited appreciable increases in BSH activity compared to the original strain, with the average specific activity of induced clones remaining ≤1‑fold relative to the original (Fig. 2b).
a In vitro induction workflow. B. animalis subsp. lactis H22B656 was serially passaged ten times in MRS-cys medium supplemented with 0.3% (w/v) conjugated bile acids. After the tenth passage, 60 single colonies were randomly isolated and assayed for BSH-specific activity. b Fold-change distribution of BSH specific activity for the n = 60 biologically independent in vitro induced clones relative to the parental strain. The green dashed line at 1.0 indicates baseline activity; gray points represent clones with unchanged or reduced activity. Data for each isolate are presented as a box plot with overlaid individual data points. Box plot elements represent the median (centre line), the 25th and 75th percentiles (bounds of the box), and the minimum to maximum range. For each isolate, the box plot represents n = 3 biologically independent samples. c In vivo induction workflow. GF C57BL/6J mice (n = 4) were mono-colonized by oral gavage with H22B656 (1 × 10⁹ CFU/day for three days) and then maintained on stepwise high-fat, high-cholesterol diets (30%, 45%, and 60% fat + 0.2% cholesterol) over seven weeks. Fecal samples were collected at weeks 1, 3, 5, and 7; at each time point, 60 colonies were randomly isolated and assayed for BSH specific activity. d Fold-change distribution of BSH specific activity for the 60 in vivo induced isolates. Data for each isolate are presented as a box plot with overlaid individual data points. Box plot elements are defined as in (b). For each isolate, the box plot is derived from n = 3 biologically independent bacterial cultures. Panels (a) and (c) were created with BioRender (Zhang, J. (2026) https://BioRender.com/3qyx3m3). Source data are provided as a Source Data file.
The limited effectiveness of the in vitro induction likely arises from its inability to replicate the complex selective pressures of the gut, particularly the immunological interactions between host and microbiota that drive adaptive evolution25. To address this, we explored an in vivo, host‑mediated induction strategy to harness selective forces at the host-microbiota interface and promote specific microbial functions (“Methods”). Using germ‑free C57BL/6J mice as a bioreactor to eliminate interference from resident microbes, we imposed a stepwise high‑fat diet to progressively intensify bile acid metabolic stress. Over seven weeks, B. animalis subsp. lactis H22B656 colonized mice (200 µL of 109 CFU administered by oral gavage once daily for three consecutive days before diet intervention) received: (1) 30% fat + 0.2% cholesterol for weeks 1–3; (2) 45% fat + 0.2% cholesterol for weeks 3–5; and (3) 60% fat + 0.2% cholesterol for weeks 5–7 (Fig. 2c). To assess the in vivo induction’s efficacy, we also isolated 60 clones from fecal samples collected at different time points and measured their BSH specific activity. Fifteen clones exhibited a significant increase in BSH activity compared to the original (fold change > 1.35), yielding a directed induction success rate of 25% (Fig. 2d). Among these, strain W5S9 exhibited the highest specific activity (fold change = 1.77).
Compared with the in vitro approach, the in vivo strategy imposed a substantially greater selective impact on bile acid metabolism, as reflected by a markedly higher variance in BSH activity (SD = 0.3440 in vivo versus 0.0784 in vitro). This suggests that host‑derived pressures can dramatically reprogram microbial metabolic traits (both positively and negatively), generating a diverse pool of adapted candidates. These results confirm the feasibility and utility of in vivo induction as a powerful approach to optimize probiotics’ beneficial effects.
Genetic adaptations in bile acid-metabolism via in vivo induction
We hypothesized that the enhanced bile acid-metabolizing capacity of strain W5S9 resulted from the accumulation of genetic variants that altered key corresponding metabolic pathways. To understand these molecular mechanisms, we conducted whole‑genome resequencing (WGS) and transcriptomic profiling (RNA‑seq) of W5S9. These analyses aimed to systematically map the single-nucleotide variants (SNVs) acquired during in vivo induction and to assess their impact on gene expression programs governing bile acid metabolism (“Methods”).
To improve the accuracy of mutation identification, we applied two independent computational pipelines (inStrain30 and BCFtools31) and used the intersection of their outputs to define a high-confidence SNV set. This approach identified 282 SNVs, which were then annotated via inStrain (Fig. 3a; Supplementary Fig. 1d; Supplementary Data 1; “Methods”). Based on genomic location, we divided these SNVs into coding and noncoding categories. Of the 244 coding-region SNVs, 39 were synonymous (affecting 38 genes), and 205 were nonsynonymous (spread across 164 genes). As for the remaining 38 SNVs that fell within noncoding regions, promoter-prediction analysis indicated that 11 lie in putative regulatory elements, potentially influencing transcription of nine downstream genes (“Methods”). In addition, given the close genomic linkage, we suspect that these 38 noncoding SNVs may collectively affect the expression of up to 32 downstream genes32 (Fig. 3b).
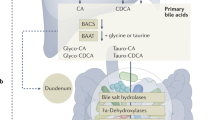
a Circos plot of 282 high-confidence SNVs in W5S9 relative to H22B656. From outer to inner rings: chromosomal coordinates (kb), GC content, sequencing depth, and SNV density (darker shading = higher local density). Key mutations in cbh and mdr are highlighted. b Genomic distribution of SNVs. Of 282 total, 244 map to coding regions, comprising 205 nonsynonymous (N, 164 genes) and 39 synonymous (S, 38 genes) changes. The remaining 38 SNVs lie in noncoding regions. Promoter-prediction (and genome linkage) analysis suggests these noncoding SNVs may regulate nine (and 32 additional downstream) genes. c, d Volcano plots of differential gene expression (RNA-seq, n = 3 independent biological replicates per strain, analyzed in DESeq2 using the two-sided Wald test with Benjamini–Hochberg multiple-testing correction) comparing W5S9 versus H22B656. Significantly upregulated and downregulated genes meet |log₂FC | ≥ 0.4 and FDR < 0.001. e Conjugated primary bile acids are taken up by bacteria and undergo deconjugation by bile salt hydrolase (cbh), releasing taurine or glycine and generating free cholic acid (CA) and chenodeoxycholic acid (CDCA). Free primary bile acids may passively diffuse across the cell membrane or be extruded by efflux transporters (e.g., mdr, lmrA). Within bacterial cells, oxidoreductive reactions mediated by 7α-hydroxysteroid dehydrogenase (hdhA) and 7β-hydroxysteroid dehydrogenase (7β-HSDH) contribute to the epimerization of CA and CDCA, generating derivatives such as UCA and UDCA. Subsequently, multi-step 7α-dehydroxylation catalyzed by the bai gene cluster (baiA/baiB/baiCD/baiE) transforms these intermediates into the secondary bile acids deoxycholic acid (DCA) and lithocholic acid (LCA). Transporters such as BaiG and BaiF mediate the export of secondary bile acids into the extracellular space. f, g RNA-seq–derived counts per million (CPM) for two key upregulated genes: (f) cbh (log₂FC = 0.4802, FDR < 0.001) and (g) mdr (log₂FC = 0.5355, FDR < 0.001). Bars show mean ± SEM derived from n = 3 biologically independent samples per strain. Statistical significance was determined by DESeq2 using the two-sided Wald test with Benjamini–Hochberg correction. Panels (b) and (e) were created with BioRender (Zhang, J. (2026) https://BioRender.com/3qyx3m3). Source data are provided as a Source Data file.
To explore the phenotypic and functional impact of these mutations, we conducted differential transcriptomic analysis between three adapted clones and three original strains (FDR < 0.001 and |log₂FC|≥0.4 were defined as significant). Among the 164 protein-coding genes harboring non‑synonymous SNVs, 16 were significantly upregulated, and 32 were significantly downregulated (Fig. 3c). Of the 41 putative target genes associated with non‑coding SNVs, six exhibited significant upregulation, and five showed significant downregulation (Fig. 3d).
We next sought to determine whether these differentially expressed genes are involved in bile acid metabolism and transport, and thus contribute to the observed phenotypic changes. The bacterial bile acid detoxification pathway is now well-characterized as a coordinated process of stepwise modification and efflux. First, intracellular bile salt hydrolase (cbh) catalyzes the deconjugation of conjugated bile acids (e.g., glycocholic acid) into free bile acids and amino acids. The liberated bile acids can then be eliminated by one of two routes: (1) direct efflux via Major Facilitator Superfamily (MFS) efflux pumps (e.g., mdr, lmrB), which actively export bile acids across the cell membrane33; or (2) deeper modification followed by efflux, in which hydroxysteroid dehydrogenase (hdhA, 7β-HSDH) mediates oxidoreductive modification of free bile acids, and 7α-dehydroxylase enzymes encoded by the bai gene cluster subsequently catalyze their conversion into secondary bile acids. These secondary bile acids are subsequently extruded by the BaiF/BaiG transport system, thereby mitigating bile acid-induced membrane permeability and cytotoxicity34(Fig. 3e). In total, fifteen key genes coordinate this detoxification cascade.
By cross-referencing the 59 (48 associated with coding SNVs and 11 associated with non-coding SNVs) differentially expressed genes with those involved in bile acid metabolism (n = 15), we identified two key candidates that were significantly upregulated in the adapted strain. The first, cbh (K01442, EC 3.5.1.24), encodes bile salt hydrolase, which catalyzes the hydrolysis of conjugated bile acids into free bile acids and amino acids (log₂ FC = 0.4802, FDR < 0.001; Fig. 3f). This gene is associated with a non-coding SNV (rs1846941), suggesting a potential regulatory variant affecting its expression. The second gene mdr (via NCBI BLASTp) was also significantly upregulated (log₂FC = 0.5355, FDR < 0.001; Fig. 3g). Mdr encodes an MFS multidrug resistance (MDR) efflux pump responsible for exporting free bile acids. We found that a non-synonymous SNV (rs402499) within its coding sequence may alter transporter function. Together, integration of genomic and transcriptomic data from adapted and original strains potentially highlights these two genes, both genetically and transcriptionally remodeled, as drivers of enhanced bile acid metabolism during host-mediated in vivo induction strategy.
