Abstract
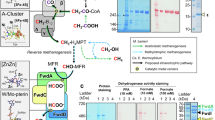
Bioelectrocatalytic CO2 reduction offers a sustainable route for CO2 bioconversion, yet remains limited by interfacial-intramolecular electron transfer and oxygen sensitivity. Here, we mine a formate dehydrogenase from Shewanella oneidensis MR-1 (SoFdhAB) featuring completely oxygen tolerant and direct-electron-transfer (DET) electrocatalytic performances. Cryo-electron microscopy (Cryo-EM) analysis reveals an intramolecular electron highway comprising five [4Fe-4S] clusters, a regional face-face contact facilitating interfacial ET, and a unique oxygen resistance mechanism different from inactivation-activation. By acquiring a beneficial variant SoFdhAB-Y94S, a direct bioelectrocatalytic CO2 reduction system is constructed, accumulating 2.88 ± 0.03 mmol formate in 64 hours with a steady rate of 45.3 ± 0.5 μmol h−1 cm−2 and a Faradaic efficiency of 93.1 ± 5.2%. The merits of oxygen tolerance and efficient (electro)catalytic property endow SoFdhAB a robust enzyme adopted in potential application scenarios, and the inherent DET capability may inspire the interfacial engineering of other oxidoreductases.
Similar content being viewed by others
Data availability
The structural model of SoFdhAB has been deposited in the Protein Data Bank (PDB) with accession codes 9VAP. Protein structures of EcFDH-H, PDB ID 1FDI, TkHDCR, PDB ID 7QV7 and DvH-FDH, PDB ID 6SDR are available from the PDB. The genomic data of Shewanella oneidensis MR-1 are available from the National Center for Biotechnology Information (NCBI) database [https://www.ncbi.nlm.nih.gov/nuccore/AE014299.2]. All relevant data generated and analyzed during this study which include enzyme activity assay and electrochemical data are included in this article and its supplementary information. Source data are provided with this paper.
References
Liu, Z., Wang, K., Chen, Y., Tan, T. & Nielsen, J. Third-generation biorefineries as the means to produce fuels and chemicals from CO2. Nat. Catal. 3, 274–288 (2020).
Gleizer, S. et al. Conversion of Escherichia coli to generate all biomass carbon from CO2. Cell 179, 1255–1263 (2019).
Cai, T. et al. Cell-free chemoenzymatic starch synthesis from carbon dioxide. Science 373, 1523–1527 (2021).
Zheng, T. T. et al. Upcycling CO2 into energy-rich long-chain compounds via electrochemical and metabolic engineering. Nat. Catal. 5, 388–396 (2022).
Cui, H. et al. Converting CO2 to single-cell protein via an integrated electrocatalytic-biosynthetic system. Appl. Catal. B Environ. 350, 123946 (2024).
Wu, R. et al. Enzymatic Electrosynthesis of glycine from CO2 and NH3. Angew. Chem. Int. Ed. 62, e202218387 (2023).
Fang, W. et al. Durable CO2 conversion in the proton-exchange membrane system. Nature 626, 86–91 (2024).
Contaldo, U. et al. Efficient electrochemical CO2/CO interconversion by an engineered carbon monoxide dehydrogenase on a gas-diffusion carbon nanotube-based bioelectrode. ACS Catal. 11, 5808–5817 (2021).
Cobb, S. J. et al. Fast CO2 hydration kinetics impair heterogeneous but improve enzymatic CO2 reduction catalysis. Nat. Chem. 14, 417–424 (2022).
Reda, T., Plugge, C. M., Abram, N. J. & Hirst, J. Reversible interconversion of carbon dioxide and formate by an electroactive enzyme. Proc. Natl. Acad. Sci. USA 105, 10654–10658 (2008).
Robinson, W. E., Bassegoda, A., Reisner, E. & Hirst, J. Oxidation-state-dependent binding properties of the active site in a Mo-containing formate dehydrogenase. J. Am. Chem. Soc. 139, 9927–9936 (2017).
Robinson, W. E., Bassegoda, A., Blaza, J. N., Reisner, E. & Hirst, J. Understanding how the rate of C–H bond cleavage affects formate oxidation catalysis by a Mo-dependent formate dehydrogenase. J. Am. Chem. Soc. 142, 12226–12236 (2020).
