Abstract
The mycotoxin deoxynivalenol (DON) poses severe threats to human and animal well-being globally. Enzymatic degradation is the most effective way to eliminate DON toxicity, yet no catalytic process for complete degradation of DON has been uncovered. Here, we show that a metabolic pathway initiated by C3-epimerization and C8-reduction is responsible for complete degradation of DON in the DON-metabolizing bacterium Nocardioides sp. S5-5. Two horizontally transferred aldo-keto reductase genes, DONepi and DONrd, have evolved to orchestrate C3-epimerization and C8-reduction respectively. Notably, the octameric-structured DONepi alone catalyzes C3-epimerization of DON by steering the rigid-body rotation of the transient 3-keto intermediate for stereoinverting reduction. Moreover, DONrd can catalyze the C8-reduction of DON and its C3-epimerized product 3-epi-DON simultaneously to form C8-hydroxyl products, which facilitates the further degradation by a potential oxidase and other putative enzymes. DONepi expression in transgenic plants confers resistance to DON, representing potential for controlling mycotoxin contamination pre- and postharvest.
Similar content being viewed by others
Introduction
Mycotoxins, which are fungal secondary metabolites, contaminate 60–80% of crops globally, posing severe threats to both humans and livestock1. Deoxynivalenol (DON) is the most common mycotoxin worldwide, occurring at higher levels in Europe and Asia2,3. Recently, due to climate change and crop residue incorporation, the increasing frequency of Fusarium head blight epidemics, which is caused by DON-producing pathogenic Fusarium, has led to more serious and recurrent risks of DON contamination in cereal raw materials and finished food products4,5,6. With stable physicochemical properties, DON cannot be efficiently removed during food and feed processing and can, therefore, enter human and animal food chains7. DON can interfere with protein translation through binding to the A-site of the peptidyl transferase center on the large subunit (60S) of the eukaryotic ribosome, leading to acute and chronic mycotoxicosis in humans and animals8,9. Microbial and enzymatic degradation represent promising approaches for managing DON contamination due to their high specificity, cost-efficiency, and environmental friendliness10. In the past few decades, several patterns for transforming DON and its structural analogs by diverse microbes and enzymes have been investigated, including: (1) reductive de-epoxidation and glutathione conjugation targeting the C12, 13 epoxide group6,11,12,13,14; (2) oxidation, epimerization, and glycosylation targeting the C3 hydroxyl15,16,17; (3) hydroxylation targeting the C1618; and (4) isomerization targeting the C9-C10 double bond19,20,21. However, the products resulting from DON transformation via these pathways exhibited varying degrees of retained toxicity22,23,24.
The final product of DON C3-epimerization, 3-epi-DON, is one of the lowest-toxicity metabolites among the currently known microbial metabolites of DON 12,23. Enzymes involved in this process have been identified in several bacterial strains, which convert DON to 3-epi-DON through two independent reactions catalyzed by distinct classes of enzymes. The initial step is the oxidation of DON to form the intermediate 3-keto-DON, carried out by pyrroloquinoline quinone (PQQ) dependent alcohol dehydrogenases (PQQ-ADHs). Subsequently, the second step entails the reduction of 3-keto-DON to produce the final product 3-epi-DON, which is accomplished by aldo-keto reductases (AKRs) 25,26. Nevertheless, PQQ-ADHs require the particular coenzyme PQQ, which cannot be produced naturally in typical expression systems employed in industrial biotechnology, including E. coli and yeast26. This incapability severely restricts the application of PQQ-ADHs within the realms of food and feed safety, since the need for exogenous addition of the coenzyme PQQ leads to prohibitively high costs. In contrast to the formation of diverse products with differential levels of retained toxicity via the well-known pathway associated with DON transformation, certain microbes have the capacity to degrade DON completely. However, to date, the isolation of strains endowed with such degradation ability has remained scarce. Moreover, the specific enzymes involved in and the precise catalytic mechanisms governing the complete degradation of DON remain unknown.
Here, we identify the enzymes and catalytic process underlying the complete degradation of DON in Nocardioides sp. S5-5, a bacterium with the capacity to degrade and assimilate DON. The process is initiated by two distinct pathways: C3-epimerization and C8-reduction. Specifically, two AKR genes, DONepi and DONrd, acquired via horizontal gene transfer mediated by genomic islands, govern the two pathways, respectively. To reveal their catalytic mechanisms, we employ Cryo-EM structural analysis and QM/MM metadynamics simulations. Furthermore, in plants, DONepi exhibits substantial activity in increasing the resistance to DON, suggesting the broad application potential in mycotoxin degradation and crop resistance improvement.