Validation of bile acid metabolic enhancement mediated by two key SNVs
To test whether the two key SNVs identified in the adapted strain indeed contribute to enhancing bile acid metabolism and associated phenotypic improvements, we performed systematic validation. Firstly, given the potential for false positives in WGS, we confirmed the presence of the two SNVs using an additional Sanger sequencing. Compared to the parental genome, we observed: (1) a C-to-A substitution at position 1,846,941 in the upstream non-coding region (likely the promoter) of cbh; and (2) an A-to-G substitution at position 402,499 in the coding region of mdr, resulting in a missense mutation that changed the 192nd amino acid of the mdr protein from asparagine (Asn) to aspartic acid (Asp, N192D). These results fully agree with our WGS findings (Fig. 4a, b; “Methods”).
a, b Sanger sequencing confirmed the presence of the two critical SNVs in the adapted strain W5S9. c Comparison of the mCherry fluorescence intensity of reporter plasmids driven by wild-type and rs1846941 mutation under DCA, CA, and CDCA conditions. Bars show mean ± SEM (n = 5 independent samples). Statistical significance was determined using multiple two-sided Mann–Whitney tests with the two-stage step-up method (Benjamini, Krieger, and Yekutieli) to control the False Discovery Rate. d, f Relative expression levels were determined by RT-qPCR in the presence of DCA, CA, and CDCA. Data were normalized to the wild-type Bifidobacterium animalis subsp. lactis H22B656 (set to 1) independently for each condition. Bars represent the mean ± SD of biological replicates (n = 3), with circles indicating individual data points. Statistical comparisons were performed using one-way ANOVA, followed by Tukey’s honest significant difference (HSD) post hoc test to correct for multiple comparisons. e BSH specific activity measured by a ninhydrin colorimetric assay using conjugated bile acids as substrate. Bars represent mean ± SD (n = 3 biologically independent samples). Statistical significance was evaluated using one-way ANOVA with Tukey’s HSD post-hoc test for multiple comparisons. g Log10 CFU of various strains grown under DCA, CA, and CDCA treatments. Bars represent mean ± SD (n = 3 biologically independent samples). Statistical significance was determined using one-way ANOVA with Tukey’s HSD post-hoc test for multiple comparisons. h Molecular docking-predicted binding affinities (kcal/mol) of WT and N192D MDR proteins with DCA, CA, and CDCA. Lower energy indicates stronger binding. Data are presented as mean ± SEM (n = 10 independent docking runs). i Structural and docking models of WT and N192D MDR proteins with DCA, CA, and CDCA. Left panels show the AlphaFold3-predicted 3D structures of the WT and N192D variants of MDR. Right panels depict the highest-scoring molecular docking poses with DCA, CA, and CDCA, with hydrogen bonds highlighted by dashed lines and red arrows. Statistical significance for docking energies in (h) was assessed using multiple two-sided Mann–Whitney tests with the two-stage step-up method (Benjamini, Krieger, and Yekutieli) to control the False Discovery Rate. Source data are provided as a Source Data file.
We next sought to confirm that the upregulation of cbh in the adapted strain was attributable to the non-coding mutation. To this end, we constructed transcriptional reporters in which either the wild-type or the rs1846941-harboring upstream sequence was fused to an mCherry cassette and introduced these plasmids into the parental B. animalis subsp. lactis H22B656. Under three different bile-salt conditions (0.3% deoxycholic acid, DCA; cholic acid, CA; and chenodeoxycholic acid, CDCA), all strains carrying the rs1846941 variant exhibited significantly higher mCherry fluorescence than those with the wild-type sequence, indicating that this non-coding mutation enhances cbh transcription during bile acid stress (fold changes of 1.30, 1.71, and 12.36 for DCA, CA, and CDCA, respectively; Multiple Mann-Whitney tests with FDR correction, q < 0.01; Fig. 4c).
To directly verify its regulatory effect within the genomic context, we knocked out the cbh promoter region containing rs1846941 and measured cbh expression levels under three different bile salt conditions (0.3% DCA, CA, and CDCA). The results showed that under these three different bile salt conditions, the cbh expression in the wild-type adapted strain was significantly higher than in the parental wild-type strain, suggesting that this non-coding mutation enhances cbh transcription under bile acid stress (fold changes of 1.65, 1.51, and 1.49 for DCA, CA, and CDCA, respectively; one-way ANOVA with Tukey’s HSD test, p < 0.01; Fig. 4d). Importantly, after knockout of the promoter sequence, cbh expression was significantly reduced, further confirming that the upregulation of cbh in the adapted strain is indeed driven by this non-coding mutation. Moreover, assessment of bile salt hydrolysis using the ninhydrin colorimetric assay revealed that the adapted strain exhibited significantly higher hydrolytic activity toward glycine-conjugated bile salts compared to the parental strain (fold change = 1.25; one-way ANOVA with Tukey’s HSD test, p < 0.01; Fig. 4e). For taurine-conjugated bile salts, hydrolytic activity was also higher in the adapted strain (fold change = 1.16). Notably, knockout of the promoter region significantly reduced hydrolytic activity toward conjugated bile salts in both strains. Together, these findings suggest that the promoter region containing rs1846941 regulates the cbh gene expression and plays a critical role in enhancing the hydrolysis of conjugated bile acids.
For rs402499, we similarly performed a knockout of the mdr gene, which carries the mutation, and assessed mdr expression under three different bile salt conditions (0.3% DCA, CA, and CDCA) to determine whether the mutation directly affects gene expression. Under these three bile acid treatment conditions, the expression of mdr in the adapted strain was significantly higher compared to the parental wild-type strain, suggesting that this mutation enhances mdr transcription in response to bile acid stress (fold changes of 1.64, 2.13, and 1.73 for DCA, CA, and CDCA, respectively; one-way ANOVA with Tukey’s HSD post-hoc test, p < 0.01; Fig. 4f). Further analysis revealed that knockout of the mdr gene resulted in a significant reduction in mdr expression, confirming that the mutation directly impacts the phenotype through the regulation of mdr expression. Consistently, heterologous expression assays confirmed that the N192D mutation enhances mdr expression at translational levels under bile salt stress (Supplementary Fig. 2). Additionally, we performed complementation experiments to assess whether the restoration of mdr expression could recover the phenotype. Under CA, DCA, and CDCA treatment conditions, the complemented adapted strain exhibited significantly higher mdr expression than the complemented wild-type strain (One-way ANOVA with Tukey’s HSD post-hoc test, p < 0.01; Fig. 4f). These findings suggest that the rs402499 (N192D) mutation drives adaptive phenotypes by increasing mdr expression under bile acid stress.
MDR, as a transporter protein, facilitates the efflux of free bile acids, thereby mitigating bile acid toxicity and improving bacterial survival35. To further validate the causal relationship between mdr gene expression and the phenotype, we assessed the viable bacterial counts under bile acid stress conditions. The results showed that the colony counts of the adapted wild-type strain were significantly higher than those of the parental wild-type strain under all three bile salt conditions (One-way ANOVA with Tukey’s HSD post-hoc test, p < 0.01; Fig. 4g). Notably, under CA and CDCA treatment, the mdr knockout strains exhibited a marked growth disadvantage, whereas the differences between knockout strains were less pronounced under DCA conditions (p = 0.5499). In complementation experiments, the complemented adapted strain showed significantly higher colony counts than the complemented wild-type strain, further supporting the functional role of mdr gene expression in bile acid stress. Together, the N192D substitution significantly impacted both the transcriptional and translational levels of the mdr gene under bile stress, conferring a robust expression advantage and increasing the strain’s survival rate.
Since rs402499 encodes a non-synonymous N192D substitution, we hypothesized that this change might also alter the protein’s structural conformation in a way that improves its ability to export free bile acids. To test this hypothesis, we conducted comparative structural modeling and molecular docking simulations for both the wild-type and N192D variant of the protein to assess their binding affinity for free bile acids (“Methods”). The simulations revealed that the mutant protein exhibited significantly lower binding free energy with all three substrates (DCA, CA, and CDCA) and formed a greater number of hydrogen bonds (two-tailed Mann-Whitney test with FDR-corrected, q < 0.001; Fig. 4h, i). These results suggest that the N192D mutation stabilizes the protein-substrate complex, thereby potentially facilitating more efficient substrate recognition and transmembrane transport.
Adapted strain outperforms the original strain in NAFLD prevention
To assess the beneficial potential of B. animalis subsp. lactis W5S9 in modulating host metabolism, we employed a diet-induced nonalcoholic fatty liver disease (NAFLD) mouse model, grounded in the established link between bile acid dysregulation and NAFLD progression36. We compared the adapted strain (W5S9) with its original strain (H22B656) in safety, bile acid metabolism, and preventive efficacy (Fig. 5a). Specific pathogen-free (SPF) C57BL/6J mice (n = 6 per group) were randomized into four cohorts: (1) Control (normal diet, “Methods”); (2) HFAD (normal diet + 60% fat); (3) HFAD-H22B656 (HFAD with daily oral gavage of 10⁹ CFU B. animalis subsp. lactis H22B656); and (4) HFAD-W5S9 (HFAD with daily oral gavage of 10⁹ CFU B. animalis subsp. lactis W5S9). Probiotics were administered in 200 µL saline by gavage throughout eight weeks, and the control or HFAD group also received empty 200 µL saline to minimize the effect of stimulation by oral gavage. At the end of the trial, we collected serum for biochemical profiling, harvested livers for histopathology, and obtained fecal samples for metagenomic sequencing and targeted bile acid metabolomics. Metagenomic analysis confirmed that there was no cross-contamination in the control and HFAD groups, and that oral gavage efficiency was equivalent in the HFAD-H22B656 and HFAD-W5S9 groups, consistent with the fecal qPCR quantification results (Fig. 5b, c and Supplementary Fig. 3a).
a Experimental design. SPF C57BL/6J mice (n = 6 per group) were assigned to: Control (standard diet), HFAD (60% fat), HFAD with B. animalis subsp. lactis H22B656 (HFAD-H22B656), or HFAD with B. animalis subsp. lactis W5S9 (HFAD-W5S9). Mice received daily oral gavage of 10⁹ CFU of the indicated strain in 200 µL saline for eight weeks (or just saline for Control and HFAD). Body weight was recorded weekly; at endpoint, serum was collected for biochemical assays, livers for histopathology (H&E and Oil Red O), and feces for WMS and targeted bile-acid metabolomics (Created in BioRender. Zhang, J. (2026) https://BioRender.com/3qyx3m3). b, c Assessment of Bifidobacterium animalis colonization in feces at week 8 via (b) relative abundance analysis (WMS) and c qPCR quantification (log CFU/g). Data are shown as bar plots with individual points (mean ± SEM, n = 6 mice per group). Kruskal-Wallis test followed by Dunn’s test with multiple comparison correction was used to assess statistical significance; d Body-weight trajectories (mean ± SEM, n = 6 per group). Statistical comparisons at week 8 were conducted by a two-tailed Mann-Whitney test. p = 0.0022 for W5S9-HFAD vs. HFAD, p = 0.0022 for Control vs. HFAD, p = 0.0411 for Control vs. W5S9-HFAD. e Bar plot displaying the net increase in body weight g across groups over the 8-week intervention period, with individual measurements shown (mean ± SEM, n = 6 per group). Group differences were evaluated using the two-tailed Mann-Whitney tests. f Boxplots of serum biomarkers at week 8 (n = 6). Elements represent the median (center line), 25th/75th percentiles (bounds of box), and 1.5× IQR (whiskers). Group differences for each biomarker were analyzed using the two-tailed Mann-Whitney tests. g, h Representative liver histology. g H&E-stained sections and h Oil Red O staining (scale bars, 50 µm) for each group. i Quantification of lipid accumulation in liver sections stained with Oil Red O. Bar graph shows the mean lipid droplet area per field across groups (n = 6 mice per group). Data are expressed as mean ± SEM. Statistical significance was determined using the Kruskal-Wallis test followed by Dunn’s multiple comparisons test. Source data are provided as a Source Data file.
