Abstract
Polar chromosomes, which initially attach to the mitotic spindle behind the pole, are prone to missegregation and micronuclear entrapment, contributing to chromosomal instability in cancer. Yet, the mechanisms ensuring their faithful segregation remain unclear. Here, we show that polar chromosomes require a unique step involving spindle elongation, which repositions chromosome-bound astral microtubules by pivoting them around the centrosome toward the spindle surface. By modulating Eg5/KIF11 activity, we demonstrate that spindle elongation determines the direction and extent of pivoting, with microtubules from the opposite spindle half facilitating final movement. Kinetochores on polar chromosomes form mono-lateral attachments, recruiting corona components and partially Mad2, but lacking Astrin. In cancer cell lines, limited spindle elongation delays polar chromosome resolution, whereas enhanced elongation accelerates it. These findings highlight the role of spindle elongation in the timely rescue of chromosomes from the “danger zone” behind the pole, providing mechanistic insight into chromosome congression errors in cancer.
Similar content being viewed by others
Introduction
Chromosomes must efficiently congress to the equatorial plane of the spindle to ensure proper segregation1. Based on their position at the nuclear envelope breakdown (NEBD), chromosomes undertake different pathways toward the equator, resulting in distinct congression efficiencies and segregation error rates2. Chromosomes behind the spindle poles are particularly at risk of missegregation, as they are shielded by the pole and require a multi-step process to reach the equator3,4. First, these chromosomes approach the spindle pole from the back through centripetal movement driven by kinetochore dynein pulling along astral microtubules and nuclear actomyosin3,5,6,7,8. Subsequently, the chromosomes travel along the main spindle surface using CENP-E at kinetochores, pulling forces by end-on attached microtubules, and polar ejection forces3,9,10,11,12,13,14,15,16. However, the mechanism by which polar chromosomes traverse the pole and shift from the astral region to the main spindle remains unknown.
Chromosomes that fail to escape the area behind the pole before the spindle fully forms risk stabilization of improper attachments, ensheathment by the endoplasmic reticulum, and missegregation in case of the spindle assembly checkpoint (SAC) weakness, bypass or delayed satisfaction4,17,18,19,20. Recent studies discovered that such unaligned (i.e., behind the spindle pole) and misaligned (i.e., near the pole on the main spindle) chromosomes represent an important source of segregation errors, with the highest probability of ending up in micronuclei, the strongest correlation with chromosomal instability, and a direct link to metastatic potential19,21. Several labs have observed mis- and unaligned chromosomes in cancer cell lines, such as ovarian, bone, colorectal and breast cancer, and in patient samples of primary and metastatic breast cancer19,21,22,23. Multiple lines of evidence also indirectly suggest that chromosome 1, whose 1q arm gain directly leads to cancer development and malignant transformation, might often be located behind the pole and thus at risk of unalignment4,24,25,26. While these implications highlight the importance of escaping the polar region on time, the reason why chromosomes occasionally fail to do so, especially in cancer cells, is still unclear.
Here, we develop approaches integrating stimulated emission depletion (STED), lattice light-sheet (LLS), and confocal microscopy with molecular perturbations to overcome spatiotemporal resolution barriers that hinder studies of polar chromosome dynamics, which occurs quickly within a spindle region that is densely populated with microtubules. We present a mechanism crucial for chromosome segregation, based on a fundamental, yet previously unexplored, property of centrosome-anchored microtubules—their ability to pivot around the centrosome. Our study reveals that spindle elongation prompts microtubules attached to polar chromosomes to pivot towards the center of the spindle, releasing the chromosomes from the “danger zone” behind the spindle pole and enabling their contact with the spindle surface. During pivoting, polar chromosomes form predominantly mono-lateral and other complex attachments with astral microtubules, which persist until the chromosome contacts the spindle surface and contribute to spindle assembly. From a functional perspective, we demonstrate a critical role of spindle elongation-driven pivoting in promoting timely mitosis and preventing chromosome alignment errors in cancer cells, suggesting potential strategies to modulate these errors.
Results
Polar chromosomes require a unique congression step to transit across the spindle pole
Given that chromosomes located behind the spindle pole at NEBD take longer to congress to the metaphase plate in comparison to other chromosomes4, we hypothesized that they require a specific congression step. Alternatively, they may exhibit slower progression of the steps shared with other chromosomes. To distinguish between these possibilities, we imaged human retinal pigment epithelium (RPE1) cells with labeled centromeres and either tubulin or centrosomes using a synergy of STED immunofluorescence (Fig. 1a) and LLS live-cell imaging (Fig. 1b). STED microscopy enabled visualization of microtubules within the polar region in super-resolution (50–100 nm, Supplementary Fig. 1a)27. This level of resolution was essential for distinguishing individual microtubules or small bundles together with their attachments to kinetochores within a microtubule-packed region, considering that human spindles contain about 6000 microtubules28. LLS microscopy enabled imaging of all kinetochores throughout mitosis at a time resolution of 6 s due to the low phototoxicity of this technique29. The high time resolution was crucial for tracking individual polar kinetochores, as they move quickly and pass closely by other kinetochores (Supplementary Movie 1).
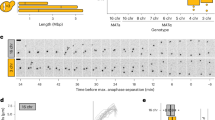
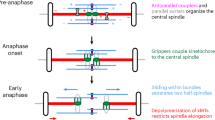
a STED superresolution images of microtubules immunostained for α-tubulin (white) in RPE1 cells stably expressing CENP-A-GFP (colorful, confocal). Images show maximum intensity projections of the whole cell. Kinetochores are color-coded for depth with the brgbcmyw LUT in ImageJ throughout 8–12 z-planes in prometaphase and 35 z-planes in metaphase, corresponding to 2.4–3.6 µm and 10.5 µm, respectively. The images represent different phases of mitosis: I – nuclear envelope breakdown (NEBD), II – a spindle starts forming and kinetochores in the polar region start their centripetal movement towards the spindle pole from the back, III – a prometaphase ring forms and centripetal movement of kinetochores ends, IV – a spindle rapidly elongates while kinetochores located at the back of the pole pass the polar region, V – spindle elongation ends and all kinetochores land onto the spindle surface, VI – a spindle is fully elongated and kinetochores travel towards the spindle equator. These stages were inferred from the arrangements of microtubules and kinetochores, see “Methods”. The polar region and the main spindle surface are denoted in image II. b Time-lapse images of RPE1 cells stably expressing CENP-A-GFP and Centrin1-GFP (colorful, lattice light-sheet (LLS)). Images show maximum intensity projections of the whole cell. Kinetochores are color-coded for depth with the brgbcmyw LUT in ImageJ throughout 32–40 z-planes, corresponding to 4.6–5.8 µm. Mitotic phases from I to VI correspond to the ones described in a. The polar region is denoted as a region behind two dashed lines perpendicular to the long spindle axes. Centrosomes are denoted with white circles. White arrows represent one polar kinetochore pair tracked from NEBD until alignment to the spindle equator. c Polar chromosome-centered (upper) and spindle-centered (lower) timelines of prometaphase events. Events are marked with roman numerals that correspond to the ones shown in a, b. The times are calculated based on 39 kinetochore pairs in 20 cells from 8 experiments. d Schematic representation of the first (end of centripetal movement) and the last (landing) known time points between which the time gap and possible passage across the polar region occur. e Three possible models for passing the spindle pole. In the first model, a motor bound to polar kinetochores brings them onto the spindle surface, representing diverse mechanisms based on kinetochore- and chromosome-bound motors. The second model proposes that spindle elongation drives pivoting of astral microtubules that consequently bring polar kinetochores in front of the spindle pole. In the last model, polar kinetochores attach to microtubules emanating from the opposite spindle half that bring them onto the spindle surface. All scale bars, 2 µm.
By using LLS movies (Fig. 1b), we found that at NEBD, 15 ± 2 chromosomes per cell (all data are given as mean ± SEM, n = 24 cells) were located behind the spindle poles (Supplementary Fig. 1b–d). Out of those 15 chromosomes, 8 ± 1 ended their centripetal movement in the region between the two poles, whereas 7 ± 1 chromosomes ended their centripetal movement in the region behind the pole (see examples in Fig. 1a and Supplementary Fig. 1b). We refer to the latter group as polar chromosomes. By constructing a timeline of the polar chromosome movements (Fig. 1b, c, see individual stages inferred from the arrangements of microtubules and kinetochores in Fig. 1a), we found that after NEBD a typical polar chromosome started its centripetal movement approaching the spindle pole from the back until it reached a relatively stable distance from the pole at 3.4 ± 0.2 min, which roughly corresponds to the formation of the prometaphase chromosome ring around the nascent spindle30. Remarkably, there was a 4-min time gap before polar chromosomes reached the spindle surface, an event we term “landing”, which occurred 7.4 ± 0.3 min after NEBD (Fig. 1b, c). During this time gap, the spindle rapidly elongated and other chromosomes have already started congressing to the metaphase plate. Once the polar chromosomes landed on the spindle surface, they immediately started traveling towards the spindle midplane, finishing aligning at 12.3 ± 0.3 min (Fig. 1b, c, and Supplementary Fig. 1e).
The observed time gap may be specific to the location of polar chromosomes behind the pole in prometaphase, but it may also be due to their large distance to the spindle midplane. To discriminate between these possibilities, we compared the timing of polar and non-polar chromosomes with a similar initial distance to the centrosome midpoint (Supplementary Fig. 1f), and found that only polar chromosomes showed a delay in landing and a time gap from the end of the centripetal movement to the attachment to the spindle surface, indicating that the gap is specific to chromosome location (Supplementary Fig. 1g, h). The other segments of the chromosome’s travel, the centripetal movement and the transfer along the spindle surface, lasted similarly for polar and non-polar chromosomes (Supplementary Fig. 1i, j), which argues against polar chromosomes having slower progression of shared steps. Thus, the delayed alignment of polar chromosomes causing their susceptibility to missegregation is due to a time gap wherein the polar chromosome traverses across the pole to reach the main spindle surface. This reveals an additional step in congression specific to polar chromosomes (Fig. 1d).
To test whether this additional step includes the attachments to microtubules, we analyzed STED images (Fig. 1a) and found that 58.1 ± 3.2% of chromosomes already had microtubules next to their kinetochores at a time close to NEBD, whereas 41.9 ± 3.2% were completely unattached (Supplementary Fig. 1k, l). By the end of the centripetal movement phase, all kinetochore pairs were attached to microtubules and remained attached throughout the subsequent phases (Supplementary Fig. 1k, l). In principle, the chromosomes that were initially located behind the pole might have remained unattached as they were passed by the moving centrosome, resulting in their first contact with spindle microtubules between the spindle poles. However, we found that polar chromosomes, i.e., those that made their first contact behind the pole, were attached to microtubules during the transit across the pole (Fig. 1a, and Supplementary Fig. 1k, l). Thus, even though these data are based on fixed cells, they suggest that pole crossing by the polar chromosomes involves the attachment of kinetochores to microtubules.
Passage across the pole occurs independently of the motor proteins implicated in chromosome congression
To uncover the mechanism of the passage across the polar region, we envisioned three models (Fig. 1e). In the first model, motor proteins located at the kinetochore (CENP-E or dynein) or on the chromosome arm (chromokinesins) of polar chromosomes attach to the microtubules of the main spindle surface and pull the polar chromosome in the direction of the spindle surface. In the second model, the spindle elongates, causing both poles to move outward, thereby repositioning chromosomes from the back of the spindle pole to the main spindle surface. In the third model, microtubules extending from the opposite side of the spindle attach to the polar chromosome and pull it towards the main spindle surface.
To test the first model, we depleted CENP-E (kinesin-7)1,3,9,11, Spindly, which promotes dynein recruitment to kinetochores in early mitosis31, or co-depleted the chromokinesins KIF4A (kinesin-4) and Kid (kinesin-10)15,32,33 by using small interfering RNAs (siRNAs) (Supplementary Fig. 2a). As an alternative approach to disrupt CENP-E activity, we used the small-molecule inhibitor GSK92329534 (Supplementary Fig. 2a). We also perturbed actomyosin contraction5 by depolymerizing actin via Latrunculin A (Supplementary Fig. 2a), thereby completing the test of motor-driven mechanisms that were previously implied in chromosome congression1. Even when those proteins were effectively depleted or inhibited, as determined by a ~90% reduction of their signal, absence from the expected location, and the characteristic phenotype (Supplementary Fig. 2b, c), nearly all polar chromosomes successfully traveled from the back of the spindle towards the main spindle surface (Supplementary Fig. 2d). A tiny fraction (~2%) of polar chromosomes after Spindly depletion remained behind the pole, as they were ejected more than 6 µm away from the pole into the region beyond the future area of the spindle (Supplementary Fig. 2d, e). In contrast, in control cells, polar kinetochores approached the pole and remained at a distance of about 3 µm, indicating the importance of proximity to the pole for successful passage across the pole (Supplementary Fig. 2e). Given that Spindly and CENP-E are essential for the expansion of the kinetochore fibrous corona35,36,37, these data show that the expanded corona is dispensable for the initial chromosome transit across the pole. Taken together, these results show that the passage across the pole occurs independently of the previously described congression drivers, but benefits from the role of dynein in keeping the chromosomes close to the spindle pole.