Bassegoda, A., Madden, C., Wakerley, D. W., Reisner, E. & Hirst, J. Reversible interconversion of CO2 and formate by a molybdenum-containing formate dehydrogenase. J. Am. Chem. Soc. 136, 15473–15476 (2014).
Li, W. et al. Direct detection of a single [4Fe-4S] cluster in a tungsten-containing enzyme: electrochemical conversion of CO2 into formate by formate dehydrogenase. Carbon Energy 5, e304 (2023).
Chen, H. et al. Fundamentals, applications, and future directions of bioelectrocatalysis. Chem. Rev. 120, 12903–12993 (2020).
Karyakin, A. A. et al. The limiting performance characteristics in bioelectrocatalysis of hydrogenase enzymes. Angew. Chem. Int. Ed. 46, 7244–7246 (2007).
Wang, Y., Kang, Z., Zhang, L. & Zhu, Z. Elucidating the interactions between a [NiFe]-hydrogenase and carbon electrodes for enhanced bioelectrocatalysis. ACS Catal. 12, 1415–1427 (2022).
Becker, J. M. et al. Bioelectrocatalytic CO2 reduction by redox polymer-wired carbon monoxide dehydrogenase gas diffusion electrodes. ACS Appl. Mater. Interfaces 14, 46421–46426 (2022).
Parkin, A., Seravalli, J., Vincent, K. A., Ragsdale, S. W. & Armstrong, F. A. Rapid and efficient electrocatalytic CO2/CO interconversions by carboxydothermus hydrogenoformans CO dehydrogenase I on an electrode. J. Am. Chem. Soc. 129, 10328–10329 (2007).
Hirst, J. Elucidating the mechanisms of coupled electron transfer and catalytic reactions by protein film voltammetry. Biochim. Biophys. Acta Bioenerg. 1757, 225–239 (2006).
Zu, Y., Fee, J. A. & Hirst, J. Breaking and re-forming the disulfide bond at the high-potential, respiratory-type rieske [2Fe-2S] center of thermus thermophilus: characterization of the sulfhydryl state by protein-film voltammetry. Biochemistry 41, 14054–14065 (2002).
Zu, Y. et al. Reduction potentials of rieske clusters: importance of the coupling between oxidation state and histidine protonation state. Biochemistry 42, 12400–12408 (2003).
Badiani, V. M. et al. Elucidating film loss and the role of hydrogen bonding of adsorbed redox enzymes by electrochemical quartz crystal microbalance analysis. ACS Catal. 12, 1886–1897 (2022).
Chemla, Y., Kaufman, F., Amiram, M. & Alfonta, L. Expanding the genetic code of bioelectrocatalysis and biomaterials. Chem. Rev. 124, 11187–11241 (2024).
Badiani, V. M. et al. Engineering electro- and photocatalytic carbon materials for CO2 reduction by formate dehydrogenase. J. Am. Chem. Soc. 144, 14207–14216 (2022).
Alvarez-Malmagro, J. et al. Bioelectrocatalytic activity of W-formate dehydrogenase covalently immobilized on functionalized gold and graphite electrodes. ACS Appl. Mater. Interfaces 13, 11891 (2021).
Liu, W. et al. Bioelectrocatalytic carbon dioxide reduction by an engineered formate dehydrogenase from Thermoanaerobacter kivui. Nat. Commun. 15, 9962 (2024).
Schwarz, F. M., Schuchmann, K. & Muller, V. Hydrogenation of CO2 at ambient pressure catalyzed by a highly active thermostable biocatalyst. Biotechnol. Biofuels 11, 237 (2018).
Dietrich, H. M. et al. Membrane-anchored HDCR nanowires drive hydrogen-powered CO2 fixation. Nature 607, 823–830 (2022).
Schuchmann, K. & Müller, V. Direct and reversible hydrogenation of CO2 to formate by a bacterial carbon dioxide reductase. Science 342, 1382–1385 (2013).
de Bok, F. A. et al. Two W-containing formate dehydrogenases (CO2-reductases) involved in syntrophic propionate oxidation by Syntrophobacter fumaroxidans. Eur. J. Biochem. 270, 2476–2485 (2003).
Böhmer, N., Hartmann, T. & Leimkühler, S. The chaperone FdsC for Rhodobacter capsulatus formate dehydrogenase binds the bis-molybdopterin guanine dinucleotide cofactor. FEBS Lett. 588, 531–537 (2014).
