Abstract
Plant roots release a wide array of metabolites into the rhizosphere, shaping microbial communities and their functions. While metagenomics has expanded our understanding of these communities, little is known about the physiology of their members in host environments. Transcriptome analysis via RNA sequencing is a common approach to learning more, but its use has been challenging because of low bacterial biomass and interference from plant RNA. To overcome this, we developed a randomly-barcoded promoter-library insertion sequencing (RB-PI-seq) combined with chassis-independent recombinase-assisted genome engineering (CRAGE). Using Pseudomonas simiae WCS417 as a model rhizobacterium, this method enabled targeted amplification of barcoded transcripts, bypassing plant RNA interference and allowing measurement of thousands of promoter activities during Arabidopsis root colonization. Our analysis revealed temporally resolved transcriptional regulation, including those associated with cell growth, chemotaxis, plant immune suppression, biofilm formation, and stress responses, reflecting the coordinated physiological adaptation to the root environment. Additionally, we discovered that transcriptional activation of xanthine dehydrogenase and a lysozyme inhibitor is crucial for evading plant immune systems. This framework is scalable to other bacterial species and provides new opportunities for understanding rhizobacterial gene regulation in native environments.
Similar content being viewed by others
Introduction
Plant roots secrete nutrient-rich compounds into the surrounding soil, creating unique habitats that support diverse microbial communities1,2. These microbial communities, in turn, play critical roles in plant physiology by promoting growth, protecting against disease, or, in some cases, causing infection3,4,5. Over the past decade, extensive metagenomic studies have identified the members of microbial communities across various plant species, genotypes, and growth environments6,7,8,9,10. The next major challenge is to develop a framework that predicts microbial gene functions, regulation, and impacts on plant physiology. Achieving this requires in-depth knowledge of the physiology of individual microbial species within their host environments. In particular, studying key organisms such as plant-growth-promoting rhizobacteria and pathogens is critical for advancing sustainable agricultural systems.
Among plant-associated bacteria, Pseudomonas species have been extensively studied because of their roles in promoting plant growth and protecting against disease. In particular, Pseudomonas simiae WCS417 has served as a model bacterium, with traits beneficial to its hosts and a remarkable ability to colonize a wide range of plant species11. Previous studies have uncovered molecular mechanisms underlying these symbiotic relationships, such as siderophore production12 and modulation of plant immune systems13,14. Additionally, transposon mutagenesis sequencing of P. simiae WCS417 identified genetic determinants that influence early-stage root colonization, including through chemotaxis and substrate utilization15. These findings suggest that bacteria engage in sophisticated chemical interactions with host plants during colonization. However, global profiling of these physiological adaptations remains limited.
Transcriptomics is a powerful tool for characterizing the temporal dynamics of global gene expression. In host plants, transcriptomic studies have revealed coordinated cascades of gene expression in response to microbial colonization or microbe-associated molecular patterns16,17,18. Although the host responses have been studied at both population and single-cell levels, the adaptations of rhizobacteria remain less explored. This gap is primarily due to technical challenges, as bacterial RNA is significantly outnumbered by host-plant RNA. This issue is common in plant-associated bacteria, and a similar issue is found in bacteria associated with other hosts19,20,21,22. To address this, various strategies have been explored, including deep RNA-sequencing23,24, depletion of host and bacterial rRNA23,24,25, enrichment of bacterial mRNA using customized probes22,26, and physical separation of bacterial cells from host tissues through techniques such as laser microdissection27, flow cytometry28, and density-gradient centrifugation26. Alternatively, in vitro cell culturing with root exudates has been a common approach. Although the choice of approach depends on the research question, profiling transcriptomes in planta with high efficiency and scalability remains the most desirable option.
Synthetic biology offers an alternative way to profile global transcriptional activities. Using DNA barcodes to report promoter activity enables quantification through a straightforward workflow involving PCR amplification and barcode sequencing. This method has been successfully used in model prokaryotic and eukaryotic organisms, revealing unique features of promoter regulatory elements29,30,31,32. In the present study, we introduced a DNA-barcoded promoter library into P. simiae WCS417, using chassis-independent recombinase-assisted genome engineering (CRAGE)33,34, a versatile strain-engineering technology developed in our group. We then established an experimental workflow to characterize in planta promoter activities by selective amplification of barcoded transcripts, overcoming interference from plant RNA. We term this workflow as “CRAGE randomly-barcoded promoter-library insertion sequencing (CRAGE-RB-PI-seq),” or PI-seq in short. Using this method, we identified a set of uniquely regulated promoters involved in plant-root colonization and validated their activities and functions through independent experiments. The framework we developed enables rapid characterization of transcriptional activities and can be applied to a wide range of bacterial species, providing new insights into gene function and regulation in native environments.
Results
Development of the CRAGE-RB-PI-seq workflow
To generate a barcoded promoter library, we first computationally extracted the 140 bp upstream sequences from the start codons of all annotated protein-coding sequences in the P. simiae WCS417 genome (Fig. 1A). The 140 bp promoter length was selected based on the maximum oligonucleotide size available from Twist Bioscience at the time (200 bp total, including 60 bp of flanking sequences for cloning). This length accommodates ~65% of transcriptional factor binding sites in P. simiae WCS417 (Supplementary Fig. S1) and the typical 20–80 bp distance between transcription start sites (TSSs) and start codons observed in bacteria29,35,36,37. To optimize versatility, we divided promoter libraries into three groups according to the distances to upstream genes: group I (≤30 bp), group II (31–139 bp), and group III (≥140 bp). This reflects typical features of bacterial promoter architecture38; Group I has short intergenic regions and are less likely to contain independent promoters, whereas groups II and III contain intermediate and longer intergenic regions that may include internal or independent promoters together with associated regulatory elements. This grouping later helped us reduce library complexity by selecting groups with active promoters.
A A total of 5541 sequences, each 140 bp sequences immediately upstream of annotated start codons, were computationally extracted from P. simiae WCS417 genome. These promoter sequences were categorized into three groups according to the distances to upstream genes: group I (≤30 bp), group II (31–139 bp), and group III (≥140 bp). B Each promoter library was synthesized, barcoded with 23 bp sequences, and cloned into the CRAGE accessory vector (pW26_mod), which included an apramycin-resistant marker (AprR) flanked by two mutually exclusive lox sites (lox2272 and lox5171), necessary for site-specific recombination. C The plasmids were transformed into an E. coli donor strain and conjugated into P. simiae WCS417 (strain SB599), which harbored a CRAGE landing pad on the chromosome. The landing pad is located between locus tags PS417_RS26490 and PS417_RS26495, without disrupting any coding sequences34. Cre-mediated recombination integrated the barcoded promoters into the landing pad. D The promoter-barcode regions were amplified from the genomic DNA and sequenced via targeted DNA-seq to associate each promoter with its corresponding unique barcodes. E The transcriptional activity of promoters was characterized by growing cells under various conditions, extracting DNA and RNA, and amplifying barcode regions from the DNA and cDNA. The cDNA was synthesized from RNA by selective reverse transcription. Promoter activities were then quantified by normalizing RNA barcode counts to DNA barcode counts for each gene.
Each library group was synthesized, barcoded, and cloned into a CRAGE accessory vector (Fig. 1B), and the resulting constructs were transformed into a conjugal E. coli donor strain. The promoter libraries were then chromosomally integrated into P. simiae WCS417 (strain SB599) using CRAGE (Fig. 1C, Supplementary Table S1). The integration site lies in the middle of an intergenic region, without disrupting any genes34, at nucleotide position 268,402 between locus tags PS417_RS26490 and PS417_RS26495 (both annotated as hypothetical proteins). Transcriptomic and Raman microscopy analyses confirmed that the engineering process did not significantly alter cellular physiology (Supplementary Figs.S2, 3).
DNA sequencing (DNA-seq) was used to link each promoter with unique barcodes (Fig. 1D). This process successfully recovered 91% of promoters (5040 out of 5541), each tagged with unique barcodes (Supplementary Data 1). The pooled cell libraries were cultured under various conditions, both in vitro and in planta (Fig. 1E). Promoter activities were then quantified using targeted RNA-seq and DNA-seq, normalizing RNA-derived barcode counts to DNA-derived counts (Fig. 1E).
Transcriptional profiling of P. simiae WCS417 under various in vitro conditions
To test the functionality of our promoter library, we first grew the pooled library (groups I–III) under five different conditions and prepared sequencing samples for both PI-seq and conventional RNA-seq (Fig. 2A). PI-seq consistently demonstrated high library coverage and reproducibility across conditions. For example, DNA barcode analysis showed that under the reference condition (20 mM glucose), 98.8% of promoters (4982 out of 5040 in the library) were covered at least 50 times per million reads, and the histogram showed a narrow distribution (Fig. 2B). The data reproducibility between biological replicates was high (Pearson’s r > 0.99) (Fig. 2C), indicating sufficient library coverage under these culture conditions.
