Abstract
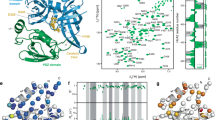
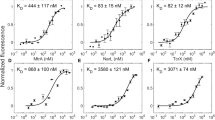
Bacterial proteasomal activator (Bpa) is a regulatory particle of the Mycobacterium tuberculosis proteasome that facilitates the recruitment of substrates and their subsequent degradation by the 20S core particle. Substrate-bound structures of Bpa are unavailable, leaving its recruitment mechanism incompletely understood. Here, we use mass spectrometry and NMR to show that Bpa reversibly assembles into dodecamers from dimers/tetramers in a temperature-dependent manner in vitro, and map the oligomerization interfaces during assembly. To overcome the limitations posed by the poor solubility of natural Bpa substrates, we establish the DNA-binding domain of hTRF1 as a model substrate. We quantify the affinity and stoichiometry of the Bpa-hTRF1 interaction using methyl-TROSY NMR, identifying a 12 Bpa subunit: 3 hTRF1 binding ratio with micromolar affinity that is modulated by salt concentration. Our work maps the Bpa-hTRF1 interface at atomic resolution, identifies determinants of substrate engagement, and introduces a tractable substrate for dissecting proteasomal recognition in mycobacteria.
Similar content being viewed by others
Data availability
Mass spectrometry data are available from the MassIVE database as entry MSV000099239 (https://massive.ucsd.edu/ProteoSAFe/dataset.jsp?task=1e313073db3545efaf0ba91f95fa9a72) (HDX raw data). Source data are provided with this paper.
Code availability
The code used for SEC, HDX, and NMR analysis in this study has been deposited in the Zenodo database under accession code [https://zenodo.org/records/18498816] (Code deposition).
References
Darwin, K. H., Ehrt, S., Gutierrez-Ramos, J.-C., Weich, N. & Nathan, C. F. The proteasome of Mycobacterium tuberculosis is required for resistance to nitric oxide. Science 302, 1963–1966 (2003).
von Rosen, T., Pepelnjak, M., Quast, J.-P., Picotti, P. & Weber-Ban, E. ATP-independent substrate recruitment to proteasomal degradation in mycobacteria. Life Sci. Alliance 6, e202301923 (2023).
Janssen, G. V. et al. Development of Tyrphostin Analogues to Study Inhibition of the Mycobacterium tuberculosis Pup Proteasome System. ChemBioChem 22, 3082–3089 (2021).
Lin, G., Tsu, C., Dick, L., Zhou, X. K. & Nathan, C. Distinct Specificities of Mycobacterium tuberculosis and Mammalian Proteasomes for N-Acetyl Tripeptide Substrates. J. Biol. Chem. 283, 34423–34431 (2008).
Lupoli, T. J., Vaubourgeix, J., Burns-Huang, K. & Gold, B. Targeting the Proteostasis Network for Mycobacterial Drug Discovery. ACS Infect. Dis. 4, 478–498 (2018).
Li, D. et al. Structural basis for the assembly and gate closure mechanisms of the Mycobacterium tuberculosis 20S proteasome. EMBO J. 29, 2037–2047 (2010).
Hu, K. et al. Proteasome substrate capture and gate opening by the accessory factor PafE from Mycobacterium tuberculosis. J. Biol. Chem. 293, 4713–4723 (2018).
Lin, G. et al. Mycobacterium tuberculosis prcBA genes encode a gated proteasome with broad oligopeptide specificity. Mol. Microbiol. 59, 1405–1416 (2006).
Kavalchuk, M., Jomaa, A., Müller, A. U. & Weber-Ban, E. Structural basis of prokaryotic ubiquitin-like protein engagement and translocation by the mycobacterial Mpa-proteasome complex. Nat. Commun. 13, 276 (2022).
Xiao, X. et al. The β-Grasp Domain of Proteasomal ATPase Mpa Makes Critical Contacts with the Mycobacterium tuberculosis 20S Core Particle to Facilitate Degradation. mSphere 7, e00274-22 (2022).
Jastrab, J. B. et al. An adenosine triphosphate-independent proteasome activator contributes to the virulence of Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA. 112, E1763–E1772 (2015).
Bai, L. et al. Structural analysis of the dodecameric proteasome activator PafE in Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. 113, E1983–E1992 (2016).
Bolten, M. et al. Structural Analysis of the Bacterial Proteasome Activator Bpa in Complex with the 20S Proteasome. Structure 24, 2138–2151 (2016).
Delley, C. L. et al. Bacterial Proteasome Activator Bpa (Rv3780) Is a Novel Ring-Shaped Interactor of the Mycobacterial Proteasome. PLoS One 9, e114348 (2014).