Pivoting of astral microtubules around the centrosome repositions polar chromosomes close to the spindle surface
In the second model of polar chromosome resolution (Fig. 1e), spindle elongation leads to repositioning of polar chromosomes attached to astral microtubules towards the main spindle surface. This model relies on two assumptions: (1) chromosome-attached microtubules are able to pivot around the centrosome, (2) chromosomes move through the cytoplasm considerably less than the centrosomes. To explore these assumptions, we tracked in 3D the polar kinetochores and their microtubules, as well as centrosomes and chromosome arms, from the end of the centripetal movement until after landing on the spindle surface (Fig. 2a–c, Supplementary Fig. 2f, and Supplementary Movies 2–4). To quantify the pivoting motion of a polar chromosome, we measured the angle between the centrosome-centrosome axis and the line connecting the sister kinetochores midpoint with the centrosome, termed “pivoting angle” (Fig. 2d). The pivoting angle showed a fast decrease during the period of rapid spindle elongation and stabilized as the spindle length stabilized (Fig. 2e, f). Moreover, the pivoting angle was strongly anti-correlated with the spindle length for individual kinetochore pairs (Fig. 2g, and Supplementary Fig. 2g). These results indicate that polar kinetochores, along with their attached microtubules, pivot around the centrosome towards the spindle center.
a Upper panel: time-lapse images of RPE1 cells stably expressing EYFP-tubulin (white, confocal) and H2B-mRFP (not imaged) with added HaloTag-CENP-A (blue, confocal). Each image represents a single central z-plane and was rotated and cropped to align the spindle horizontally and position it in the center of the frame. The orange arrow denotes the kinetochore pair attached to the pivoting astral microtubule. Lower panel: Schematic representation of pivoting (red arrow) based on the images from the upper panel, showing the current and previous microtubule positions. b Time-lapse images of RPE1 cells stably expressing EYFP-tubulin (white, confocal) and H2B-mRFP (not imaged) with added HaloTag-CENP-A (blue, confocal). Moving and stagnant spindle poles are denoted in the first image. The blue arrow represents the microtubule attached to the polar kinetochore (KC) tracked over time. Yellow and orange arrows represent microtubules without attached polar kinetochores on the moving and stagnant spindle pole, respectively. Insets and schematic representations of the previously described microtubule types are shown below the images of the whole spindle; spindle poles are depicted as white circles. Images are maximum intensity projections of 3 central z-planes, corresponding to 3 µm. c Images of RPE1 cells stably expressing EB3-GFP (colorful, confocal, upper image) and H2B-mCherry (colorful, confocal, lower image). Images are temporal maximum intensity projections of the centrosome (upper) and chromosome (lower) movements over 3 central z-planes (3 µm) and a time period of 3.3 min, color-coded for time with the Spectrum LUT in ImageJ. Spindle pole movement is depicted with white arrow. d Schematic representation of measured parameters for polar kinetochores along with the starting (end of centripetal movement) and last (landing) tracking point (t = 0 min). Spindle length was determined as the distance between the two spindle poles. The pivoting angle was determined as an angle that the midpoint of the kinetochore pair closes with the long spindle axis. e The pivoting angle of polar kinetochore pairs and f corresponding spindle length in time. The landing point of each kinetochore pair is set to zero. Values are shown as mean (dark line) and SEM (shaded areas). N = 39 kinetochore pairs in 20 cells from 8 experiments. g Anticorrelation of the pivoting angle and spindle length for the same polar kinetochores as in e. Each color represents one polar kinetochore pair and each dot represents its pivoting angle and the corresponding spindle length in time. Linear regression. N = 39 kinetochore pairs in 20 cells from 8 experiments. h The pivoting angle of the microtubule attached to the kinetochores on the moving pole (blue), microtubule without attached kinetochores on the moving pole (yellow) and microtubule without attached kinetochores on the stagnant pole (orange) in time. Values are shown as mean (dark line) and SEM (shaded areas). N = 16 microtubules with polar kinetochores attached to them on the moving pole in 10 cells from 4 experiments. N = 16 microtubules without polar kinetochores attached to them on the moving pole in 10 cells from 4 experiments. N = 14 microtubules without polar kinetochores attached to them on the stagnant pole in 6 cells from 4 experiments. The fourth group, microtubules with polar kinetochores attached to them on the stagnant pole, could not be found, as poles were stagnant only when they were already at the nuclear border at NEBD and, therefore, had no polar chromosomes to begin with. i Schematic representation of the spindle pole movement and pivoting angle for each microtubule marked in b in time (from red to blue). Note that only the microtubule attached to polar kinetochores pivots around the spindle pole. j Schematic representation of the centrosome, astral microtubule and chromosome movements during pivoting. Note that the centrosome moves significantly while microtubules pivot under the weight of a chromosome which moves only slightly. All scale bars, 2 µm.
To directly see the pivoting motion of astral microtubules, which are undetectable in typical videos of mitosis, we developed a confocal microscopy protocol with adjusted pixel size (See “Methods”) to image RPE1 cells expressing EYFP-α-tubulin with added HaloTag-CENP-A (Fig. 2a, Supplementary Movies 2 and 3). The kinetochores of a polar chromosome were found to be adjacent to a microtubule signal representing a single astral microtubule, or a few in a bundle. We found that this astral microtubule pivoted around the centrosome in the direction of the spindle surface, while maintaining its connection to the polar chromosome, during the centrosome’s movement outwards (Fig. 2a, h, i). The microtubules remained straight instead of bending (Supplementary Fig. 2h), indicating that the entire microtubule pivots around the centrosome, as opposed to solely the plus-end-proximal segment gliding along the cell cortex or organelles. Overall, these direct observations of microtubule pivoting support the second model and indicate that the force driving the pivoting of chromosome-attached microtubules exceeds the force preserving their extension direction at the centrosome.
In contrast to the chromosome-carrying astral microtubules, the microtubules that were not attached to a chromosome did not exhibit directed pivoting but instead pivoted around the centrosome in a random manner (Fig. 2b, h, i, and Supplementary Fig. 2i, j). Their pivoting was similar in the cases when their centrosome moved significantly and when it appeared stationary (Fig. 2b, h, i, and Supplementary Fig. 2i–k). Stationary poles were observed in cells where one pole was positioned at the nuclear border during NEBD, resulting in the absence of polar chromosomes and the presence of only chromosome-free microtubules at this pole. Whereas chromosome-carrying microtubules pivoted by ~36°, on average, in 3 min, the chromosome-free microtubules changed their angle by less than ~8° irrespective of the pole movement (Supplementary Fig. 2i). These findings suggest that the force driving the pivoting of chromosome-free microtubules is independent of the centrosome movement and smaller than the force maintaining their direction at the centrosome.
For spindle elongation to reposition polar chromosomes towards the main spindle surface, chromosomes must move less than centrosomes during this process. This was confirmed by temporal maximum-intensity projections of spindles labeled with EB3-GFP and H2B-mCherry, which showed that chromosomes remained relatively stationary while the centrosomes moved outward (Fig. 2c, and Supplementary Movie 4). Collectively, these findings support the model in which spindle elongation induces pivoting of astral microtubules attached to stationary polar chromosomes (Fig. 2j), enabling their interaction with the spindle surface.
Direction of the centrosome movement dictates the direction of microtubule pivoting
The pivoting model provides a straightforward prediction: if the spindle dynamics reverse from elongation to shortening, the direction of microtubule pivoting should also reverse, implying that microtubules should pivot towards the region behind the spindle pole instead of towards the main spindle surface. Similarly, a constant spindle length should be associated with negligible pivoting. To test these predictions, we designed a set of manipulations of Eg5/KIF11 (kinesin-5), the main motor protein driving spindle elongation38,39, to either reverse, restore, or block spindle elongation (Fig. 3a–d, and Supplementary Movies 5 and 6).
a-d Time-lapse images of RPE1 cells stably expressing CENP-A-GFP and Centrin1-GFP (colorful, confocal) for a DMSO control, b STLC (S-Trityl-L-Cysteine)-, c FCPT (2-(1-(4-fluorophenyl)cyclopropyl)-4-(pyridin-4-yl)thiazole)-, and d monastrol-treated cells. White arrows represent one polar kinetochore (KC) pair over time. White circles represent spindle poles, horizontal dashed lines represent the long spindle axis and lines perpendicular to it denote the borders of the polar region. Note that the STLC treatment caused spindle shortening, FCPT treatment blocked spindle elongation, whereas monastrol treatment caused spindle shortening that could be reversed by washout. Kinetochores are color-coded for depth with the brgbcmyw LUT in ImageJ throughout 14–21 z-planes, corresponding to 5.6–8.4 µm. e Spindle length and f pivoting angle in time for polar kinetochores in control (gray), STLC (yellow) and FCPT (blue) treatment. Values are shown as mean (dark line) and SEM (shaded areas). t = 0 min corresponds to inhibitor addition and activity. N = 7 kinetochore pairs in 7 cells from 7 experiments (DMSO), N = 5 kinetochore pairs in 3 cells from 3 experiments (STLC), N = 9 kinetochore pairs in 7 cells from 7 experiments (FCPT). g Spindle length and h pivoting angle in time during the monastrol washout experiment, for the spindle and kinetochore pair marked in d. Dashed lines denote the time required for the addition or washout of monastrol together with the time required for the spindle length to start changing. i Number of chromosomes behind the pole at the indicated times in the monastrol washout experiment for Cell 1 (shown in (d)) and Cell 2 (shown in Supplementary Fig. 3d). j The experimental correlation of the pivoting angle change and spindle length change for the control (orange), DMSO control (dark green), monastrol washout (light green), FCPT (blue), monastrol (purple) and STLC (yellow) treatments along with the theoretical curve calculated based on the experimental parameters from DMSO and STLC experiments (See Supplementary Fig. 3). Linear regression. k Schematic representation of pivoting angle change based on the spindle length change. All scale bars, 2 µm.
To reverse spindle elongation into shortening, we added the Eg5 inhibitor STLC (S-Trityl-L-Cysteine)40 during prometaphase at a point when the spindle had formed, and the kinetochores of polar chromosomes approached the main spindle surface but had not yet attached to it. After the addition of STLC, the spindle started shortening, and the kinetochores of the polar chromosomes that were already close to the spindle surface started pivoting towards the back of the spindle (Fig. 3b, and Supplementary Movie 5). This reversed pivoting was evident as an increase in the pivoting angle during spindle shortening in STLC-treated cells, in contrast to the decrease of the pivoting angle during spindle elongation in untreated cells (Fig. 3e, f, and Supplementary Fig. 3a–c). Remarkably, the pivoting direction was reversible, moving back and forth (Fig. 3d, Supplementary Fig. 3d, and Supplementary Movie 6). The washout of the Eg5 inhibitor monastrol41 reversed the inhibitor-induced spindle shortening into elongation, which triggered kinetochore pivoting towards the main spindle surface (Fig. 3g, h). The number of kinetochore pairs behind the pole decreased before monastrol addition, increased during monastrol treatment, and decreased again after monastrol washout (Fig. 3i), in agreement with the predictions from the model.
To block spindle elongation, we added the ATP-competitive Eg5 inhibitor FCPT (2-(1-(4-fluorophenyl)cyclopropyl)-4-(pyridin-4-yl)thiazole)42,43, which induces tight binding of Eg5 onto microtubules. Subsequently, the spindle length remained constant, and the kinetochores of polar chromosomes did not pivot significantly, providing further support to the model (Fig. 3c, e, f, Supplementary Fig. 3a, b, e, Supplementary Movie 5). Finally, to exclude the possibility that the observed effects of Eg5 inhibitors were due to roles of Eg5 other than driving spindle elongation, we employed an orthogonal approach. This involved adding a low dose of nocodazole (20 nM), which induces mild spindle shortening after NEBD while preserving the attachments of chromosomes to astral microtubules44. In agreement with the results from Eg5 inhibition, spindle shortening caused an increase in the number of kinetochores behind the pole (Supplementary Fig. 3f–h). Collectively, these data show that the centrosome movement due to spindle elongation or shortening dictates the direction of kinetochore pivoting.
To quantitatively test our model, we explored whether the extent of pivoting can be explained by the extent of spindle elongation or shortening. A strong correlation between the change in spindle length and the simultaneous change in the pivoting angle supports this notion (Fig. 3j). Moreover, we calculated the expected change in the pivoting angle for a given change in the spindle length, assuming immobile chromosomes (see “Methods” and Supplementary Fig. 3i). The resulting theoretical prediction closely matched the linear fit of the data (Fig. 3j). Taken together, our results reveal spindle elongation as a necessary and sufficient driver of chromosome passage across the spindle pole, dictating both the direction and magnitude of pivoting of chromosome-carrying astral microtubules (Fig. 3k).
Polar chromosomes form complex attachments yet remain bound to the same microtubule during pivoting
Prior to establishing end-on attachments, kinetochores initially bind to microtubules laterally14,30,45. To explore the types of attachments that polar kinetochores form with astral microtubules during their pivoting, and to determine if they transition between different microtubules, we imaged microtubules by STED microscopy in fixed RPE1 cells expressing CENP-A-GFP, at various stages between the end of chromosome centripetal movement and landing (Fig. 4a). Surprisingly, rather than primarily observing lateral attachments, we found a wide variety of attachment types, including lateral, monotelic, and syntelic configurations, together with more complex and previously unidentified types (Fig. 4a). We define them as: (1) bilateral, when two sister kinetochores attach laterally to two astral microtubules, (2) mono-lateral, when one sister kinetochore has an end-on attachment in addition to both sisters being laterally attached to the same astral microtubule, (3) mono-bilateral, as a combination of bilateral and one end-on attachment, and (4) syn-lateral, as a combination of syntelic and one lateral attachment. Mono-lateral attachments were the most frequent, followed by lateral and bilateral ones (Fig. 4b).
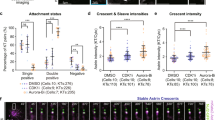
a STED superresolution images of microtubules immunostained for α-tubulin (white) in RPE1 cells stably expressing CENP-A-GFP (colorful, confocal). Images show maximum intensity projections of the whole cell. Kinetochores (KCs) are color-coded for depth with the brgbcmyw LUT in ImageJ throughout 8–17 z-planes, corresponding to 2.4–5.1 µm. White arrows represent a kinetochore pair attached to microtubules shown in insets below the image. Insets show single z-planes or maximum intensity projections of 2–3 z-planes where the microtubule of interest is located, smoothed with 0.75-mm-sigma Gaussian blur. The schematic representations of different attachment types are shown next to the associated insets. b Percentages of different attachment types for polar chromosomes between the end of the centripetal movement and landing quantified from STED images. N = 60 kinetochore pairs in 24 cells from 5 experiments. c Pie chart for the number of microtubule changes during pivoting measured in live RPE1 cells stably expressing EYFP-tubulin with added HaloTag-CENP-A (See Supplementary Fig. 4). N = 19 kinetochore pairs in 10 cells from 4 experiments. d Pie chart for attachment changes on further (yellow), closer (blue) and both (green) polar kinetochores measured in the same cells as in c. N = 13 kinetochore pairs in 5 cells from 3 experiments. e Schematic representations of types of attachment changes that kinetochores (blue circles) of polar chromosomes (chrom.) undergo during pivoting, along with their frequencies. The orange arrows denote which kinetochore in a pair changes microtubules during pivoting. All scale bars, 2 µm.