A A pooled library of groups I–III was grown under five different conditions. PI-seq and RNA-seq libraries were created for each condition (n = 2 biological replicates). B–E Analysis of PI-seq data for the cells grown in 20 mM glucose. DNA barcode counts showed a narrow distribution and high coverage of the designed library (B) with high reproducibility (C). RNA barcode counts normalized to DNA barcode counts showed a large variation in promoter activity. Groups II and III promoters (intergenic distance ≥31 bp) were more active than group I promoters (intergenic distance ≤ 30 bp). log₂ promoter activity is shown in arbitrary units (A.U.) (D). The biological replicates for groups II and III showed high reproducibility of promoter activity, while group I promoter activity was weaker (E). Pearson’s r correlation values for each group are shown. F In silico prediction of operon structure identified 1149 operons and 2094 single genes. Of these 3243 genes, 2880 promoters were cloned into the library. G Comparison of promoter activity under two conditions. PI-seq data under 20 mM glucose (x-axis; blue points) and 20 mM citrate (y-axis; red points) highlights upregulated promoters. Differentially activated promoters were selected based on fold changes (FCs) (log2FC > 2.5) after filtering out low-activity promoters (log2 promoter activity <2). H Comparison of gene expression under 20 mM glucose and 20 mM citrate conditions using RNA-seq data. Genes driven by the promoters identified by PI-seq in (G) are highlighted. I Comparison of log2 fold changes between PI-seq (x-axis) and RNA-seq (y-axis) shows strong quantitative agreement, particularly for promoters with large fold changes (log2FC > 2.5). The fold changes were calculated by analyzing expression under 20 mM citrate condition divided by that under 20 mM glucose condition for each method. Pearson’s r correlation values are provided (all genes: dashed gray line, genes with log2FC > 2.5: solid black line). J Illustrations of the most upregulated promoters and their associated downstream genes. 140 bp promoter regions are highlighted in pink. Source data are provided as a Source Data file.
Promoter activity, determined by normalizing RNA barcode counts to DNA barcode counts, showed a large variation within and between groups I–III, with the activity levels differing by several orders of magnitude (Fig.2D). In particular, group II and III promoters (green and magenta bars in the figure) exhibited overall higher activity than group I promoters (blue bars). The results of biological replicates were highly reproducible for group II and III libraries (Pearson’s r > 0.83), while the results of biological replicates for group I were less reproducible (Pearson’s r ~ 0.52) (Fig.2E), a pattern observed across all growth conditions (Supplementary Fig. S4).
We hypothesized that the lower activity of group I promoters (derived from ≤30 bp intergenic regions) reflected that many of these promoters were located within operons lacking active transcription start sites38. To investigate this, we used Rockhopper39 to predict operon structures in silico, incorporating P. simiae WCS417 genome sequence data and RNA-seq data. This analysis identified 1149 operons comprising 3447 genes, as well as 2094 single isolated genes (Fig.2F, Supplementary Data 2). As expected, most group I promoters were part of operons, while all group III promoters corresponded to either operon start sites or isolated genes. Of the 3243 genes (1149 operonic + 2094 single), 2880 promoters were successfully cloned into our libraries. Subsequent analysis focused on these 2880 promoters to further validate their activities.
Analysis of differentially induced promoters from PI-seq data
We next compared PI-seq data from cells grown under 20 mM glucose conditions with the data from 20 mM citrate conditions. These two carbon sources are abundant components of plant root exudates40 and require distinct transporters and metabolic pathways41, allowing us to assess the promoter library’s performance. Differential analysis identified 32 promoters that were uniquely upregulated under glucose conditions, and 10 promoters that were uniquely upregulated under citrate conditions (Fig. 2G). RNA expression driven by these upregulated promoters was similarly upregulated, as RNA-seq verified these patterns (Fig. 2H). Comparison of fold changes in promoter activity and RNA expression demonstrated a strong quantitative agreement between PI-seq and RNA-seq, as illustrated in Fig. 2I. Notably, there was a strong correlation between the two methods for differentially induced promoters (Fig. 2I, blue and red points, fitted with a solid line: Pearson’s r = 0.91). A subset of promoters did not match RNA-seq results, which may reflect missing regulatory elements (Supplementary Fig. S1) or post-transcriptional regulation42,43 in our library design.
Additional analysis of other culture conditions confirmed similar trends for both PI-seq and RNA-seq (Supplementary Fig. S5, Supplementary Data 3). Unsupervised clustering analysis of the PI-seq data revealed distinct clustering depending on culture conditions, consistent with the pattern in RNA-seq data (Supplementary Fig. S6). Importantly, PI-seq captured physiologically relevant changes in promoter activity (Fig. 2J). For example, under glucose conditions, the two most upregulated promoters (annotated by arrows in Fig. 2G) drive genes (PS417_RS06235, PS417_RS06240) involved in glucose binding and transport, while under citrate conditions, the most upregulated promoter drives an operon that encodes porin and tripartite-tricarboxylate transporter-substrate-binding protein (PS417_RS20775, PS417_RS20780), both functional for citrate uptake44,45. These findings align with expected differences corresponding to the different culture conditions, and the large fold changes in RNA expression were confirmed by RNA-seq (annotated in Fig. 2H). Overall, these analyses suggest that PI-seq is effective at identifying a core set of promoters that are reprogrammed in response to environmental changes.
Transcriptional profiling of P. simiae WCS417 during Arabidopsis root colonization
Using the engineered strains, we investigated P. simiae WCS417 promoter activity during colonization of Arabidopsis roots (Fig. 3A). Pooled promoter libraries from groups II and III were grown in M9–glucose medium and subsequently inoculated with 10-day-old seedlings on phytagel plates supplemented with 0.5X Murashige & Skoog (MS) basal salts. Group I promoters were omitted because of their low activity. 10-day-old seedlings were chosen, because this developmental stage is widely used in other agar-based root colonization studies4,15,46. Whole root samples (~30 seedlings per replicate) were collected at 10 min, 3 h, 24 h, and up to 7 days after inoculation. Promoter activity was quantified based on barcode counts from extracted DNA and RNA.
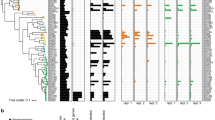
A P. simiae WCS417 populations containing promoter libraries from groups II and III were grown in M9-glucose medium and inoculated onto phytagel plates with 10-day-old Arabidopsis seedlings. Group I was excluded because of low promoter activity (Fig.2). Whole-root samples were collected at 10 min, 3 h, 24 h and up to 7 days after bacterial inoculation. Barcode sequencing libraries were prepared by targeted PCR amplification from extracted RNA and DNA to assess promoter activity (n = 4 biological replicates per time point). PI-seq data from root-colonizing cells (day-7) demonstrated a high library coverage (B) and reproducibility (C), a large variation (D) and high reproducibility of promoter activity across biological replicates (E). F Promoters differentially expressed relative to the M9-glucose condition were identified (|log2FC|> 1 and BH-adjusted p-value < 0.05) with a total of 1009 promoters. Log2 promoter activity was z-score normalized per promoter and visualized as a heat map. Rows were hierarchically clustered and split into five clusters based on their activity profiles. G Principal component analysis of promoter activity across biological replicates. Points are colored by the sampling time point. H Cluster-mean log2 promoter activity values ± SD were plotted across sampling points. Colors match cluster assignments in the heat map. I Predicted operon structures (from Fig.2F) were linked to promoters, and the corresponding genes were subjected to KEGG pathway analysis. J Top-5 enriched KEGG pathways in each cluster. Dashed lines indicate statistical significance thresholds (adjusted p-value < 10−5). Examples of promoter activities: (K) plant immune suppression genes in cluster 4, and (L, M) ROS response and other defense-related genes in cluster 5. Each data point represents the mean log₂ promoter activity ± SD across 4 biological replicates. Source data are provided as a Source Data file.
Despite having ~100 times fewer cells in planta (~2 × 107) than the number of cells in liquid cultures (1.6 × 109) and the presence of ~20 mg of root tissue per sample, PI-seq achieved high library coverage and reproducible results (Fig.3B–E). This demonstrates the utility of our approach, as conventional RNA-seq often requires deep sequencing to overcome plant RNA contamination and obtain sufficient bacterial reads (Supplementary Table S2). Furthermore, host transcriptome analysis confirmed that Arabidopsis inoculated with the PI-seq library strain exhibited transcriptional responses indistinguishable from those inoculated with wild-type P. simiae WCS417 (Supplementary Fig. S7).
To understand colonization dynamics, we compared promoter activities at each sampling point against the M9–glucose condition and identified a total of 1009 differentially expressed promoters. These promoters were grouped into five clusters by hierarchical clustering based on their activity profiles (Fig.3F, Supplementary Data 4). Promoter activities changed most dynamically within the first 24 h, after which they remained relatively stable. This pattern was also evident in principal component analysis (Fig.3G). Each cluster displayed distinct temporal patterns of activation and inactivation (Fig.3H). Promoters in clusters 1 and 2 were highly active in M9–glucose, although cluster 2 exhibited a transient reactivation at 3 h. Cluster 3 and cluster 4 showed transient activation at 10 min and 3 h, respectively. Cluster 5 displayed a gradual increase in activity after 24 h, marking the late stage of colonization. The activities of these promoters were further validated by deep RNA-seq (Supplementary Fig. S8, Supplementary Data 5).
We next examined the genes driven by these promoters using Rockhopper operon structure analysis (Fig.3I). Functional insights from KEGG pathway enrichment revealed distinct pathways in each cluster (Fig.3J). Clusters 1 and 2 were enriched in diverse metabolic pathways, including ABC transporters, biosynthesis of cofactors and amino acids, and ribosome biogenesis, consistent with their preferential expression under M9–glucose conditions. Cluster 3 was enriched for flagellar assembly and bacterial chemotaxis, whose strong induction as early as 10 min implies their importance in early root colonization, consistent with previous reports15,47,48. Cluster 4 was enriched for glycine, serine, and threonine metabolism. Its activation coincided with cluster 2, which included ribosome and amino acid biosynthesis pathways, suggesting that the 3-h induction reflects initiation of cell growth by taking up root exudates. Cluster 5 was enriched for microbial metabolism in diverse environments, two-component systems, and biofilm formation. The functional distinction between cluster 5 and clusters 1–2 suggests a physiological shift from a metabolically active state to adhesive and protective states during colonization, in line with prior studies23,49,50. The detailed analysis of the enriched “metabolic pathways” in clusters 1–2 and 5 further revealed activation of distinct metabolic programs engaged under each condition (Supplementary Fig. S9).
