Abstract
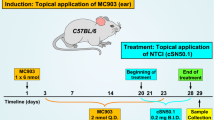
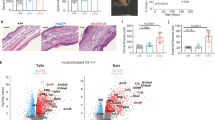
Interleukin-2-inducible T cell kinase is expressed by T cells and amplifies T cell receptor-dependent signals. Interleukin-2-inducible T cell kinase deletion or inhibition reduces production of interleukin-4 and interleukin-13, key drivers of atopic dermatitis. Nerve growth factor signals via the receptor tropomyosin-related kinase A and may promote pruritus in atopic dermatitis lesions. Here we describe PF-07245303, a compound which potently inhibits interleukin-2-inducible T cell kinase and tropomyosin-related kinase family kinases capable of inhibiting T cell receptor-mediated cytokine production from CD4 and CD8 T cells and suppressing nerve growth factor-induced human basophil activation. In human skin explants, PF-07245303 demonstrates inhibition of tropomyosin-related kinase A phosphorylation, suppresses cytokine expression from T cell receptor-activated resident T cells and reverses the expression of atopic dermatitis associated genes. Topical application of PF-07245303 reduces proinflammatory and epidermal changes in a dermatitis model using female mice. By inhibiting both pathogenic inflammatory mechanisms, PF-07245303 may have therapeutic value for patients with atopic dermatitis.
Similar content being viewed by others
Data availability
Source data has been provided, and all primary data generated for this study will be readily shared with researchers for reproducibility purposes upon request. The diffraction data and coordinates of ITK bound to the PF-07245303 compound have been deposited in the Protein Data Bank (PDB) under the accession code 9NWX. The RNA sequencing data generated has been deposited in the GEO repository under accession code GSE297645. Source data are provided with this paper.
References
Brunner, P. M., Guttman-Yassky, E. & Leung, D. Y. The immunology of atopic dermatitis and its reversibility with broad-spectrum and targeted therapies. J. Allergy Clin. Immunol. 139, S65–S76 (2017).
Brar, K. K., Nicol, N. H. & Boguniewicz, M. Strategies for successful management of severe atopic dermatitis. J. Allergy Clin. Immunol. Pract. 7, 1–16 (2019).
Otsuka, A. et al. The interplay between genetic and environmental factors in the pathogenesis of atopic dermatitis. Immunol. Rev. 278, 246–262 (2017).
Sabat, R., Wolk, K., Loyal, L., Docke, W. D. & Ghoreschi, K. T cell pathology in skin inflammation. Semin. Immunopathol. 41, 359–377 (2019).
Sahu, N. & August, A. ITK inhibitors in inflammation and immune-mediated disorders. Curr. Top. Med. Chem. 9, 690–703 (2009).
Andreotti, A. H., Schwartzberg, P. L., Joseph, R. E. & Berg, L. J. T-cell signaling regulated by the Tec family kinase, Itk. Cold Spring Harb. Perspect. Biol. 2, a002287 (2010).
Felices, M., Falk, M., Kosaka, Y. & Berg, L. J. Tec kinases in T cell and mast cell signaling. Adv. Immunol. 93, 145–184 (2007).
Au-Yeung, B. B., Katzman, S. D. & Fowell, D. J. Cutting edge: Itk-dependent signals required for CD4+ T cells to exert, but not gain, Th2 effector function. J. Immunol. 176, 3895–3899 (2006).
Lin, T. A. et al. Selective Itk inhibitors block T-cell activation and murine lung inflammation. Biochemistry 43, 11056–11062 (2004).
Andreotti, A. H., Joseph, R. E., Conley, J. M., Iwasa, J. & Berg, L. J. Multidomain control over TEC kinase activation state tunes the T cell response. Annu. Rev. Immunol. 36, 549–578 (2018).
August, A. & Ragin, M. J. Regulation of T-cell responses and disease by Tec kinase Itk. Int. Rev. Immunol. 31, 155–165 (2012).