To explore if chromosomes can switch from one microtubule to another during pivoting, we imaged live cells expressing EYFP-α-tubulin and H2B-mRFP with HaloTag-CENP-A (Supplementary Fig. 4b). We found that 63.1 ± 11.1% of polar chromosomes remained attached to a single microtubule during pivoting, 21.1 ± 9.4% made one change between microtubules, whereas multiple changes were rare (Fig. 4c). Most of the observed changes included only a change for the further kinetochore, whereas the closer one remained attached to the same microtubule during the entire process (Fig. 4d, and Supplementary Fig. 4b). Thus, in a typical case with 7 polar chromosomes, 4 of them remain attached to a single microtubule during pivoting, 2 change the attachment of the further kinetochore, e.g., from lateral to bilateral, whereas 1 shows a more complex transition (Fig. 4e). In sum, polar chromosomes make a variety of attachments with astral microtubules but typically remain attached to the same microtubules while crossing the polar region.
Kinetochores on polar chromosomes show molecular signatures of lateral and immature end-on attachments
To characterize the molecular nature of polar kinetochore-microtubule attachments together with STED imaging of microtubules during intact early mitosis, we tested the levels of Mad2, a checkpoint protein that marks kinetochores with missing or unstable end-on attachments46,47. Among sister kinetochores, the one positioned further from the pole contained higher levels of Mad2 (Fig. 5a, b and Supplementary Fig. 4a). This suggests that the further kinetochore typically forms a lateral attachment, whereas the closer one forms an end-on attachment in addition to the lateral one, in agreement with microtubule localization observed by STED microscopy (Fig. 5a). Mad2 levels at kinetochores with end-on attachments (Fig. 5a) were lower than at those with lateral attachments, as expected, but higher than at end-on attached kinetochores in metaphase (Fig. 5c), suggesting that the end-on attachments on polar chromosomes are immature.
a STED superresolution images of polar chromosomes attached to microtubules immunostained for α-tubulin (white) in RPE1 cells stably expressing CENP-A-Cerulean (blue) and Mad2-mRuby (yellow). Images show a single z-plane, with kinetochores and microtubules smoothed with 0.75-mm-sigma Gaussian blur. Intensities of CENP-A and Mad2 are equally adjusted throughout the images. b Univariate scatter plot of the relative Mad2/CENP-A signal for closer (blue) and further (yellow) polar kinetochore (KC). Measurements were done on STED images as from a. N = 42 kinetochore pairs in 29 cells from 3 experiments. Statistical test: two-sided Mann–Whitney U test. c Univariate scatter plot of the relative Mad2/CENP-A signal for closer polar kinetochore in metaphase (blue), in prometaphase when the kinetochore makes an end-on attachment (orange) and in prometaphase when the kinetochore makes a lateral attachment (yellow). Measurements are done on STED images as from a. N = 10 kinetochores from 5 kinetochore pairs in 2 cells from 1 experiment for metaphase, N = 26 end-on attached kinetochores and 12 laterally attached kinetochores in 29 cells from 3 experiments for prometaphase. Statistical test: Kruskal–Wallis with post-hoc Dunn’s test. d–j Confocal images of mitotic spindles in early prometaphase cells expressing CENP-A-GFP and centrin-1-GFP (cyan, d, g) or CENP-A-GFP alone (cyan, i), immunostained for Astrin (yellow, d) and Mad2 (not shown, d), CENP-E (yellow, g), and ZW10 (yellow, i), smoothed with 1-mm-sigma Gaussian blur. Individual fluorescence channels are shown on the right, along with the merged enlargements of the polar region and the corresponding individual channels. Circles denote centrosome position and arrowheads point to polar kinetochores. Univariate scatter plots of mean Astrin (e), Mad2 (f), CENP-E (h), and ZW10 (j) signal intensities, normalized to mean CENP-A levels at kinetochores, for the specified spindle positions and mitotic phases. Note: Mad2 levels at closer polar and non-polar kinetochores are intermediate compared to those at further polar and aligned kinetochore pairs. N = 116 kinetochore pairs in 20 cells (e, f) from 3 experiments; 92 kinetochore pairs in 8 cells (h), and 115 kinetochore pairs in 10 cells (j), from 2 experiments. Statistical test in (e), (f), (h), (j): ANOVA (Analysis of Variance) with post-hoc Tukey test. On all univariate scatter plots, colored points represent individual cells; black lines show the mean, with light and dark grey areas marking 95% confidence intervals for the mean and standard deviation, respectively. k Scheme of the observed protein levels (red: high, orange: medium, yellow: low, white: absent) on polar and non-polar kinetochores (circles to the upper left of the spindle and in the spindle center, respectively) in prometaphase and metaphase, as indicated. All scale bars, 2 µm.
To assess a positive marker of end-on attachments, we examined Astrin, which localizes to aligned kinetochores with low Aurora B activity, but is absent from unaligned kinetochores48,49. Consistent with these findings, we observed that Astrin was absent from both polar kinetochores within a pair, irrespective of their relative distance to the centrosome (Fig. 5d, e). Astrin was also absent from non-polar kinetochores located on the spindle surface in early prometaphase (Fig. 5e), likely reflecting the immature state of end-on attachments at this stage of mitosis16. Accordingly, these kinetochores displayed intermediate Mad2 levels, lower than those on distant polar kinetochores but higher than those on aligned kinetochores at the metaphase plate, which were devoid of Mad2 (Fig. 5f). This suggests that end-on attachments on non-polar kinetochores in early prometaphase can partially remove Mad2, but are not yet competent to recruit Astrin16. As congression progresses and end-on attachments stabilize, Astrin is subsequently recruited to kinetochores16,50.
The levels of CENP-E and ZW10, components of the fibrous corona that promotes lateral attachments51, were high at polar kinetochores, including both sisters, indicating the presence of a large corona (Fig. 5g–j). In comparison with polar kinetochores, non-polar kinetochores on prometaphase spindles recruited less CENP-E and ZW10, whereas the lowest signal was at aligned kinetochores in metaphase (Fig. 5g–j). Taken together, these results suggest that in early intact mitosis, both sister kinetochores on polar chromosomes typically form lateral attachments, while the sister closer to the pole additionally establishes immature end-on attachments characterized by the presence of checkpoint and corona proteins and the absence of Astrin. These attachments mature as the kinetochores interact with the spindle body and congress, recruiting Astrin and losing corona and checkpoint proteins16 (Fig. 5k).
Microtubules from the opposite spindle half help the movement of polar chromosomes near the spindle surface
To explore the contribution of microtubules extending from the other spindle half to the movement of polar chromosomes (Fig. 1e), we used STED microscopy and found that in phases between the end of centripetal movement and landing, 30.0 ± 5.9% of kinetochore pairs in the polar region (i.e., those attached to an astral microtubule and within ~3 μm from the spindle pole) were located next to a microtubule extending from the opposite side of the spindle (Fig. 6a), whereas 70.0 ± 5.9% had no such microtubule nearby (Fig. 6b). Attachments to microtubules from the opposite spindle half were rare behind the pole, seen in only 20.5 ± 6.5% kinetochore pairs, but were more frequent in the region in front of the pole, where 47.6 ± 10.9% of kinetochore pairs showed attachments before joining the main spindle surface (Fig. 6b). Although we could not determine whether these microtubules originated at the opposite pole or along the microtubules of the opposite spindle half, their origin at the opposite side indicates that their plus end was facing the kinetochore. In agreement with this orientation, these microtubules were attached to kinetochores predominantly in an end-on configuration (Fig. 6c, d, and Supplementary Fig. 5a). Most of these microtubules had tubulin intensity similar to that of astral microtubules, suggesting they are single microtubules (Supplementary Fig. 5b).
a STED superresolution images of microtubules (MTs) immunostained for α-tubulin (white) in RPE1 cells stably expressing CENP-A-GFP (colorful, confocal). Images show maximum intensity projections of the whole cell. Kinetochores are color-coded for depth with the brgbcmyw LUT in ImageJ throughout 16 and 9 z-planes, corresponding to 4.8 and 2.7 µm, respectively. White boxes represent regions containing polar kinetochores attached to the microtubule from the opposite side of the spindle, which are enlarged to the right and represented schematically. White arrows denote the microtubule coming from the opposite side of the spindle. Enlargements are maximum intensity projections of 2 and 3 z-planes where the microtubule of interest is located, smoothed with 0.75-mm-sigma Gaussian blur. b Percentages of kinetochores that are unattached (blue) or attached to the microtubule from the opposite side of the spindle (yellow) measured in cells as in a. The shaded area represents kinetochores located at the angle above 90° with respect to the long spindle axis. N = 18 kinetochore pairs in 15 cells from 5 experiments. Statistical test: two-sided Fisher’s exact test. c Pie chart representing the percentage of different attachment types with the microtubule from the opposite side. N = 18 kinetochore pairs in 15 cells from 5 experiments. d Pie chart representing the percentage of polar kinetochores from LLS videos that initiate their movement towards the opposite pole after landing (blue), before landing at an angle below 90° (yellow) and before landing at an angle above 90° (orange), indicative of a force other than pivoting. N = 39 kinetochore pairs in 20 cells from 8 experiments. e Schematic representation of percentages of polar chromosomes that pivot without forming an attachment to the microtubule from the opposite side before landing (left), that pivot and attach to microtubule from the opposite side once they reach the angle below 90° (middle), and the ones that attach to microtubule from the opposite side at the angle above 90° (right). Scale bars, 2 µm.
If microtubules from the opposite side pull the chromosome across the pole, this is expected to increase the distance between the kinetochores and the closer pole, as well as the interkinetochore distance, assuming that these microtubules attach to the sister kinetochore that is not end-on attached to the closer pole. We found such changes within 1 min before landing (Supplementary Fig. 5c, d), when the kinetochores are typically close to the main spindle surface (at an angle <60°, see Fig. 2e), suggesting that the pulling by microtubules from the opposite side contributes mainly to the last part of the chromosome movement towards the spindle surface. Live-cell STED microscopy provided direct visualization of the process in which such kinetochores attain end-on attachment to a microtubule extending from the opposite spindle half, followed by an increase in interkinetochore distance and movement towards the opposite pole (Supplementary Fig. 5e). These findings show that microtubules extending from the opposite side of the spindle can pull the chromosome, especially when it is already close to the spindle surface. Together, our results suggest that of the 7 polar chromosomes, around 5 cross the polar region and reach the main spindle surface only by elongation-driven pivoting, 1–2 are pulled by microtubules from the opposite side after the initial pivoting, whereas <1 chromosome per cell interacts with microtubules from the opposite side before pivoting (Fig. 6e).
Incomplete spindle elongation leads to delayed chromosome passage across the pole and missegregation
To explore the functional relevance of the pivoting model, we asked whether incomplete spindle elongation may lead to missegregation of polar chromosomes. To obtain a system with chromosome segregation errors in RPE1 cells, we inhibited the mitotic kinase Mps1, a key element of the SAC and the correction of erroneous attachments, by the small-molecule inhibitor AZ314652 (Fig. 7a, and Supplementary Movie 7). We reasoned that due to the premature anaphase onset, spindle elongation may be incomplete in a subset of cells. This should in turn lead to incomplete passage of polar chromosomes across the pole in those cells, allowing us to explore the consequences of perturbed pole crossing on the segregation fidelity of these chromosomes.
a Time-lapse images of RPE1 cells stably expressing CENP-A-GFP and Centrin1-GFP (colorful, confocal) and treated with Mps1 inhibitor AZ3146. The images show maximum intensity projections of the imaged cells. The upper panel represents cells without unaligned chromosomes and the lower panel represents cells with unaligned chromosomes. Kinetochores are color-coded for depth with the brgbcmyw LUT in ImageJ throughout 14–21 z-planes, corresponding to 5.6–8.4 µm. b Percentage of polar (left bar) and non-polar (right bar) kinetochore pairs, as determined at NEBD, that segregated normally (no error, light blue) or ended up as lagging (green), misaligned (dark blue) or unaligned (yellow) kinetochores in anaphase. Each group of kinetochores is depicted on the scheme below the graph. N = 52 polar kinetochore pairs in 7 cells from 4 experiments. N = 54 non-polar kinetochore pairs in 7 cells from 4 experiments. Statistical test: two-sided Fisher’s exact test. c Spindle length in time for cells treated with AZ3146 divided into those that have (orange) or do not have (blue) unaligned chromosomes at anaphase onset. Values are shown as mean (dark line) and SEM (shaded areas). N = 3 cells with unaligned chromosomes from 2 experiments. N = 4 cells without unaligned chromosomes from 2 experiments. d RPE1 cell imaged using lattice light-sheet microscopy at the end of centripetal movement. The dashed oval denotes the approximate position of the nucleus in the preceding interphase (see Supplementary Fig. 6). The straight white line denotes the position of the long nuclear axis in the preceding interphase, the blue line is the spindle axis in prometaphase at the end of the centripetal movement, and β is the angle between them. The image is a maximum intensity projection of 42 z-planes covering the whole cell, corresponding to 6.1 µm. e Scheme showing the angle β as in d, grey areas denote β < 45°. f Percentage of cells with β < 45° (yellow) and β > 45° (blue). Error bars represent SEM. N = 24 cells from 8 experiments. g Images show nuclei in interphase in fixed RPE1 cells stably expressing dCas9-3xGFP (orange) and stained with anti-α-tubulin (gray) and DAPI (blue). Cells are transduced with lentiviruses containing sgRNA for Chr1-telo or Chr9-cen, as indicated. Images are maximum intensity projections of 30 z-planes, corresponding to 15 µm. The diagram on the right illustrates the “danger zone,” which consists of two nuclear caps. Each cap covers one-third of the nuclear long axis. Chromosomes positioned in these caps during interphase are more likely to end up behind the centrosome in early mitosis. h Percentage of cells with chromosome 1 or 9 located in the nuclear danger zone (yellow) or outside the nuclear danger zone (blue). N = 266 chromosomes in 133 cells from 4 experiments for chromosome 1. N = 72 chromosomes in 36 cells from 3 experiments for chromosome 9. Statistical test: two-sided Chi-squared test. All scale bar, 2 µm.
