Abstract
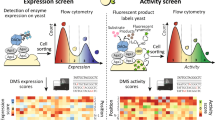
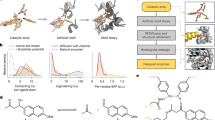
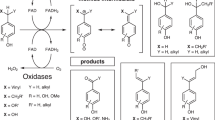
Substrate specificity is a defining feature of enzyme function, but its molecular underpinnings remain difficult to decode and engineer. Here, we leverage enzyme proximity sequencing (EP-Seq) to systematically map how single-point and combinatorial mutations reshape the substrate preferences of D-amino acid oxidase (DAOx) from Rhodotorula gracilis, a model promiscuous enzyme. We generate ~40,000 sequence–phenotype pairs, enabling us to profile the activities of ~6,500 unique DAOx variants against five D-amino acid substrates with distinct physicochemical properties. Our analysis reveals that substrate-specific mutations are distributed throughout the enzyme structure. Mutations near the active site drive strong specificity shifts but also incur catalytic penalties, while distal mutations subtly rewire intramolecular contacts in order to modulate specificity with minimal loss of activity. We identify and validate positional hotspots that act allosterically to influence specificity, and characterize key variants that acquire exclusive substrate specificity or exhibit up to 230-fold changes in substrate preference. Combining mutations with complementary effects further sharpens substrate discrimination, enabling rational design of highly selective biocatalysts. This work establishes a powerful framework for decoding enzyme specificity and provides foundational datasets to advance AI-guided enzyme engineering.
Similar content being viewed by others
Data availability
All data required to replicate this study and generated in this study are publicly available. DNA sequencing reads can be accessed through the NCBI Sequence Read Archive under BioProject accession PRJNA1289092. Raw data for all figures, and processed output data are available at https://doi.org/10.5281/zenodo.15846928. The PDB code of the previously published structure used in this study is 1C0P. Source Data are provided as a Source Data file. Source data are provided with this paper.
Code availability
All custom code and scripts used in this study are publicly available in the Zenodo repository at https://doi.org/10.5281/zenodo.15846928
References
Fischer, E. Einfluss der Configuration auf die Wirkung der Enzyme. Ber. Dtsch. Chem. Ges. 27, 2985–2993 (1894).
Khersonsky, O. & Tawfik, D. S. Enzyme promiscuity: a mechanistic and evolutionary perspective. Annu. Rev. Biochem. 79, 471–505 (2010).
Tawfik, D. S. Accuracy-rate tradeoffs: how do enzymes meet demands of selectivity and catalytic efficiency?. Curr. Opin. Chem. Biol. 21, 73–80 (2014).
Tawfik, D. S. & Gruic-Sovulj, I. How evolution shapes enzyme selectivity - lessons from aminoacyl-tRNA synthetases and other amino acid utilizing enzymes. FEBS J 287, 1284–1305 (2020).
Cornish-Bowden, A. & Cárdenas, M. L. Specificity of non-Michaelis-Menten enzymes: necessary information for analyzing metabolic pathways. J. Phys. Chem. B 114, 16209–16213 (2010).
Peracchi, A. The limits of enzyme specificity and the evolution of metabolism. Trends Biochem. Sci. 43, 984–996 (2018).
Matreyek, K. A. et al. Multiplex assessment of protein variant abundance by massively parallel sequencing. Nat. Genet. 50, 874–882 (2018).
Fowler, D. M., Stephany, J. J. & Fields, S. Measuring the activity of protein variants on a large scale using deep mutational scanning. Nat. Protoc. 9, 2267–2284 (2014).
Araya, C. L. & Fowler, D. M. Deep mutational scanning: assessing protein function on a massive scale. Trends Biotechnol. 29, 435–442 (2011).
Amorosi, C. J. et al. Massively parallel characterization of CYP2C9 variant enzyme activity and abundance. Am. J. Hum. Genet. 108, 1735–1751 (2021).
Markin, C. J. et al. Revealing enzyme functional architecture via high-throughput microfluidic enzyme kinetics. Science 373, eabf8761 (2021).
Vanella, R., Kovacevic, G., Doffini, V., de Santaella, J. F. & Nash, M. A. High-throughput screening, next generation sequencing and machine learning: advanced methods in enzyme engineering. Chem. Commun. 58, 2455–2467 (2022).
Gantz, M., Neun, S., Medcalf, E. J., van Vliet, L. D. & Hollfelder, F. Ultrahigh-throughput enzyme engineering and discovery in in vitro compartments. Chem. Rev. 123, 5571–5611 (2023).