Weeks, S., Harris, R. & Karimi, M. Targeting ITK signaling for T cell-mediated diseases. iScience 24, 102842 (2021).
Ferrara, T. J., Mueller, C., Sahu, N., Ben-Jebria, A. & August, A. Reduced airway hyperresponsiveness and tracheal responses during allergic asthma in mice lacking tyrosine kinase inducible T-cell kinase. J. Allergy Clin. Immunol. 117, 780–786 (2006).
Jain, N. et al. CD28 and ITK signals regulate autoreactive T cell trafficking. Nat. Med. 19, 1632–1637 (2013).
Mueller, C. & August, A. Attenuation of immunological symptoms of allergic asthma in mice lacking the tyrosine kinase ITK. J. Immunol. 170, 5056–5063 (2003).
Lechner, K. et al. Targeting of the Tec kinase ITK drives resolution of T cell-mediated colitis and emerges as potential therapeutic option in ulcerative colitis. Gastroenterology 161, 1270–1287 (2021).
Kannan, A. K., Kim, D. G., August, A. & Bynoe, M. S. Itk signals promote neuroinflammation by regulating CD4+ T-cell activation and trafficking. J. Neurosci. 35, 221–233 (2015).
Gomez-Rodriguez, J. et al. Itk is required for Th9 differentiation via TCR-mediated induction of IL-2 and IRF4. Nat. Commun. 7, 10857 (2016).
Deakin, A. et al. Characterisation of a K390R ITK kinase dead transgenic mouse-implications for ITK as a therapeutic target. PLoS ONE 9, e107490 (2014).
Bonin, A. von et al. Inhibition of the IL-2-inducible tyrosine kinase (Itk) activity: a new concept for the therapy of inflammatory skin diseases. Exp. Dermatol. 20, 41–47 (2011).
Otake, S. et al. Topical application of BMS-509744, a selective inhibitor of interleukin-2-inducible T cell kinase, ameliorates imiquimod-induced skin inflammation in mice. Biol. Pharm. Bull. 44, 528–534 (2021).
Nadeem, A. et al. Inhibition of interleukin-2-inducible T-cell kinase causes reduction in imiquimod-induced psoriasiform inflammation through reduction of Th17 cells and enhancement of Treg cells in mice. Biochimie 179, 146–156 (2020).
Sun, Y. et al. Inhibition of the kinase ITK in a mouse model of asthma reduces cell death and fails to inhibit the inflammatory response. Sci. Signal. 8, ra122 (2015).
Matsumoto, Y. et al. Identification of highly expressed genes in peripheral blood T cells from patients with atopic dermatitis. Int. Arch. Allergy Immunol. 129, 327–340 (2002).
Kannan, A. et al. Allele-sensitive mutant, Itkas, reveals that Itk kinase activity is required for Th1, Th2, Th17, and iNKT-cell cytokine production. Eur. J. Immunol. 45, 2276–2285 (2015).
Mollanazar, N. K., Smith, P. K. & Yosipovitch, G. Mediators of chronic pruritus in atopic dermatitis: getting the itch out? Clin. Rev. Allergy Immunol. 51, 263–292 (2016).
Benecke, H., Lotts, T. & Stander, S. Investigational drugs for pruritus. Expert Opin. Investig. Drugs 22, 1167–1179 (2013).
Botchkarev, V. A. et al. Neurotrophins in skin biology and pathology. J. Investig. Dermatol. 126, 1719–1727 (2006).
Rukwied, R. R., Main, M., Weinkauf, B. & Schmelz, M. NGF sensitizes nociceptors for cowhage- but not histamine-induced itch in human skin. J. Investig. Dermatol. 133, 268–270 (2013).
Yamaguchi, J., Aihara, M., Kobayashi, Y., Kambara, T. & Ikezawa, Z. Quantitative analysis of nerve growth factor (NGF) in the atopic dermatitis and psoriasis horny layer and effect of treatment on NGF in atopic dermatitis. J. Dermatol. Sci. 53, 48–54 (2009).