The chromosomes that were polar at NEBD missegregated more frequently than non-polar chromosomes that were similarly positioned with respect to the nucleus and the centrosome midpoint in Mps1-inhibited cells (Fig. 7b)4. Whereas polar and non-polar chromosomes had a similar tendency to end up as misaligned and lagging chromosomes in anaphase, there was a stark difference in the occurrence of unaligned chromosomes (Fig. 7b). About 20% of polar chromosomes remained unaligned throughout mitosis, leading to aneuploid daughter cells. In contrast, such cases were absent in the non-polar group (Fig. 7b, and Supplementary Movie 7). These results suggest that the missegregation bias towards polar chromosomes is largely caused by their persistent unalignment.
Among the chromosomes with erroneous segregation, the persistently unaligned chromosomes were the last ones to pass from the back to the front of the spindle pole (Supplementary Fig. 6a), suggesting that the cause for their unalignment lies in perturbed passage around the pole. Based on the pivoting model, we expect that the cells that entered anaphase with persistently unaligned chromosomes had less extensive spindle elongation than those without unaligned chromosomes. Indeed, we found that among the Mps1-inhbited spindles, those that had persistently unaligned chromosomes elongated more slowly, entering anaphase at a ~4 μm shorter length than the spindles without such chromosomes (Fig. 7c, and Supplementary Fig. 6b, c). Different timing of anaphase was not the cause of persistent unalignment, as the spindles with and without unaligned chromosomes entered anaphase at a similar time (17.8 ± 2.5 vs. 17.9 ± 0.9 min). These data show that the propensity of polar chromosomes to remain unaligned is higher in cells with slower spindle elongation, suggesting the importance of effective chromosome passage across the pole.
The pivoting model may be linked with chromosome identity, as certain chromosomes missegregate more frequently than others across various treatments and cancer types4,53. We hypothesize that there is a zone within the interphase nucleus, related to the axis of the future spindle, that would predispose chromosomes to missegregate as persistently unaligned, which we term “danger zone”. In most RPE1 cells, centrosome separation occurs before NEBD, positioning one centrosome above and the other below the nucleus, followed by spindle elongation and rotation that aligns the spindle parallel to the coverslip surface30. Within this plane, we observed that in prometaphase, the centrosome-centrosome axis aligns closely with the long nuclear axis from the preceding interphase, as reflected by an angle of 26.1° ± 4.2° between the two axes (N = 24 cells, Fig. 7d–f, Supplementary Fig. 6d, e, and Supplementary Movie 8). Furthermore, the nuclear axis in interphase is typically aligned with the long cellular axis (Supplementary Fig. 6f, and Supplementary Movie 8)54. This suggests that the danger zone is situated in the nuclear caps, defined as regions near the ends of the major axis of the nucleus (Fig. 7g, diagram), because the chromosomes located there in interphase typically end up behind the centrosome in early mitosis.
We then compared interphase positions of chromosomes 1 and 9 with respect to the identified danger zone, which show a missegregation bias and random missegregation, respectively4 (Fig. 7g). Remarkably, we found that chromosome 1 resides in the danger zone in 71.00 ± 2.78% of cases. In contrast, chromosome 9 resides in the danger zone in only 45.83 ± 5.87% of cases (Fig. 7h). These findings support the idea that the missegregation bias of chromosome 1 arises from its preferential position in interphase within the nuclear caps. Chromosomes located in this danger zone are susceptible to failure in crossing the pole, leading to their persistent unalignment during mitosis4.
Spindle elongation helps to resolve polar chromosomes in cancer cell lines
Finally, we set out to investigate the universality of the pivoting model by testing cancer-derived cell lines. To determine whether microtubules attached to polar chromosomes pivot around the spindle pole in cancer cells, we used the osteosarcoma U2OS cell line expressing GFP-tagged EB355 and CENP-A-mCherry. We visualized microtubule contours using time projections of EB3 spots and observed that microtubules associated with polar kinetochores pivoted around the pole as it moved outward during spindle elongation (Fig. 8a, and Supplementary Movie 9). Tracking of kinetochores from NEBD to landing in live U2OS cells expressing CENP-A-GFP and tubulin-mCherry showed that polar kinetochores transitioned from the back to the front of the spindle pole, reflected by a decrease in their pivoting angle, during spindle elongation (Fig. 8b–d, Supplementary Fig. 7a, b, and Supplementary Movie 10). These findings indicate that microtubule pivoting occurs during spindle elongation in U2OS cells similarly to RPE1 cells.
a Time-lapse images of live U2OS cells stably expressing EB3-GFP (white, confocal) and CENP-A-mCherry (blue, confocal) showing astral microtubule (MT) attached to the polar kinetochore pair and pivoting over time. Images are temporal maximum intensity projections of a single z-plane in 3–5 sequential time frames during which the EB3 spot was visible. Astral microtubule contours during the pivoting are shown to the right, and a scheme showing the imaged region within the spindle is at the far right. b Time-lapse confocal images of live U2OS cells stably expressing CENP-A-GFP (colored for depth) and tubulin mCherry (white). Circles mark the poles and dashed lines mark the spindle axis and the edge of the polar region. c, d Pivoting angle for 5 polar kinetochore pairs (c) and spindle length (d) over time for the cell shown in b. e Univariate scatter plot of the time from NEBD until anaphase onset in live U2OS cells as in b, comparing cells with aligned chromosomes (left, N = 111) and those that had one or more chromosomes that were not aligned (right, N = 73) at the end of spindle elongation. Statistical test: unpaired, two-tailed t-test. Colored points represent individual cells; black lines show the mean, with light and dark grey areas marking 95% confidence intervals for the mean and standard deviation, respectively. f Time-lapse images of control (left) and Kif4A siRNA-treated (right) live OVSAHO cells stained with SPY-555-DNA (blue) and SiR-tubulin (white), imaged using lattice light-sheet microscopy. Dashed lines denote the edge of the polar regions; arrowhead points to chromosomes near the pole. g Spindle length over time for OVSAHO (top; N = 16, 29, 20 cells for the indicated treatments) and MDA-MB-231 cells (bottom; N = 18, 22, 5, 3 cells for the indicated treatments) treated as stated in the legend. Lines, mean; shaded areas, SEM. h Polar transit time for OVSAHO (top) and MDA-MB-231 cells (bottom) as a function of the spindle length measured 5 min after NEBD. Small circles show individual cells, and large circles with error bars are mean and SEM for each treatment. Linear regression (solid line) on individual data points is shown. Statistics: two-sided t-test. The number of cells is the same as in g, MDA-MB-231 cells treated with monastrol were pooled from the two concentrations shown in g. i Images of fixed OVSAHO cells, control (left) and KIF4A siRNA-treated (right), stained with anti-α-tubulin antibody (white), anti-CREST (orange), and DAPI (blue). The white arrow marks the chromosome at the pole in metaphase. j Univariate scatter plot of the spindle length in untreated (N = 319, 6 independent experiments) and KIF4A siRNA-treated (N = 157, 3 independent experiments) fixed OVSAHO cells. Statistical test: unpaired, two-tailed t-test. Dispersion measures as in e. k Percentage of metaphase cells with polar chromosomes in untreated (untrea.) and KIF4A siRNA-treated fixed OVSAHO cells from j. Error bars represent SEM. Statistical test: two-sided Chi-squared test. In b, f, i, images are maximum intensity projections of the whole cell. All scale bars, 2 µm. l A model of how polar chromosomes pass across the pole in normal cells and fail to cross the pole when spindle elongation is impaired, such as in certain cancer cells. In healthy cells, polar chromosomes are attached to astral microtubules. While the chromosomes remain largely immobile, the spindle elongates, thereby driving the pivoting of astral microtubules around the spindle pole. Consequently, the chromosomes with their microtubules get close to the main spindle surface, where they attach and move to the spindle midplane. In cells with impaired spindle elongation, this elongation is insufficient to drive microtubule pivoting from the back of the spindle pole all the way towards the main spindle surface, leaving polar chromosomes persistently unaligned.
To evaluate the efficiency of the pivoting mechanism in this U2OS cell line, which has chromosome congression defects4, we tracked and counted polar kinetochores over time (see “Methods”). We found that 96.1% of polar kinetochores successfully crossed the pole and congressed to the metaphase plate, whereas 3.9% remained unaligned, i.e., behind the pole, at the end of spindle elongation (N = 44 out of 1127 polar kinetochore pairs from 101 cells). These results suggest that the pivoting mechanism is highly efficient in U2OS cells and facilitates successful congression of most polar kinetochores. Tracking of kinetochores that remained unaligned revealed that they eventually moved to the main spindle surface (Supplementary Fig. 7c), likely through capture by microtubules from the opposite spindle half. However, this elongation-independent movement was significantly delayed, as indicated by a threefold increase in mitotic duration in cells retaining an unaligned chromosome after spindle elongation (Fig. 8e). Thus, failure of timely polar chromosome transit during spindle elongation prolongs mitosis in a cancer cell line.
According to our model, slower spindle elongation should delay the incorporation of polar chromosomes into the spindle body, whereas enhanced elongation should promote it. To test this prediction, we perturbed spindle elongation in two highly chromosomally unstable cancer cell lines, OVSAHO (ovarian carcinoma) and MDA-MB-231 (triple-negative breast cancer), both characterized by congression defects21,23 (Supplementary Fig. 7d). We used low-dose monastrol to slow spindle elongation and KIF4A depletion to enhance it56 (Fig. 8f, Supplementary Fig. 7e–g, and Supplementary Movies 11 and 12). Monastrol concentrations were titrated in each line to reduce spindle elongation post-NEBD without inducing monopolar arrest in most cells (see “Methods”). Live-cell imaging confirmed that monastrol-treated spindles were shorter and KIF4A-depleted spindles were longer than untreated ones throughout early mitosis in OVSAHO and MDA-MB-231 cells (Fig. 8g). The duration of polar chromosome transit, defined as the time when the last polar chromosome moved from the rear to the front side of the spindle pole, was inversely correlated with spindle length in early prometaphase in both cell lines (Fig. 8h), in agreement with our model.
To extend our findings to larger cell populations, we analyzed fixed, immunostained metaphase cells treated with either KIF4A siRNA to increase spindle length or low-dose nocodazole (10 ng/mL) to reduce spindle length (Supplementary Fig. 7h). Nocodazole was selected over monastrol for the fixed-cell assay, as monastrol-treated spindles reached near-normal length by metaphase (Fig. 8g). In addition to OVSAHO and MDA-MB-231, we analyzed the Caco-2 cell line (colorectal adenocarcinoma)57. These experiments revealed an overall trend where cells with longer spindles had fewer polar chromosomes than those with shorter spindles (Supplementary Fig. 7i). In OVSAHO cells, KIF4A depletion increased spindle length from 11.16 ± 0.09 μm (N = 319) to 12.74 ± 0.15 μm (N = 157), and this was associated with a fourfold reduction in the fraction of cells harboring polar chromosomes (from 13.2 ± 2.1% to 3.2 ± 1.4%) (Fig. 8i–k). The distance of polar chromosomes from the nearest pole was comparable between untreated and KIF4A-depleted cells (Supplementary Fig. 7j), indicating that polar ejection forces58,59 were not significantly altered. Conversely, nocodazole treatment shortened spindle length and increased the number of polar chromosomes relative to controls (Supplementary Fig. 7i), likely due to both spindle shortening and microtubule stabilization44. MDA-MB-231 and Caco-2 cells exhibited changes in the same direction as OVSAHO following KIF4A depletion and nocodazole treatment, though the effect of KIF4A was not statistically significant (Supplementary Fig. 7i).
In conclusion, our results demonstrate that the spindle elongation-driven pivoting model applies across multiple cancer cell lines and plays a key role in ensuring rapid mitosis. Treatments that reduce spindle elongation impair the resolution of polar chromosomes, whereas enhanced elongation facilitates their timely incorporation into the spindle.
Discussion
Our study uncovers an essential step in chromosome congression that safeguards genome integrity by rescuing polar chromosomes, which are at the highest risk of missegregation4, from the “danger zone” behind the spindle pole. During early prometaphase, once the centripetal movement of chromosomes ends, the spindle begins to elongate30. During this phase, the poles push outward, moving beyond the polar chromosomes, which are relatively stationary. As the chromosomes remain attached to astral microtubules throughout the process, these microtubules passively pivot around the spindle pole from the outer to the inner spindle region, ultimately reaching the spindle surface and enabling polar chromosomes to interact with it and undergo congression (Fig. 8l).
Our findings indicate that, once the centripetal chromosome movement ends, polar chromosomes usually remain attached to the same microtubule bundle extending from the centrosome until landing on the spindle surface. If the microtubules were to extend at a constant angle from the centrosome without the ability to pivot, the chromosome would move along with the centrosome, away from the spindle center. However, instead of moving, the chromosome remains stationary, and it is the angle of the microtubules that changes, repositioning the chromosome from behind the centrosome to its front side where it can congress to the spindle midplane. Taking an evolutionary perspective, since a similar type of microtubule rotational movement has been observed in yeast60, it will be valuable to assess how conserved microtubule pivoting, its underlying mechanisms, and functions are across various species.
The mechanism we propose relies on a biomechanical property of the microtubule-centrosome connection that enables the microtubules to pivot under force. Given that the pericentriolar material anchors the microtubules at the centrosome61, it will be interesting to explore how mechanical properties of the pericentriolar material affect microtubule pivoting and, in turn, the rescue of polar chromosomes. Remarkably, the old and the young centrosomes behave differently in mitosis, and the old one retains more unaligned chromosomes62. It may be that microtubule pivoting is different on the old and young centrosome and that this leads to a bias in chromosome alignment errors.
The pivoting of microtubules attached to polar chromosomes may also play a role in spindle assembly. We found that typical polar chromosomes arrive at the spindle surface with mono-lateral attachments. The microtubules in the “mono” part of the attachment may represent an early kinetochore fiber, which, upon chromosome arrival to the spindle surface, becomes incorporated into the growing spindle. Thus, kinetochore fibers of polar chromosomes may start to form already when the chromosome is behind the pole, which is much earlier than previously thought and can explain the observed fast chromosome biorientation occurring as soon as the chromosome interacts with the spindle surface13,16.