Jastrab, J. B., Samanovic, M. I., Copin, R., Shopsin, B. & Darwin, K. H. Loss-of-Function Mutations in HspR Rescue the Growth Defect of a Mycobacterium tuberculosis Proteasome Accessory Factor E (pafE) Mutant. J. Bacteriol. 199, https://doi.org/10.1128/jb.00850-16 (2017).
Parijat, P. & Batra, J. K. Role of DnaK in HspR-HAIR interaction of ycobacterium tuberculosis. IUBMB Life 67, 816–827 (2015).
Bucca, G., Brassington, A. M., Schönfeld, H. J. & Smith, C. P. The HspR regulon of Streptomyces coelicolor: a role for the DnaK chaperone as a transcriptional co-repressordagger. Mol. Microbiol 38, 1093–1103 (2000).
Striebel, F., Hunkeler, M., Summer, H. & Weber-Ban, E. The mycobacterial Mpa–proteasome unfolds and degrades pupylated substrates by engaging Pup’s N-terminus. EMBO J. 29, 1262–1271 (2010).
Pearce, M. J. et al. Identification of substrates of the Mycobacterium tuberculosis proteasome. EMBO J. 25, 5423–5432 (2006).
Elharar, Y. et al. Posttranslational regulation of coordinated enzyme activities in the Pup-proteasome system. Proc. Natl. Acad. Sci. USA 113, E1605–E1614 (2016).
von Rosen, T. et al. Substrates bind to residues lining the ring of asymmetrically engaged bacterial proteasome activator Bpa. Nat. Commun. 16, 3042 (2025).
Davis, B. T. V., Velyvis, A. & Vahidi, S. Fluorinated Ethylamines as Electrospray-Compatible Neutral pH Buffers for Native Mass Spectrometry. Anal. Chem. 95, 17525–17532 (2023).
Liu, R., Li, Q. & Smith, L. M. Detection of large ions in time-of-flight mass spectrometry: effects of ion mass and acceleration voltage on microchannel plate detector response. J. Am. Soc. Mass Spectrom. 25, 1374–1383 (2014).
Chernushevich, I. V. & Thomson, B. A. Collisional cooling of large ions in electrospray mass spectrometry. Anal. Chem. 76, 1754–1760 (2004).
Morgan, C. R. & Engen, J. R. Investigating solution-phase protein structure and dynamics by hydrogen exchange mass spectrometry. Curr. Protoc. Protein Sci. Chapter 17, 17.6.1–17.6.17 (2009).
James, E. I., Murphree, T. A., Vorauer, C., Engen, J. R. & Guttman, M. Advances in Hydrogen/Deuterium exchange mass spectrometry and the pursuit of challenging biological systems. Chem. Rev. 122, 7562–7623 (2022).
Konermann, L., Pan, J. & Liu, Y.-H. Hydrogen exchange mass spectrometry for studying protein structure and dynamics. Chem. Soc. Rev. 40, 1224–1234 (2011).
Konermann, L., Rodriguez, A. D. & Sowole, M. A. Type 1 and Type 2 scenarios in hydrogen exchange mass spectrometry studies on protein–ligand complexes. Analyst 139, 6078–6087 (2014).
Bai, Y., Milne, J. S., Mayne, L. & Englander, S. W. Primary structure effects on peptide group hydrogen exchange. Proteins 17, 75–86 (1993).
Nguyen, D., Mayne, L., Phillips, M. C. & Walter Englander, S. Reference parameters for protein hydrogen exchange rates. J. Am. Soc. Mass Spectrom. 29, 1936–1939 (2018).
Vorauer, C. et al. Rapid assessment of pepsin column activity for reliable HDX-MS studies. J. Am. Soc. Mass Spectrom. 32, 2386–2390 (2021).
Yang, M. et al. Recombinant Nepenthesin II for Hydrogen/Deuterium Exchange Mass Spectrometry. Anal. Chem. 87, 6681–6687 (2015).
Peterle, D., DePice, D., Wales, T. E. & Engen, J. R. Increase the flow rate and improve hydrogen deuterium exchange mass spectrometry. J. Chromatogr. A 1689, 463742 (2023).
Hodge, E. A., Benhaim, M. A. & Lee, K. K. Bridging protein structure, dynamics, and function using hydrogen/deuterium-exchange mass spectrometry. Protein Sci. 29, 843–855 (2020).
Vosper, K. R., Velyvis, A. & Vahidi, S. HDgraphiX: A Web-Based Tool for Visualization of Hydrogen/Deuterium Exchange Mass Spectrometry Data. J. Am. Soc. Mass Spectrom. 36, 1200–1203 (2025).