Results
Complete degradation of DON by strain Nocardioides sp. S5-5
To isolate the strain capable of completely neutralizing the toxicity of DON, the screening strategy based on toxicity assessment using Lemna minor was employed in the initial selection process. After incubation with DON, the culture filtrate from soil sample S5 showed no inhibitory effect on the growth of L. minor, while most of the other samples showed strong toxicity to L. minor similar to that of DON (Fig. 1a). Further analysis using GC-MS indicated that DON was either not degraded or converted into slightly less toxic metabolites, such as 3-keto-DON, in these culture filtrates that retained toxicity to L. minor (Fig. 1b and Supplementary Fig. 1). Notably, no DON or DON-derived metabolites were detected in the culture filtrate of sample S5, suggesting a complete degradation of DON (Fig. 1b). Moreover, a time-dependent detection by HPLC demonstrated that 3-epi-DON served as the intermediate product during DON degradation by consortium S5 (Supplementary Fig. 2). To isolate single colonies capable of completely degrading DON from S5 subcultures, the subcultures were treated with antibiotics. 16S rRNA high-throughput sequencing revealed that treatment with the antibiotic Penicillin G (PenG) enhanced the relative abundance of the Nocardioides genus. In contrast, this genus was absent in Erythromycin-treated subcultures, which lacked DON-degrading activity, indicating the potential role of the Nocardioides genus in DON degradation (Fig. 1c). Eventually, a bacterial strain (termed S5-5) that showed similar DON-degrading activity to the S5 consortium was obtained from the PenG-treated subculture (Supplementary Fig. 3). Based on morphological and phylogenetic analysis, it was identified as Nocardioides sp. (Fig. 1d and Supplementary Fig. 4).
a Screening of enrichment cultures from soils following seven days of incubation with DON using toxicity assessment with Lemna minor. The growth and morphology of L. minor in response to the culture filtrates were investigated at 1 day, 3 days, and 6 days after treatment. b GC-MS detection of DON and DON-derived metabolites in the enrichment cultures following seven days of incubation with DON. The right inserts indicate the structures of DON and DON-derived metabolites. TIC, total ion chromatogram. c The relative abundance of consortium S5 at the genus level and corresponding degradation activity responding to different treatments of Penicillin G (PenG) and Erythromycin (Ery). ND not detected. d The phylogeny of Nocardioides sp. strain S5-5. A neighbor-joining phylogenetic tree on the basis of partial 16S rRNA sequences of strain S5-5 and related microorganisms was constructed. The bar indicates a 0.5% sequence divergence. e Degradation and assimilation of DON by strain S5-5. The growth of strain S5-5 in the MM medium containing DON as the only carbon source was investigated. n = 3 biological replicates per condition. Data are presented as mean ± SD, and analyzed by two-tailed t-test. Numbers in color represent the p-values from the statistical analysis relative to the control group without DON addition. NS, no significant difference. f GC-MS profiles of the intermediate products at 0 to 24 h during DON degradation by strain S5-5. The right panel indicates the enlarged view of the retention time, ranging from 9.2 min to 10.8 min. g A line graph showing DON depletion and transient accumulation of various intermediates as shown in (f). The relative abundance was calculated by comparing to the peak area of DON at 0 h. n = 3 biological replicates per condition. Data are presented as mean ± SD. h Mass spectra of the intermediates M1, M2 and M3 detected by GC-MS. The inserts indicate the corresponding molecular structures of these intermediates. EI-MS, electron ionization-mass spectrum.