We propose that microtubules laterally attached to the polar chromosome also integrate into the spindle, forming antiparallel overlaps with microtubules extending from the opposite side. This antiparallel bundle likely facilitates the formation of end-on attachments on the sister kinetochore facing the further pole by guiding the growth of new microtubules towards this kinetochore. As end-on attachments of both sister kinetochores mature and lateral attachments diminish30, the previously laterally attached microtubules may turn into bridging fibers that connect sister kinetochore fibers63,64. Thus, polar chromosomes bring along components of their fibers to the growing spindle.
Our findings show that incomplete or slow spindle elongation, accompanied by reduced microtubule pivoting, can lead to chromosome missegregation due to the delayed or unsuccessful arrival of polar chromosomes to the spindle surface. Previous work has shown that HeLa cells following the prophase pathway of spindle assembly, where centrosomes quickly separate to the opposite sides of the nucleus before NEBD, exhibit fewer chromosome segregation errors than those following the prometaphase pathway, where centrosome separation is slower and continues during prometaphase65. Our model suggests that the differences in segregation errors between the two pathways arise from the presence of polar chromosomes2. In the prophase pathway, rapid spindle elongation ensures that no chromosomes are left behind the pole at NEBD, resulting in error-free segregation. In contrast, the slower centrosome separation in the prometaphase pathway leads to the presence of polar chromosomes, whose delayed arrival at the metaphase plate may increase the likelihood of segregation errors.
The sustained persistence of polar chromosomes may elevate Aurora A activity at the centrosome through Mps1-dependent signaling at kinetochores66. This, in turn, can hyperactivate Aurora B at kinetochores, thereby inhibiting the formation of stable end-on attachments67. Thus, the delayed passage of polar chromosomes across the spindle pole further diminishes their likelihood of establishing stable end-on attachments and successfully congressing to the metaphase plate.
The spatial arrangement of chromosomes and centrosomes at the start of mitosis may explain why certain chromosomes are more prone to missegregation4. As the chromosomes that are found behind the spindle pole in early mitosis are typically positioned at the nuclear periphery during the preceding interphase4, our proposed model identifies the biomechanical principles underlying the biased missegregation of peripheral chromosomes.
As a potential link to disease, we demonstrate in several cancer cell lines that insufficient spindle elongation, leading to impaired microtubule pivoting, delays the alignment of polar chromosomes. Conversely, enhancing spindle elongation accelerates polar chromosome alignment. However, additional mechanisms may contribute to efficient chromosome alignment. For instance, in cell lines with short spindles but minimal accumulation of unaligned chromosomes, such as the ovarian cancer cell line OVKATE23,68, pulling forces exerted by microtubules from the opposite spindle half may facilitate the integration of polar chromosomes into the spindle. The relative contribution of these alternative mechanisms across different cell lines and over time remains an interesting question for future investigation. We further propose that the failure of polar chromosomes to cross the pole during spindle elongation, observed in a subset of cancer cells and after nocodazole treatment, may be linked to increased stabilization of end-on attached microtubules44,69,70, giving rise to thick microtubule bundles with limited pivoting capacity. The resulting delay in the movement of polar chromosomes may contribute to prolonged mitosis in cancer cell lines with chromosome congression defects.
Finally, we speculate that the spindle elongation-driven microtubule pivoting may be relevant to explain and modify errors in chromosome alignment in cancer cells. This idea is supported by the fact that cells without PTEN, the second most frequently mutated tumor suppressor after p5371,72,73, recruit less Eg5 to their spindles, resulting in both shorter spindles and misaligned chromosomes74. Thus, a PTEN mutation might be one of the indicators that a particular type of cancer has unaligned chromosomes that stem from impaired spindle elongation and might be rescued by restoring PTEN function. More broadly, we propose that manipulating spindle elongation in certain cancer cells could be used to influence mitotic errors, either by correcting them to maintain genome stability or by worsening them to a degree that compromises cell viability.
Methods
Cell lines and plasmids
The experiments were performed using the following cell lines: human hTERT-RPE1 (hTERT-immortalized retinal pigment epithelium) cells stably expressing CENP-A-GFP, human hTERT-RPE1 cells stably expressing CENP-A-GFP and Centrin1-GFP30, and human hTERT-RPE1 cells stably expressing CENP-A-GFP and Mis12-mCherry, a gift from Alexey Khodjakov (Wadsworth Center, New York State Department of Health, Albany, NY, USA); human hTERT-RPE1 cells stably expressing CENP-A-Cerulean and Mad2-mRuby75, a gift from Jonathon Pines (Institute of Cancer Research, London, UK); human hTERT-RPE1 cells stably expressing tubulin-EYFP and H2B-mRFP76, a gift from Andrew Holland (Johns Hopkins University School of Medicine, Baltimore, MD, USA); human hTERT-RPE1 cells stably expressing dCas9-GFP-3xFKBP77, a gift from Susanne Lens (Utrecht University, Utrecht, Netherlands); human hTERT-RPE1 cells stably expressing EB3-GFP and H2B-RFP, a gift from Patrick Meraldi (University of Geneva, Geneva, Switzerland) and generated by Will Krek; unlabeled human OVSAHO cell line (JCRB1046 Cell Bank, Tebubio, Le Perray-en-Yvelines, France); unlabeled human MDA-MB-231 cell line, a gift from Dragomira Majhen (Ruđer Bošković Institute, Zagreb, Croatia); unlabeled human Caco-2 cell line (86010202, Sigma Aldrich); and human U2OS cells stably expressing EB3-2xGFP and CENP-A-mCherry63.
The plasmids used for the experiments were: pHTN HaloTag-CENP-A78, a gift from Patrick Meraldi (University of Geneva, Geneva, Switzerland); psPAX2 (Addgene, No. 12260); psMD2.G (Addgene, No. 12259); pLH sgRNA Chr1-telo79 and pLH sgRNA Chr9-cen77, both gifts from Susanne Lens (Utrecht University, Utrecht, Netherlands).
Cell culture
All cell lines, except an OVSAHO cell line, were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM) containing 1 g/L D-glucose, pyruvate and L-glutamine (Gibco), supplemented with 10% (vol/vol) heat-inactivated Fetal Bovine Serum (FBS, Sigma Aldrich, St. Louis, MO, USA) and penicillin (100 IU/mL)/streptomycin (100 mg/mL) solution (Lonza, Basel, Switzerland). The OVSAHO cell line was maintained in RPMI 1640 medium containing L-glutamine and sodium bicarbonate (Sigma-Aldrich, St. Louis, MO, USA), complemented with the same supplements used in the previously described DMEM medium. All cells were kept at 37 °C and 5% CO2 in a Galaxy 170R humidified incubator (Eppendorf, Hamburg, Germany). All cell lines have been routinely examined for mycoplasma contamination by using SPY-555-DNA (1:100,000, Spirochrome, Stein am Rhein, Switzerland) and by checking if any extracellular DNA is present. All cell lines used for experiments have been confirmed to be mycoplasma free.
RNAi treatment
Cells for RNAi transfection were seeded on 35 mm uncoated dishes with the 0.17 mm (1.5 coverslip) glass thickness (Ibidi GmbH, Gräfeling, Germany) in 1 mL of the previously described DMEM medium to reach ~60% confluence the following day. siRNA constructs were diluted in OPTI-MEM medium (Life Technologies, Waltham, MA, US) and RNAi treatments were performed using Lipofectamine RNAiMAX Reagent (Life Technologies, Waltham, MA, US) according to the instructions provided by the manufacturer.
The siRNA constructs and their final concentrations were: 100 nM CENP-E siRNA (L-003252-000010; Dharmacon, Lafayette, CO, USA), 100 nM Spindly/CCDC99 siRNA (L-016970-00-0010, Dharmacon, Lafayette, CO, USA), 100 nM KIF4A siRNA (sc-60888; Santa Cruz Biotechnology, Dallas, TX, USA), 100 nM Kid/KIF22 siRNA (4392420; Ambion, Austin, TX, USA). Cells were incubated for 4 h with the transfection mixture followed by medium replacement into the previously described DMEM medium. All experiments were performed 24 h after siRNA transfection.
Plasmid transfection
To transiently express CENP-A in an RPE1 cell line stably expressing tubulin-EYFP and H2B-mRFP, 5 µg of pHTN HaloTag-CENP-A plasmid was used for transfection performed on Nucleofector Kit R with the Nucleofector 2b Device (Lonza) following manufacturer’s instructions. The Nucleofactor Y-001 program was used for the transfection of the RPE1 cell line. After transfection, around 500,000 cells were seeded on 35 mm uncoated dishes with the 0.17 mm (1.5 coverslip) glass thickness (Ibidi GmbH, Gräfeling, Germany) in 1 mL of the previously described DMEM medium. Transfection was performed 48 h before imaging.
Lentivirus transduction and cell line generation
To produce lentiviruses, HEK293T cells were seeded on 10 cm Petri dishes to reach 80% confluence the next day. Plasmid constructs were diluted in OPTI-MEM medium (Life Technologies, Waltham, MA, US) and transfections were performed using Lipofectamine 3000 Transfection Reagent (Life Technologies, Waltham, MA, US) according to the manufacturer’s protocol. The plasmid constructs and their final masses were as follows: 10 µg of psPAX2, 5 µg of psMD2.G, 15 µg of pLH sgRNA Chr1-telo or 15 µg of pLH sgRNA Chr9-cen. The cell medium was replaced with the previously described DMEM medium 24 h after transfection. Lentiviruses containing sgRNA for Chr1-telo or Chr9-cen were harvested 48 h after transfection by filtering the cell medium through 45 µm filters (No. 99745, TPP, Switzerland). RPE1 cells stably expressing dCas9-GFP-3xFKBP were seeded in 6-well plates (120,000 cells/well, Thermo Fisher Scientific, Waltham, MA, USA) the day before lentivirus transduction. Lentiviruses were added to cells together with 6 µg/ml of polybrene (TR-1003-G, Merck Milipore, Darmstadt, Germany) and incubated for 24 h.
Treatments with inhibitors
The following Eg5 inhibitors were used on a “one cell per dish” principle: STLC (final concentration 40 µM, 164739, Sigma-Aldrich, St. Louis, MO, USA) was added ~30 s before landing to instantly cause spindle shortening upon addition while preserving bipolarity; FCPT (final concentration 40 µM, a gift from Timothy Mitchison) was added at the end of centripetal approaching to block the spindle elongation; monastrol (final concentration 100 µM in RPE1 cells, HY-101071A, MedChemExpress, Monmouth Junction, USA) was added 30 s before landing and then washed out with a pre-warmed DMEM medium when spindle stopped shortening. Nocodazole (final concentration 20 nM in RPE1 cells, HY-13520, MedChemExpress, Monmouth Junction, USA), Mps1 inhibitor AZ3146 (final concentration 3 µM, HY-14710, MedChemExpress, Monmouth Junction, USA), Mps1 inhibitor Cpd-5 (final concentration 62.5 nM, a gift from R. Medema via G. J. P. L. Kops lab4,80) and actin polymerization inhibitor latrunculin A (final concentration 2 µM, 428021, Sigma-Aldrich, St. Louis, MO, USA) were added ~45 min prior to imaging and cells within the dish were subsequently imaged for 4 h. CENP-E inhibitor GSK923295 (MedChemExpress, Monmouth Junction, USA, IC50 value 3.2 nM) was added at a final concentration of 80 nM 1–4 h before imaging.
For live-cell imaging of monastrol-treated OVSAHO cells, 15 µM monastrol was chosen as it was sufficient to produce slower spindle elongation without inducing monopolar arrest in the majority of cells. In MDA-MB-231 cells, 30 and 50 µM monastrol produced a similar result. Only the cells that formed a bipolar spindle within the first 10 min after NEBD were analyzed. For fixed-cell imaging of nocodazole-treated cancer cells, 10 ng/mL (~33 nM) of nocodazole was added 2 h prior to fixation.
Live-cell dyes
To identify NEBD in RPE1 cells stably expressing CENP-A-GFP and Centrin1-GFP, the SPY-555 or SPY-595 DNA probe (1:100,000, Spirochrome, Stein am Rhein, Switzerland) was added to the cells 45 min to 2 h prior to imaging. When microtubules were visualized during live-cell imaging in the same cell line, the SPY-650-tubulin probe (1:50,000, Spirochrome, Stein am Rhein, Switzerland) was added to the cells 45 min to 2 h prior to imaging. For the live-cell imaging of OVSAHO, MDA-MB-231, and Caco-2 cell lines, the SPY-555 DNA probe (1:50,000, Spirochrome, Stein am Rhein, Switzerland) was added to the cells along with the SiR-tubulin probe (50 nM, Spirochrome, Stein am Rhein, Switzerland;81) 45 min prior to imaging. The cells were subsequently imaged for 1–3 h. For STED live-cell imaging, the SiR-tubulin probe (100 nM, Spirochrome, Stein am Rhein, Switzerland81) was added to the cells 15 min prior to imaging and cells were subsequently imaged for 45 min. To avoid efflux of the dyes, a broad-spectrum efflux pump inhibitor verapamil (Spirochrome, Stein am Rhein, Switzerland) was added at a final concentration of 1 μM to cells along with tubulin and/or DNA probes. For the live-cell imaging of RPE1 cells stably expressing EYFP-α-tubulin and H2B-mRFP, cells were incubated for 30 min with Janelia Fluor 635 HaloTag ligand (a final concentration of 20 nM, Promega, Madison, WI, USA) 24 h before imaging to visualize the transiently expressed CENP-A-HaloTag.