Additionally, several promoters linked to genes critical for bacterial root colonization were upregulated. Cluster 4 included pyrroloquinoline quinone (PQQ) dependent genes (PS417_RS10765, PS417_RS10785, PS417_RS10720) that suppress root immune responses via gluconic acid production14 (Fig. 3K). Cluster 5 included promoters driving catalase (PS417_RS27325) and DNA repair enzymes (PS417_RS21405, PS417_RS14920), both involved in mitigating reactive oxygen species (ROS) likely produced by plants as a defense response51,52 (Fig. 3L). Other upregulated promoters in cluster 5, potentially important for colonization, included those driving a xanthine dehydrogenase (PS417_RS0404) and a hypothetical protein (PS417_RS02110) predicted to be a member of the MliC (membrane-bound lysozyme inhibitor of c-type lysozyme) family protein (Fig. 3M); we experimentally validated the latter two functions, as shown in Fig.4.
A Growth rates of the wild-type and four insertion mutants were compared in M9 media with 20 mM glucose. Growth measurements were performed using n = 3 independent biological replicates. Statistical comparisons were performed using one-way ANOVA, comparing each mutant to wild-type. Mutants are identified by their locus tags with “PS417_” omitted. The bars represent mean ± SD across biological replicates. B Root colonization competition assay. Wild-type and individual mutant strains were mixed 1:1 and applied to seedling plates. After 5 days, wild-type and mutant strain colonies were counted from root samples. The log2 ratios of mutant to wild-type colonies are shown in the graph. Each experiment consisted of independent biological replicates (n = 6 for RS10720, RS18295; n = 7 for RS02110, RS04040), where each replicate corresponds to an independent seedling plate. The bars represent mean ± SD across biological replicates. Statistical significance was assessed using two-sided one-sample t-tests against 0. Exact p-values are shown in the figure (*p < 0.05; **p < 0.01). C Lysozyme assay of the wild-type and a mutant of the MliC protein. Cultures were incubated in PBS, either without (left) or with (right) lysozyme. Viable cell numbers were determined by CFU counts. The assay was performed using n = 3 independent biological replicates. The bars represent mean ± SD across biological replicates. Statistical significance was assessed using two-sided one-sample t-tests comparing log₂ ratios to 0. Exact p-values are shown in the figure (*p < 0.05; ns not significant). D Oxidative stress tolerance assay to examine the function of XDH. Wild-type and mutant strains were grown in M9 media with 20 mM glucose and challenged with 5 mM H2O2 (n = 3 biological replicates: bars represent mean ± SD). E The legend table lists the mutant locus tags and their corresponding gene products. Log₂ fold changes in promoter activity at 24 h and day 7, relative to the M9-glucose condition, are shown based on PI-seq data (Fig.3). Adjusted p-values are provided in parentheses. Source data are provided as a Source Data file.
Characterization of mutant phenotypes
To examine the functions of genes driven by upregulated promoters, we obtained transposon insertion mutants for three genes generated by Cole et al.15 (see Fig.4E legend table) and characterized their metabolic and root colonization phenotypes. In addition, we included a mutant of PS417_RS18295, encoding an undecaprenyl-phosphate glucose phosphotransferase involved in polysaccharide-mediated biofilm formation. Although the promoter of this gene was not identified in the differential expression analysis, the corresponding pathway was enriched in cluster 5 and therefore selected for further phenotypic analysis.
When grown individually, all mutants displayed growth rates comparable to the wild-type strain in M9-glucose minimal medium (Fig.4A). However, in competitive root colonization assays, the wild-type strain significantly outcompeted the mutants (Fig.4B). The reduced colonization observed for mutants of the PQQ-dependent catabolism-associated protein (PS417_RS10720) and the undecaprenyl-phosphate glucose phosphotransferase (PS417_RS18295) is consistent with previous findings that highlight the importance of plant immune suppression14 and biofilm formation53,54, respectively, in root colonization.
The wild-type strain also outcompeted two other mutants carrying insertion mutations in genes PS417_RS02110 and PS417_RS04040 whose roles in the root colonization ability of P. simiae WCS417 had not been previously reported. According to the RefSeq annotation PS417_RS02110 encodes an MliC family protein. The MliC protein from Pseudomonas aeruginosa and Escherichia coli is known to function as a c-type lysozyme inhibitor, aiding in host colonization of humans and other animals55,56. Although P. simiae WCS417’s MliC shares limited protein sequence identity with these homologs, AlphaFold 3 predicted significant structural similarities57 (Supplementary Fig. S10), suggesting a similar lysozyme-inhibiting function. This is especially relevant given recent studies showing that lysozyme-like hydrolases are produced by Arabidopsis58 and epiphytic fungi59, potentially influencing microbial communities. While Pseudomonas species are generally lysozyme-resistant, the role of MliC may be critical in the presence of agents that destabilize the outer membrane of the bacterium, thereby increasing its susceptibility to external lysozymes60,61. To investigate whether MliC increases lysozyme tolerance in P. simiae WCS417, we examined the effect of lysozyme on the wild-type strain and a mutant carrying an insertion in mliC. The mutant showed reduced cell numbers compared to the wild-type in the presence of lysozyme, whereas no difference was observed between them in the absence of lysozyme (Fig. 4C), indicating that MliC contributes to P. simiae WCS417’s protection against lysozymes from plants and from other microbes.
The second gene, PS417_RS04040, encodes xanthine dehydrogenase (XDH), whose transcriptional activation has been linked to ROS tolerance and host infection in Ralstonia solanacearum62 and in Borrelia burgdorferi63. Since ROS generation is a common plant immune response, we hypothesized that XDH in P. simiae WCS417 plays a role in neutralizing plant-generated ROS. To test this, we examined the effect of ROS stress on the wild-type strain and a mutant carrying an insertion in xdh during exponential growth in M9 minimal medium with glucose. Upon exposure to 5 mM hydrogen peroxide, both wild-type and mutant strains experienced growth arrest, while the mutant showed a longer arrest by ~60 min compared to the wild-type strain (Fig. 4D), supporting our hypothesis. These findings suggest XDH may contribute to root colonization by working in concert with other ROS-responsive enzymes such as catalases and DNA ligases.
Application of PI-seq to Arabidopsis grown in a soil-like system
To test whether PI-seq is applicable to plants grown in a more natural system, we inoculated pooled promoter libraries from groups II and III (grown in M9–glucose medium) into the rhizosphere of 10-day-old Arabidopsis seedlings cultivated in clay (particle size ~4 mm) supplemented with 0.5× MS basal salts. Root samples were collected after 7 and 21 days (Fig.5A), extending the colonization period compared to the phytagel plate experiments. Clay was selected as the substrate because it resembles the physical properties of soil while also permitting gentle separation from Arabidopsis roots that are fine and fragile (Supplementary Fig. S11).
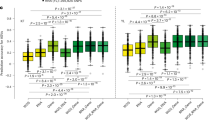
A P. simiae WCS417 populations containing promoter libraries from groups II and III were grown in M9–glucose medium and inoculated into clay pots with 10-day-old Arabidopsis seedlings. Root samples were collected after 7 and 21 days. Barcode sequencing libraries were prepared by targeted PCR amplification from extracted RNA and DNA to quantify promoter activity (n = 3 biological replicates per time point). B Library coverage based on DNA barcode data at day 7. C Reproducibility of DNA barcode counts across two biological replicates at day 7. D Distribution of promoter activity at day 7, showing ~30% inactive promoters. E Reproducibility of promoter activity across two biological replicates at day 7. F–I Same analyses as in panels (B–E), but using day 21 samples. J Volcano plot comparing promoter activity between clay and phytagel at day 7; promoters driving genes in central carbon metabolism are highlighted. K Volcano plot comparing promoter activity between day 7 and day 21 in clay; promoters driving stress response and maintenance genes are highlighted. For panels (J, K), statistical significance was assessed using empirical Bayes–moderated two-sided t-tests with Benjamini–Hochberg correction for multiple comparisons. Source data are provided as a Source Data file.
As in the phytagel experiments, DNA barcode analysis confirmed high library representation: 98% of the 3382 promoters in the groups II and III library were detected at least 50 times per million reads in both day 7 and 21 samples, demonstrating sufficient library coverage (Fig. 5B, F). Biological replicates of DNA barcode counts were reproducible (Fig. 5C, G), although correlations were lower than those observed in phytagel plates. In contrast, RNA barcodes were recovered from a subset of the library: promoter activity was detected in 70% of promoters at day 7 and 69% at day 21, with the remaining ~30% inactive (Fig.5D,H). This reduced activity resulted in weaker correlations across biological replicates compared to phytagel experiments (Fig. 5E, I). Deep RNA-seq on these samples also revealed that only 0.02% of reads mapped to P. simiae in clay-grown samples, compared to ~0.10% in phytagel plates (Supplementary Table S2). This lower RNA recovery likely results from 1) adsorption of root exudates to clay particles64, 2) reduced nutrient availability in the open environment, and 3) decreased metabolic activity during the late-stage colonization (Fig. 3), all contributing to reduced transcriptional activity.
We next compared promoter activity at day 7 between phytagel plates and clay, filtering out promoters with zero activity in the clay dataset. Differential expression analysis identified 3 promoters upregulated and 81 downregulated in clay relative to phytagel (Fig. 5J, Supplementary Data 6). The downregulated set in clay included genes associated with central carbon metabolism, consistent with lower metabolic activity in soil-like conditions. Comparing day 7 and day 21 in clay samples revealed 31 upregulated and 4 downregulated promoters at the later time point. The upregulated group included promoters driving genes for stress response and cellular maintenance such as peroxiredoxin, XDH, ClpS, and Fe-S biogenesis protein NfuA (Fig. 5K, Supplementary Data 6), consistent with the phytagel experiments, where the late-stage colonizers further shifted toward stress tolerance and long-term survival.