We then compared the effects of the adapted and original strains on NAFLD progression across three parameters: body weight, physiological markers, and histopathologic changes. Over the eight-week intervention, all HFAD-fed groups (e.g., HFAD, HFAD-H22B656, and HFAD-W5S9) gained more weight than the control group on standard chow. Within the HFAD-fed groups, HFAD-W5S9 mice exhibited the smallest weight gain (two-tailed Mann-Whitney test, p = 0.0931), followed by HFAD-H22B656 (p = 0.0411), while the HFAD group showed the greatest increase (p = 0.0022). By week 8, body weight in the HFAD-W5S9 cohort was significantly lower than in the HFAD group (p = 0.0022), whereas there was no significant difference between HFAD-H22B656 and HFAD at week 8 (p = 0.1905; Fig. 5d). To quantify this therapeutic effect, we analyzed the net weight gain. W5S9 intervention significantly prevented HFAD-induced weight gain by 33.6% (mean gain: 7.76 g vs. 11.68 g in the HFAD group; Two-tailed Mann-Whitney test, p = 0.0022), exhibiting the smallest weight gain among the HFAD-fed groups. In contrast, the original strain showed a less pronounced reduction with higher variability (mean gain: 8.14 g vs. 11.68 g in the HFAD group; inhibition rate = 30.3%, p = 0.1320, Fig. 5e). No mortality occurred in mice receiving either H22B656 or W5S9 throughout the eight weeks. In vivo and in vitro safety and stability assessments further demonstrated that the adapted strain W5S9 retains parental-like safety and genetic stability without detectable off-target effects (Supplementary Figs. 4 and 5).
Regarding serum physiological metrics, both the original strain and the adapted strain significantly reduced levels of liver injury marker AST, low-density lipoprotein cholesterol (LDL-C), and pro-inflammatory cytokines (IL-6 and TNF-α) by week 8 compared to the HFAD group (Two-tailed Mann-Whitney test, HFAD vs. HFAD-H22B656 or HFAD vs. HFAD-W5S9, p < 0.05). Notably, compared to the HFAD-H22B656 group, the HFAD-W5S9 group exhibited further improvements in high-density lipoprotein cholesterol (HDL-C), malondialdehyde (MDA), and total cholesterol (TC). For example, while both strains modestly increased serum HDL-C levels, the increase was significantly greater in the HFAD-W5S9 group than in the HFAD-H22B656 group (p = 0.041). For MDA levels, H22B656 did not significantly reduce lipid peroxidation, whereas HFAD-W5S9 had a marked decrease (p = 0.016). A similar trend was observed for TC, with W5S9 showing a significant reduction compared to H22B656 (p = 0.041; Fig. 5f).
Histopathological analysis further corroborated the physiological improvements. Hematoxylin and eosin (H&E) staining of liver sections showed that HFAD-H22B656 mice had moderate alleviation of hepatic steatosis relative to HFAD alone, although scattered microvesicular lipid droplets persisted. In contrast, livers from the HFAD-W5S9 group largely maintained normal lobular architecture, exhibiting only mild, focal microvesicular steatosis (Fig. 5g). Oil Red O staining confirmed a marked reduction in hepatic lipid accumulation in HFAD-W5S9 mice, with more pronounced histological recovery than in the HFAD-H22B656 mice (Fig. 5h). Quantitative image analysis of lipid droplet area (n = 6 per group) further reinforced this observation, revealing a significantly greater reduction in hepatic lipid content in the HFAD-W5S9 group compared to the HFAD group (p = 0.0225, Kruskal-Wallis with Dunn’s test), approaching levels observed in the healthy control group (Fig. 5i). Together, these results demonstrate that the in vivo-induced and directed-evolved probiotic W5S9 provides significantly greater protection against NAFLD progression than the original strain.
Adapted strain’s impact on bile acid-metabolism and NAFLD prevention
Given the strong interplay between gut microbiota and bile acid metabolism34, we sought to determine how W5S9 enhances this interaction under HFAD conditions. We compared the effects of the original H22B656 and adapted W5S9 on gut microbial composition and intestinal bile acid metabolism using metagenomic sequencing and targeted bile acid profiling. In terms of taxonomic structure, significant increases in microbial α-diversity were observed in both HFAD-H22B656 and HFAD-W5S9 mice relative to HFAD alone, as reflected by higher richness and Shannon indices (two-tailed Mann–Whitney test, p < 0.05; Fig. 6a). For β-diversity, principal coordinate analysis (PCoA) of species-level Bray-Curtis dissimilarity showed that, after eight weeks, the gut microbiota of both HFAD-H22B656 and HFAD-W5S9 mice diverged markedly from those of the HFAD and control groups (Fig. 6b). However, there was substantial overlap between the microbiota profiles of the HFAD-H22B656 and HFAD-W5S9 mice, with no significant difference observed between them (p = 0.73; Fig. 6c). Moreover, among 423 species shared by the two groups, no significant differences in relative abundance were detected (multiple Mann–Whitney tests, q > 0.05; Supplementary Fig. 3e), indicating that the adapted strain exerts a broadly similar impact on community structure as the original strain, and that W5S9’s functional enhancement did not come at the expense of major shifts in gut microbiota (Supplementary Fig. 3b, c, d).
a Microbial α-diversity at week 8 (species richness and Shannon index at the species level, WMS data, n = 6 mice per group). Significance is determined by a two-tailed Mann–Whitney test. b PCoA based on Bray-Curtis dissimilarity at the species level (WMS data from n = 6 mice per group). c Pairwise Bray–Curtis dissimilarity between the Control and other groups. Box plots show the median (centre line), 25th and 75th percentiles (box limits) and 1.5× interquartile range (whiskers) for n = 6 mice per group; individual data points are overlaid. Statistical comparisons were evaluated using two-tailed Mann–Whitney tests. d–g OPLS-DA based on the targeted bile-acid metabolic profiles (n = 6 mice per group; OPLS-DA models were fitted using the R package ropls). Score plots (left) and heat maps (right) depict differential metabolites for HFAD vs. Control (d), HFAD-H22B656 (e), HFAD-W5S9 (f), and (g) HFAD-H22B656 vs. HFAD-W5S9. Differential bile acids were defined as those with variable importance in projection (VIP) > 1 and |log₂FC | > 1 for the indicated pairwise comparisons. Top 10 bile acids ranked by VIP score. h Venn diagram of significantly altered bile acids (VIP > 1, |log₂FC | > 1) shared among HFAD vs. Control, HFAD vs. HFAD-H22B656, and HFAD vs. HFAD-W5S9 comparisons. i KEGG pathway enrichment of differential metabolites. Bubble size indicates the richness factor. j Schematic of primary bile-acid biosynthesis (ko00120). Hepatic TCDCA, GCDCA, TCA, and GCA are deconjugated in the gut by BSH (cbh) to regenerate free CDCA and CA (Created in BioRender. Zhang, J. (2026) https://BioRender.com/3qyx3m3). k Concentration of the six key bile acids highlighted in (j). Log₂FC versus HFAD is annotated above each panel. Data are presented as mean ± SEM (n = 6). All shown metabolites met the significance criteria of VIP > 1 and |log2FC | > 1 compared to the HFAD group. Source data are provided as a Source Data file.
Given the minimal impact of W5S9 on overall taxonomic composition relative to its original strain, we next examined functional alterations, focusing on bile acid metabolism. We performed pairwise comparisons of targeted fecal bile acid profiles across the four groups using orthogonal partial least squares discriminant analysis (OPLS-DA). We found that: (1) HFAD and control groups showed clear separation, with 31 significantly altered bile acid metabolites (VIP > 1, |log₂FC | > 1; Fig. 6d); (2) HFAD and HFAD-H22B656 also diverged distinctly, identifying 28 differential metabolites (Fig. 6e); (3) HFAD-W5S9 profile deviated even further from HFAD, with 25 significantly altered bile acids. Notably, glycochenodeoxycholic acid (GCDCA) and taurochenodeoxycholic acid (TCDCA) were markedly reduced in B. animalis subsp. lactis W5S9-treated mice (Fig. 6f); (4) A clear separation between HFAD-H22B656 and HFAD-W5S9 emerged, with eight differential metabolites identified (Fig. 6g). Together, these metabolomic data demonstrate that W5S9 exerts a stronger modulatory effect on bile acid-related pathways than the original strain. Consistent with these metabolomic changes, HUMAnN 3-based functional profiling revealed higher total bile salt hydrolase (EC 3.5.1.24) gene abundance in HFAD-W5S9 than HFAD-H22B656 group, with stratified analysis indicating that this enhancement is primarily driven by W5S9 colonization and enzymatic activity rather than broad remodeling of the resident microbiota (Supplementary Fig. 6).
We next mapped the significantly altered metabolites from our OPLS-DA comparisons onto the primary bile acid biosynthesis pathway (KEGG KO00120). To compare the metabolic modulation capabilities of the original and adapted strains, we generated a Venn diagram highlighting shared and unique pathway metabolites. Both B. animalis subsp. lactis H22B656 and W5S9 significantly altered levels of GCDCA, glycocholic acid (GCA), chenodeoxycholic acid (CDCA), and cholic acid (CA). Notably, B. animalis subsp. lactis W5S9 additionally regulated TCDCA and taurocholic acid (TCA), indicating broader engagement with primary bile acid biosynthesis (Fig. 6h). Furthermore, KEGG enrichment analysis revealed that differential metabolites in the HFAD vs. Control group comparison, as well as HFAD vs. HFAD-W5S9, were significantly enriched in KO00120 (p = 0.0357). In contrast, metabolites distinguishing HFAD from HFAD-H22B656 mice did not show significant enrichment (p = 0.2143; Fig. 6i).