Immunofluorescence
To determine the RNAi efficiency, RPE1 cells stably expressing CENP-A-GFP and Centrin1-GFP were grown on 35 mm uncoated dishes with the 0.17 mm (1.5 coverslip) glass thickness (Ibidi GmbH, Gräfeling, Germany) and fixed by 1 ml of ice-cold methanol for 1 min at –20 °C, except for Spindly/CCD99 immunostaining where cells were pre-extracted with 0,2% Triton in PHEM buffer (60 mM PIPES, 10 mM EGTA, 25 mM HEPES, 4 mM MgCl2) for 1 min and subsequently fixed with 4% pre-warmed to 37 °C for 10 min. Cells for CENP-E and ZW10 immunostainings were pre-extracted for 20 s with 0.5% Triton in PEM buffer (0.1 M PIPES, 1 mM EGTA, 1 mM MgCl2) followed by fixation by 1 ml of ice-cold methanol for 1 min. Following fixation, cells were washed three times for 5 min with 1 ml of PBS and permeabilized with 0.5% Triton-X-100 in PBS for 15 min at room temperature. To block unspecific binding, cells were incubated in 1 ml of blocking buffer (1% normal goat serum (NGS)) for 1 h at 4 °C. Cells were then washed three times for 5 min with 1 ml of PBS and incubated with 250 µl of primary antibody solution overnight at 4 °C. The primary antibodies were used as follows: rabbit anti-CENP-E (1:100 for determination of RNAi efficiency, and 1:500 for staining of outer corona, C7488; Sigma-Aldrich), rabbit anti-Spindly/CCD99 (1:1000, A301_354A, Bethyl), mouse anti-KIF4A (1:100, sc-365144; Santa Cruz Biotechnology), mouse anti-Kid (1:100, sc-390640; Santa Cruz Biotechnology), rabbit polyclonal anti-ZW10 (1:500, ab21582, Abcam), rabbit polyclonal anti-MAD2 (1:250, 924601, BioLegend), mouse monoclonal anti-Astrin, clone C-1 (1:250, MABN2487, Merck). After the incubation with a primary antibody, cells were washed three times for 5 min with 1 ml of PBS and then incubated with 250 µl of secondary antibody for 1 h at room temperature. Alexa Fluor 594 (Abcam, ab150108 or ab150076) was used as a secondary antibody at a 1:250 dilution for CENP-E, Kif4A, and Kid staining and at a 1:1000 dilution for Spindly staining in experiments where RNAi efficiency was tested. Alexa Fluor 647 (Abcam, ab150107) was used as a secondary antibody at a 1:500 dilution for Astrin staining. Alexa Fluor 594 was used as a secondary antibody at a 1:500 dilution for staining of CENP-E and Mad2 on polar kinetochores. Alexa Fluor 647 (Abcam, ab150075) was used as a secondary antibody at a 1:1000 dilution for ZW10 staining.
To image polar chromosomes in cancer cell lines OVSAHO, MDA-MB-231, and Caco-2, cells were fixed with 1 ml of 4% paraformaldehyde pre-warmed to 37 °C for 10 min followed by all the previously described steps for immunostaining. The primary antibodies were used as follows: rat anti-tubulin (1:500, MA1-80017, Invitrogen) and human anti-centromere protein (1:300, 15-234, Antibodies Incorporated, CA, USA). The secondary antibodies were: donkey anti-rat Alexa Fluor 488 (1:1000, Abcam, ab150153) and goat anti-human DyLight 594 (1:500, Abcam, ab96909). DAPI (1 µg/mL) was used for chromosome visualization in all cell lines.
For STED imaging, RPE1 cells stably expressing CENP-A-GFP and RPE1 cells stably expressing CENP-A-Cerulean and Mad2-mRuby were grown on 35 mm uncoated dishes with the 0.17 mm (1.5 coverslip) glass thickness (Ibidi GmbH, Gräfeling, Germany). Cell medium was removed and 0.5% Triton in PEM buffer (0.1 M PIPES, 1 mM EGTA, 1 mM MgCl2) was added for 15 s to remove the components of the cytoplasm. Following extraction, cells were fixed in 3% paraformaldehyde and 0.1% glutaraldehyde solution for 10 min. To reduce the background fluorescence, quenching (100 mM glycine in PBS) and reduction (0.1% sodium borohydride in PBS) solutions were added for 7 and 10 min, respectively. To prevent non-specific binding, cells were incubated in blocking/permeabilization buffer (2% NGS and 0.5% Triton-X-100 in PBS) for 2 h at 4 °C. Microtubules were then stained using the rat anti-tubulin primary antibody (1:500 in blocking/permeabilization buffer, MA1-80017, Invitrogen) with a 4 °C overnight incubation. The next day, cells were washed with PBS three times for 5 min. After washing, the secondary antibody donkey anti-rat Alexa Fluor 594 (1:1000 in blocking/permeabilization buffer, Abcam, ab150156) or donkey anti-rat Alexa Fluor 647 (1:1000 in blocking/permeabilization buffer, Abcam, ab150155) was added and incubated for 1 h at room temperature. DAPI (1 µg/mL) or SPY-505-DNA (1:10 000, Spirochrome, Stein am Rhein, Switzerland) was added to visualize chromosomes. The protocol was originally modified for82 based on the protocols from83,84. For validation of actin depolymerization by Latrunculin A, RPE1 cells stably expressing CENP-A-GFP and Centrin1-GFP were fixed following the same fixation protocol as above, and SiR-actin (200 nM in PBS, Spirochrome, Stein am Rhein, Switzerland) was incubated for 20 min at room temperature.
To analyze the position of chromosomes 1 and 9 in interphase cells, RPE1 cells stably expressing Chr1-telo-GFP or Chr9-cen-GFP were stained for tubulin and centrin-3 following the same immunostaining protocol used for STED imaging. Rabbit anti-centrin-3 primary antibody (1:1000, ab228690, Abcam) and secondary donkey anti-rabbit Alexa Fluor 647 (1:1000, Abcam, ab150075) were used for immunostaining.
Microscopy
STED microscopy of fixed cells, live-cell imaging of pivoting microtubules, live-cell imaging of EB3 comets, and live-cell imaging of cells treated with Cpd-5 were performed using an Expert Line easy3D STED microscope system (Abberior Instruments, Göttingen, Germany) with the 100 x/1.4NA UPLSAPO100x oil objective (Olympus, Tokio, Japan) and an avalanche photodiode (APD) detector, as previously described27. The 405 nm, 488 nm, 561 nm and 647 nm laser lines were used for the excitation of Cerulean, GFP, Alexa Fluor 594/mRuby/mCherry, and Alexa Fluor 647/Janelia Fluor 635/SiR-tubulin, respectively. The 775 nm laser line was used for the depletion of red and far-red lines during the STED superresolution imaging. Images were acquired using the Imspector software. The xy pixel size for fixed RPE1 cells stably expressing CENP-A-GFP was 20 nm and 20–30 focal planes were acquired with the 300 nm distance between them to cover the whole cell. The xy pixel size for fixed RPE1 cells stably expressing CENP-A-Cerulean and Mad2-mRuby was 25 nm and 1 focal plane containing the kinetochores of polar chromosomes was acquired. For live-cell STED imaging of pivoting microtubules, the xy pixel size was set to 30 nm, and one focal plane was acquired every 60 s. For confocal live-cell imaging of pivoting microtubules in RPE1 cells on the same microscope, the xy pixel size was set to 55 nm and 3 focal planes were acquired, with the 1 µm distance between the planes and 15 s time intervals between different time frames. For confocal live-cell imaging of EB3 comets in RPE1 cells, the xy pixel size was set to 80 nm and 3 focal planes were acquired, with the 1 µm distance between the planes and 20 s time intervals between different time frames. For confocal live-cell imaging of EB3 comets in U2OS cells, the xy pixel size was set to 80 nm and one focal planes was acquired, with 1 s time intervals between different time frames. For confocal live-cell imaging of cells treated with Cpd-5, the 488 nm laser line was used to excite the GFP, the 647 nm laser line was used to excite the SPY-650-tubulin, the xy pixel size was 80 nm and 6–7 focal planes were acquired, with 1 µm distance between the planes and 20 s time intervals between different time frames, as previously described4. Movies of U2OS cells expressing CENP-A-GFP and tubulin-mCherry were acquired as previously described4.
Confocal microscopy of OVSAHO cells and RPE1 cells immunostained to check the respective RNAi efficiencies was performed using a previously described microscope setup85, consisting of a Bruker Opterra Multipoint Scanning Confocal Microscope (Bruker Nano Surfaces, Middleton, WI, USA), mounted on a Nikon Ti-E inverted microscope with a Nikon CFI Plan Apo VC 100 ×/1.4 numerical aperture oil objective (Nikon, Tokyo, Japan). To excite DAPI, GFP, Alexa Fluor 594 or DyLight 594, a 405 nm, 488 nm, and 561 nm laser lines were used, respectively. Opterra Dichroic and Barrier Filter Set 405/488/561 enabled the separation of excitation light from the emitted fluorescence. Images were acquired using the Evolve 512 Delta Electron Multiplying Charge Coupled Device (EMCCD) Camera (Photometrics, Tuscon, AZ, USA), with a camera readout mode of 20 MHz. The xy pixel size was 83 nm and the slit aperture was 22 µm. In all experiments, the whole spindle stack was imaged. The z-stacks were acquired using the unidirectional xyz scan mode with the 1 μm distance between the planes to cover the entire cell.
Confocal live cell imaging of RPE1 cells stably expressing CENP-A-GFP and Centrin1-GFP treated with STLC, FCPT, monastrol and AZ3146 was performed on a Dragonfly spinning disk confocal microscope system (Andor Technology, Belfast, UK) using the 100×/1.47 HC PL APO oil objective (Nikon, Tokyo, Japan) and Sona 4.2B-6 Back-illuminated sCMOS camera (acquisition mode set to high speed, Andor Technology) linked with iXon Life 888 EMCCD Camera (EM gain set to 208, Andor Technology). To simultaneously image two channels on two independent cameras, the 565 LP image splitter was used. Images were acquired using the Fusion software. During imaging, cells were maintained at 37 °C and 5% CO2 within the heating chamber (Okolab, Pozzuoli, NA, Italy). For the excitation of GFP and SPY-650-tubulin, the 488-nm, and 637-nm laser lines with corresponding filters were used, respectively. Image acquisition was done every 0.4 µm with 21 focal planes and a time interval of 10 s.
The Lattice Lightsheet 7 system (Carl Zeiss, Germany) was used for the live cell imaging of control RPE1 cells expressing CENP-A-GFP and Centrin1-GFP and for the imaging of cells with CENP-E, Spindly/CCD99, and Kid/KIF4A depletion, inhibition of actin polymerization, or nocodazole addition. The same system was used for live-cell imaging of cancer cell lines. The system includes an illumination objective lens 13.3×/0.4 (at a 30° angle to cover the glass) with a static phase element and a detection objective lens 44.83×/1.0 (at a 60° angle to cover the glass) with an Alvarez manipulator, as previously described16. For the live-cell imaging of RPE1 cells, the GFP was excited using the 488-nm diode laser (power output 10 Mw), with the laser power set to 2.5%, LBF 405/488/561/642 emission filter and 12 ms exposure time. The detection module consisted of a Hamamatsu ORCA-Fusion sCMOS camera. The width of the imaging area was set to 32–36 µm with the x interval of 0.3 µm. Images were acquired every 6 s. For the live-cell imaging of cancer cell lines, the SPY-555-DNA was excited using the 561-nm diode laser, with the laser power set to 2.0–4.0%. SiR-tubulin was excited using the 561-nm diode laser, with the laser power set to 1.0–1.5%. The exposure time was 15–20 ms. The width of the imaging area was set to 36 µm with the x interval of 0.5 µm. Images were acquired every minute. During the imaging, cells were kept at 37 °C and at 5% CO2 in a Zeiss stage incubation chamber system (Carl Zeiss, Germany). All movies were deskewed with the ZEN 3.7 software using the “Linear Interpolation” and “Cover Glass Transformation” settings. Deconvolution was performed using the “Constrained Iterative” algorithm with automatic normalization and strength set to 4.0. The imaging parameters for CENP-E inhibition experiments were described previously16.
Confocal imaging of chromosomes 1 and 9 in RPE1 cells, as well as fixed cancer cell lines OVSAHO, MDA-MB-231, and Caco-2, was performed on the Airyscan Zeiss LSM800 confocal laser scanning microscope with the 40x/0.95 Corr M27 objective (Carl Zeiss, Germany) for RPE1 cells, or 63x/1.4 Oil DIC M27 objective (Carl Zeiss) for cancer cell lines, and LSM 800 GaAsP photomultiplier detectors. To excite DAPI, GFP, Alexa Fluor 594 and 647, the 405 nm, 488 nm, 561 nm and 647 nm laser lines were used, respectively. Images were acquired in ZEN blue 3.5 software using the Tiles recording mode, with the 0.5 µm distance between the planes to cover the whole cells.
Image analysis
In LLS and confocal videos, kinetochores as markers for chromosomes and centrosomes as markers for spindle poles were tracked in a z-stack using the Point Tool in Fiji/ImageJ. When the signal covered several z-planes, the central z-plane was used for tracking. Spindle length, pivoting angle (i.e., the angle between the long spindle axis and the line that connects the kinetochore midpoint with the closer centrosome), cTilt (i.e., the angle between the long spindle axis and the line that connects sister kinetochores13), interkinetochore distance, distance to closer and further centrosome, and distance to centrosome midpoint were subsequently calculated from the coordinates in 3D using a home-written MatLab script. The end of centripetal movement was determined as a time point when the distance of the kinetochore midpoint to the closer centrosome started varying instead of continuously becoming smaller. The landing on the main spindle surface was determined as a time point when the pivoting angle was below 60° and cTilt was below 22.5°13, corresponding to the moment when kinetochores first align along the microtubules of the main spindle surface, even if they experience subsequent detachments. The landing was additionally verified visually by checking that a kinetochore pair was adjacent to the microtubules of the main spindle body. The alignment was defined as the moment when the kinetochore pair was equidistant (±1–2 µm) to both poles. The formation of the prometaphase ring was visually determined as a time point when most kinetochores finish their centripetal movement. The end of spindle elongation was defined as a time point when spindle length starts varying and/or decreasing instead of continuously increasing. The anaphase onset was visually determined as a time point when sister kinetochores first start separating. Polar chromosomes were defined as those that had a pivoting angle of >90° at NEBD and subsequently made their first contact with the spindle behind the spindle pole. The position of the first contact was determined at the end of centripetal movement, when the distance of the kinetochore pair to the spindle contour was usually <3 µm. Pivoting was then analyzed between the end of centripetal movement until after landing. Only kinetochores that could be reliably tracked from NEBD to alignment, remaining distinguishable from neighboring kinetochores throughout this period, were included in the analysis. In Eg5 manipulation experiments, analysis was done from the first frame when the inhibitor started working, i.e., when the spindle length started increasing or decreasing.