The growth curve of strain S5-5 in a mineral medium (MM) containing DON at concentrations ranging from 10 to 1000 μM demonstrated that strain S5-5 was capable of utilizing DON as the sole carbon and energy source (Fig. 1e). Additionally, strain S5-5 showed high degradation efficiency for DON in the culture medium and agricultural matrix under diverse conditions (Supplementary Fig. 5). Besides, it can also degrade multiple type A and B trichothecene mycotoxins that often co-contaminate grains with DON, such as Neosolaniol, 3-Acetyl deoxynivalenol, Nivalenol, and Fusarenon-X (Supplementary Fig. 6). To comprehensively explore the detailed process of DON complete degradation by S5-5, we meticulously carried out a time-dependent mass spectrometry assay targeting metabolites. Intriguingly, this assay unveiled that, in addition to 3-epi-DON, three hitherto-unknown intermediate metabolites (designated as M1, M2, and M3) emerged during the DON degradation process by S5-5 (Fig. 1f). These metabolites could not be detected using the HPLC-UV technique. Moreover, the increase of metabolites was disproportionate to the decrease of DON in relative abundance, and all of them disappeared at 24 hai (hours after inoculation) (Fig. 1g). Compared to the other three metabolites, M3 was the latest to reach its highest relative abundance (Fig. 1g). The further assay using these metabolites as substrates for strain S5-5 indicated that M3 was actually the catalytic product of 3-epi-DON. (Supplementary Fig. 7).
Notably, GC-MS profiles showed that the corresponding TMSI (N-(trimethylsilyl)imidazole) derivatives of M1, M2, and M3 have the same molecular ion at m/z 586.3 (Fig. 1h), leaving a 74-dalton (Da) difference compared to derivatized DON and 3-epi-DON (512.3). The displacement of a hydroxyl hydrogen atom with a trimethylsilyl group increases the molecular mass by 72 Da. This suggests that the formation of M1 and M2 from DON, and M3 from 3-epi-DON, involves the reduction of a ketone group to a hydroxyl group—leading to a 2 Da mass increase (Fig. 1h). We further confirmed this 2 Da mass difference between the three intermediates and their respective parent compounds via liquid chromatography-high-resolution mass spectrometry (LC-HRMS) (Supplementary Fig. 8). Since DON and 3-epi-DON each contain only one ketone group, located at the C8 position, the ketone reduction reaction must occur at C8. To further elucidate the chemical structure, nuclear magnetic resonance (NMR) analysis was performed. Compared to the ¹H-NMR spectrum of DON, the spectrum of 8-OH-DON showed a new proton signal at approximately δ 4.27 ppm (a region characteristic of CH–OH groups). Additionally, significant chemical shift perturbations and multiplicity changes were observed for protons at C7 and C10 (H-7, H-10) (Supplementary Fig. 9; Table 1). These spectral changes are consistent with the expected electronic and structural effects of C8 ketone reduction. Thus, M1 and M2 are a pair of epimers, designated α- and β-8-hydroxyl-DON (8-OH-DON), respectively. Likewise, M3 is designated 8-hydroxyl-3-epi-DON (8-OH-3-epi-DON). These intermediate metabolites were further degraded by S5-5, and no signal typical of trichothecenes was detected by LC-HRMS under full scan mode at 24 hai (Supplementary Fig. 10a), while the culture filtrates from S5-5 post-DON degradation had no toxic effect on L. minor (Supplementary Fig. 10b, c). Moreover, obvious production of CO2 and 13CO2 was observed after degradation of DON by strain S5-5, where DON and stable carbon isotope (13C)-labelled DON were used as the sole carbon sources in MM medium (Supplementary Fig. 11). Taken together, these data indicated that DON was completely degraded and assimilated, serving as the carbon source for the growth of strain S5-5. Notably, this process was initiated by two distinct metabolic pathways: C3-epimerization and C8-reduction.