Tugarinov, V., Hwang, P. M., Ollerenshaw, J. E. & Kay, L. E. Cross-correlated relaxation enhanced 1H−13C NMR Spectroscopy of Methyl Groups in Very High Molecular Weight Proteins and Protein Complexes. J. Am. Chem. Soc. 125, 10420–10428 (2003).
Rosenzweig, R. & Kay, L. E. Bringing dynamic molecular machines into focus by Methyl-TROSY NMR. Ann. Rev. Biochem. 83, 291–315 (2014).
Schütz, S. & Sprangers, R. Methyl TROSY spectroscopy: A versatile NMR approach to study challenging biological systems. Prog. Nucl. Magn. Reson. Spectrosc. https://doi.org/10.1016/j.pnmrs.2019.09.004 (2019).
Sprangers, R., Velyvis, A. & Kay, L. E. Solution NMR of supramolecular complexes: providing new insights into function. Nat. Methods 4, 697–703 (2007).
Evans, R., Dal Poggetto, G., Nilsson, M. & Morris, G. A. Improving the Interpretation of Small Molecule Diffusion Coefficients. Anal. Chem. 90, 3987–3994 (2018).
Valencia, D. P. & González, F. J. Estimation of diffusion coefficients by using a linear correlation between the diffusion coefficient and molecular weight. J. Electroanal. Chem. 681, 121–126 (2012).
Goodhill, G. J. Mathematical guidance for axons. Trends Neurosci. 21, 226–231 (1998).
Nishikawa, T., Nagadoi, A., Yoshimura, S., Aimoto, S. & Nishimura, Y. Solution structure of the DNA-binding domain of human telomeric protein, hTRF1. Structure 6, 1057–1065 (1998).
Sekhar, A., Rosenzweig, R., Bouvignies, G. & Kay, L. E. Mapping the conformation of a client protein through the Hsp70 functional cycle. Proc. Natl. Acad. Sci. 112, 10395–10400 (2015).
Rosenzweig, R., Sekhar, A., Nagesh, J. & Kay, L. E. Promiscuous binding by Hsp70 results in conformational heterogeneity and fuzzy chaperone-substrate ensembles. Elife 6, e28030 (2017).
Sekhar, A., Nagesh, J., Rosenzweig, R. & Kay, L. E. Conformational heterogeneity in the Hsp70 chaperone-substrate ensemble identified from analysis of NMR-detected titration data. Protein Sci. 26, 2207–2220 (2017).
Sekhar, A., Rosenzweig, R., Bouvignies, G. & Kay, L. E. Hsp70 biases the folding pathways of client proteins. Proc. Natl. Acad. Sci. USA 113, E2794–E2801 (2016).
Harkness, R. W. et al. Competing stress-dependent oligomerization pathways regulate self-assembly of the periplasmic protease-chaperone DegP. Proc. Natl. Acad. Sci. USA 118, e2109732118 (2021).
Harkness, R. W., Ripstein, Z. A., Di Trani, J. M. & Kay, L. E. Flexible Client-Dependent Cages in the Assembly Landscape of the Periplasmic Protease-Chaperone DegP. J. Am. Chem. Soc. 145, 13015–13026 (2023).
Turner, M. et al. Structural basis for allosteric modulation of M. tuberculosis proteasome core particle. Nat. Commun. 16, 3138 (2025).
Yuwen, T., Sekhar, A., Baldwin, A. J., Vallurupalli, P. & Kay, L. E. Measuring diffusion constants of invisible protein conformers by triple-quantum 1H CPMG relaxation dispersion. Angew. Chem. Int. Ed. 57, 16777–16780 (2018).
Kramer, A., Schwebke, I. & Kampf, G. How long do nosocomial pathogens persist on inanimate surfaces? A systematic review. BMC Infect. Dis. 6, 130 (2006).
Qi, D. & Scholthof, K.-B. G. A one-step PCR-based method for rapid and efficient site-directed fragment deletion, insertion, and substitution mutagenesis. J. Virol. Methods 149, 85–90 (2008).
Tugarinov, V., Hwang, P. M. & Kay, L. E. Nuclear magnetic resonance spectroscopy of high-molecular-weight proteins. Annu. Rev. Biochem 73, 107–146 (2004).
Laemmli, U. K. Cleavage of Structural Proteins during the Assembly of the Head of Bacteriophage T4. Nature 227, 680–685 (1970).