Matsumura, S., Terao, M., Murota, H. & Katayama, I. Th2 cytokines enhance TrkA expression, upregulate proliferation, and downregulate differentiation of keratinocytes. J. Dermatol. Sci. 78, 215–223 (2015).
Tominaga, M., Tengara, S., Kamo, A., Ogawa, H. & Takamori, K. Psoralen-ultraviolet A therapy alters epidermal Sema3A and NGF levels and modulates epidermal innervation in atopic dermatitis. J. Dermatol. Sci. 55, 40–46 (2009).
Bürgi, B. et al. Basophil priming by neurotrophic factors. Activation through the trk receptor. J. Immunol. 157, 5582–5588 (1996).
Bischoff, S. C. & Dahinden, C. A. Effect of nerve growth factor on the release of inflammatory mediators by mature human basophils. Blood 79, 2662–2669 (1992).
Bryan, M. C. & Rajapaksa, N. S. Kinase Inhibitors for the treatment of immunological disorders: recent advances. J. Med. Chem. 61, 9030–9058 (2018).
Zapf, C. W. et al. Covalent inhibitors of interleukin-2 inducible T cell kinase (itk) with nanomolar potency in a whole-blood assay. J. Med. Chem. 55, 10047–10063 (2012).
Han, S. et al. Selectively targeting an inactive conformation of interleukin-2-inducible T-cell kinase by allosteric inhibitors. Biochem. J. 460, 211–222 (2014).
Proschak, E., Stark, H. & Merk, D. Polypharmacology by design: a medicinal chemist’s perspective on multitargeting compounds. J. Med. Chem. 62, 420–444 (2019).
Roberts, M. S. et al. Topical drug delivery: history, percutaneous absorption, and product development. Adv. Drug Deliv. Rev. 177, 113929 (2021).
Di, L. et al. Development of a new permeability assay using low-efflux MDCKII cells. J. Pharm. Sci. 100, 4974–4985 (2011).
Aprile, S., Serafini, M. & Pirali, T. Soft drugs for dermatological applications: recent trends. Drug Discov. Today 24, 2234–2246 (2019).
Ryu, S. et al. Evaluation of fraction unbound across 7 tissues of 5 species. J. Pharm. Sci. 109, 1178–1190 (2020).
Yu, S. et al. A novel liquid chromatography/tandem mass spectrometry based depletion method for measuring red blood cell partitioning of pharmaceutical compounds in drug discovery. Rapid Commun. Mass Spectrom. 19, 250–254 (2005).
Bagal, S. K. et al. Discovery of potent, selective, and peripherally restricted Pan-Trk kinase inhibitors for the treatment of pain. J. Med. Chem. 61, 6779–6800 (2018).
Rashighi, M. & Harris, J. E. Vitiligo pathogenesis and emerging treatments. Dermatol. Clin. 35, 257–265 (2017).
Olayinka, J. J. T. & Richmond, J. M. Immunopathogenesis of alopecia areata. Curr. Res. Immunol. 2, 7–11 (2021).
Ito, Y. et al. Basophil recruitment and activation in inflammatory skin diseases. Allergy 66, 1107–1113 (2011).
Lourenço, F. D. et al. Activated status of basophils in chronic urticaria leads to interleukin-3 hyper-responsiveness and enhancement of histamine release induced by anti-IgE stimulus. Br. J. Dermatol. 158, 979–986 (2008).
Borriello, F., Granata, F. & Marone, G. Basophils and skin disorders. J. Investig. Dermatol. 134, 1202–1210 (2014).
Yan, W. et al. Insights into current tropomyosin receptor kinase (TRK) inhibitors: development and clinical application. J. Med. Chem. 62, 1731–1760 (2019).
Price, E. A., Krasowska-Zoladek, A., Nanda, K. K., Stachel, S. J. & Henze, D. A. Development of a pharmacodynamic biomarker to measure target engagement from inhibition of the NGF-TrkA pathway. J. Neurosci. Methods 282, 34–42 (2017).
Smith, S. H. et al. Development of a topical treatment for psoriasis targeting rorgamma: from bench to skin. PLoS ONE 11, e0147979 (2016).