In the primary bile acid biosynthesis pathway, primary bile acids are conjugated with glycine or taurine in the liver to yield TCDCA, GCDCA, TCA, and GCA, which are then secreted into the intestine via the biliary tract. Approximately 95 % of these conjugated bile acids undergo active reabsorption by intestinal epithelial cells, while the remaining fraction is deconjugated by microbial BSHs, producing free bile acids such as CDCA and CA (Fig. 6j). To comprehensively evaluate the regulatory magnitude of the strains, we analyzed the log2 fold changes (log2FC) relative to the HFAD model (Supplementary Fig. 7). While the parental strain H22B656 elicited a moderate reduction in pathological bile acid accumulation, the adapted strain W5S9 exhibited a significantly more profound restorative effect. Specifically, W5S9 treatment resulted in a substantial decrease in both conjugated bile acids (TCDCA, log₂FC = –1.14; TCA, log₂FC = –1.19; GCDCA, log₂FC = –∞; GCA, log₂FC = –2.68;) and free bile acids (e.g., CDCA, log₂FC = –2.09; CA, log₂FC = –2. 02) in the intestinal lumen (Fig. 6k). Notably, bile acid concentrations in HFAD-W5S9 mice approached those of the Control group more closely than in the HFAD-H22B656 mice, indicating a more robust reestablishment of bile acid homeostasis. Given the central role of primary bile acid biosynthesis in maintaining enterohepatic equilibrium and regulating hepatic lipid metabolism, and its dysregulation as a key driver of NAFLD, these data suggest that the adapted strain enhances bile acid transformation and clearance, thereby helping to reestablish metabolic balance and mitigate disease progression.
Discussion
In this study, we propose an in vivo, host-mediated adaptation strategy that emulates natural intestinal selective pressures to direct probiotic evolution. Specifically, by exposing B. animalis subsp. lactis to a dynamically shifting bile acid milieu, which was induced through a high-fat, high-cholesterol diet, we isolated an adapted strain with markedly enhanced bile acid-metabolizing capacity. WGS and transcriptome analyses identified two key SNVs underpinning this phenotype: one variant elevated deconjugation of bile acids, while the other augmented efflux of free bile acids. Subsequently, in a diet-induced NAFLD mouse model that recapitulates key features of non-alcoholic steatohepatitis, dietary supplementation with the adapted strain more effectively restored bile-acid homeostasis, as indicated by a stronger reduction and normalization of both conjugated (TCDCA, GCDCA, TCA, and GCA) and free primary bile acids (CDCA, CA), and was accompanied by lower inflammatory markers and more pronounced attenuation of hepatic steatosis and liver injury than observed with the parental strain.
Current in vitro approaches, such as ALE and genome editing, have been broadly applied to enhance microbial growth37,38, optimize metabolic pathways39, and improve stress tolerance40. However, these techniques face significant technical challenges. ALE’s efficacy is highly dependent on controlled laboratory conditions and often yields phenotypes that fail to translate to real-world applications41. Moreover, many adaptive traits emerge only after prolonged evolution, necessitating repeated dilution and passaging to maintain selection pressure, which is labor-intensive, difficult to scale, and prone to cross-contamination42,43. Conversely, although genome editing provides precise genetic modifications, its effectiveness in complex microbial consortia is curtailed by low editing efficiencies44,45, limited transformation systems (particularly for genetically recalcitrant anaerobes)46,47, and the challenge of concurrently optimizing multiple loci involved in coordinated regulation48,49. These constraints make it challenging to satisfy current industrial and clinical needs. By contrast, the host-mediated in vivo induction strategy we propose offers several key advantages. First, the gut microenvironment promotes high-frequency spontaneous mutations, enabling rapid accumulation of genetic diversity and the development of a high-throughput, high-intensity screening system that greatly outperforms existing genome editing approaches in evolutionary efficiency. Second, this method leverages natural selection pressures within the host gut to drive spontaneous mutation and phenotypic optimization, ensuring that resulting strains are functionally adapted to realistic, biologically relevant settings. Third, because no exogenous genetic material is introduced, this approach complies with non-GMO standards, providing clear benefits in terms of industrial applicability, biosafety, and regulatory compliance.
Our previous work demonstrated that host-mediated adaptation can substantially improve probiotic functionality. For example, multiple rounds of gut passage of Lactiplantibacillus plantarum HNU082 yielded a set of 12 adapted strains bearing insertions within the Cro/C1-type HTH DNA-binding domain50. These insertions enabled the adapted strains to outcompete the endogenous antitoxin module for binding to the cognate toxin under bile acid stress, thereby markedly enhancing bile tolerance and improving both colonization density and persistence in the host gut. Building on these observational results, we sought to test whether defined host-derived selective pressures could be used to drive targeted functional enhancement. In this study, we imposed a controlled bile acid metabolic challenge to actively direct strain adaptation in vivo and compared this approach with traditional in vitro induction. Through systematic multi-omics analysis, we dissected the molecular basis of enhanced bile acid metabolism and ultimately validated the clinical potential of our adapted strains in an animal model to combat NAFLD (Supplementary Fig. 8).
Importantly, this host-mediated in vivo induction strategy is not restricted to NAFLD but can be broadly generalized to other probiotic functions and disease indications. By rationally tuning host background, diet, and environmental stressors, the gut can be used as a programmable evolutionary bioreactor to enrich for strains that are optimized for distinct therapeutic tasks. For instance, strains could be adapted in inflammatory or epithelial-injury models to bolster gut barrier restoration and mucosal immune regulation, thereby potentially benefiting inflammatory bowel disease, post-antibiotic gut dysfunction, or complications following chemotherapy and radiotherapy51. Likewise, when evolved under dietary fiber-enriched or prebiotic-supplemented conditions, probiotics might acquire high short-chain fatty acid-producing capacity and refined signaling properties, making them attractive adjuncts for metabolic, cardiovascular, or neuropsychiatric disorders in which gut-brain and gut-liver axes are involved52,53. Furthermore, applying selective pressures linked to xenobiotic degradation or drug metabolism could yield strains with enhanced capabilities for therapeutic compound modulation or detoxification, broadening their utility in pharmaco-microbiomics and microbiome-based drug-response optimization54. Beyond single-trait enhancement, combining this platform with multi-strain consortia or personalized dietary regimens may ultimately enable patient-specific, functionally tailored live biotherapeutic products that extend well beyond liver disease and provide a generalizable route to next-generation microbiome therapeutics.
Despite its great potential applications, our study has several limitations. First, the current in vivo induction model relies on GF mice, and the extent to which these results translate to other hosts or human populations remains untested. To bridge the translational gap inherent in murine models, future studies will validate the strain’s colonization stability and metabolic efficacy using humanized gnotobiotic mice and human intestinal organoid systems. Second, although the adapted strain demonstrates enhanced bile acid metabolism, the long-term genetic stability of the key SNVs identified requires further validation, especially under conditions of persistent colonization or within complex microbial communities. Nevertheless, this study not only introduces a host microenvironment-directed in vivo adaptation strategy for functional probiotic engineering but also provides a technological pathway for NAFLD prevention. These findings offer proof-of-concept for a next-generation paradigm in functionally tailored microbiome therapeutics.
Methods
Animal ethics and housing conditions
All animal experiments were approved by the Animal Ethics Committee of Hainan University (Approval No. HNUAUCC-2023-00190) and were conducted in strict accordance with the Guidelines for the Care and Use of Experimental Animals of Hainan University. Germ-free mice were maintained in sterile isolators, while specific pathogen-free (SPF) mice were housed in SPF animal facilities. All mice were kept under a 12 h light/12 h dark cycle and housed individually, with a controlled ambient temperature of 25 ± 2 °C and relative humidity of 55 ± 5%.
Calcium precipitation assay for bile salt hydrolase activity
Bifidobacterium animalis subsp. lactis (B. animalis subsp. lactis) strains were obtained from the Tropical Probiotics Development and Utilization Team at Hainan University, which were originally isolated from the intestinal tracts of healthy infants.
To screen for B. animalis subsp. lactis strains with bile acid-metabolizing capacity, 12 randomly selected strains were subjected to a calcium precipitation assay to evaluate their bile salt hydrolase activity. Briefly, 50 µL of each overnight culture was spotted onto MRS-cysteine (MRS-cys) agar supplemented with 0.1% cysteine·HCl, 3.0 g/L taurodeoxycholic acid sodium salt, 3.0 g/L glycodeoxycholic acid sodium salt, 2.0 g/L sodium thioglycolate, and 0.37 g/L anhydrous CaCl₂. Plates were incubated anaerobically at 37 °C for 48 h, after which the diameter of the calcium precipitation halo (indicative of bile salt hydrolase activity) was measured. MRS-cys agar without bile acids served as the negative control. All assays were performed in triplicate.
Bile salt hydrolase specific activity assay
To determine the BSH-specific activity of the candidate strains, we applied the classical ninhydrin colorimetric assay to quantify released amino acids55, a well-established method for amino acid measurement that indirectly reflects BSH hydrolysis efficiency. Specifically, overnight cultures of candidate strains were harvested at mid-log phase and thoroughly mixed. A 1 mL aliquot of culture was centrifuged at 3000 × g for 10 min at 4 °C, and the supernatant was discarded. The cell pellet was washed twice with 200 μL of ice-cold 0.1 M PBS (pH 6.0) and then resuspended in 500 μL of PBS (pH 6.0) to prepare a concentrated cell suspension.
BSH-specific activity was then assayed using glycine-conjugated bile acids (10 mM) as the substrate. In each reaction, 100 μL of the prepared cell suspension was mixed with 100 μL of substrate solution, and the final volume was adjusted to 1 mL with PBS (pH 6.0). Mixtures were incubated at 37 °C for 30 min. In parallel, blank controls contained 100 μL of cell suspension and 900 μL of PBS (no substrate). At the end of the incubation, 500 μL of 15% trichloroacetic acid was added to terminate the reaction, followed by 3 min of incubation at room temperature. Samples were then centrifuged at 14,000 × g for 10 min, and the supernatant was transferred to a 15 mL tube. To each supernatant, 1.5 mL of ninhydrin reagent was added and gently mixed before heating in a boiling water bath for 15 min. Tubes were immediately cooled on ice for 3 min, and absorbance was measured at 570 nm. Released amino acid concentrations were calculated against a concurrently prepared standard curve.
Simultaneously, total protein concentration in the crude lysate was determined using the Bradford assay with bovine serum albumin (BSA) as the standard. Specific activity (U/mg) was defined as the micromoles of amino acid released per milligram of protein per minute, and calculated using the following formula:
To assess functional enhancement in evolved strains relative to their parental counterparts, we calculated the fold change in BSH activity as follows:
where Cn is the Specific activity by the nth induced strain, and C₀ is the corresponding value for its original strain (n = 1, 2, …, 60).