Two horizontally transferred aldo-keto reductases trigger the degradation of DON
To identify candidate genes responsible for DON degradation, the genome of strain S5-5 was sequenced (GenBank accession No. CP199913), and a BAC library of strain S5-5 consisting of 1536 clones with an average insert size of about 140 kb was screened based on the enzymatic activity assay (Fig. 2a and Supplementary Figs. 12 and 13). Paired-end sequencing of BAC clones and alignment to the reference genome of S5-5 revealed that thirteen genes were commonly shared among the inserts of 25 clones with 3-epi-DON forming activity (Supplementary Fig. 14), and eleven genes were commonly shared among the inserts of 29 clones with 8-OH-DON forming activity (Supplementary Fig. 15). A total of 24 genes were isolated from the strain S5-5 and expressed in E. coli. Among them, two recombinant proteins exhibited stable DON catalytic activity. One protein encoded by gene0108 (GenBank accession No. PX396128) catalyzes C3-epimerization of DON and is designated as DON 3-epimerase (DONepi) (Fig. 2b, e, and Supplementary Fig. 16a). Kinetic analysis revealed that the kcat, Km, and kcat/Km of DONepi are 0.33 ± 0.04 min−1, 2.71 ± 0.85 mM, and 121.77 min−1 M−1, respectively (Supplementary Fig. 16b). Another protein encoded by gene4454 (GenBank accession No. PX396129) catalyzes C8-reduction of DON and is designated as DON 8-reductase (DONrd) (Fig. 2c, f, and Supplementary Fig. 17a). Kinetic analysis revealed that the kcat, Km, and kcat/Km of DONrd are 32.07 ± 2.84 min−1, 153.38 ± 29.06 μM, and 209.12 min−1 mM−1, respectively (Supplementary Fig. 17b). DONrd also contributes to the reduction of 3-epi-DON to produce 8-OH-3-epi-DON (Fig. 2d, g). Various chemical and metal ions affect the enzymatic activity of DONepi and DONrd to varying degrees (Supplementary Fig. 18).
a Process of screening candidate genes for DON degradation. A bacterial artificial chromosome (BAC) library was constructed, consisting of 1536 clones with an average insert fragment size of 140 kb. The DON degradation activity of these clones was assayed. BAC end sequencing was employed to analyze the common genes present in clones with DON-degrading capability. b Epimerization of DON at C3 by the recombinant DON epimerase (DONepi). DON and its catalyzed product by DONepi were detected using HPLC. c Reduction of DON at C8 by the recombinant DON reductase (DONrd). DON and its catalyzed product by DONrd were detected using GC-MS. d Reduction of 3-epi-DON at C8 by the recombinant DONrd. 3-epi-DON and its catalyzed product by DONrd were detected using GC-MS. TIC, total ion chromatogram. Influence of pH and temperature on the epimerization activity of recombinant DONepi towards DON (e), and the reduction activity of recombinant DONrd towards DON (f) and 3-epi-DON (g). n = 3 biological replicates per condition. Data are presented as mean ± SD. h Protein sequence similarity network (SSN) analysis of DONepi, DONrd, and AKR superfamily members. A 45% amino acid sequence identity threshold was employed to partition different AKRs into distinct clusters. Enzymes within each color-coded cluster are hypothesized to exhibit homogeneous biochemical activity, based on sequence-function correlations. Nodes containing all AKRs from the strain S5-5 genome and their directly connected neighbors by edges were enlarged and colored in red. The inserts indicate the sources and GenBank accession numbers of these AKRs. i DONepi and DONrd are situated within the genomic islands (GIs) of the genome of strain S5-5. The GIs were predicted using the IslandViewer4 webserver. A circular visualization of all predicted genomic islands is presented. The blocks are colored based on the prediction method: integrated results in dark red, IslandPath-DIMOB in blue, SIGI-HMM in orange, and IslandPick in green. The inner black circle represents the GC content of the S5-5 genome. The inward-facing part indicates a lower GC content relative to the average GC content of the entire genome, whereas the outward-facing part indicates a higher GC content.
DONepi and DONrd, which share a 25% amino acid identity, are most closely related to families AKR12 and AKR15, respectively (Supplementary Fig. 19). However, both of them have an identity of less than 40% relative to the members of these families. This suggests that they belong to new AKR families. Furthermore, a sequence similarity network (SSN) showed that most AKRs from both prokaryotes and eukaryotes were grouped into a primary cluster. This cluster included all AKRs from strain S5-5, with the exception of DONepi and DONrd, as well as other AKRs known for their activities in DON oxidation and 3-keto-DON reduction. (Fig. 2h). In contrast, DONrd was grouped into an additional smaller cluster, while DONepi had not been mapped to any cluster (Fig. 2h), which revealed the functional divergence of DONepi and DONrd.