Iliuta, I. & Larachi, F. Direct-air capture conversion of CO2 in fixed-bed microreactors with immobilized formate dehydrogenase and glucose dehydrogenase: Concept feasibility. Chem. Eng. Res. Des. 193, 306–319 (2023).
Min, K. et al. Elevated conversion of CO2 to versatile formate by a newly discovered formate dehydrogenase from Rhodobacter aestuarii. Bioresour. Technol. 305, 123155 (2020).
Brouwer, E.-M. et al. Characterization of the oxygen-tolerant formate dehydrogenase from Clostridium carboxidivorans. Front. Microbiol. 15, 1527626 (2025).
Yoshikawa, T. et al. Multiple electron transfer pathways of tungsten-containing formate dehydrogenase in direct electron transfer-type bioelectrocatalysis. Chem. Commun. 58, 6478–6481 (2022).
Jeon, B. W. et al. Enzymatic conversion of CO2 to formate: the potential of tungsten-containing formate dehydrogenase in flow reactor system. J. CO2 Util. 82, 102754 (2024).
Cha, J., Lee, J., Jeon, B. W., Kim, Y. H. & Kwon, I. Real flue gas CO2 hydrogenation to formate by an enzymatic reactor using O2- and CO-tolerant hydrogenase and formate dehydrogenase. Front. Bioeng. Biotechnol. 11, 1265272 (2023).
Lee, J. et al. Molar-scale formate production via enzymatic hydration of industrial off-gases. Nat. Chem. Eng. 1, 354–364 (2024).
Fera, M. -C. et al. Bioelectrocatalytic CO2 reduction to formate by Candida boidinii formate dehydrogenase overcoming NADH dependence with tailored amino-viologen redox polymers. J. CO2 Util. 93, 103041 (2025).
Oliveira, A. R. et al. Toward the mechanistic understanding of enzymatic CO2 reduction. ACS Catal. 10, 3844–3856 (2020).
Kuk, S. K. et al. NADH-free electroenzymatic reduction of CO2 by conductive hydrogel-conjugated formate dehydrogenase. ACS Catal. 9, 5584–5589 (2019).
Oliveira, A. R. et al. An allosteric redox switch involved in oxygen protection in a CO2 reductase. Nat. Chem. Biol. 20, 111–119 (2024).
Yu, T. et al. Enzyme function prediction using contrastive learning. Science 379, 1358–1363 (2023).
Wang, Z. et al. Robust enzyme discovery and engineering with deep learning using CataPro. Nat. Commun. 16, 2736 (2025).
Li, Y. et al. Metabolic regulation of Shewanella oneidensis for microbial electrosynthesis: From extracellular to intracellular. Metab. Eng. 80, 1–11 (2023).
Brondino, C. D. et al. Incorporation of either molybdenum or tungsten into formate dehydrogenase from Desulfovibrio alaskensis NCIMB 13491: EPR assignment of the proximal iron-sulfur cluster to the pterin cofactor in formate dehydrogenases from sulfate-reducing bacteria. J. Biol. Inorg. Chem. 9, 145–151 (2004).
Maia, L. B., Fonseca, L., Moura, I. & Moura, J. J. G. Reduction of carbon dioxide by a molybdenum-containing formate dehydrogenase: a kinetic and mechanistic study. J. Am. Chem. Soc. 138, 8834–8846 (2016).
Cobb, S. J., Dharani, A. M., Oliveira, A. R., Pereira, I. A. C. & Reisner, E. Carboxysome-inspired electrocatalysis using enzymes for the reduction of CO2 at low concentrations. Angew. Chem., Int. Ed. 62, e202218782 (2023).
Yang, J. Y., Kerr, T. A., Wang, X. S. & Barlow, J. M. Reducing CO2 to HCO2− at mild potentials: lessons from formate dehydrogenase. J. Am. Chem. Soc. 142, 19438–19445 (2020).
Steinhilper, R., Höff, G., Heider, J. & Murphy, B. J. Structure of the membrane-bound formate hydrogenlyase complex from Escherichia coli. Nat. Commun. 13, 5395 (2022).
Page, C. C., Moser, C. C., Chen, X. X. & Dutton, P. L. Natural engineering principles of electron tunnelling in biological oxidation-reduction. Nature 402, 47–52 (1999).