Discussion
Plant-growth-promoting rhizobacteria offer a promising avenue for enhancing plant biomass and crop yields without high chemical fertilizer use. To harness their full potential, it is critical to understand their in planta physiology, including gene functions and regulatory mechanisms. However, studying these rhizobacteria in host-associated environments is challenging because of their low biomass. Traditional RNA-seq approaches for plant-colonizing bacteria often require large sample sizes and extensive rRNA depletion, followed by deep-sequencing. Even then, less than 1% of reads map to bacterial gene coding sequences, with a large proportion mapping to plants23,26 Further optimizations such as improved combinatorial rRNA removals24 and physical separation of cells prior to lysis26 are necessary to enrich bacterial mRNA when RNA-seq approaches are used.
In this study, we developed CRAGE-RB-PI-seq in the model rhizobacterium P. simiae WCS417 to profile transcriptional activities during Arabidopsis root colonization. PI-seq overcomes the inherent limitations of conventional RNA-seq by amplifying RNA barcodes from a pre-defined promoter library, thereby enriching bacterial transcripts by nearly 100% and enabling high-resolution transcriptional profiling from low input materials. In our experiments, promoter activities were quantifiable from as few as ~107 bacterial cells collected from ~20 mg of plant root tissue and the input of bacterial cell amount could be reduced even further by increasing the number of PCR cycles. Beyond sensitivity, PI-seq offers several practical advantages. First, by focusing on sequencing short barcode amplicons, it markedly decreases the required sequencing depth, reagent consumption, and data volume. Second, this approach is also broadly applicable, as CRAGE facilitates integration of barcoded promoter libraries into diverse bacterial hosts. Our current strain collection encompasses ~50 species across multiple phyla, many of which were isolated from soil environments (Supplementary Table S3). Finally, the use of unique barcodes allows clear differentiation of homologous genes across species, addressing a common challenge in metatranscriptomic analyses.
For library design, we selected the 140 bp sequences that were immediately upstream of individual genes as promoters because of the technical limitations of the oligo pools available at the time from Twist Biosciences. Despite this constraint, our results showed high correlation between PI-seq and RNA-seq, particularly in identifying differentially expressed genes (Fig.2I, J, Supplementary Figs. S5, S8). We expect that extending the promoter regions would result in even greater accuracy as it covers a broader range of transcriptional factor binding sites (Supplementary Fig. S1); this is now possible with new oligo pools that can harbor 240 bp sequences (Twist Biosciences). Advances in oligo synthesis technologies such as enzymatic oligo synthesis and DropSynth will also allow construction of longer promoter sequences65,66, improving library quality. Additionally, results could be further enhanced by refining library design to capture entire promoter regions, including regulatory elements and transcriptional start sites, which could be facilitated by 5’-enriched RNA-seq methods such as differential RNA-seq67 and CAGE68. The success of such approaches will also depend on high-quality genome annotations and operon predictions.
Using PI-seq, we identified temporally resolved promoter activations during root colonization, including those associated with chemotaxis, plant immune suppression, biofilm formation, and ROS response, consistent with previous genetic studies14,15,47,48. In addition, we uncovered previously unrecognized transcriptional changes, such as the late-stage activation of the xdh and mliC genes, which further enhance the mutualistic interactions between P. simiae WCS417 and Arabidopsis through ROS neutralization and lysozyme inhibition, respectively (Fig.4). Because plant responses and root exudates strongly influence bacterial states, future studies could examine how specific metabolites and signaling molecules regulate bacterial gene expression. For example, Arabidopsis roots-derived coumarins dynamically regulate genes involved in motility and biofilm formation in P. simiae WCS41769.
Notably, we observed significant dynamics in promoter activity during the first 24 h, highlighting this period as critical for successful bacterial adaptation to the root environment. By contrast, relatively little change was observed at later time points (>24 h) in both phytagel and soil experiments, likely reflecting reduced metabolic activity as cells enter a dormant phase after depleting locally available plant-derived nutrients. While our current analyses used whole-root samples, future studies employing spatially resolved sampling or image-based transcriptional readouts could uncover finer spatial regulation of bacterial gene expression. For example, root tips secrete large amounts of metabolites, which can locally boost bacterial activity70. Although Arabidopsis roots are too small and fragile for such spatial dissection, plants with larger root systems, such as Brachypodium, may be suitable models for future studies.
Among all promoters, the strongest upregulation occurred for those driving PQQ-related systems (PS417_RS10765 and PS417_RS10720), showing 206-fold and 24-fold increases, respectively, at 3 h of colonization (Fig. 3K). Notably, PS417_RS10720 controls a downstream response regulator transcription factor (PS417_RS10715), which shares 86% sequence identity with AgmR (also known as PedR1) of Pseudomonas aeruginosa PAO1, a key regulator of PQQ biosynthesis71. Although PI-seq analysis did not detect activation of the downstream PQQ biosynthesis operon (pqqBCDE), likely due to missing upstream regulatory sites, we confirmed its strong induction through deep RNA-seq (Supplementary Fig. S12). PQQ is a known cofactor involved in sugar and alcohol oxidation, with the metabolic byproducts that contribute to plant immune suppression by modulating local pH14. PQQ can also function as a growth-promoting factor when directly applied to plants72,73, and it stimulates production of natural products such as antibiotics in a broad range of Actinobacteria74. These findings underscore the vital role of PQQ production in the rhizosphere and its potential impact on microbial community structure and plant physiology.
Although PI-seq requires further refinement, this method provides a valuable tool for identifying regulatory elements associated with host-microbe interactions, particularly in non-model bacteria. Given the diverse roles that bacteria play in host physiology, such as biofertilization, biostimulation, and biocontrol, there is ample opportunity to uncover new molecular mechanisms. This knowledge will be instrumental in advancing the use of biologics, as well as in developing new biosensors and genetic circuits that can mediate interactions between microbes and their hosts75,76,77,78.
Methods
Bacterial strains
We based our study on Pseudomonas simiae WCS417, originally obtained from Dr. Corne Pieterse (Utrecht University). This strain was domesticated by integrating a landing pad, resulting in strain SB599, as detailed by Wang et al.34. The barcoded transposon library was generated by Cole et al.15. Individual insertion mutants were isolated at the Joint Genome Institute. E. coli WM3064 was obtained from Dr. William Metcalf (University of Illinois).
Design of barcoded promoter library
We used the annotated genome of Pseudomonas simiae WCS417 (NCBI: NZ_CP007637) for a promoter library design. Individual genes were categorized into three groups based on the distances to upstream genes: group I (≤30 bp), group II (31–139 bp), and group III (≥140 bp). We extracted the 140 bp regions immediately upstream of the start codon for all genes using Geneious software (Dotmatics). To facilitate cloning, 30 bp priming sites were added to both ends of the 140 bp sequences. The three groups of 200 bp DNA libraries, referred to as promoter pools, were then synthesized by Twist Bioscience.
For the DNA barcodes, we designed 23 bp sequences, with the first three nucleotides set to CGT, followed by GGA, AGG, or GAG, depending on the group (I, II, or III). The remaining 17 bp were randomized. The first three nucleotides were designed to distinguish different strains for future studies, while the second set corresponded to the groups I-III, allowing for clear identification between them. We added 30 bp priming sites to both ends of the barcode sequences for cloning purposes, and the three groups of 83 bp DNA libraries were synthesized by Integrated DNA Technologies. These are referred to as barcode pools.
Library synthesis, assembly, transformation, and conjugation
The following procedures were conducted in parallel for group I-III libraries. The oligo pools for the promoter and barcode pools were resuspended to 10 ng/µl and 10 µM, respectively. dsDNA pools were generated via PCR, using 1 µl of promoter libraries, 2.5 µl of barcode libraries, and 2.5 µl of 10 µM 5’-forward primers. PCR reactions were performed for 7 cycles in a total volume of 50 µl using Q5® Hot Start High-Fidelity DNA Polymerase (NEB: M0493L). The amplified products were subsequently gel-purified.
For the conjugal vector, we used a modified version of pW2634 (pW26_mod: available upon request), which contains two BsaI cloning sites and sfGFP, flanked by two mutually exclusive lox sites (lox2272 and lox5171). The vector was digested with BsaI (NEB: R3733S) overnight and dephosphorylated using a quick dephosphorylation kit (NEB: M0525S). The digested vector was gel-purified and Gibson assembly was performed using ~23 ng of the digested vector and ~12 ng of dsDNA pools in a 10 µl reaction volume with a NEBuilder® HiFi DNA Assembly Master Mix (NEB: E2621S).
The assembled plasmids were transformed into E. coli EC100 pir+ competent cells (Lucigen: ECP09500) for amplification: After diluting the plasmids 5-fold in water, 1 µl was gently mixed with 20 µl of competent cells on ice. Following electroporation, a small aliquot of the recovery cultures was diluted and plated to determine cloning efficiency, while 500 µl of the cultures were diluted in 3 ml LB and spread onto bioassay dishes (VWR: 73520-774) containing LB agar and 50 µg/ml apramycin. The coverage of all group I-III libraries was determined to be >200×, calculated by dividing the number of colony-forming units by the size of the designed library. Plasmid DNA was then extracted from the colonies using a midiprep kit (Promega: A2492).