Interestingly, the homologous proteins with the highest identity to these two specialized AKRs across all kingdoms of life were found in the genera Rhodococcus and Agromyces, sharing 75% and 55% amino acid identities with DONepi and DONrd, respectively. Based on the phylogenetic analysis, DONepi and its close homologs from the genera Nocardia, Rhodococcus, and Pseudonocardia were clustered together and distributed in a completely distinct branch separate from the Nocardioides clade, which is composed of nearly all homologs of DONepi from the Nocardioides genus (Supplementary Fig. 20). Similarly, DONrd was also clustered together with its close homologs from the genera Microbacterium, Agromyces, and Arthrobacter in an independent branch distinct from the clade composed of DONrd homologs from the Nocardioides genus (Supplementary Fig. 21). Moreover, synteny analysis revealed that the genomic sequences flanking the DONepi and DONrd genes respectively constitute a microsyntenic region that is conserved among bacterial species within the DONepi and DONrd clades. (Supplementary Figs. 20b and 21b). Thus, the occurrence of the DONepi and DONrd genes in the genome of strain S5-5 could be caused by horizontal gene transfer (HGT) events. Further analysis indicated that the DONepi and DONrd genes were located in two genomic islands (GIs) of the strain S5-5 genome, where the GC content was significantly different compared to the adjacent region of the genome. The predicted GIs, including DONepi and DONrd, have a large size of 13.2 kb and 18.9 kb, and contain fourteen and nineteen genes, respectively, which, according to the COG annotation, are mainly related to metabolism (Fig. 2i, Supplementary Fig. 22, and Supplementary Tables 2, 3). Additionally, the microsyntenic regions related to DONepi and DONrd in the genomes of other bacteria from the DONepi and DONrd clades were also located in the GIs (Supplementary Figs. 23–26). In conclusion, these results revealed that DONepi and DONrd, acquired via GIs-mediated HGT, initiate the degradation of DON in two distinct metabolic pathways by epimerizing DON at the C3 position and reducing DON at the C8 position.
Cryo-EM structure of DONepi and the catalytic mechanism of C3-epimerization
To determine the structure of DONepi, single-particle cryo-EM was performed. Micrographs showed that particles with diverse angular coverage and secondary structural features could be observed in two-dimensional (2D) class averages (Supplementary Fig. 27). Further data processing resulted in a 3D reconstruction of DONepi with the dihedral symmetry (D4) and an overall resolution of 2.56 Å (Supplementary Fig. 27 and Table 4). DONepi assembles into an octameric structure (115 × 82 Å) (PDB ID: 9WK1). It is composed of two tightly-packed homotetramers with a 30° offset (Fig. 3a, b). Each protomer of DONepi contains a typical AKR signature structure-(α/β)8-barrel, and one coenzyme NADP+, which takes on an extended conformation. The nicotinamide ring on NADP+ is π–π stacked with Trp207, and the carboxamide group interacts with Thr155. Moreover, the adenine ring on NADP⁺ makes contact with Ser216 and Gln291 (Supplementary Fig. 28). The C-terminal tail loops of four protomers within one homotetramer act as anchors and mutually insert into the corresponding protomers in the other homotetramer, consequently forming an octameric quaternary structure (Fig. 3b). Although DONepi adopts a similar octameric architecture, the insert direction of anchors in DONepi is obviously different from that of other known AKRs in the family (Supplementary Fig. 29).
a Single-particle cryo-EM reconstruction of the DONepi octamer showing top and side views. Each protomer is presented in a distinct color. b Cartoon representation of the DONepi octamer. DONepi constitutes an octameric structure made up of two tightly-packed homotetramers. Chains A–D are part of one tetramer, while chains E–H belong to the other. The two homotetramers are clasped with a 30° offset, and the C-terminal tail loops serve as anchors to stabilize the structure, as indicated in the enlarged view. c Close-up view of the binding model of the ternary complex of DONepi-NADP+-DON. Amino acids involved in DON coordination are shown as cyan sticks. DON and NADP+ molecules are shown as red and orange sticks, respectively. Dashed yellow lines indicate the hydrogen bonding interactions between DONepi and DON. d Variation and conservation in prospective activity sites among DONepi homologs. The phylogenetic tree was constructed using the maximum likelihood method with MEGA X. The numbers on top of the sequence alignment represent the positions of amino acids in DONepi. Sequence logos denote the relative frequency of each amino acid at that position, which was calculated using the top 1000 BLASTp hits. The red stars below indicate DON-binding residues in DONepi and conserved catalytic tetrad active sites. e DON-epimerizing activity of wild-type DONepi, DONepi homologs, and DONepi mutants of prospective activity sites. Amino acid residues in the catalytic tetrad were mutated into Alanine (Ala, A). Y27 and R86 were mutated to W and H or G based on (d). n = 3 biological replicates per group. Data are presented as mean ± SD, and analyzed by two-tailed t-test. ** indicate the significant difference at P < 0.01 compared with DONepi.