Assessment of tolerance and growth performance
To comprehensively evaluate the gut-survival potential of these 12 candidate strains, we assessed their acid tolerance, bile acid tolerance, and growth performance. For acid and bile acid tolerance assays, strains were grown to mid-log phase and then transferred (2% v/v) into modified MRS-cys broth: one set adjusted to pH 2.0 (acid tolerance) and another supplemented with 0.3% (w/v) bovine bile (bile acid tolerance). Acid-treated cultures were incubated anaerobically at 37 °C for 2 h, and bile acid-treated cultures for 4 h, with untreated cultures in MRS-cys as controls. After incubation, OD₆₀₀ was measured in both treated and control groups, and survival rates (%) were calculated. As for growth performance, activated cultures were inoculated at 2% (v/v) into MRS-cys broth and incubated anaerobically at 37 °C for 36 h. OD₆₀₀ measurements were taken every 4 h, with uninoculated medium serving as the control.
In vitro induction experiment
To specifically enhance bile acid-metabolizing capacity, each candidate strain was first inoculated into 25 mL of MRS-cys broth and incubated anaerobically at 37 °C until reaching the logarithmic growth phase. At that point, 2% (v/v) of the culture was transferred into an equal volume of fresh MRS-cys medium supplemented with 0.3% (w/v) conjugated bile acids (a 1:1 mixture of sodium glycodeoxycholate and sodium taurodeoxycholate) and incubated anaerobically for 24 h. This serial passaging (using a 2% (v/v) inoculum each time) was repeated for ten consecutive rounds. After the tenth passage, the evolved population was diluted and plated onto solid MRS-cys agar. Subsequently, 60 colonies were randomly selected and assayed for BSH-specific activity to evaluate the effectiveness of the in vitro induction.
Host-mediated in vivo induction
To implement the in vivo adaptation strategy, we employed GF mice as an evolution bioreactor to eliminate interference from endogenous microbial strains. Male GF C57BL/6J mice (7 weeks old; n = 4) were obtained from Shenzhen Jingtai Biotech (Guangdong, China) and maintained in sterile isolators on irradiated standard chow for 7 days. To establish colonization by B. animalis subsp. lactis H22B656, each mouse received a 200 μL oral gavage of a 10⁹ CFU suspension once daily for three consecutive days. Following colonization, mice were subjected to a stepwise high-fat high-cholesterol dietary regimen over a 7-week induction period: weeks 1–3 on chow containing 30% fat and 0.2% cholesterol; weeks 3–5 on 45% fat and 0.2% cholesterol; and weeks 5–7 on 60% fat and 0.2% cholesterol.
To isolate adapted strains from GF mice, fresh stool samples were aseptically collected at the end of weeks 1, 3, 5, and 7, immediately suspended in 1.5 mL of sterile saline, and thoroughly homogenized under anaerobic conditions. Homogenates were serially diluted to 10-⁴ to 10-⁶ and plated on MRS-cys agar. After anaerobic incubation at 37 °C for 24 h, colonies from an appropriate dilution were randomly picked. To ensure a representative evaluation of the adaptation process, 15 isolates were randomly selected from the plates of each sampling time point (Weeks 1, 3, 5, and 7), resulting in a total library of 60 induced isolates. These isolates were inoculated into MRS-cys liquid medium, labeled sequentially, and stored at –80 °C for subsequent analyses. Finally, 60 induced isolates were assayed alongside the original B. animalis subsp. lactis H22B656 for BSH-specific activity to evaluate the efficacy of the host-mediated in vivo induction.
Intervention diets
The normal diet (ND) was designed in accordance with established nutritional guidelines and comprised casein, corn starch, lard, vitamins, minerals, and fiber, providing a balanced mix of protein, fat, and carbohydrates. To generate an intestinal high-bile-acid environment in GF mice, we formulated a high-fat/high-cholesterol diet by increasing lard content to supply 30%, 45%, or 60% of energy from fat and supplementing with 0.2% cholesterol. For the NAFLD intervention study, a high-fat diet (HFAD) was similarly prepared from the normal diet to provide 60% energy from fat, without cholesterol supplementation. All diets were sterilized by γ irradiation and provided by Anhui Kuibu Shuyu Biotechnology Co., Ltd. Full dietary compositions are listed in Supplementary Data 2.
Genome assembly and annotation of the original strain
To obtain the complete reference genome of the original strain B. animalis subsp. lactis H22B656, genomic DNA was extracted from pure cultures of B. animalis subsp. lactis H22B656 using the QIAamp DNA Stool Mini Kit (Qiagen), following the manufacturer’s protocol. DNA concentration and integrity were assessed by NanoDrop spectrophotometry (A260/A280 ≈ 1.8) and 0.8% agarose gel electrophoresis. Samples meeting these quality criteria were sent to Beijing Novogene Bioinformatics Technology Co., Ltd. for third-generation long-read sequencing on the Oxford Nanopore Technologies (ONT) PromethION platform.
Nanopore libraries were prepared using the ONT Ligation Sequencing Kit (Oxford Nanopore Technologies) as follows: Genomic DNA integrity was assessed by gel electrophoresis and quantified using a Qubit fluorometer. The DNA underwent end-repair and A-tailing reactions, after which sequencing adapters and motor proteins were ligated to both ends. Unligated molecules were removed via purification steps, and libraries meeting quality standards were sequenced on the PromethION platform. Sample B. animalis subsp. lactis H22B656 generated 433,564 raw reads with a total output of 2.4 Gb.
For genome assembly, raw reads were first demultiplexed and adapter-trimmed to generate subreads, which were then filtered to retain only those ≥50 bp in length. De novo assembly of the filtered subreads was performed with Canu56 (v2.2). The resulting draft contigs were polished through three rounds of alignment and consensus correction by coupling Minimap257 (v2.22) and Racon58 (v1.4.13, command: minimap2 -t 40 assembly.fasta pass.fastq.gz > round1.paf; racon -t 4 pass.fastq.gz round1.paf assembly.fasta > round1.fasta).
Gene prediction and functional annotation of the complete genome were performed with Prokka59 v1.14.6 (command: prokka --outdir test --prefix test test.fa). Predicted proteins were further annotated via the KEGG Automatic Annotation Server60 (KAAS v4.0; http://www.genome.jp/tools/kaas/), using the bi-directional best hit method for pathway assignment. To further annotate hypothetical proteins, BLASTp searches were also conducted with NCBI BLAST+ v2.16.0 against the non-redundant (nr) protein database (e-value ≤ 1e-5, identity ≥ 30%, query coverage ≥ 70%, up to 20 alignments; command: blastp -query query.faa -db nr -out blast.tsv -outfmt 6 -evalue 1e-5 -num_threads 24 -num_alignments 20).
Whole genome sequencing of the adapted strains
To investigate the genetic basis of its enhanced bile acid-metabolizing capacity, we performed whole-genome sequencing (WGS) of the adapted strain W5S9. Genomic DNA from purified W5S9 cultures was extracted and assessed for its purity (A260/A280 = 1.8-2.0) and fragment integrity ( > 20 kb). DNA was then randomly sheared to the desired fragment size using a Covaris ultrasonicator. Fragmented DNA underwent end repair and 3’ A-tailing, followed by ligation of Illumina sequencing adapters. Adapter-ligated libraries were size-selected to remove off-target fragments, PCR-amplified to enrich adapter-containing molecules, and purified to eliminate residual contaminants. Equimolar libraries were pooled and sequenced by Novogene Co., Ltd. (Beijing, China) using an Illumina HiSeq 2500 platform (2 × 150 bp paired-end). Raw reads were stringently filtered for quality using fastp61 v0.23.2. Sample W5S9 yielded 8,310,066 raw reads, of which 8,289,624 high-quality reads were retained after filtering.
SNVs calling
To accurately capture genetic variants acquired during in vivo adaptation, we performed SNV calling on WGS data using two complementary pipelines (inStrain30 v1.0.0 and BCFtools31 v 1.21) and retained only the intersection of their call sets for downstream analysis. First, WGS data were aligned to the original H22B656 reference genome with Bowtie262 v2.5.4. Alignments were converted to coordinate-sorted, indexed BAM files using SAMtools31 v1.22. SNV detection was then carried out independently with: (1) inStrain (command: inStrain profile sample.bam ref.fna -c 100 -f 0.49 -o sample.profile -p 24 -g ref_genes.fna); (2) BCFtools (command: bcftools mpileup -C 50 -Ou -m 3 -F 0.0002 -f ref.fa sorted.bam | bcftools call -mv -Ob -o variants.bcf). High-confidence SNVs were defined as variants identified by both methods. We identified a total of 282 high-confidence SNVs in the adapted strain W5S9. The distribution of SNVs across the genome was plotted using the R package circlize (version 0.4.15, Fig. 3a).
Promoter prediction for non-coding SNVs
To determine whether SNVs in non-coding regions might influence downstream gene expression, we conducted promoter prediction and target-gene association analyses for all intergenic SNVs. First, each SNV’s nearest gene was identified using BEDTools63 v2.18, and the distance to the gene’s transcription start site (TSS) was calculated. Genes located on the same strand, with SNVs positioned within 0–300 bp upstream of their TSS, were selected as candidate regulatory targets. Promoter regions encompassing each candidate SNV were then predicted using BPROM (www.softberry.com), and the predicted promoter elements were inspected for canonical −10/ − 35 motifs to evaluate their regulatory potential.
Transcriptome sequencing and data analysis
To identify key genes with expression changes caused by genetic variants acquired during in vivo adaptation, we performed transcriptome sequencing (RNA-seq) on the original strain H22B656 and the adapted strain W5S9. Briefly, each strain was cultured anaerobically for 12 h in MRS-cys broth supplemented with 0.3% conjugated bile acid (sodium glycodeoxycholate and sodium taurodeoxycholate). Cultures were centrifuged at 1500 × g for 30 min, and pellets were immediately processed for RNA extraction.
Total RNA was isolated using a Qiagen RNeasy kit, and integrity (RIN > 8) and purity (A260/A280:1.8-2.0) were confirmed by agarose gel electrophoresis and NanoDrop spectrophotometry. Ribosomal RNA was depleted, and the remaining RNA was purified by ethanol precipitation. First-strand cDNA synthesis was primed with random hexamers, and second-strand synthesis incorporated dUTP to preserve strand specificity. The resulting double-stranded cDNA underwent end repair, 3’ A-tailing, adapter ligation, size selection, USER enzyme digestion to remove the dUTP-containing strand, PCR enrichment, and bead-based purification to yield strand-specific libraries. Library concentration was measured by Qubit and qPCR, and fragment size distribution was assessed on an Agilent Bioanalyzer. Libraries meeting quality thresholds were pooled equimolarly and sequenced (2 × 150 bp) on an Illumina HiSeq 2500 by Novogene Co., Ltd. (Beijing, China).