The extracted plasmids were subsequently transformed into E. coli WM3064 competent cells to be used as the donor strain for conjugation. After transformation, the cells were recovered in LB supplemented with 0.3 mM diaminopimelic acid (DAP) for 1.5 h at 30 °C, and spread onto bioassay dishes containing LB agar, 50 µg/ml apramycin, and 0.3 mM DAP. The following day, cells were scraped from the plates, resuspended in 10 ml LB medium, and prepared for conjugation.
Library integration into the chromosome of P. simiae WCS417 was performed using CRAGE via conjugation, as described by Wang et al.33 and Wang et al.34. We used P. simiae strain SB599, which harbors a Cre recombinase gene and a kanamycin-resistant marker (KmR) flanked by two mutually exclusive lox sites (lox2272 and lox5171) in the landing pad, as the recipient strain. The landing pad is located at nucleotide position 268,402 between locus tag PS417_RS26490 and PS417_RS26495 without disrupting any genes34. For recombination, both donor and recipient strains were grown overnight in LB medium at 28 °C. The next morning, the cultures were washed twice with LB medium and mixed at a 4:1 (donor:recipient) ratio based on OD600 to a total volume of 1 ml. Then the mixed cultures were washed once again and resuspended in 100 µl of LB + 0.3 mM DAP medium. The mixture was plated onto LB agar plates containing 0.3 mM DAP and incubated overnight at 28 °C.
The following day, all colonies were collected using a loop, resuspended in 1 ml LB medium, washed once, and spread onto bioassay dishes containing LB agar and 50 µg/ml apramycin. One day later, colonies were scraped from the plates, resuspended in 10 ml LB containing 10% glycerol, and stored at −80 °C for later experiments. The generated cell library population is available upon request.
Bacterial cell culturing in liquid media
A 10 µl glycerol stock containing each group I-III library was inoculated into 3 ml of fresh LB medium supplemented with 100 µg/ml apramycin and grown to saturation at 28 °C in a shaking incubator at 200 rpm. After pooling the group I-III cultures in equal ratios, 30 µl of the cultures were transferred into 3 ml of fresh M9-based growth media containing different nutrients and grown overnight. The following day, mid-log phase cultures were diluted to an OD600 = 0.05–0.1 in pre-warmed growth media and cultured again at 28 °C (or 37 °C for heat stress conditions).
When the OD600 reached ~1.0 (~8 × 108 cells/ml), 2 ml of the cultures were pelleted, and the supernatant was removed. The pellets were resuspended in 750 µl DNA/RNA Shield (Zymo Research: R1100), and the cells were lysed for 10 min using 0.1 mm and 0.5 mm beads at maximum speed. DNA and RNA were extracted in parallel using the ZymoBIOMICS DNA/RNA Miniprep kit (Zymo Research: R2002).
The growth media were based on M9 minimal medium, containing 1× M9 minimal salts (Gibco: A1374401), 2 mM magnesium sulfate, 0.1 mM calcium chloride, and 10 µM ferrous sulfate. The primary carbon sources used were 20 mM glucose, 40 mM glycerol, or 20 mM citrate. For high osmolarity conditions, sodium chloride was added to a final concentration of 300 mM.
Plant growth conditions
Arabidopsis thaliana Columbia-0 (Col-0) seeds were surface-sterilized in 70% ethanol for 5 min, followed by treatment with 50% bleach plus 0.1% Triton-X100 for another 5 min. The sterilized seeds were washed 5 times with sterile water and stratified in the dark for at 4 °C for 2–4 days. After stratification, ~100 seeds were plated on a nylon mesh filter (100 µm pore size [Genesee Scientific: 57-103], cut to ~8 cm2) placed on top of plant growth media solidified with phytagel. The media consisted of 0.5× Murashige and Skoog basal salts (PhytoTech Labs: M404), 2.5 mM MES (Sigma-Aldrich: M3671), and 0.6% phytagel (Sigma-Aldrich: P8169), with the pH adjusted to 5.7 using potassium hydroxide (Sigma-Aldrich: 319376), poured in a 10 cm square petri dish (Carolina Biological supply Company: 741470). The phytagel plates were sealed with micropore surgical tape (VWR: 56222-182) and grown upright in a Percival incubator (Geneva Scientific: CU36L5) for 10 days under long-day mode (16 h light and 8 h dark cycle) at 22 °C until exposure to bacterial cells. No humidity control was performed during the incubation.
Bacterial root colonization assay
A 10 µl glycerol stock of the group II and III promoter libraries was inoculated into 3 ml of fresh LB medium supplemented with 100 µg/ml apramycin and grown for 5–6 h at 28 °C in a shaking incubator (200 rpm) until the late exponential phase. Equal volumes of group II and III cultures were then pooled, and 3 µl of the mixed culture was transferred into 3 ml of M9 medium containing 20 mM glucose as the carbon source. After overnight incubation to mid-log phase, cells were harvested by centrifugation (3000 × g, 1 min), washed twice by pelleting and resuspension in 0.5× MS liquid medium, and finally resuspended to an OD600 of 0.5.
For phytagel plate experiments, 50 µl of the suspension was spread onto fresh phytagel plates using 5–10 sterile glass beads. Ten-day-old Arabidopsis seedlings grown on nylon mesh filters were transferred onto the plates using sterile tweezers. For clay experiments, 200 µl of the suspension was applied directly to the rhizosphere of Arabidopsis transplanted from germination plates to 130 g of clay supplied with 110 ml of 0.5× MS liquid medium in a vented vessel (PhytoTech Labs: C2110). Details of the clay substrate are described in Sasse et al.64.
Plates and vessels were incubated upright in a Percival incubator at 22 °C under a 16-h light and 8-h dark cycle. Root samples were collected at 10 min, 3 h, 24 h, and up to 7 days for phytagel experiments, and at 7 days and 21 days for clay experiments, following bacterial inoculation.
Raman microscopy
Wild-type and group I–III library strains were pre-cultured in M9 glucose media. After overnight growth, 1.4 ml of cell culture was collected, washed twice with sterile water, and resuspended in 10 µl of sterile water. An aliquot (1.5 µl) of the suspension was spotted onto quartz slides (Ted Pella: 26012) and air-dried at room temperature for 40 min. Raman spectra of dried cells were collected using a Horiba Jobin Yvon LabRAM ARAMIS confocal Raman microscope equipped with a 532 nm excitation laser (50 mW; Laser Quantum), a 532 nm long-pass filter (Semrock), a 1200 grooves/mm grating, a 200 µm pinhole, and a 100× 0.9 NA objective (Olympus). To minimize photodamage and ensure representative sampling, 285 spectra per sample were acquired by scanning the laser spot across a 19 × 15 pixel grid, with a pitch of 3 µm between pixels and an acquisition time of 1 s per pixel. Principal Component Analysis (PCA) was applied, when necessary, to distinguish spectra corresponding to cells from those lacking signal or corresponding to the quartz substrate. Cell spectra were then averaged.
Library preparation to associate promoters and barcodes
A 10 µl glycerol stock containing each group I-III library was inoculated into 3 ml of fresh LB media supplemented with 100 µg/ml apramycin and grown overnight at 28 °C in a shaking incubator at 200 rpm. After overnight growth, 300 µl of the saturated cultures were pelleted, and the supernatant was removed. The pellets were resuspended in 750 µl DNA/RNA Shield, and the cells were lysed for 10 min using 0.1 mm and 0.5 mm beads at maximum speed. DNA was extracted from each group I-III sample using a miniprep kit (Zymo Research: R2002).
For the first PCR, a region spanning the promoter to the barcode was amplified using specific primers (Supplementary Table S4: map_fwd/rev). The same PCR settings as described below for the barcode amplifications were used, except the number of cycles was set to 15. A second PCR was then performed to add indexes and Illumina P5 and P7 adaptors. The resulting amplicons were sequenced on Illumina NovaSeq platforms using 250 bp paired-end sequencing.
Nucleic acid extraction from root samples
DNA and RNA were extracted from root samples using the ZymoBIOMICS DNA/RNA Miniprep Kit (Zymo Research: R2002) with a modified protocol to ensure efficient isolation and lysis of bacterial cells. To facilitate this, two different sizes of bashing beads were used. Seedlings were cut below the root/shoot junction, and the isolated roots were placed into 2 ml tubes. Roots from one plate (~30 seedlings, ~20 mg) were pooled into a single sample.
The pooled roots were vortexed for 5 s in 800 µl of M9 buffer to remove loosely adhered cells (and also clay when used) from the root surface. After the buffer was removed, 800 µl of DNA/RNA Shield and 2 mm beads (Zymo Research: S6003-50) were added to the tubes. The tubes were then placed onto an adapter (Zymo Research: S5001-7) attached to a Vortex Genie2, and the samples were ground for 10 min at maximum speed. Once the root tissues were disrupted, the lysed samples were transferred to tubes containing 0.1 mm and 0.5 mm beads from the R2002 kit and ground for another 40 min at maximum speed to lyse the bacterial cells. Following this, we used the parallel DNA and RNA extraction procedure, including DNAse treatment, as described in the manufacturer’s protocol (Zymo Research: R2002). DNA and RNA were eluted in 70 µl of ddH2O and stored at −20 °C.
Library preparation for barcode amplicon
To create sequencing libraries, barcoded regions of genomic DNA or cDNA were amplified in a two-step PCR process. For genomic DNA samples, DNA was amplified using primers flanking the barcode region (Supplementary Table S4: barcode_fwd/rev). Six tubes of 50 µl reactions were prepared for each sample, with the following components: l of Q5 reaction buffer (New England Biolabs: B9027S); 1 µl of dNTP (New England Biolabs: N447L); µl of 10 µM forward primer (barcode_fwd); 2.5 µl of 10 µM reverse primer (barcode_rev); 10 µl of purified DNA; 0.5 µl of Q5 polymerase (New England Biolabs: M0493L); 10 µl of betaine solution (Sigma-Aldrich: B-0300); 13.5 µl of ddH2O.