Jones, P. et al. Design and synthesis of a pan-janus kinase inhibitor clinical candidate (PF-06263276) suitable for inhaled and topical delivery for the treatment of inflammatory diseases of the lungs and skin. J. Med. Chem. 60, 767–786 (2017).
Brunner, P. M. et al. Early-onset pediatric atopic dermatitis is characterized by T(H)2/T(H)17/T(H)22-centered inflammation and lipid alterations. J. Allergy Clin. Immunol. 141, 2094–2106 (2018).
Choi, J. et al. Translational relevance of mouse models of atopic dermatitis. J. Clin. Med. 10, 613 (2021).
Guttman-Yassky, E. et al. Dupilumab progressively improves systemic and cutaneous abnormalities in patients with atopic dermatitis. J. Allergy Clin. Immunol. 143, 155–172 (2019).
Patrick, G. J., Archer, N. K. & Miller, L. S. Which way do we go? Complex interactions in atopic dermatitis pathogenesis. J. Investig. Dermatol. 141, 274–284 (2021).
Guttman-Yassky, E. & Krueger, J. G. Atopic dermatitis and psoriasis: two different immune diseases or one spectrum? Curr. Opin. Immunol. 48, 68–73 (2017).
Lechner, K. S., Neurath, M. F. & Weigmann, B. Role of the IL-2 inducible tyrosine kinase ITK and its inhibitors in disease pathogenesis. J. Mol. Med. 98, 1385–1395 (2020).
Riether, D. et al. 5-Aminomethylbenzimidazoles as potent ITK antagonists. Bioorg. Med. Chem. Lett. 19, 1588–1591 (2009).
Trani, G. et al. Design, synthesis and structure-activity relationships of a novel class of sulfonylpyridine inhibitors of Interleukin-2 inducible T-cell kinase (ITK). Bioorg. Med. Chem. Lett. 24, 5818–5823 (2014).
Burch, J. D. et al. Tetrahydroindazoles as interleukin-2 inducible T-cell kinase inhibitors. Part II. Second-generation analogues with enhanced potency, selectivity, and pharmacodynamic modulation in vivo. J. Med. Chem. 58, 3806–3816 (2015).
Zhong, Y. et al. Targeting interleukin-2-inducible T-cell kinase (ITK) and resting lymphocyte kinase (RLK) using a novel covalent inhibitor PRN694. J. Biol. Chem. 290, 5960–5978 (2015).
Zhou, D., Zuo, Y. & Pan, Z. Discovery of potent and highly selective interleukin-2-inducible T-cell kinase degraders with in vivo activity. J. Med. Chem. 66, 4979–4998 (2023).
Hsu, L.-Y. et al. Synthesis and characterization of soquelitinib a selective ITK inhibitor that modulates tumor immunity. NPJ Drug Discov. 1, 2 (2024).
Colgan, J. D. & Hankel, I. L. Signaling pathways critical for allergic airway inflammation. Curr. Opin. Allergy Clin. Immunol. 10, 42–47 (2010).
Gomez-Rodriguez, J. et al. Differential expression of interleukin-17A and −17F is coupled to T cell receptor signaling via inducible T cell kinase. Immunity 31, 587–597 (2009).
Gomez-Rodriguez, J. et al. Itk-mediated integration of T cell receptor and cytokine signaling regulates the balance between Th17 and regulatory T cells. J. Exp. Med. 211, 529–543 (2014).
Neil, J. E., Brown, M. B. & Williams, A. C. Human skin explant model for the investigation of topical therapeutics. Sci. Rep. 10, 21192 (2020).
Takano, N., Sakurai, T., Ohashi, Y. & Kurachi, M. Effects of high-affinity nerve growth factor receptor inhibitors on symptoms in the NC/Nga mouse atopic dermatitis model. Br. J. Dermatol. 156, 241–246 (2007).