Bozkurt, E. U., Ørsted, E. C., Volke, D. C. & Nikel, P. I. Accelerating enzyme discovery and engineering with high-throughput screening. Nat. Prod. Rep. https://doi.org/10.1039/d4np00031e (2024).
Melnikov, A., Rogov, P., Wang, L., Gnirke, A. & Mikkelsen, T. S. Comprehensive mutational scanning of a kinase in vivo reveals substrate-dependent fitness landscapes. Nucleic Acids Res. 42, e112 (2014).
Steinberg, B. & Ostermeier, M. Environmental changes bridge evolutionary valleys. Sci. Adv. 2, e1500921 (2016).
Chen, J. Z., Fowler, D. M. & Tokuriki, N. Comprehensive exploration of the translocation, stability and substrate recognition requirements in VIM-2 lactamase. Elife 9, e56707 (2020).
Wrenbeck, E. E., Azouz, L. R. & Whitehead, T. A. Single-mutation fitness landscapes for an enzyme on multiple substrates reveal specificity is globally encoded. Nat. Commun. 8, 15695 (2017).
Vanella, R. et al. Understanding activity-stability tradeoffs in biocatalysts by enzyme proximity sequencing. Nat. Commun. 15, 1807 (2024).
Vanella, R., Bazin, A., Ta, D. T. & Nash, M. A. Genetically encoded stimuli-responsive cytoprotective hydrogel capsules for single cells provide novel genotype–phenotype linkage. Chem. Mater. https://doi.org/10.1021/acs.chemmater.8b04348. (2019)
Küng, C., Vanella, R. & Nash, M. A. Directed evolution of Rhodotorula gracilisd-amino acid oxidase using single-cell hydrogel encapsulation and ultrahigh-throughput screening. React. Chem. Eng. 8, 1960–1968 (2023).
Vanella, R., Ta, D. T. & Nash, M. A. Enzyme-mediated hydrogel encapsulation of single cells for high-throughput screening and directed evolution of oxidoreductases. Biotechnol. Bioeng. 116, 1878–1886 (2019).
Heiniger, M., Vanella, R., Walsh-Korb, Z. & Nash, M. A. Functionalized polysaccharides improve sensitivity of tyramide/peroxidase proximity labeling assays through electrostatic interactions. ACS Biomater. Sci. Eng. 10, 5869–5880 (2024).
Kueng, C., Dalkiran, A., Vanella, R., Oyarzun, D. & Nash, M. A. Discovery of electron hole-hopping redox mutations in myoglobin by deep mutational learning. bioRxiv https://doi.org/10.1101/2025.08.27.672588. (2025).
Pollegioni, L. et al. Properties and applications of microbial D-amino acid oxidases: current state and perspectives. Appl. Microbiol. Biotechnol. 78, 1–16 (2008).
Rosini, E., Pollegioni, L., Ghisla, S. & Orru, R. Optimization of d-amino acid oxidase for low substrate concentrations–towards a cancer enzyme therapy. FEBS J. 276, 4921–4932 (2009).
Bava, A. et al. D-amino acid oxidase-nanoparticle system: a potential novel approach for cancer enzymatic therapy. Nanomedicine 8, 1797–1806 (2013).
Pollegioni, L. et al. Yeast D-amino acid oxidase: structural basis of its catalytic properties. J. Mol. Biol. 324, 535–546 (2002).
Castro-Fernandez, V. et al. Reconstructed ancestral enzymes reveal that negative selection drove the evolution of substrate specificity in ADP-dependent kinases. J. Biol. Chem. 292, 15598–15610 (2017).
Peracchi, A. & Polverini, E. Using steady-state kinetics to quantitate substrate selectivity and specificity: a case study with two human transaminases. Molecules 27, 1398 (2022).
Carlson, J. C., Badran, A. H., Guggiana-Nilo, D. A. & Liu, D. R. Negative selection and stringency modulation in phage-assisted continuous evolution. Nat. Chem. Biol. 10, 216–222 (2014).
Lipovsek, D. et al. Selection of horseradish peroxidase variants with enhanced enantioselectivity by yeast surface display. Chem. Biol. 14, 1176–1185 (2007).
Yi, L. et al. Engineering of TEV protease variants by yeast ER sequestration screening (YESS) of combinatorial libraries. Proc. Natl. Acad. Sci. USA. 110, 7229–7234 (2013).
Miller, S. R. An appraisal of the enzyme stability-activity trade-off. Evolution 71, 1876–1887 (2017).
Siddiqui, K. S. Defying the activity-stability trade-off in enzymes: taking advantage of entropy to enhance activity and thermostability. Crit. Rev. Biotechnol. 37, 309–322 (2017).