Successful passage across the pole was defined as the passage across the 90° mark within 15 min from NEBD, not taking into account any subsequent failures to form stable attachments to microtubules of the main spindle surface. Misaligned chromosomes were defined as those located within the two spindle poles and less than 2 µm away from the centrosome/spindle pole in metaphase. Unaligned chromosomes were defined as those located behind the spindle pole or, alternatively, in front of the pole but not attached to microtubules of the main spindle surface in metaphase. Lagging chromosomes were defined as those located in the central part of the spindle with a stretched kinetochore during anaphase.
When chromosomes were labeled instead of kinetochores, two points were put in the centromeric region during the tracking. When poles were marked with tubulin or EB3 instead of Centrin1, the point was put in the center of tubulin/EB3 signal. When astral microtubules were tracked instead of chromosomes or kinetochores, their movement was tracked by putting two points along the central part of the microtubule contour, and their shape was determined by putting two points at the spindle poles and subsequent five along the entire microtubule contour. Kinetochore switching between microtubules was investigated using visual analysis.
In STED images of fixed cells, NEBD was determined as the stage when the centrosomes were one above the other, or when the centrosomes were positioned sideways, but microtubules did not occupy the whole area between the centrosomes. The beginning of the centripetal movement was identified as the stage when >90% kinetochores were in close proximity to microtubules, regardless of their distance from the centrosome. The end of the centripetal movement was determined as the stage when kinetochores were arranged in a prometaphase ring at similar distances, smaller than 4 µm, to the centrosomes. Passage across the pole was identified as the stage when the spindle was almost fully formed, with the majority of kinetochores on the spindle body and a minority at the front side of a centrosome, but not in the immediate proximity of the spindle surface. Landing was determined as the stage when all kinetochore pairs were found on the spindle body at locations >4 µm away from a centrosome.
The length of long xy-, short xy- and z- nuclear axes was measured on maximum intensity projections of the whole cell just before NEBD using the Line Tool in ImageJ. The long xy axis was defined as the axis with the largest distance between the two opposite sides of the nuclear border, the short xy axis was defined as perpendicular to the middle of the long xy axis, and z axis was defined as the distance from the first focal plane in which the signal was present until the last. The spindle axis, used to determine the danger zone, was measured using the Line Tool in ImageJ on maximum intensity projections of the whole cell when the prometaphase ring formed and most kinetochores finished their centripetal movement. The angle between the long nuclear xy axis just before NEBD and the long spindle axis in prometaphase was then calculated to determine which chromosomes are most likely to attach behind the pole. This angle was also compared to the angle between the long cellular xy axis at NEBD, determined using the Line Tool in ImageJ, and the long spindle axis in prometaphase.
The long axis of interphase RPE1 dCas9-GFP cells with labeled subtelomeric region of chromosome 1 or pericentromeric region of chromosome 9 was measured on maximum intensity projections of the whole cell. The long axis was then divided into three equal parts that represent the borders of three cellular compartments—two polar zones and the central zone. The position of the labeled chromosomes was determined as polar if they were located in any of the polar zones and as central if they were located in the central zone.
To determine the position of kinetochore pairs relative to the pole in STED images, kinetochores and poles were marked in 3D using a Point Tool in ImageJ and the angle that the line connecting the kinetochore pair midpoint with the closer centrosome forms with the long spindle axis (i.e., pivoting angle) was subsequently calculated using a home-written MatLab script. Kinetochore pairs in fixed-cell STED images were considered polar if they were located above 90° or, alternatively, were located within ~3 μm from the pole but not attached to the microtubules of the main spindle surface during later stages of prometaphase when the spindle was already formed.
Attachment types were determined from STED images of fixed cells. Polar chromosomes were considered attached to microtubules when there was a microtubule either directly ending at the kinetochore or in very close proximity to the kinetochore. Polar chromosomes were considered unattached when there was no microtubule in their proximity. The type of attachment was determined visually. A limitation of this method is that attachment cannot always be distinguished from mere proximity.
For fixed-cell confocal images of CENP-A together with Mad2, Astrin, CENP-E, or ZW10, protein levels were measured using the Polygon Tool in ImageJ. The same polygon was used for measuring the signal in both channels for the same kinetochore, as well as for the background in the vicinity of the kinetochore in both channels. The final Mad2/CENP-A signal on the kinetochore was calculated using the formula: Mad2/CENP-A signal = (Integrated density of Mad2 on the kinetochore – (Kinetochore area x Mean fluorescence of the background))/(Integrated density of CENP-A on the kinetochore – (Kinetochore area x Mean fluorescence of the background)). The other 3 proteins were analyzed in the same manner. In fixed-cell confocal images, kinetochore pairs were classified as polar if their pivoting angle exceeded 90°. For each cell, three to ten kinetochores located between the centrosomes were analyzed and classified as non-polar. Additionally, three to ten kinetochores per metaphase cell were analyzed. Metaphase cells were identified by the presence of fully formed metaphase plates.
The intensity of microtubules from the opposite side of the spindle, the intensity of astral microtubules and the accompanying background were measured in a single z-plane using the 20 × 20 pixel Square Tool in ImageJ. The final intensity of the microtubules was then calculated using the formula: Microtubule intensity = Mean intensity of the microtubule – Mean intensity of the background.
In the validation experiments for protein depletion by siRNA, CENP-E, Spindly, KIF4A and Kid intensities were measured in sum intensity projections of the entire cell, covering 33 z-planes or 33 µm. The total signal of the protein of interest within the spindle was marked by using the Polygon Tool in ImageJ. Background in the cytoplasm was measured using the 25 × 25 pixel Square Tool. The total intensities were calculated using the following formula: Protein intensity = Integrated density within the spindle – (Area of the spindle × Mean fluorescence of the background). The total intensities were then normalized to the mean total intensity obtained from control cells.
Tracking of kinetochores in U2OS cells expressing CENP-A-GFP and tubulin-mCherry was performed using 2D maximum intensity projections and the “TrackMate” plugin in ImageJ, with the LoG detector86. The estimated blob diameter was set to match the size of a single kinetochore labeled with CENP-A, and thresholding was adjusted until all visible kinetochores were detected, with no false positives outside the cell. To avoid interference from neighboring cells, the field of view was cropped accordingly. Using this approach, the average number of tracked kinetochore pairs at NEBD was 42, fewer than the reported modal chromosome number for the U2OS cell line (n = 74, ATCC). This discrepancy is likely due to insufficient spatial resolution in 2D projections, which may prevent the complete separation of overlapping kinetochore pairs. The number of polar chromosomes that successfully passed the spindle pole and reached the metaphase plate by the end of spindle elongation in live-imaged U2OS cells was calculated by subtracting the number of identified polar kinetochores remaining behind the pole at the end of spindle elongation from the total number of polar kinetochores counted at NEBD. Kinetochore pairs at NEBD and at the end of spindle elongation were classified as polar if their pivoting angle in 2D projection exceeded 90°. The end of spindle elongation was estimated as described previously4. The majority of kinetochores remaining behind the pole were continuously backtracked frame by frame to NEBD, enabling clear distinction from those that successfully traversed the spindle pole.
For live-cell measurements of spindle elongation and polar chromosome transit in cancer cell lines, the spindle length was measured each minute as the 2D distance between the two spindle poles using the Line Tool in ImageJ. Coupled with this were the measurements of the polar transit time for each cell, defined as the time point when the last chromosome of those that were initially polar moved from the back side to the front side of the centrosome.
Statistics and reproducibility
All measurements were performed in ImageJ/Fiji (National Institutes of Health), followed by data analysis in MATLAB (MathWorks). Figures and schematic representations were assembled in Adobe Illustrator CC (Adobe Systems). Generally, the Shapiro–Wilk test was used to determine normality of the sample data. T-test (for two samples unless stated otherwise) or ANOVA (Analysis of Variance) with post-hoc Tukey test (for more than two samples) were then used for data that followed a normal distribution, and Mann–Whitney U test (for two samples) or Kruskal–Wallis with posthoc Dunn’s test (for more than two samples) were used otherwise. To analyze categorical data, Chi-squared test was used for larger samples (N > 50) and Fisher’s exact test was used for smaller samples. Relationship between variables was modeled using Ordinary Least Squares linear regression. The significance of the slope was evaluated using a two-sided Student’s t-test and all analyzes were performed in MATLAB using the “fitlm” function. No adjustments were made for multiple comparisons. The result was considered statistically significant when the p-value was equal to or less than 0.05. Statistical analysis was performed as noted in the respective Figure captions. The calculations and data in Supplementary Figs. 2j and 7d were taken from previous works87,88,89,90,91,92,93,94,95,96.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The source data used for the main figures have been deposited in the Figshare database (https://doi.org/10.6084/m9.figshare.29126831)97. All other data supporting the findings of this study are available from the corresponding author.
Code availability
Codes used to analyze and plot the data are available from the corresponding author on request.
References
Maiato, H., Gomes, A. M., Sousa, F. & Barisic, M. Mechanisms of chromosome congression during mitosis. Biology 6, 13 (2017).
Vukušić, K. & Tolić, I. M. Polar chromosomes-challenges of a risky path. Cells 11, 1531 (2022).
Barisic, M., Aguiar, P., Geley, S. & Maiato, H. Kinetochore motors drive congression of peripheral polar chromosomes by overcoming random arm-ejection forces. Nat. Cell Biol. 16, 1249–1256 (2014).
Klaasen, S. J. et al. Nuclear chromosome locations dictate segregation error frequencies. Nature 607, 604–609 (2022).
Booth, A. J. et al. Contractile acto-myosin network on nuclear envelope remnants positions human chromosomes for mitosis. eLife 8, e46902 (2019).
Li, Y., Yu, W., Liang, Y. & Zhu, X. Kinetochore dynein generates a poleward pulling force to facilitate congression and full chromosome alignment. Cell Res. 17, 701–712 (2007).
Rieder, C. L. & Alexander, S. P. Kinetochores are transported poleward along a single astral microtubule during chromosome attachment to the spindle in newt lung cells. J. Cell Biol. 110, 81–95 (1990).
Yang, Z., Tulu, U. S., Wadsworth, P. & Rieder, C. L. Kinetochore dynein is required for chromosome motion and congression independent of the spindle checkpoint. Curr. Biol. 17, 973–980 (2007).
Cai, S., O’Connell, C. B., Khodjakov, A. & Walczak, C. E. Chromosome congression in the absence of kinetochore fibres. Nat. Cell Biol. 11, 832–838 (2009).
Craske, B. & Welburn, J. P. I. Leaving no-one behind: how CENP-E facilitates chromosome alignment. Essays Biochem. 64, 313–324 (2020).
Kapoor, T. M. et al. Chromosomes can congress to the metaphase plate before biorientation. Science 311, 388–391 (2006).
McEwen, B. F. et al. CENP-E is essential for reliable bioriented spindle attachment, but chromosome alignment can be achieved via redundant mechanisms in mammalian cells. Mol. Biol. Cell 12, 2776–2789 (2001).
Renda, F. et al. Non-centrosomal microtubules at kinetochores promote rapid chromosome biorientation during mitosis in human cells. Curr. Biol. 32, 1049–1063.e4 (2022).
Shrestha, R. L. & Draviam, V. M. Lateral to end-on conversion of chromosome-microtubule attachment requires kinesins CENP-E and MCAK. Curr. Biol. 23, 1514–1526 (2013).
Wandke, C. et al. Human chromokinesins promote chromosome congression and spindle microtubule dynamics during mitosis. J. Cell Biol. 198, 847–863 (2012).
Vukušić, K. & Tolić, I. M. CENP-E initiates chromosome congression by opposing Aurora kinases to promote end-on attachments. Nat. Commun. 16, 8537 (2025).
Dick, A. E. & Gerlich, D. W. Kinetic framework of spindle assembly checkpoint signalling. Nat. Cell Biol. 15, 1370–1377 (2013).
Ferrandiz, N., Downie, L., Starling, G. P. & Royle, S. J. Endomembranes promote chromosome missegregation by ensheathing misaligned chromosomes. J. Cell Biol. 221, e202203021 (2022).
Gomes, A. M. et al. Micronuclei from misaligned chromosomes that satisfy the spindle assembly checkpoint in cancer cells. Curr. Biol. 32, 4240–4254.e5 (2022).
Kabeche, L. & Compton, D. A. Cyclin A regulates kinetochore microtubules to promote faithful chromosome segregation. Nature 502, 110–113 (2013).
Tucker, J. B. et al. Misaligned chromosomes are a major source of chromosomal instability in breast cancer. Cancer Res. Commun. 3, 54–65 (2023).
Ertych, N. et al. Increased microtubule assembly rates influence chromosomal instability in colorectal cancer cells. Nat. Cell Biol. 16, 779–791 (2014).
Tamura, N. et al. Specific mechanisms of chromosomal instability indicate therapeutic sensitivities in high-grade serous ovarian carcinoma. Cancer Res. 80, 4946–4959 (2020).
Bolzer, A. et al. Three-dimensional maps of all chromosomes in human male fibroblast nuclei and prometaphase rosettes. PLoS Biol. 3, e157 (2005).
Girish, V. et al. Oncogene-like addiction to aneuploidy in human cancers. Science 381, eadg4521 (2023).
Tovini, L. & McClelland, S. E. Impaired CENP-E function renders large chromosomes more vulnerable to congression failure. Biomolecules 9, 44 (2019).
Koprivec, I., Štimac, V. & Tolić, I. M. Super-resolution imaging of mitotic spindle microtubules using STED microscopy. Methods Mol. Biol. 2872, 3–19 (2025).
Kiewisz, R. et al. Three-dimensional structure of kinetochore-fibers in human mitotic spindles. eLife 11, e75459 (2022).