Kalambet, Y., Kozmin, Y., Mikhailova, K., Nagaev, I. & Tikhonov, P. Reconstruction of chromatographic peaks using the exponentially modified Gaussian function. J. Chemometr. 25, 352–356 (2011).
Glasoe, P. K. & Long, F. A. Use of glass electrodes to measure acidities in deuterium oxide1,2. J. Phys. Chem. 64, 188–190 (1960).
Vahidi, S., Bi, Y., Dunn, S. D. & Konermann, L. Load-dependent destabilization of the γ-rotor shaft in FOF1 ATP synthase revealed by hydrogen/deuterium-exchange mass spectrometry. Proc. Natl. Acad. Sci. 113, 2412–2417 (2016).
Sørensen, L. & Salbo, R. Optimized workflow for selecting peptides for HDX-MS data analyses. J. Am. Soc. Mass Spectrom. 29, 2278–2281 (2018).
Meng, E. C. et al. UCSF ChimeraX: Tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Marty, M. T. et al. Bayesian deconvolution of mass and ion mobility spectra: from binary interactions to polydisperse ensembles. Anal. Chem. 87, 4370–4376 (2015).
Todd, A. R. & Jarrold, M. F. Dramatic improvement in sensitivity with pulsed mode charge Detection Mass Spectrometry. Anal. Chem. 91, 14002–14008 (2019).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Kay, L. E., Keifer, P. & Saarinen, T. Pure absorption gradient enhanced heteronuclear single quantum correlation spectroscopy with improved sensitivity. J. Am. Chem. Soc. 114, 10663–10665 (1992).
Choy, W.-Y. et al. Distribution of molecular size within an unfolded state ensemble using small-angle X-ray scattering and pulse field gradient NMR techniques. J. Mol. Biol. 316, 101–112 (2002).
Bouvignies, G. & Kay, L. E. A 2D 13C-CEST experiment for studying slowly exchanging protein systems using methyl probes: an application to protein folding. J. Biomol. NMR 53, 303–310 (2012).
Yuwen, T., Kay, L. E. & Bouvignies, G. Dramatic decrease in CEST measurement times using multi-site excitation. ChemPhysChem 19, 1707–1710 (2018).
Korzhnev, D. M., Kloiber, K., Kanelis, V., Tugarinov, V. & Kay, L. E. Probing slow dynamics in high molecular weight proteins by methyl-TROSY NMR Spectroscopy: Application to a 723-Residue Enzyme. J. Am. Chem. Soc. 126, 3964–3973 (2004).
Tanford, C. Contribution of hydrophobic interactions to the stability of the globular conformation of proteins. J. Am. Chem. Soc. 84, 4240–4247 (1962).
Acknowledgements
B.T.V.D. acknowledges support from a Graduate Tuition Scholarship from the University of Guelph. Financial support was provided by Canadian Institutes of Health Research Project Grant PJT451412 to S.V. and a Foundation Grant FND-503573 to L.E.K., and by Discovery Grants from the Natural Sciences and Engineering Council of Canada RGPIN-2021-02843 to S.V and 024-03872 to L.E.K. MS data were recorded at the Mass Spectrometry Facility of the Advanced Analysis Centre, University of Guelph. We thank Dr. Dyanne Brewer (University of Guelph) for assistance with MS measurements. We thank Prof. Ashok Sekhar (Indian Institute of Science, Bangalore) for providing backbone resonance assignments of TRF1. We thank Dr. Taylor Forrester (University of Guelph) for help with the generation of the protein topology diagrams. We thank Dr. Algirdas Velyvis (University of Guelph) for guidance and helpful discussions.
Author information
Authors and Affiliations
Contributions
B.T.V.D., E.R., L.E.K., and S.V. initiated the project; B.T.V.D., E.R., L.E.K., and S.V. designed research; B.T.V.D., E.R., A.H., J.U., L.E.K., and S.V. performed research; B.T.V.D., E.R., A.H., J.U., D.B., K.R., K.G., L.E.K., and S.V. contributed new reagents/analytic tools; B.T.V.D., E.R., A.H., and S.V. analyzed data; B.T.V.D., E.R., L.E.K., and S.V. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
A.H, J.U, D.B, K.R, and K.G. are employees of Waters Corporation, whose instrumentation was used in this study. Waters provided technical support, including access to software and, in the case of CD-MS, the prototype instrument itself, enabling nonconventional experiments to be performed. However, the company did not participate in experimental design, data interpretation, or manuscript preparation. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Davis, B.T.V., Rennella, E., Haris, A. et al. Structural heterogeneity and substrate engagement mechanism of the bacterial proteasome activator Bpa. Nat Commun (2026). https://doi.org/10.1038/s41467-026-69978-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-69978-w