Goldenzweig, A. & Fleishman, S. J. Principles of protein stability and their application in computational design. Annu. Rev. Biochem. 87, 105–129 (2018).
Yu, H. & Dalby, P. A. Exploiting correlated molecular-dynamics networks to counteract enzyme activity-stability trade-off. Proc. Natl. Acad. Sci. USA. 115, E12192–E12200 (2018).
Küng, C., Protsenko, O., Vanella, R. & Nash, M. A. Deep mutational scanning reveals a de novo disulfide bond and combinatorial mutations for engineering thermostable myoglobin. Protein Sci. 34, e70112 (2025).
Starr, T. N. et al. Deep mutational scanning of SARS-CoV-2 receptor binding domain reveals constraints on folding and ACE2 binding. Cell 182, 1295–1310.e20 (2020).
Klesmith, J. R., Bacik, J.-P., Wrenbeck, E. E., Michalczyk, R. & Whitehead, T. A. Trade-offs between enzyme fitness and solubility illuminated by deep mutational scanning. Proc. Natl Acad. Sci. 114, 2265–2270 (2017).
Sacchi, S. et al. Engineering the substrate specificity of D-amino-acid oxidase. J. Biol. Chem. 277, 27510–27516 (2002).
Ball, J., Gannavaram, S. & Gadda, G. Structural determinants for substrate specificity of flavoenzymes oxidizing d-amino acids. Arch. Biochem. Biophys. 660, 87–96 (2018).
Yang, G., Miton, C. M. & Tokuriki, N. A mechanistic view of enzyme evolution. Protein Sci. 29, 1724–1747 (2020).
Karamitros, C. S. et al. Leveraging intrinsic flexibility to engineer enhanced enzyme catalytic activity. Proc. Natl. Acad. Sci. USA. 119, e2118979119 (2022).
Fasan, R., Meharenna, Y. T., Snow, C. D., Poulos, T. L. & Arnold, F. H. Evolutionary history of a specialized p450 propane monooxygenase. J. Mol. Biol. 383, 1069–1080 (2008).
van der Meer, J.-Y. et al. Using mutability landscapes of a promiscuous tautomerase to guide the engineering of enantioselective Michaelases. Nat. Commun. 7, 10911 (2016).
Tokuriki, N. & Tawfik, D. S. Protein dynamism and evolvability. Science 324, 203–207 (2009).
Duran, C., Casadevall, G. & Osuna, S. Harnessing conformational dynamics in enzyme catalysis to achieve nature-like catalytic efficiencies: the shortest path map tool for computational enzyme redesign. Faraday Discuss 252, 306–322 (2024).
Röthlisberger, D. et al. Kemp elimination catalysts by computational enzyme design. Nature 453, 190–195 (2008).
Jiang, L. et al. De novo computational design of retro-aldol enzymes. Science 319, 1387–1391 (2008).
Otten, R. et al. How directed evolution reshapes the energy landscape in an enzyme to boost catalysis. Science 370, 1442–1446 (2020).
Sacchi, S., Rosini, E., Molla, G., Pilone, M. S. & Pollegioni, L. Modulating D-amino acid oxidase substrate specificity: production of an enzyme for analytical determination of all D-amino acids by directed evolution. Protein Eng. Des. Sel. 17, 517–525 (2004).
Gietz, R. D. & Woods, R. A. Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. Methods Enzymol 350, 87–96 (2002).
Bushnell, B. BBMap: A Fast, Accurate, Splice-Aware Aligner. https://www.semanticscholar.org (2014).
Jurcik, A. et al. CAVER Analyst 2.0: analysis and visualization of channels and tunnels in protein structures and molecular dynamics trajectories. Bioinformatics 34, 3586–3588 (2018).
Acknowledgements
This work was supported by the University of Basel, ETH Zurich, the SNF-NCCR in Molecular Systems Engineering, and an SNSF Grant (200021_191962 and 10004516) to M.A.N.
Author information
Authors and Affiliations
Contributions
R.V. and M.A.N. conceived the study and drafted the manuscript. R.V. carried out the practical work and computational analyses. S.B. optimized and performed the expression and purification of the soluble DAOx variants. C.K. contributed to the conceptualization and optimization of the workflow. M.A.N. secured funding and administered the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Shoji Takahashi and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Vanella, R., Boult, S., Küng, C. et al. Decoding the substrate specificity landscape of a promiscuous enzyme through multi-substrate mutational scanning. Nat Commun (2026). https://doi.org/10.1038/s41467-026-69913-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-69913-z