The PCR conditions were as follows: (1) initial heating at 98 °C for 30 s, (2) 14 cycles of 10 s at 98 °C and 60 s at 72 °C, (3) A final extension at 72 °C for 60 s.
The six reactions were pooled into a single sample and purified using the DNA Clean & Concentrate Kit (Zymo Research: D4013), eluted into 20 µl of ddH2O, and quantified using a Qubit fluorometer (Thermo Scientific).
For the second PCR, we added indexes and Illumina P5 and P7 adaptors to the samples. A 50 µl reaction was prepared as follows: l of Q5 reaction buffer (New England Biolabs: B9027S); 1 µl of dNTP (New England Biolabs: N447L); 2.5 µl of 10 µM indexed P5 adapter primer; 2.5 µl of 10 µM indexed P7 adapter primer; ~30 ng of diluted DNA samples; 0.5 µl of Q5® Hot Start High-Fidelity DNA Polymerase (New England Biolabs: M0493L); ddH2O to a total volume of 50 µl.
The PCR conditions were as follows: (1) Initial heating at 98 °C for 30 s, (2) 5 cycles of 10 s at 98 °C, 15 s at 60 °C, and 30 s at 72 °C, (3) A final extension at 72 °C for 60 s.
The samples were then gel-purified and eluted into 30 µl of ddH2O using a cleanup kit (Macherey-Nagel: 740609.050). The quality of the amplicon samples was validated using Bioanalyzer DNA Analysis (Agilent: 5067-1504).
For RNA samples, 70 µl of purified RNA was treated with DNase (Thermo Scientific: AM1907) to remove any trace genomic DNA. RNA quality was checked using Bioanalyzer RNA Analysis (Agilent: 5067-1511), and samples were concentrated with the RNA Clean & Concentrate Kit (Zymo Research: D1013). Selective reverse transcription was carried out using SuperScript II Reverse Transcriptase (invitrogen: 18064022) with a 1st strand synthesis primer (Supplementary Table S4: cDNA_syn), following the manufacturer’s protocol. Synthesized cDNA was amplified by PCR in eight tubes of 50 µl reactions, each containing the same components as for genomic DNA amplification, except using 2 µl of cDNA and 21.5 µl of ddH2O. The PCR conditions were as follows: (1) Initial heating at 98 °C for 30 s; (2) 22 cycles of 10 s at 98 °C and 60 s at 72 °C, (3) A final extension at 72 °C for 60 s.
The eight reactions were pooled, purified using the DNA Clean & Concentrate Kit (Zymo Research: D4013) and eluted in 20 µl of ddH2O. The quality of the PCR products was validated using Bioanalyzer High-sensitivity DNA Analysis (Agilent: 5067-4626). Sequencing libraries were prepared by adding indexes and Illumina P5 and P7 adaptors.
For samples from liquid cultures, the cycle numbers of the first PCR were reduced to 12 for genomic DNA and 20 for cDNA. Otherwise, the same procedures were followed. Amplicon samples were sequenced on Illumina NovaSeq platforms using 150 bp paired-end sequencing, targeting 10–20 million reads per sample.
RNA-seq library preparation
For samples from liquid culture, ribosomal RNA (rRNA) was removed from 1 μg of total RNA using an rRNA depletion kit (New England Biolabs: E7860S), and the treated RNA was eluted in 6.5 µl of ddH2O. Sequencing libraries were prepared using the TruSeq Stranded mRNA Library Prep kit (Illumina: 20020594). This procedure involved adding 3 µl of rRNA-depleted RNA into 15 µl of fragment, prime, and finish (FPF) buffer, followed by the manufacturer’s protocol. Library quality was validated using Bioanalyzer DNA Analysis (Agilent: 5067-1504). Samples were sequenced on Illumina NovaSeq platforms using 150 bp paired-end sequencing, targeting 5 million reads per sample.
Additional RNA-seq experiments from liquid culture were later performed to compare the wild-type and promoter library strains (Supplementary Fig. S2). The library preparation and sequencing were carried out by Azenta Life Sciences using their standard bacterial RNA-seq protocol. Deep RNA-seq on root samples (Supplementary Fig. S8) were also processed by Azenta Life Sciences, incorporating dual rRNA depletion for both plant and bacterial RNA. For a subset of RNA samples, rRNA depletion was omitted to evaluate the effect of the treatment (Supplementary Table S2). These samples were sequenced on an Illumina NovaSeq platform using 150 bp paired-end reads, targeting 150 million reads per sample.
Promoter and barcode association
A custom shell script using BBtools (https://sourceforge.net/projects/bbmap/), along with an R script incorporating the Biostrings (https://bioconductor.org/packages/Biostrings) and dplyr (https://dplyr.tidyverse.org/) packages, was used to map barcode sequences to their corresponding promoter sequences. Briefly, 140 bp promoter regions and 20 bp barcodes (excluding the initial CGT specific to P. simiae WCS417) were extracted from read 1 sequences. Promoter-barcode pairs were retained if they were detected more than 100 times in the sequencing reads. Barcodes mapped to two or more distinct promoter sequences were excluded from further analysis. Promoter sequences that exactly matched their native counterparts were then selected. These steps were repeated for each group (I-III). Overall, 91% (5040 promoters) of the total 5541 promoters were successfully assigned unique barcodes, with each promoter associated with an average of 50 unique barcodes.
Barcode quantification
Barcode sequences were extracted and counted from the sequencing data using a custom shell script with BBtools and an R script utilizing Biostrings and dplyr packages. Barcode counts were normalized to sequencing depth for each sample and then summed for each promoter. Library coverage was assessed by normalizing DNA barcode counts to counts per million (CPM). To calculate individual promoter activity (in arbitrary units, A.U.), the barcode counts from RNA-derived cDNA were divided by the barcode counts from genomic DNA.
Differential expression analysis on PI-seq data
For the root colonization assay, statistical analysis was carried out using edgeR and limma packages, with custom modifications based on Law et al.79. Briefly, raw barcode counts were normalized and transformed to log2 counts per million (log2-CPM) with associated precision weights using voom. Linear models were then fitted for each promoter, and empirical Bayes moderation was applied with the trend and robust options enabled. Differential activity was quantified as log2 fold-changes with associated p-values, and p-values were adjusted for multiple testing using the Benjamini–Hochberg method. For the liquid culture experiments, differentially expressed promoters were selected based on |Log2FC|> 2.5, with inactive promoters (log2 promoter activity <2) filtered out. In addition, promoters with zero RNA barcode counts in glucose conditions were excluded from the analysis.
Functional analysis
Operon structures were predicted using Rockhopper39, using the P. simiae WCS417 genome sequence and RNA sequencing reads from five different liquid cultures as input. Before analysis, RNA reads mapped to rRNA were filtered out using BBsplit function from BBtools. KEGG pathway analysis was performed using the kegga function from the limma package80 in R. For promoters identified as driving operons (based on the Rockhopper results), clusters of genes driven by the promoter were used as input for pathway analysis. KEGG IDs of the genes were obtained from IMG81.
RNA-seq data analysis
RNA-seq data were processed using a custom shell script with BBtools and R scripts using dplyr, edgeR and limma packages. In this pipeline, reads mapped to 5S, 16S, and 23S rRNA were filtered out using the BBsplit function. The remaining reads were mapped to the P. simiae WCS417 genome using HISAT282. Mapping results were used to calculate gene-level read counts with FeatureCounts83. For sequencing data generated by Azenta Life Sciences, read alignment and count generation were performed by Azenta using the Bowtie2 aligner and FeatureCounts, following their standard bacterial RNA-seq analysis workflow. Count data from all samples were normalized to counts per million (CPM) and subsequently log₂-transformed for downstream analyses.
Bacterial competition assay for root colonization
Wild-type and individual mutant strains (kanamycin-resistance) were cultured, using 1x M9 minimal medium with 20 mM glucose (no antibiotics) for pre-cultures, and 1x M9 minimal salts (Gibco: A1374401) for washing. Cell cultures were normalized to an OD600 of 0.5, and wild-type and mutant strains were mixed at a 1:1 ratio. A 50 μl aliquot of this mixture was spread onto phytagel plates. 10-day-old Arabidopsis seedlings, grown on nylon mesh filters, were then transferred onto the inoculated plates using sterile tweezers. The plates were incubated upright in a Percival incubator at 22 °C with a 16-h light and 8-h dark cycle.
After 5 days of incubation, root samples were collected and washed once with 1x M9 buffer. The buffer was removed, and 800 µl of fresh buffer along with 2 mm beads (Zymo Research: S6003-50) were added to the tubes. The tubes were placed on a vortex adapter, and the roots were ground at maximum speed for 10 min. The lysed samples were diluted 10,000-fold, and 100 µl were spread onto LB plates containing 200 µg/ml kanamycin. After incubation at room temperature for two days, colony numbers were determined. Mutant cell counts were calculated based on colonies growing on LB plates with kanamycin, while wild-type counts were determined by subtracting the mutant colony numbers from the colony numbers on LB plates. Typically, around 20 mg of root tissues was collected per experiment. Statistical analysis was performed using GraphPad Prism (version 10.6.0).
Lysozyme assay
To investigate the function of the MliC family protein (PS417_RS02110), we tested the sensitivity of both the wild-type and the mutant strain to lysozyme. Cells were cultured in LB medium and then transferred to M9 minimal medium with 20 mM glucose at 28 °C. After overnight incubation, cells were washed three times with phosphate-buffered saline (PBS) and resuspended to an OD600 of 0.05. Lysozyme (Roche: 10837059001) was then added to a final concentration of 25 mg/ml. Control samples were incubated in PBS without lysozyme. After 24 h of incubation, cells were serially diluted and plated onto LB agar plates. Colony counts were determined after 48 h of incubation at 28 °C. Statistical analysis was performed using GraphPad Prism (version 10.6.0).