Raw data were quality-filtered with fastp v0.23.2 to remove adapter contamination, trim low-quality bases (Q < 20), and discard reads containing ambiguous nucleotides (N), resulting in high-quality clean reads for downstream analysis. Sequencing of six transcriptomic samples yielded an average of 11.95 ± 0.15 million raw reads and 11.79 ± 0.15 million clean reads per sample (mean ± SEM). Transcript-level quantification was performed with Salmon64 v1.9.0 against the reference transcriptome (salmon quant -i mg -l A −1 test_1.fastq.gz −2 test_2.fastq.gz -o test.quant). Differential expression analysis between the original and adapted strains was performed in R (v4.4.1) using the DESeq2 (v1.44.0) package (n = 3 per strain, Benjamini–Hochberg FDR correction; FDR < 0.001 and |log₂FC| > 0.4), and the resulting differential features were visualized as volcano plots using OmicShare (https://www.omicshare.com/; Fig. 3c, d). Gene expression differences between adapted and original strains for cbh and mdr were shown as bar charts in GraphPad Prism v10 (Fig. 3f, g).
Validation of identified SNVs by targeted Sanger sequencing
To validate the candidate SNVs (rs1846941 and rs402499) identified by WGS, we designed locus-specific primer pairs (mrd-F/mrd-R and rs1846941-f/ rs1846941-r; Supplementary Data 3) to amplify 200 bp regions centered on each variant (100 bp upstream and downstream). Purified PCR products were submitted to Beijing Qinke Biotechnology Co., Ltd. for Sanger sequencing. Following sequencing, the raw reads were aligned against the B. animalis subsp. lactis H22B656 reference genome to verify the target mutations. Sequence visualization and variant confirmation were performed using Geneious Prime v2022.2 (Fig. 4a, b).
Determination of the genetic stability of key SNVs
To verify the genetic stability of key SNVs in the adapted strain, we conducted a rigorous long-term continuous passage experiment followed by whole genome sequencing (WGS). Specifically, the adapted strain W5S9 was subjected to daily serial passaging for 4 weeks under two conditions: (i) a non-selective medium (MRS broth supplemented with L-cysteine hydrochloride), used to assess stability in the absence of selection pressure; and (ii) a selective medium (MRS broth supplemented with bile salts and L-cysteine hydrochloride), used to mimic gut environmental pressure. Target strains were re-isolated weekly, genomic DNA was extracted, and WGS was performed to monitor potential genetic changes (Supplementary Fig. 4).
Reporter plasmid construction and fluorescence assays
To assess the regulatory impact of SNVs rs1846941 and rs402499 on target gene expression, we constructed mCherry and EGFP reporter plasmids harboring either the wild-type or variant sequences.
For SNV rs1846941 located in the non-coding region, we PCR-amplified 100 bp fragments (50 bp upstream and downstream) carrying either the wild-type or rs1846941 allele from genomic DNA of the original H22B656 and the adapted W5S9 strains, respectively. Each non-coding fragment was fused to mCherry by overlap-extension PCR, and the resulting expression cassettes were cloned into the EcoRI site of pLH01. Recombinant plasmids were introduced into B. animalis subsp. lactis H22B656 by electroporation. Single colonies were then grown anaerobically at 37 °C in MRS-cys medium (supplemented with 2% glucose, 10 µg/mL chloramphenicol, and 0.3% (w/v) conjugated sodium deoxycholate, sodium cholate, or sodium chenodeoxycholate) to an OD₆₀₀ of ~0.5. Cells were then harvested, washed twice with PBS, and fluorescence was measured on a BioTek Synergy H1 multimode plate reader.
For SNV rs402499 located in the coding region (in mdr), wild-type and N192D gene-egfp fusions were assembled by overlap-extension PCR and cloned into SalI/XhoI–linearized pET28a. Plasmids were transformed into E. coli BL21(DE3) by heat shock. Transformants were cultured in LB medium (with 0.3% conjugated sodium deoxycholate, conjugated sodium cholate, or conjugated sodium chenodeoxycholate and 50 µg/mL kanamycin) at 37 °C to OD₆₀₀ = 0.6–0.8. After harvesting and two PBS washes, fluorescence was recorded on the Synergy H1. Statistical analyses and data visualization were performed using GraphPad Prism v10. For Fig. 4c and Supplementary Fig. 2, multiple two-tailed Mann–Whitney tests were used to compare H22B656 and W5S9 (n = 3 independent samples per group), and multiple comparisons were controlled using a two-stage step-up false discovery rate (FDR) procedure with Q = 1%.
Construction of gene knockout and complemented strains
To establish a causal relationship between the rs1846941 and rs402499 mutations and the phenotype of interest, we employed a homologous recombination strategy using the temperature-sensitive suicide plasmid pKO403 to generate deletion mutants targeting the mdr and cbh promoter regions in both the parental strain Bifidobacterium animalis subsp. lactis H22B656 and the adapted strain B. animalis subsp. lactis W5S9. Genomic DNA from the respective strains was used as the template to amplify the upstream and downstream homologous arms flanking the mdr gene and cbh promoter regions by PCR using primers listed in Supplementary Data 3. Amplicons were resolved by agarose gel electrophoresis and purified. The pKO403 plasmid was linearized by double digestion with SalI and BamHI and seamlessly assembled with the purified homologous arms using 2×Ezmax® Universal CloneMix (37 °C, 30 min). Assembly products were transformed into E. coli DH5α competent cells, and recombinant plasmids were identified by colony PCR. Plasmids confirmed by PCR were isolated and electroporated into electrocompetent H22B656 and W5S9 cells (2.1 kV, 200 Ω, 25 μF). Following electroporation, cells were recovered in 1 mL MRS-cys medium under anaerobic conditions at 37 °C for 4 h and plated on MRS-cys agar containing chloramphenicol (10 μg/mL) to select for single-crossover integrants. Chloramphenicol-resistant colonies were cultured in MRS-cys broth supplemented with chloramphenicol (10 μg/mL) at 37 °C for 48 h and subsequently plated on MRS-cys agar with chloramphenicol for anaerobic incubation at 42 °C for 48 h to promote double-crossover recombination. Successful deletion of the target region was confirmed by PCR, and double-crossover deletion mutants were obtained. To eliminate the pKO403 plasmid, verified mutants were passaged for three consecutive generations in antibiotic-free MRS-cys medium.
For complementation of the Δmdr mutant, a 1648-bp fragment containing the full-length mdr gene and its native promoter was PCR-amplified from the genomic DNA of H22B656 or W5S9 using primers listed in Supplementary Data 3 and cloned into SalI/BamHI-digested pKO403. The resulting plasmid was introduced into the Δmdr strain by electroporation, and transformants were selected on MRS-cys agar containing chloramphenicol (10 μg/mL). Successful construction of complemented strains was confirmed by PCR.
RT-qPCR gene expression analysis
To assess the impact of the rs402499 and rs1846941 variants on gene expression, bacterial cultures were grown in MRS-cys medium supplemented with 0.3% (w/v) conjugated sodium deoxycholate, sodium cholate, or sodium chenodeoxycholate. After 12 h of incubation, bacterial cells were collected for subsequent analysis. Total RNA was extracted with TRIzol reagent and treated with DNase I to remove genomic DNA. cDNA synthesis was performed using the PrimeScript™ RT kit according to the manufacturer’s instructions. The cycle threshold (CT) values were normalized to the reference gene rplx to obtain ΔCT values. Relative gene expression was calculated using the comparative CT method (2-ΔΔCT), with the wild-type Bifidobacterium animalis subsp. lactis H22B656 strain serving as the calibrator (relative expression = 1) for each bile salt condition independently. Statistical analyses were performed using R v4.4.1 software with the rstatix (v0.7.2) and ggpubr (v0.6.0) packages. Data are presented as mean ± SD, with individual data points from biological replicates overlaid (n = 3). Comparisons between strains under each bile salt condition were analyzed using one-way analysis of variance (ANOVA) followed by Tukey’s Honest Significant Difference (HSD) post-hoc test for multiple comparisons. Statistical significance was defined as p < 0.05 (Fig. 4d, f).
Bile acid stress survival assay for mdr phenotypic validation
To validate the contribution of mdr expression to bile acid tolerance, survival under bile acid stress was assessed by viable colony counting. The parental wild-type strain H22B656, the adapted wild-type strain W5S9, their isogenic mdr knockout mutants, and the corresponding complemented strains were grown overnight under standard culture conditions as described above. Cultures were harvested and adjusted to the same optical density, and equal aliquots were inoculated into fresh medium supplemented with 0.3% (w/v) cholic acid (CA), deoxycholic acid (DCA), or chenodeoxycholic acid (CDCA). After incubation under bile acid stress, cultures were serially diluted and plated on bile acid–free agar. Colony-forming units (CFU) were counted after incubation, and survival was expressed as CFU per mL. Each condition was tested in three independent biological replicates, and data were analyzed by one-way ANOVA followed by Tukey’s HSD post hoc test (Fig. 4g).
Molecular docking analysis
To investigate the impact of the nonsynonymous variant rs402499 on MDR structure and its binding affinity for free bile acids, we used the AlphaFold server (https://alphafoldserver.com) to predict the three-dimensional structures of both wild-type and mutant MDR from their amino acid sequences, exporting the resulting models as PDB files for downstream analyses. The three-dimensional structures of deoxycholic acid (CID: 222528, https://pubchem.ncbi.nlm.nih.gov/compound/Deoxycholic-acid), cholic acid (CID: 221493, https://pubchem.ncbi.nlm.nih.gov/compound/Cholic-acid), and chenodeoxycholic acid (CID: 10133, https://pubchem.ncbi.nlm.nih.gov/compound/Chenodeoxycholic-acid) were retrieved from PubChem, converted from SDF to PDBQT format with Open Babel (v3.1.1), and assigned Gasteiger charges.
Both wild-type and mutant MDR models were prepared for docking with AutoDockTools (v1.5.7) by removing crystallographic water molecules, adding hydrogens, and assigning Kollman charges before exporting them as PDBQT files. Potential ligand-binding pockets were identified using DoGSiteScorer (https://proteins.plus/). A ligand-centered docking grid was then defined (box size: 126³ ų; grid spacing: 0.368 Å) and centered at coordinates x = −0.855, y = 10.657, z = −4.238 (dimensions: 20.33 × 20.33 × 20.33 ų). Targeted molecular docking was performed with AutoDock Vina65 (v1.5.7). Statistical analyses and data visualization were performed using GraphPad Prism v10 (Fig. 4h). The binding pose with the lowest predicted binding energy was visualized in PyMOL (v2.5.4), and key ligand-receptor interactions, including hydrogen bonding, hydrophobic contacts, and shape complementarity, were analyzed (Fig. 4i).