Roblin, D. et al. Topical TrkA kinase inhibitor CT327 is an effective, novel therapy for the treatment of pruritus due to psoriasis: results from experimental studies, and efficacy and safety of CT327 in a phase 2b clinical trial in patients with psoriasis. Acta Derm. Venereol. 95, 542–548 (2015).
Kleinman, E., Laborada, J., Metterle, L. & Eichenfield, L. F. What’s new in topicals for atopic dermatitis? Am. J. Clin. Dermatol. 23, 595–603 (2022).
Wang, F. et al. A basophil-neuronal axis promotes itch. Cell 184, 422–440 (2021).
Sin, A. Z., Roche, E. M., Togias, A., Lichtenstein, L. M. & Schroeder, J. T. Nerve growth factor or IL-3 induces more IL-13 production from basophils of allergic subjects than from basophils of nonallergic subjects. J. Allergy Clin. Immunol. 108, 387–393 (2001).
Simpson, E. L. et al. Efficacy and safety of dupilumab in adolescents with uncontrolled moderate to severe atopic dermatitis: a phase 3 randomized clinical trial. JAMA Dermatol 156, 44–56 (2020).
Folkersen, L. et al. Integration of known DNA, RNA and protein biomarkers provides prediction of anti-TNF response in rheumatoid arthritis: results from the COMBINE study. Mol. Med. 22, 322–328 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Chen, Y., Chen, L., Lun, A. T. L., Baldoni, P. L. & Smyth, G. K. edgeR v4: powerful differential analysis of sequencing data with expanded functionality and improved support for small counts and larger datasets. Nucleic Acids Res. 53, gkaf018 (2025).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43, e47 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Wickham, H. Ggplot2: Elegant Graphics for Data Analysis (Springer, 2009).
Acknowledgements
We thank the staff of the Pfizer Flow Cytometry and Next-Generation Sequencing Technology Centers for their support. We thank Alexia Scaros and Ying Zhang for outsourcing support. We thank Isac Lee and Samir Lal with assistance with the submission of the RNAseq gene expression data and figure formatting.
Author information
Authors and Affiliations
Contributions
A.L.B., K.F., L.J., J.B., J.H.S., K.K.C., C.A., Y.Z., T.A., R.T.S., W.L., A.R., and J.V. performed biochemical, cell-based, ex vivo skin experiments or in vivo study, analyzed the data, and prepared the methods sections. K.H. performed microscopic analysis of tissue sections from in vivo model. N.L.N., S.L. performed crystallography studies and determined the structures. J.E.D., J.W.S., A.C-G., G.M.C., F.E.L., R.A.D., S.D., R.H., M.P., J.I.T., S.W.B., and B.S.G. designed and synthesized PF-07245303. S.A.J., Y.L. conducted transcriptomics and the corresponding quantitative analysis and data visualization. J.G. and S.A.J. performed statistical analysis. J.L.D., S.J., T.L.F., S.J.S., K.N., M.H., F.V., K.L.L., B.S.G., and M.J.P. were involved in the experimental design and data interpretation. J.L.D., S.W.J., J.B., S.L., S.W.B., B.S.G., and M.J.P. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
J.L.D., K.K.C., L.J., A.B., K.F., J.B., S.J., Y.L., J.H.S., Y.Z. T.A., K.H., C.A., R.T.S., J.G., S.B., A.C-G, G.C., J.D., R.A.D., S.D., T.F., R.W.H., S.L., F.L., N.N., M.P., J.S., J.I.T., S.S., K.N., M.H., F.V., K.L.L., B.S.G., and M.J.P. were employees of Pfizer Inc. at the time this work was performed. W.L., A.R., and J.V. were employees of MedPharm Ltd. at the time this work was performed. The authors declare no other competing interests.
Peer review
Peer review information
Nature Communications thanks Asier Unciti-Broceta, Avery August, Michihiro Hide, and Balázs István Tóth for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Duffen, J.L., Crouse, K.K., Ji, L. et al. Discovery of an ITK and TRK kinase inhibitor for the potential topical treatment of atopic dermatitis. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70000-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70000-6