Oxidative stress assay
The sensitivity of both wild-type and mutant strains of xanthine dehydrogenase (PS417_RS04040) to hydrogen peroxide (H2O2) was determined by monitoring growth curves. Cells cultured overnight in LB medium were transferred into M9-glucose medium at a 1:1000 dilution and incubated at 28 °C overnight. The following day, cells were transferred to a 96-well plate containing 200 μL of M9-glucose medium, adjusted to a starting OD600 of 0.1. The cells were grown in a microplate spectrophotometer (Infinite 200 PRO, Tecan, Switzerland) with absorbance at 600 nm measured every 30 min. H2O2 (final concentration of 5 mM) was added after 7 h of incubation, which was set as time 0 in Fig.4D.
Statistics & reproducibility
No statistical method was used to predetermine sample size. Sample sizes are consistent with standards in the field. The number of biological replicates for each experiment is indicated in the corresponding figure legends. No data were excluded from the analyses unless explicitly stated. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment. All experiments were independently replicated, and reproducibility was assessed by consistency across biological replicates, as indicated in the figures.
Inclusion & ethics statement
All authors contributed to the study in accordance with their institutional ethical and authorship guidelines.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All sequencing data generated in this study have been deposited in the NCBI Short Read Archive under accession code PRJNA1221951. The genome-scale datasets are provided as Supplementary Data 1–6. Source data are provided with this paper.
Code availability
Custom scripts used for PI-seq analysis are available on GitHub at https://github.com/tomoyahonda/CRAGE-RB-PI-seq and archived on Zenodo at https://doi.org/10.5281/zenodo.18462023.
References
Zhalnina, K. et al. Dynamic root exudate chemistry and microbial substrate preferences drive patterns in rhizosphere microbial community assembly. Nat. Microbiol. 3, 470–480 (2018).
McLaughlin, S., Zhalnina, K., Kosina, S., Northen, T. R. & Sasse, J. The core metabolome and root exudation dynamics of three phylogenetically distinct plant species. Nat. Commun. 14, 1649 (2023).
Jansson, J. K., McClure, R. & Egbert, R. G. Soil microbiome engineering for sustainability in a changing environment. Nat. Biotechnol. 1–13, https://doi.org/10.1038/s41587-023-01932-3 (2023).
Haney, C. H., Samuel, B. S., Bush, J. & Ausubel, F. M. Associations with rhizosphere bacteria can confer an adaptive advantage to plants. Nat. Plants 1, 15051 (2015).
Wang, N. R. et al. Commensal pseudomonas fluorescens strains protect arabidopsis from closely related pseudomonas pathogens in a colonization-dependent manner. mBio 13, e02892–21 (2022).
Acharya, S. M. et al. Fine scale sampling reveals early differentiation of rhizosphere microbiome from bulk soil in young Brachypodium plant roots. ISME Commun. 3, 1–9 (2023).
Huang, A. C. et al. A specialized metabolic network selectively modulates Arabidopsis root microbiota. Science 364, eaau6389 (2019).
Levy, A. et al. Genomic features of bacterial adaptation to plants. Nat. Genet. 50, 138–150 (2018).
Lundberg, D. S. et al. Defining the core Arabidopsis thaliana root microbiome. Nature 488, 86–90 (2012).
Lebeis, S. L. et al. Salicylic acid modulates colonization of the root microbiome by specific bacterial taxa. Science 349, 860–864 (2015).
Pieterse, C. M. J. et al. Pseudomonas simiae WCS417: star track of a model beneficial rhizobacterium. Plant Soil 461, 245–263 (2021).
Berendsen, R. L. et al. Unearthing the genomes of plant-beneficial Pseudomonas model strains WCS358, WCS374 and WCS417. BMC Genomics 16, 539 (2015).
Van Wees, S. C., Van der Ent, S. & Pieterse, C. M. Plant immune responses triggered by beneficial microbes. Curr. Opin. Plant Biol. 11, 443–448 (2008).
Yu, K. et al. Rhizosphere-associated pseudomonas suppress local root immune responses by gluconic acid-mediated lowering of environmental pH. Curr. Biol. 29, 3913–3920.e4 (2019).
Cole, B. J. et al. Genome-wide identification of bacterial plant colonization genes. PLOS Biol. 15, e2002860 (2017).
Stringlis, I. A. et al. Root transcriptional dynamics induced by beneficial rhizobacteria and microbial immune elicitors reveal signatures of adaptation to mutualists. Plant J. 93, 166–180 (2018).
Bjornson, M., Pimprikar, P., Nürnberger, T. & Zipfel, C. The transcriptional landscape of Arabidopsis thaliana pattern-triggered immunity. Nat. Plants 7, 579–586 (2021).
Verbon, E. H. et al. Cell-type-specific transcriptomics reveals that root hairs and endodermal barriers play important roles in beneficial plant-rhizobacterium interactions. Mol. Plant 16, 1160–1177 (2023).
Humphrys, M. S. et al. Simultaneous transcriptional profiling of bacteria and their host cells. PLOS One 8, e80597 (2013).
Marsh, J. W., Humphrys, M. S. & Myers, G. S. A. A laboratory methodology for dual RNA-sequencing of bacteria and their host cells in vitro. Front. Microbiol. 8, 1830(2017).
Westermann, A. J., Barquist, L. & Vogel, J. Resolving host–pathogen interactions by dual RNA-seq. PLOS Pathog 13, e1006033 (2017).
Donaldson, G. P. et al. Spatially distinct physiology of Bacteroides fragilis within the proximal colon of gnotobiotic mice. Nat. Microbiol. 5, 746–756 (2020).
Andrés-Barrao, C. et al. Coordinated bacterial and plant sulfur metabolism in Enterobacter sp. SA187–induced plant salt stress tolerance. Proc. Natl. Acad. Sci. 118, e2107417118 (2021).
Vannier, N. et al. Genome-resolved metatranscriptomics reveals conserved root colonization determinants in a synthetic microbiota. Nat. Commun. 14, 8274 (2023).
Kumar, N. et al. Efficient enrichment of bacterial mRNA from host-bacteria total RNA samples. Sci. Rep. 6, 34850 (2016).
Nobori, T. et al. Transcriptome landscape of a bacterial pathogen under plant immunity. Proc. Natl. Acad. Sci. 115, E3055–E3064 (2018).
Roux, B. et al. An integrated analysis of plant and bacterial gene expression in symbiotic root nodules using laser-capture microdissection coupled to RNA sequencing. Plant J 77, 817–837 (2014).
Ma, Q., Bücking, H., Gonzalez Hernandez, J. L. & Subramanian, S. Single-cell RNA sequencing of plant-associated bacterial communities. Front. Microbiol. 10, 2452 (2019).
Johns, N. I. et al. Metagenomic mining of regulatory elements enables programmable species-selective gene expression. Nat. Methods 15, 323–329 (2018).
Jones, E. M. et al. A scalable, multiplexed assay for decoding GPCR-ligand interactions with RNA sequencing. Cell Syst. 8, 254–260.e6 (2019).
Crook, N., Ferreiro, A., Condiotte, Z. & Dantas, G. Transcript barcoding illuminates the expression level of synthetic constructs in E. coli nissle residing in the mammalian gut. ACS Synth. Biol. 9, 1010–1021 (2020).
Jores, T. et al. Synthetic promoter designs enabled by a comprehensive analysis of plant core promoters. Nat. Plants, https://doi.org/10.1038/s41477-021-00932-y (2021).
Wang, G. et al. CRAGE enables rapid activation of biosynthetic gene clusters in undomesticated bacteria. Nat. Microbiol. 4, 2498–2510 (2019).
Wang, B. et al. CRAGE-duet facilitates modular assembly of biological systems for studying plant–microbe interactions. ACS Synth. Biol. 9, 2610–2615 (2020).
Cho, B.-K. et al. The transcription unit architecture of the Escherichia coli genome. Nat. Biotechnol. 27, 1043–1049 (2009).
Wurtzel, O. et al. The single-nucleotide resolution transcriptome of pseudomonas aeruginosa grown in body temperature. PLOS Pathog 8, e1002945 (2012).
Filiatrault, M. J. et al. Genome-wide identification of transcriptional start sites in the plant pathogen Pseudomonas syringae pv. tomato str. DC3000. PLOS ONE 6, e29335 (2011).
Salgado, H., Moreno-Hagelsieb, G., Smith, T. F. & Collado-Vides, J. Operons in Escherichia coli: genomic analyses and predictions. Proc. Natl. Acad. Sci. 97, 6652–6657 (2000).
Tjaden, B. A computational system for identifying operons based on RNA-seq data. Methods 176, 62–70 (2020).
Lugtenberg, B. J. J., Dekkers, L. & Bloemberg, G. V. Molecular determinants of rhizosphere colonization by pseudomonas. Annu. Rev. Phytopathol. 39, 461–490 (2001).
Muñoz-Elías, E. J. & McKinney, J. D. Carbon metabolism of intracellular bacteria. Cell. Microbiol. 8, 10–22 (2006).
Chen, H., Shiroguchi, K., Ge, H. & Xie, X. S. Genome-wide study of mRNA degradation and transcript elongation in Escherichia coli. Mol. Syst. Biol. 11, 781 (2015).
Waters, L. S. & Storz, G. Regulatory RNAs in bacteria. Cell 136, 615–628 (2009).