Cellini, A. et al. Directed ultrafast conformational changes accompany electron transfer in a photolyase as resolved by serial crystallography. Nat. Chem. 16, 24 (2024).
Winkler, J. R. & Gray, H. B. Long-range electron tunneling. J. Am. Chem. Soc. 136, 2930–2939 (2014).
Kurisu, G. et al. Structure of the electron transfer complex between ferredoxin and ferredoxin-NADP+ reductase. Nat. Struct. Biol. 8, 117–121 (2001).
Volbeda, A., Darnault, C., Renoux, O., Nicolet, Y. & Fontecilla-Camps, J. C. The crystal structure of the global anaerobic transcriptional regulator FNR explains its extremely fine-tuned monomer-dimer equilibrium. Sci. Adv. 1, e1501086 (2015).
Cui, H. et al. Rapid and oriented immobilization of laccases on electrodes via a methionine-rich peptide. ACS Catal. 11, 2445–2453 (2021).
Boyington, J. C., Gladyshev, V. N., Khangulov, S. V., Stadtman, T. C. & Sun, P. D. Crystal structure of formate dehydrogenase H: catalysis involving Mo, molybdopterin, selenocysteine, and an Fe4S4 cluster. Science 275, 1305–1308 (1997).
Edwardes Moore, E. et al. Understanding the local chemical environment of bioelectrocatalysis. Proc. Natl. Acad. Sci. USA 119, e2114097119 (2022).
Nguyen, L. -T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2014).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, 78–82 (2024).
Jin, M. et al. Unique organizational and functional features of the cytochrome c maturation system in Shewanella oneidensis. PLoS ONE 8, e75610 (2013).
Melchor, S. & Dobado, J. A. CoNTub: An algorithm for connecting two arbitrary carbon nanotubes. J. Chem. Inf. Comput. Sci. 44, 1639–1646 (2004).
Li, C. & Chou, T. -W. Elastic moduli of multi-walled carbon nanotubes and the effect of van der Waals forces. Compos. Sci. Technol. 63, 1517–1524 (2003).
Wang, H., Michielssens, S., Moors, S. L. C. & Ceulemans, A. Molecular dynamics study of dipalmitoylphosphatidylcholine lipid layer self-assembly onto a single-walled carbon nanotube. Nano Res. 2, 945–954 (2009).
Zhang, M. et al. Engineering a binding peptide for oriented immobilization and efficient bioelectrocatalytic oxygen reduction of multicopper oxidases. ACS Appl. Mater. Interfaces 17, 2355–2364 (2025).
Edwardes Moore, E. et al. A Semi-artificial photoelectrochemical tandem leaf with a CO2-to-formate efficiency approaching 1%. Angew. Chem. Int. Ed. 60, 26303 (2021).
Miller, M. et al. Interfacing formate dehydrogenase with metal oxides for the reversible electrocatalysis and solar-diven reduction of carbon dioxide. Angew. Chem. Int. Ed. 58, 4601–4605 (2019).
Acknowledgements
This study was supported by Strategic Priority Research Program of the Chinese Academy of Sciences (XDC0120103 to L. Z.), Coal-Major Project (2025ZD1701600 to L. Z.), the Tianjin Science Fund for Distinguished Young Scholars (22JCJQJC00100 to Z. Z.), and the International Partnership Program of Chinese Academy of Sciences, Grant No. 306GJHZ2025003BS to L. Z. We are grateful to Professor Shenghai Chang for his supervision in structural analysis, and to Dr. Tailin Wang for insightful discussions.
Author information
Authors and Affiliations
Contributions
Experiment, data analysis and writing the initial manuscript were performed by W.L. P.Z. preformed the bioinformatics analysis. Molecular dynamics (MD) simulation was performed by X.W. and H.C. (Haiyang Cui) provided supervision. Cryo-EM analysis was supervised by K.Z., W.Y., and C.Y. Electrochemical experiments were supervised by H.C. (Huijuan Cui) Data analysis and biochemistry experiments were supervised by C.Y., Z.Z., and J.S. J.L. provide experimental materials. Project conception, fund support, overall supervision and writing paper were performed by L.Z.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Cunduo Tang, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, W., Zhang, P., Wang, X. et al. An interfacial-intramolecular electron highway for accelerated electrocatalytic CO2 reduction by an O2-tolerant formate dehydrogenase. Nat Commun (2026). https://doi.org/10.1038/s41467-026-69827-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-69827-w