Diet-induced NAFLD mouse model and probiotic gavage
To assess the beneficial potential of the adapted B. animalis subsp. lactis W5S9 in modulating host metabolism, we employed a diet-induced NAFLD mouse model in male SPF C57BL/6J mice (7 weeks old), based on the established link between bile acid dysregulation and NAFLD progression66. Mice were randomized into four groups (n = 6 per group): (1) Control (n = 6, normal diet, daily gavage with 200 µL sterile saline); (2) HFAD (n = 6, high-fat diet, daily gavage with 200 µL sterile saline); (3) HFAD-H22B656 (n = 6, high-fat diet, daily gavage with 200 µL sterile saline containing 109 CFU of original strain B. animalis subsp. lactis H22B656); and (4) HFAD-W5S9 (n = 6, high-fat diet, daily gavage with 200 µL sterile saline containing 109 CFU of adapted strain B. animalis subsp. lactis W5S9). The experiment lasted for 8 weeks. Body weight was measured weekly. At the end of the study, serum and liver tissues were collected for biochemical measurements and histopathological analysis to assess the preventive effects of the adapted and original strains on NAFLD. Additionally, fecal samples were collected for metagenomic sequencing and targeted bile acid metabolomics analysis.
Physiological and histopathological assessments
For the physiological markers assay, the levels of high-density lipoprotein (HDL-C), low-density lipoprotein (LDL-C), total triglycerides (TG), total cholesterol (TC), aspartate aminotransferase (AST), malondialdehyde (MDA), IL-6 and TNF-α in serum were measured according to the instructions of the ELISA kit (Shanghai Xinyu Biotechnology Co., Ltd.).
For histopathological analysis, liver specimens were rinsed in ice-cold sterile saline, and portions were either fixed in 4% paraformaldehyde or snap-frozen in liquid nitrogen. Sample processing, including paraffin embedding, sectioning, staining, and image capture, was performed by Wuhan Sewell Biotechnology Co., Ltd. (China). Semi-quantitative analysis of lipid droplet content was performed on Oil Red O-stained sections using Fiji (v2.17.0). Data visualization and statistical analyses were conducted in GraphPad Prism v10 using a Kruskal–Wallis test followed by Dunn’s multiple comparisons test (n = 6 mice per group, Fig. 5i).
Whole metagenome sequencing and data analysis
To compare the effects of the adapted versus original strains on the gut microbiota composition in the diet-induced NAFLD mouse model, fresh stool samples were collected at the end of week 8 and immediately stored at −80 °C for subsequent whole metagenome sequencing (WMS). Genomic DNA was extracted using the QIAamp DNA Stool Mini Kit (Qiagen) and evaluated for integrity by 0.8% agarose gel electrophoresis, purity via A260/A280 ratios on a NanoDrop spectrophotometer, and concentration with a Qubit 2.0 fluorometer. Paired-end libraries (2 × 150 bp) were prepared and sequenced on an Illumina HiSeq 2500 platform at Beijing Novogene Bioinformatics Technology Co., Ltd. Raw reads were quality-filtered using fastp v0.23.2 to remove adapter sequences and exclude read pairs with more than 50% of bases at Q ≤ 5 or over 10% ambiguous nucleotides (N bases). On average, each sample yielded 45.28 ± 0.56 million raw reads and 44.08 ± 0.57 million clean reads (mean ± SEM). Taxonomic profiling of the WMS data was performed using MetaPhlAn (v 4.0.6)67 to identify microbial species and estimate their relative abundances. To specifically distinguish the direct enzymatic contribution of the supplemented strains from potential regulatory effects on the resident microbiota, we performed functional profiling of the metagenomic data using HUMAnN (v3.9) with the uniref90_ko regrouping option to quantify the abundance of bile salt hydrolase (EC 3.5.1.24)–encoding genes (Supplementary Fig. 6).
qPCR quantification of B. animalis in fecal samples
To quantitatively assess the abundance of the original strain H22B656 and the adapted strain W5S9 in fecal samples, we designed H22B656-specific primers using the NCBI Primer-BLAST tool (H22BF: ACGGCACGATTCATGTGTCT; H22BR: CCAGATTCCATCCTCCTCGC). Fecal samples (0.1 g) were then subjected to DNA extraction using the TIANGEN Bacterial Genomic DNA Extraction Kit, and quantitative real-time PCR (qPCR) was performed with the above primers. qPCR data were analyzed and visualized using GraphPad Prism v10, and statistical significance was evaluated using the Kruskal–Wallis test followed by Dunn’s multiple comparisons test (Fig. 5c).
Targeted metabolomic profiling and data analysis of fecal bile acids
A total of 24 fecal samples (n = 6 biological replicates per group) were analyzed. Fecal bile acid extraction was performed using HPLC-grade acetonitrile and methanol (Merck) as solvents. Analytical standards (CNW, IsoReag, and Sigma-Aldrich) were prepared as 1 mg/mL stock solutions in methanol and stored at −20 °C. Working solutions were diluted to appropriate concentrations for calibration curves prior to analysis. Approximately 20 mg of ball-milled fecal sample was homogenized in 495 μL of methanol, spiked with 5 μL of internal standard solution (10 μg/mL), and subjected to protein precipitation at −20 °C. Following centrifugation at 13,200 × g for 10 min (4 °C) and filtration through a protein precipitation plate, supernatants were analyzed. To monitor instrument stability, quality control (QC) samples were prepared by pooling equal aliquots of all supernatants and analyzed every 10 samples.
Chromatographic separation was performed on an ExionLC™ AD UHPLC system (AB Sciex) equipped with a Waters HSS T3 C18 column (100 × 2.1 mm, 1.8 μm) maintained at 40 °C. Mobile phases consisted of (A) 0.01% acetic acid with 5 mmol/L ammonium acetate in water and (B) 0.01% acetic acid in acetonitrile. A linear gradient elution was performed at a flow rate of 0.35 mL/min: 5% to 40% B in 0–1 min; 40% to 50% B in 1–7 min; 50% to 75% B in 7–12 min; 75% to 95% B in 12–14 min; maintained at 95% B for 2 min (14–16 min); and re-equilibrated at 5% B from 16 to 17.5 min.
Mass spectrometric detection was conducted on an AB Sciex 6500 + QTRAP system coupled to an ExionLC™ AD UHPLC via electrospray ionization (ESI) in negative mode. Key parameters included: ion source temperature 550 °C, spray voltage −4500 V, and curtain gas 35 psi. Data acquisition employed scheduled multiple reaction monitoring (MRM) with Analyst software (v1.6.3), and quantification was performed using MultiQuant software (v3.0.3). Metabolites were identified by matching retention times and MS/MS spectra with the MetWare Database (MWDB) and authentic standards. Quantification was performed using MultiQuant software (v3.0.3) based on linear calibration curves. Fecal bile acid profiling was performed by MetWare (http://www.metware.cn/) based on the AB Sciex QTRAP 6500 LC-MS/MS platform. A detailed list of MRM transitions, collision energies, retention times, and mass spectra for all targeted bile acids is provided in Supplementary Data 4.
For targeted fecal bile acid metabolomics, orthogonal partial least squares discriminant analysis (OPLS-DA) was performed using the R (v4.4.1) package ropls (v1.36.0) for pairwise comparisons between the indicated groups (n = 6 per group). Differential metabolites were defined as those with a variable importance in projection (VIP) > 1 and |log₂ fold change | > 1 between the indicated groups. Score plots and heatmaps were generated with ggplot2 (v3.5.2) and pheatmap (v1.0.12), respectively (Fig. 6d–g). Intersection analysis of differential metabolites was conducted with the Evenn online tool (https://www.bic.ac.cn/test/venn/#/) and visualized as a Venn diagram (Fig. 6h). KEGG pathway enrichment results were displayed as bubble plots using ggplot2 (v3.5.2) (Fig. 6i), and quantitative data for key differential metabolites were plotted as bar graphs in GraphPad Prism v10 (Fig. 6k).
Statistics and reproducibility
Statistical analyses were performed using GraphPad Prism v10 and R software (v4.4.1). Data are presented as mean ± standard deviation (SD) or mean ± standard error of the mean (SEM), as indicated in the figure legends. Data distribution was assessed for normality. Comparisons between two groups were conducted using unpaired two-tailed Student’s t-tests or Mann–Whitney U-tests. For comparisons among more than two groups, one-way analysis of variance (ANOVA) followed by Tukey’s post-hoc test or Kruskal–Wallis tests followed by Dunn’s multiple comparisons test were applied. To correct for multiple testing in large-scale datasets (e.g., metabolomics and transcriptomics), the False Discovery Rate (FDR) was controlled using the Benjamini–Krieger–Yekutieli two-stage step-up method (or the Benjamini–Hochberg method for RNA-seq).
Specific R packages were employed for data visualization and analysis, including ggplot2 (v3.5.2) for general plotting, ggradar (v0.2) for radar plots, pheatmap (v1.0.12) for heatmaps, ggsignif (v0.6.4) for significance annotation, and vegan (v2.6-10) for diversity analysis. All schematic diagrams were created using BioRender (Zhang, J. (2026) https://BioRender.com/3qyx3m3). All experiments were performed with at least three independent biological replicates to ensure reproducibility.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All the data that support the findings of this study are available within the paper and its Supplementary Data. All third-generation long-read, whole-genome resequencing, transcriptomic (RNA-seq), and metagenomic sequencing data generated in this study have been deposited in the NCBI database under the following BioProject accession numbers: PRJNA1281804, PRJNA1281807, PRJNA1380636, PRJNA1281849, and PRJNA1283298. Metagenomic data for sample H22B656-6 are not obtained due to DNA extraction failure. The predicted structures of the wild-type protein and the N192D mutant have been deposited in Figshare (https://doi.org/10.6084/m9.figshare.31272826). Targeted metabolomics data for bile acids are available in the MetaboLights database under identifier MTBLS13718. Source data are provided with this paper.
References
Beck-Sickinger, A. G. Directed evolution in drug and antibody development: From the Nobel Prize to broad clinical application]. Internist 60, 1014–1020 (2019).
Scott, B. M. et al. Self-tunable engineered yeast probiotics for the treatment of inflammatory bowel disease. Nat. Med. 27, 1212–1222 (2021).
Karpinski, J. et al. Directed evolution of a recombinase that excises the provirus of most HIV-1 primary isolates with high specificity. Nat. Biotechnol. 34, 401–409 (2016).
Chatterjee, R. & Yuan, L. Directed evolution of metabolic pathways. Trends Biotechnol. 24, 28–38 (2006).
Phaneuf, P. V., Gosting, D., Palsson, B. O. & Feist, A. M. ALEdb 1.0: a database of mutations from adaptive laboratory evolution experimentation. Nucleic Acids Res. 47, D1164–D1171 (2019).
Bennett, A. F. & Hughes, B. S. Microbial experimental evolution. Am. J. Physiol. Regul. Integr. Comp. Physiol. 297, R17–R25 (2009).