Winnen, B., Hvorup, R. N. & Saier, M. H. The tripartite tricarboxylate transporter (TTT) family. Res. Microbiol. 154, 457–465 (2003).
Brocker, M., Schaffer, S., Mack, C. & Bott, M. Citrate utilization by Corynebacterium glutamicum is controlled by the CitAB two-component system through positive regulation of the citrate transport genes citH and tctCBA. J. Bacteriol. 191, 3869 (2009).
Finkel, O. M. et al. A single bacterial genus maintains root growth in a complex microbiome. Nature 587, 103–108 (2020).
De Weger, L. A. et al. Flagella of a plant-growth-stimulating Pseudomonas fluorescens strain are required for colonization of potato roots. J. Bacteriol. 169, 2769–2773 (1987).
Sivakumar, R. et al. Evaluation of INSeq To Identify Genes Essential for Pseudomonas aeruginosa PGPR2 Corn Root Colonization. G3 GenesGenomesGenetics 9, 651–661 (2019).
Pankievicz, V. C. S. et al. RNA-seq transcriptional profiling of Herbaspirillum seropedicae colonizing wheat (Triticum aestivum) roots. Plant Mol. Biol. 90, 589–603 (2016).
Nobori, T. et al. Dissecting the cotranscriptome landscape of plants and their microbiota. EMBO Rep. 23, e55380 (2022).
WOJTASZEK, P. Oxidative burst: an early plant response to pathogen infection. Biochem. J. 322, 681–692 (1997).
Ebel, J. & Mithöfer, A. Early events in the elicitation of plant defence. Planta 206, 335–348 (1998).
Danhorn, T. & Fuqua, C. Biofilm formation by plant-associated bacteria. Annu. Rev. Microbiol. 61, 401–422 (2007).
Rudrappa, T., Biedrzycki, M. L. & Bais, H. P. Causes and consequences of plant-associated biofilms. FEMS Microbiol. Ecol. 64, 153–166 (2008).
Callewaert, L. et al. A new family of lysozyme inhibitors contributing to lysozyme tolerance in gram-negative bacteria. PLOS Pathog. 4, e1000019 (2008).
Yum, S. et al. Structural basis for the recognition of lysozyme by MliC, a periplasmic lysozyme inhibitor in Gram-negative bacteria. Biochem. Biophys. Res. Commun. 378, 244–248 (2009).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Liu, X. et al. Host-induced bacterial cell wall decomposition mediates pattern-triggered immunity in Arabidopsis. eLife 3, e01990 (2014).
Eitzen, K., Sengupta, P., Kroll, S., Kemen, E. & Doehlemann, G. A fungal member of the Arabidopsis thaliana phyllosphere antagonizes Albugo laibachii via a GH25 lysozyme. eLife 10, e65306 (2021).
Lee, K.-M. et al. A genetic screen reveals novel targets to render pseudomonas aeruginosa sensitive to lysozyme and cell wall-targeting antibiotics. Front. Cell. Infect. Microbiol. 7, 59 (2017).
Torrens, G. et al. Targeting the permeability barrier and peptidoglycan recycling pathways to disarm Pseudomonas aeruginosa against the innate immune system. PLOS One 12, e0181932 (2017).
Flores-Cruz, Z. & Allen, C. Ralstonia solanacearum encounters an oxidative environment during tomato infection. Mol. Plant Microbe Interactions 22, 773–782 (2009).
Phelan, J. P., Bourgeois, J. S., McCarthy, J. E. & Hu, L. T. A putative xanthine dehydrogenase is critical for Borrelia burgdorferi survival in ticks and mice. Microbiology 169, 001286 (2023).
Sasse, J. et al. Root morphology and exudate availability are shaped by particle size and chemistry in Brachypodium distachyon. Plant Direct 4, e00207 (2020).
Palluk, S. et al. De novo DNA synthesis using polymerase-nucleotide conjugates. Nat. Biotechnol. 36, 645–650 (2018).
Plesa, C., Sidore, A. M., Lubock, N. B., Zhang, D. & Kosuri, S. Multiplexed gene synthesis in emulsions for exploring protein functional landscapes. Science 359, 343–347 (2018).
Sharma, C. M. et al. The primary transcriptome of the major human pathogen Helicobacter pylori. Nature 464, 250–255 (2010).
Adiconis, X. et al. Comprehensive comparative analysis of 5′-end RNA-sequencing methods. Nat. Methods 15, 505–511 (2018).
Yu, K. et al. Transcriptome signatures in Pseudomonas simiae WCS417 shed light on role of root-secreted coumarins in arabidopsis-mutualist communication. Microorganisms 9, 575 (2021).
Canarini, A., Kaiser, C., Merchant, A., Richter, A. & Wanek, W. Root exudation of primary metabolites: mechanisms and their roles in plant responses to environmental stimuli. Front. Plant Sci. 10, 157 (2019).
Gliese, N., Khodaverdi, V., Schobert, M. & Görisch, H. AgmR controls transcription of a regulon with several operons essential for ethanol oxidation in Pseudomonas aeruginosa ATCC 17933. Microbiology 150, 1851–1857 (2004).
Choi, O. et al. Pyrroloquinoline quinone is a plant growth promotion factor produced by Pseudomonas fluorescens B16. Plant Physiol. 146, 657–668 (2008).
Carreño-López, R., Alatorre-Cruz, J. M. & Marín-Cevada, V. Pyrroloquinoline quinone (PQQ): role in plant-microbe interactions. In Secondary Metabolites of Plant Growth Promoting Rhizomicroorganisms: Discovery and Applications (eds Singh, H. B., Keswani, C., Reddy, M. S., Sansinenea, E. & García-Estrada, C.) 169–184 (Springer, Singapore, 2019). https://doi.org/10.1007/978-981-13-5862-3_9.
Wang, X. et al. Elucidation of genes enhancing natural product biosynthesis through co-evolution analysis. Nat. Metab. 1–14, https://doi.org/10.1038/s42255-024-01024-9 (2024).
Boo, A. et al. Synthetic microbe-to-plant communication channels. Nat. Commun. 15, 1817 (2024).
Pini, F. et al. Bacterial biosensors for in vivo spatiotemporal mapping of root secretion. Plant Physiol. 174, 1289–1306 (2017).
Haskett, T. L. et al. Engineered plant control of associative nitrogen fixation. Proc. Natl. Acad. Sci. 119, e2117465119 (2022).
Toju, H. et al. Core microbiomes for sustainable agroecosystems. Nat. Plants 4, 247–257 (2018).
Law, C. W. et al. RNA-seq analysis is easy as 1-2-3 with limma, Glimma and edgeR. F1000Research 5, ISCB Comm J–1408 (2018).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47–e47 (2015).
Chen, I.-M. A. et al. IMG/M v.5.0: an integrated data management and comparative analysis system for microbial genomes and microbiomes. Nucleic Acids Res. 47, D666–D677 (2019).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Acknowledgements
We thank Zhiying Zhao, Robert Evans, William Kauffman, Jan-Fang Cheng, Benjamin Cole, Yuko Yoshinaga, Vlastimil Novak, and Jeffery Dangle for helpful discussions and technical suggestions. We also thank QB3 Genomics Center at UC Berkeley and Azenta Life Sciences for sequencing. T.H. acknowledges a JSPS overseas research fellowship from the Japan Society for the Promotion of Science. Lawrence Berkeley National Laboratory is managed by University of California for DOE under contract number DE-AC02-05CH11231 (T.H. S.Y., D.M., L.B., Y.Y.). Argonne National Laboratory is managed by UChicago Argonne, LLC for DOE under contract number DE-AC02-06CH11357 (GB). This work was funded by the secure biosystem design (S.Y., G.B., Y.Y.) and the bioimaging programs (T.H., G.B., Y.Y.) of the U.S. Department of Energy, Office of Biological and Environmental Research under Contract No. DE-AC02-05CH11231 and DE-AC02-06CH11357. This work (proposal: 10.46936/10.25585/60001279), conducted by the U.S. Department of Energy Joint Genome Institute (https://ror.org/04xm1d337), a DOE Office of Science User Facility, is supported by the Office of Science of the U.S. Department of Energy operated under Contract No. DE-AC02-05CH11231 (T.H., D.M., L.B., Y.Y.). This project was also supported in part by the U. S. Department of Energy, Office of Science, through the Biomolecular Characterization and Imaging Sciences Program, Office of Biological and Environmental Research, under FWP 39156 (T.H., S.Y., Y.Y., G.B.). Work at the Molecular Foundry was supported by the Office of Science, Office of Basic Energy Sciences, of the U.S. Department of Energy under Contract No. DE-AC02-05CH11231.
Author information
Authors and Affiliations
Contributions
T.H. and Y.Y. designed the study. T.H. performed the experiments, with contribution by S.Y., who helped prepare sequencing libraries and characterize mutant phenotypes. T.H. developed the computational pipeline. D.M. performed protein structural analysis. L.B. contributed to a plasmid construction and provided DAP-seq data. E.C. and S.Y. developed and performed Raman microscopy experiments. Y.Y. and G.B. supervised the study and acquired funding. T.H. and Y.Y. wrote the manuscript. All authors reviewed and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
Lawrence Berkeley National Laboratory has filed an international patent application related to high-throughput characterization of bacterial promoters from their host environments on behalf of the Regents of the University of California, on which T.H. and Y.Y. are the named inventors (PCT/US2023/031771). Y.Y. is a co-founder and has financial interest in Quorum Bio, Inc. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Yuting Liang and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Honda, T., Yu, S., Mai, D. et al. CRAGE-RB-PI-seq reveals transcriptional dynamics of plant-associated bacteria during root colonization. Nat Commun 17, 3021 (2026). https://doi.org/10.1038/s41467-026-69903-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69903-1