Shi, Y. et al. Smart lattice light-sheet microscopy for imaging rare and complex cellular events. Nat. Methods 21, 301–310 (2024).
Magidson, V. et al. The spatial arrangement of chromosomes during prometaphase facilitates spindle assembly. Cell 146, 555–567 (2011).
Griffis, E. R., Stuurman, N. & Vale, R. D. Spindly, a novel protein essential for silencing the spindle assembly checkpoint, recruits dynein to the kinetochore. J. Cell Biol. 177, 1005–1015 (2007).
Mazumdar, M., Sundareshan, S. & Misteli, T. Human chromokinesin KIF4A functions in chromosome condensation and segregation. J. Cell Biol. 166, 613–620 (2004).
Yajima, J. et al. The human chromokinesin Kid is a plus end-directed microtubule-based motor. EMBO J. 22, 1067–1074 (2003).
Wood, K. W. et al. Antitumor activity of an allosteric inhibitor of centromere-associated protein-E. Proc. Natl. Acad. Sci. USA. 107, 5839–5844 (2010).
Pereira, C. et al. Self-assembly of the RZZ complex into filaments drives kinetochore expansion in the absence of microtubule attachment. Curr. Biol. 28, 3408–3421.e8 (2018).
Sacristan, C. et al. Dynamic kinetochore size regulation promotes microtubule capture and chromosome biorientation in mitosis. Nat. Cell Biol. 20, 800–810 (2018).
Wu, J. et al. A farnesyl-dependent structural role for CENP-E in expansion of the fibrous corona. J. Cell Biol. 223, e202303007 (2023).
Kapitein, L. C. et al. The bipolar mitotic kinesin Eg5 moves on both microtubules that it crosslinks. Nature 435, 114–118 (2005).
Kashina, A. S., Rogers, G. C. & Scholey, J. M. The bimC family of kinesins: essential bipolar mitotic motors driving centrosome separation. Biochim. Biophys. Acta 1357, 257–271 (1997).
Skoufias, D. A. et al. S-trityl-L-cysteine is a reversible, tight binding inhibitor of the human kinesin Eg5 that specifically blocks mitotic progression. J. Biol. Chem. 281, 17559–17569 (2006).
Mayer, T. U. et al. Small molecule inhibitor of mitotic spindle bipolarity identified in a phenotype-based screen. Science 286, 971–974 (1999).
Groen, A. C. et al. A novel small-molecule inhibitor reveals a possible role of kinesin-5 in anastral spindle-pole assembly. J. Cell Sci. 121, 2293–2300 (2008).
Rickert, K. W. et al. Discovery and biochemical characterization of selective ATP competitive inhibitors of the human mitotic kinesin KSP. Arch. Biochem. Biophys. 469, 220–231 (2008).
Rajendraprasad, G., Eibes, S., Boldú, C. G. & Barisic, M. TH588 and low-dose nocodazole impair chromosome congression by suppressing microtubule turnover within the mitotic spindle. Cancers 13, 5995 (2021).
Alexander, S. P. & Rieder, C. L. Chromosome motion during attachment to the vertebrate spindle: initial saltatory-like behavior of chromosomes and quantitative analysis of force production by nascent kinetochore fibers. J. Cell Biol. 113, 805–815 (1991).
Kuhn, J. & Dumont, S. Spindle assembly checkpoint satisfaction occurs via end-on but not lateral attachments under tension. J. Cell Biol. 216, 1533–1542 (2017).
Waters, J. C., Chen, R. H., Murray, A. W. & Salmon, E. D. Localization of Mad2 to kinetochores depends on microtubule attachment, not tension. J. Cell Biol. 141, 1181–1191 (1998).
Mack, G. J. & Compton, D. A. Analysis of mitotic microtubule-associated proteins using mass spectrometry identifies astrin, a spindle-associated protein. Proc. Natl. Acad. Sci. USA. 98, 14434–14439 (2001).
Schmidt, J. C. et al. Aurora B kinase controls the targeting of the Astrin-SKAP complex to bioriented kinetochores. J. Cell Biol. 191, 269–280 (2010).
Shrestha, R. L. et al. Aurora-B kinase pathway controls the lateral to end-on conversion of kinetochore-microtubule attachments in human cells. Nat. Commun. 8, 150 (2017).
Yao, X., Anderson, K. L. & Cleveland, D. W. The microtubule-dependent motor Centromere–associated Protein E (CENP-E) is an integral component of kinetochore corona fibers that link centromeres to spindle microtubules. J. Cell Biol. 139, 435–447 (1997).
Hewitt, L. et al. Sustained Mps1 activity is required in mitosis to recruit O-Mad2 to the Mad1–C-Mad2 core complex. J. Cell Biol. 190, 25–34 (2010).
Worrall, J. T. et al. Non-random mis-segregation of human chromosomes. Cell Rep. 23, 3366–3380 (2018).
Stiff, T. et al. Prophase-specific perinuclear actin coordinates centrosome separation and positioning to ensure accurate chromosome segregation. Cell Rep. 31, 107681 (2020).
Stepanova, T. et al. Visualization of microtubule growth in cultured neurons via the use of EB3-GFP (end-binding protein 3-green fluorescent protein). J. Neurosci. 23, 2655–2664 (2003).
Jagrić, M., Risteski, P., Martinčić, J., Milas, A. & Tolić, I. M. Optogenetic control of PRC1 reveals its role in chromosome alignment on the spindle by overlap length-dependent forces. eLife 10, e61170 (2021).
Thompson, S. L. & Compton, D. A. Examining the link between chromosomal instability and aneuploidy in human cells. J. Cell Biol. 180, 665–672 (2008).
Chong, M. K., Rosas-Salvans, M., Tran, V. & Dumont, S. Chromosome size-dependent polar ejection force impairs mammalian mitotic error correction. J. Cell Biol. 223, e202310010 (2024).
Novais-Cruz, M. et al. Mitotic DNA damage promotes chromokinesin-mediated missegregation of polar chromosomes in cancer cells. Mol. Biol. Cell 34, ar47 (2023).
Kalinina, I. et al. Pivoting of microtubules around the spindle pole accelerates kinetochore capture. Nat. Cell Biol. 15, 82–87 (2013).
Woodruff, J. B., Wueseke, O. & Hyman, A. A. Pericentriolar material structure and dynamics. Philos. Trans. R. Soc. Lond. B Biol. Sci. 369, 20130459 (2014).
Gasic, I., Nerurkar, P. & Meraldi, P. Centrosome age regulates kinetochore–microtubule stability and biases chromosome mis-segregation. eLife 4, e07909 (2015).
Kajtez, J. et al. Overlap microtubules link sister k-fibres and balance the forces on bi-oriented kinetochores. Nat. Commun. 7, 10298 (2016).
Simunić, J. & Tolić, I. M. Mitotic spindle assembly: building the bridge between sister K-fibers. Trends Biochem. Sci. 41, 824–833 (2016).
Kaseda, K., McAinsh, A. D. & Cross, R. A. Dual pathway spindle assembly increases both the speed and the fidelity of mitosis. Biol. Open 1, 12–18 (2011).
Leça, N. et al. Proximity-based activation of AURORA A by MPS1 potentiates error correction. Curr. Biol. 35, 1935–1947.e8 (2025).
Vukušić, K. & Tolić, I. M. Kinetochore-centrosome feedback linking CENP-E and Aurora kinases controls chromosome congression. Nat. Commun. 16, 9097 (2025).
Risteski, P. et al. Microtubule poleward flux as a target for modifying chromosome segregation errors. Proc. Natl. Acad. Sci. USA. 121, e2405015121 (2024).
Bakhoum, S. F., Genovese, G. & Compton, D. A. Deviant kinetochore microtubule dynamics underlie chromosomal instability. Curr. Biol. 19, 1937–1942 (2009).
Vasquez, R. J., Howell, B., Yvon, A. M., Wadsworth, P. & Cassimeris, L. Nanomolar concentrations of nocodazole alter microtubule dynamic instability in vivo and in vitro. Mol. Biol. Cell 8, 973–985 (1997).
Li, J. et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 275, 1943–1947 (1997).
Turnham, D. J., Bullock, N., Dass, M. S., Staffurth, J. N. & Pearson, H. B. The PTEN Conundrum: how to target PTEN-deficient prostate cancer. Cells 9, 2342 (2020).
Yin, Y. & Shen, W. H. PTEN: a new guardian of the genome. Oncogene 27, 5443–5453 (2008).
He, J. et al. PTEN regulates EG5 to control spindle architecture and chromosome congression during mitosis. Nat. Commun. 7, 12355 (2016).
Marcozzi, C. & Pines, J. Assays for the spindle assembly checkpoint in cell culture. Methods Cell Biol. 144, 1–13 (2018).
Holland, A. J. et al. Polo-like kinase 4 controls centriole duplication but does not directly regulate cytokinesis. Mol. Biol. Cell 23, 1838–1845 (2012).
Truong, M. A. et al. A kinesin-based approach for inducing chromosome-specific mis-segregation in human cells. EMBO J. 42, e111559 (2023).
Castrogiovanni, C. et al. Evidence for a HURP/EB free mixed-nucleotide zone in kinetochore-microtubules. Nat. Commun. 13, 4704 (2022).
Dumont, M. et al. Human chromosome-specific aneuploidy is influenced by DNA-dependent centromeric features. EMBO J. 39, e102924 (2020).
Koch, A., Maia, A., Janssen, A. & Medema, R. H. Molecular basis underlying resistance to Mps1/TTK inhibitors. Oncogene 35, 2518–2528 (2016).
Lukinavičius, G. et al. Fluorogenic probes for live-cell imaging of the cytoskeleton. Nat. Methods 11, 731–733 (2014).
Štimac, V., Koprivec, I., Manenica, M., Simunić, J. & Tolić, I. M. Augmin prevents merotelic attachments by promoting proper arrangement of bridging and kinetochore fibers. eLife 11, e83287 (2022).
Ponjavić, I., Vukušić, K. & Tolić, I. M. Expansion microscopy of the mitotic spindle. Methods Cell Biol. 161, 247–274 (2021).
Zhang, C., Kang, J. S., Asano, S. M., Gao, R. & Boyden, E. S. Expansion microscopy for beginners: visualizing microtubules in expanded cultured HeLa cells. Curr. Protoc. Neurosci. 92, e96 (2020).
Buđa, R., Vukušić, K. & Tolić, I. M. Dissection and characterization of microtubule bundles in the mitotic spindle using femtosecond laser ablation. Methods Cell Biol. 139, 81–101 (2017).
Ershov, D. et al. TrackMate 7: integrating state-of-the-art segmentation algorithms into tracking pipelines. Nat. Methods 19, 829–832 (2022).
Berg, H. C. Random Walks in Biology: New and Expanded Edition (Princeton University Press, Princeton, N.J, 1993).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Bird, A. W. & Hyman, A. A. Building a spindle of the correct length in human cells requires the interaction between TPX2 and Aurora A. J. Cell Biol. 182, 289–300 (2008).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Choi, S. H. & McCollum, D. A role for metaphase spindle elongation forces in correction of merotelic kinetochore attachments. Curr. Biol. 22, 225–230 (2012).
de Bruijn, I. et al. Analysis and visualization of longitudinal genomic and clinical data from the AACR Project GENIE Biopharma Collaborative in cBioPortal. Cancer Res. 83, 3861–3867 (2023).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Ghandi, M. et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature 569, 503–508 (2019).
Kletter, T. et al. Volumetric morphometry reveals spindle width as the best predictor of mammalian spindle scaling. J. Cell Biol. 221, e202106170 (2022).
Reinhold, W. C. et al. CellMiner: a web-based suite of genomic and pharmacologic tools to explore transcript and drug patterns in the NCI-60 cell line set. Cancer Res. 72, 3499–3511 (2012).
Koprivec, I. et al. Polar chromosomes are rescued from missegregation by spindle elongation-driven microtubule pivoting. figshare https://doi.org/10.6084/m9.figshare.29126831
Acknowledgements
The authors thank Alexey Khodjakov, Jonathon Pines, Andrew Holland, Will Krek, Marin Barišić, Susanne Lens and Julie Welburn for the plasmids and cell lines; Oliver Vugrek and Filip Rokić for help with lentiviral work, Marko Šprem and the Abberior team for help with the Expert Line easy3D STED microscope system setup and developing microscopy protocols; Ivana Šarić for help with the drawings, members of the Tolić, Pavin, Kops, and Meraldi groups for helpful discussions and advice. The authors acknowledge funding by the European Research Council (ERC Synergy Grant, GA Number 855158), the Croatian Science Foundation (HRZZ) through Swiss-Croatian Bilateral Projects (project IPCH-2022-10-9344), HRZZ (project IP-2024-05-5336), and Cooperation Programme with Croatian Scientists in the Diaspora “Research Cooperability” (project PZS-2019-02-7653), and projects co-financed by the Croatian Government and the European Union through the European Regional Development Fund—the Competitiveness and Cohesion Operational Programme: IPSted (Grant KK.01.1.1.04.0057) and QuantiXLie Center of Excellence (Grant KK.01.1.1.01.0004).
Author information
Authors and Affiliations
Contributions
I.K. and V.Š. performed all experiments and analyzed the data, except for the following: experiments on Astrin and the cancer cell lines MDA-MB-231, Caco-2, and a subset of OVSAHO cells were conducted by M.Đ., experiments involving CENP-E inhibition, corona proteins, and U2OS cells with labeled kinetochores and microtubules were carried out by K.V., and experiments on chromosomes 1 and 9 were performed by P.M. I.K., V.Š., and I.M.T. wrote the manuscript with contribution from M.Đ. and K.V. I.M.T. supervised the project.
Corresponding author
Ethics declarations
Competing interests
All authors declare no competing interests at a time when the experiments and analyzes were conducted. At a time of the final revision round, I.K. served as a founder and CEO of Eupraxia (Eupraxia Scientific Ltd.), a science and medical communications company.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Koprivec, I., Štimac, V., Đura, M. et al. Polar chromosomes are rescued from missegregation by spindle elongation-driven microtubule pivoting. Nat Commun 17, 2049 (2026). https://doi.org/10.1038/s41467-026-69830-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69830-1