Sandberg, T. E., Salazar, M. J., Weng, L. L., Palsson, B. O. & Feist, A. M. The emergence of adaptive laboratory evolution as an efficient tool for biological discovery and industrial biotechnology. Metab. Eng. 56, 1–16 (2019).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Gu, H. et al. Physiological mechanism of improved tolerance of Saccharomyces cerevisiae to lignin-derived phenolic acids in lignocellulosic ethanol fermentation by short-term adaptation. Biotechnol. Biofuels 12, 268 (2019).
Li, J. et al. Biofortified tomatoes provide a new route to vitamin D sufficiency. Nat. Plants 8, 611–616 (2022).
Zhang, J. et al. Probiotics maintain the intestinal microbiome homeostasis of the sailors during a long sea voyage. Gut Microbes 11, 930–943 (2020).
Trager, C. et al. A probiotic approach identifies a Treg-centred immunoregulation via modulation of gut microbiota metabolites in people with multiple sclerosis and healthy individuals. EBioMedicine 116, 105743 (2025).
Murray, E. et al. Probiotic consumption during puberty mitigates LPS-induced immune responses and protects against stress-induced depression- and anxiety-like behaviors in adulthood in a sex-specific manner. Brain Behav. Immun. 81, 198–212 (2019).
Zhang, J. P. et al. High-throughput screening of probiotics in fermented foods and their potential application in alleviating alcohol-induced damage. Food Funct. 16, 2564–2576 (2025).
Keshri, J. et al. Phenotypic characterization and draft genome sequence analyses of two novel endospore-forming sporosarcina spp. isolated from Canada Goose (Branta canadensis) feces. Microorganisms 12, 70 (2023).
Zhang, C. et al. A key genetic factor governing arabinan utilization in the gut microbiome alleviates constipation. Cell Host Microbe 31, 1989–2006 e1988 (2023).
Fletcher, E. et al. Evolutionary engineering reveals divergent paths when yeast is adapted to different acidic environments. Metab. Eng. 39, 19–28 (2017).
Tilloy, V., Ortiz-Julien, A. & Dequin, S. Reduction of ethanol yield and improvement of glycerol formation by adaptive evolution of the wine yeast Saccharomyces cerevisiae under hyperosmotic conditions. Appl. Environ. Microbiol. 80, 2623–2632 (2014).
Baym, M. et al. Spatiotemporal microbial evolution on antibiotic landscapes. Science 353, 1147–1151 (2016).
Pan, C. et al. Ameba-inspired strategy enhances probiotic efficacy via prebound nutrient supply. Nat. Commun. 16, 1827 (2025).
Chang, Z. et al. Bacterial immunotherapy leveraging IL-10R hysteresis for both phagocytosis evasion and tumor immunity revitalization. Cell 188, 1842–1857.e1820 (2025).
Lee, J. S. et al. E. coli genetically modified for purine nucleobase release promotes butyrate generation and colonic wound healing during DSS insult. Gut Microbes 17, 2490211 (2025).
Charbonneau, M. R., Isabella, V. M., Li, N. & Kurtz, C. B. Developing a new class of engineered live bacterial therapeutics to treat human diseases. Nat. Commun. 11, 1738 (2020).
Ford, S. A. & King, K. C. In vivo microbial coevolution favors host protection and plastic downregulation of immunity. Mol. Biol. Evol. 38, 1330–1338 (2021).
Huang, S. et al. Candidate probiotic Lactiplantibacillus plantarum HNU082 rapidly and convergently evolves within human, mice, and zebrafish gut but differentially influences the resident microbiome. Microbiome 9, 151 (2021).
Zhao, S. et al. Adaptive evolution within gut microbiomes of healthy people. Cell Host Microbe 25, 656–667.e658 (2019).
Li, W. et al. Comparative genomics of in vitro and in vivo evolution of probiotics reveals energy restriction not the main evolution driving force in short term. Genomics 113, 3373–3380 (2021).
Aydin, O. et al. An integrated analysis of bile acid metabolism in humans with severe obesity. Hepatology 81, 19–31 (2025).
Kim, T. Y. et al. A high-fat diet activates the BAs-FXR axis and triggers cancer-associated fibroblast properties in the colon. Cell Mol. Gastroenterol. Hepatol. 13, 1141–1159 (2022).
Olm, M. R. et al. inStrain profiles population microdiversity from metagenomic data and sensitively detects shared microbial strains. Nat. Biotechnol. 39, 727–736 (2021).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Knight, J. C. Functional implications of genetic variation in non-coding DNA for disease susceptibility and gene regulation. Clin. Sci. 104, 493–501 (2003).
Bustos, A. Y., Font de Valdez, G., Fadda, S. & Taranto, M. P. New insights into bacterial bile resistance mechanisms: the role of bile salt hydrolase and its impact on human health. Food Res. Int. 112, 250–262 (2018).
Funabashi, M. et al. A metabolic pathway for bile acid dehydroxylation by the gut microbiome. Nature 582, 566–570 (2020).
Gueimonde, M., Garrigues, C., van Sinderen, D., de los Reyes-Gavilan, C. G. & Margolles, A. Bile-inducible efflux transporter from Bifidobacterium longum NCC2705, conferring bile resistance. Appl. Environ. Microbiol. 75, 3153–3160 (2009).
Smirnova, E. et al. Metabolic reprogramming of the intestinal microbiome with functional bile acid changes underlie the development of NAFLD. Hepatology 76, 1811–1824 (2022).
Pfeifer, E., Gatgens, C., Polen, T. & Frunzke, J. Adaptive laboratory evolution of Corynebacterium glutamicum towards higher growth rates on glucose minimal medium. Sci. Rep. 7, 16780 (2017).
Wannier, T. M. et al. Adaptive evolution of genomically recoded Escherichia coli. Proc. Natl. Acad. Sci. USA 115, 3090–3095 (2018).
Yu, T. et al. Reprogramming Yeast Metabolism from Alcoholic Fermentation to Lipogenesis. Cell 174, 1549–1558.e1514 (2018).
Li, X., Pei, G., Liu, L., Chen, L. & Zhang, W. Metabolomic analysis and lipid accumulation in a glucose tolerant Crypthecodinium cohnii strain obtained by adaptive laboratory evolution. Bioresour. Technol. 235, 87–95 (2017).
Li, J. et al. Advances in adaptive laboratory evolutionary engineering to microbial breeding. Sheng Wu Gong. Cheng Xue Bao 37, 130–141 (2021).
Jian, X. et al. Microbial microdroplet culture system (MMC): An integrated platform for automated, high-throughput microbial cultivation and adaptive evolution. Biotechnol. Bioeng. 117, 1724–1737 (2020).
Tenaillon, O. et al. The molecular diversity of adaptive convergence. Science 335, 457–461 (2012).
Vento, J. M., Crook, N. & Beisel, C. L. Barriers to genome editing with CRISPR in bacteria. J. Ind. Microbiol. Biotechnol. 46, 1327–1341 (2019).
Nethery, M. A., Hidalgo-Cantabrana, C., Roberts, A. & Barrangou, R. CRISPR-based engineering of phages for in situ bacterial base editing. Proc. Natl. Acad. Sci. USA 119, e2206744119 (2022).
Yan, Q. & Fong, S. S. Challenges and advances for genetic engineering of non-model bacteria and uses in consolidated bioprocessing. Front. Microbiol. 8, 2060 (2017).
Hong, W. et al. An inexpensive anaerobic chamber for the genetic manipulation of strictly anaerobic bacteria. Anaerobe 69, 102349 (2021).
Thompson, D. B. et al. The future of multiplexed eukaryotic genome engineering. ACS Chem. Biol. 13, 313–325 (2018).
Abdelrahman, M., Wei, Z., Rohila, J. S. & Zhao, K. Multiplex genome-editing technologies for revolutionizing plant biology and crop improvement. Front. Plant Sci. 12, 721203 (2021).
Jiang, S. et al. Gut microbiota drives structural variation of exogenous probiotics to enhance colonization. Gut Microbes 17, 2503371 (2025).
Barbaro, M. R. et al. Lactiplantibacillus plantarum (CECT7484 and CECT7485) and Pedioccoccus acidilactici (CECT7483) enhance actin cytoskeleton and CYP1A1 expression restoring epithelial permeability alterations induced by irritable bowel syndrome mediators. Gut Microbes 17, 2452235 (2025).
Ke, X. et al. Synbiotic-driven improvement of metabolic disturbances is associated with changes in the gut microbiome in diet-induced obese mice. Mol. Metab. 22, 96–109 (2019).
Haghikia, A. et al. Dietary fatty acids directly impact central nervous system autoimmunity via the small intestine. Immunity 43, 817–829 (2015).
Shen, S. et al. Understanding the “individual drug reaction” from the perspective of the interaction between probiotics and lovastatin in vitro and in vivo. Microbiome 11, 209 (2023).
Tanaka, H., Hashiba, H., Kok, J. & Mierau, I. Bile salt hydrolase of Bifidobacterium longum-biochemical and genetic characterization. Appl. Environ. Microbiol. 66, 2502–2512 (2000).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Vaser, R., Sovic, I., Nagarajan, N. & Sikic, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746 (2017).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Moriya, Y., Itoh, M., Okuda, S., Yoshizawa, A. C. & Kanehisa, M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 35, W182–W185 (2007).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Eberhardt, J., Santos-Martins, D., Tillack, A. F. & Forli, S. AutoDock Vina 1.2.0: new docking methods, expanded force field, and Python bindings. J. Chem. Inf. Model 61, 3891–3898 (2021).
Arab, J. P., Karpen, S. J., Dawson, P. A., Arrese, M. & Trauner, M. Bile acids and nonalcoholic fatty liver disease: Molecular insights and therapeutic perspectives. Hepatology 65, 350–362 (2017).
Blanco-Miguez, A. et al. Extending and improving metagenomic taxonomic profiling with uncharacterized species using MetaPhlAn 4. Nat. Biotechnol. 41, 1633–1644 (2023).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 32525049) awarded to J.Z.
Author information
Authors and Affiliations
Contributions
J.Z. and Z.S. conceptualized the research framework and formulated the experimental design. The experimental work and sample preparation were conducted by Z.H., X.L., D.Z., Q.G., L.Z., and S.S. Data analysis was performed by Z.H. in collaboration with Z.S. Z.H., Z.S., S.J., and Z.Z. contributed to figure preparation. The manuscript was drafted by Z.S. and Z.H., and J.Z. and Z.S. provided critical review and substantially refined the final version. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Qi Wang, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Han, Z., Sun, Z., Liu, X. et al. Harnessing a germ‑free mouse gut bioreactor for directed evolution of probiotics to combat non-alcoholic fatty liver disease. Nat Commun 17, 3133 (2026). https://doi.org/10.1038/s41467-026-69823-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69823-0