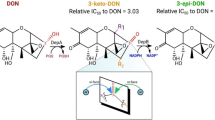
Regrettably, despite multiple attempts, we have not been able to obtain protein structure data with bound substrates, which may be attributed to weak binding affinity between the protein and substrate, among other potential factors. To examine the proposed mechanisms of DONepi catalysis, molecular docking analysis was conducted to characterize the putative interaction between the enzyme and substrate. Twenty interaction models of DONepi and DON with variances in their energies were predicted, and the model with a reasonable binding orientation, a lower binding energy, and a larger number of hydrogen bonds was selected (Supplementary Table 5). DON, acting as a ligand, was located in the predicted active pocket of DONepi and interacted with three DONepi residues via hydrogen bonds. Specifically, Tyr 27 (Y27) bound to the O8 and C7-OH of DON, Arg 86 (R86) to the C12,13-epoxy group, and His 124 (H124) to the C3-OH (Fig. 3c). Sequence alignment indicated that the residues Y27 and R86 represent a very rare amino-acid-substitution variation. Only 0.8% and 0.5% of the sequences in the top 1000 DONepi homologs possess such substitutions at these two positions (Fig. 3d). Moreover, most of the substitutions with Y and R at these two positions are distributed among the homologs in the DONepi clade (Fig. 3d). Two homologs with these specific substitutions from Rhodococcus (WP283336260) and Nocardia (WP063010560) exhibited similar DON-epimerizing activity to DONepi, while the homologs with one substitution of Tyr or without these specific substitutions, from Pseudonocardia (WP269804706) and Cellulomonas (WP291810687), showed very weak epimerization activity towards DON (Fig. 3e and Supplementary Fig. 30). We further performed reciprocal mutations between the DONepi and its homologs lacking DON-epimerizing activity. Replacing residue Y27 in DONepi with the corresponding residue W, or replacing residue R86 with H and G resulted in a significant reduction of DON-epimerizing activity (Fig. 3e). Additionally, H124 is the key residue making up the typical catalytic tetrad of AKRs coupled with D51, Y56, and K83, universally conserved among species (Fig. 3d). The mutation of each of them in DONepi could lead to the loss of DON-epimerizing activity (Fig. 3e), suggesting that the catalytic tetrad plays a crucial role in forming the active center of DONepi.
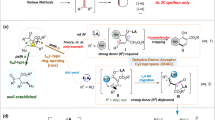
To investigate in depth the epimerization process, we analyzed the catalytic conversion of DON by DONepi using multiscale simulation methods, including molecular dynamics (MD), quantum mechanics/molecular mechanics (QM/MM) MD, and QM/MM well-tempered metadynamics (WT-MetaD). In a 600-ns MD simulation of the enzyme-ligand complex, it was observed that the C3 hydrogen atom of DON consistently remained close to the nicotinamide C4’ atom of the cofactor NADP+, with an average distance of approximately 2.5 Å (Supplementary Fig. 31). Additionally, the C3-OH group of DON formed a stable hydrogen bond with the phenolic hydroxyl group of the residue Y56 (H3···OTyr = 1.9 Å) (Supplementary Fig. 31). The proposed function of the conserved Y56 is to act as a general base in oxidation, assisting proton transfer and thereby promoting hydride transfer27,28. A snapshot of a dynamically equilibrated structure from the MD simulation was selected for subsequent QM/MM MD and WT-MetaD simulations to explore the catalytic mechanism in detail. The QM region included the entire DON molecule, part of the NADP+ molecule, and residue Y56, totaling 83 QM atoms. To describe the proton transfer (CV1) and hydride abstraction (CV2) involved in the oxidation process, two collective variables were defined: CV1 as the distance O3···H3 minus Y56O···H3, and CV2 as the distance C3···H minus C4···H (Fig. 4a). The evolution of the collective variables (CV1 and CV2) is shown in Supplementary Fig. 32. The results in Fig. 4b, c show that the reaction efficiently evolved from the initial DONepi-DON-NADP+ complex (denoted as MC) to the enzyme complex containing the 3-keto intermediate and NADPH (denoted as I). The free-energy minima of the MC and I states were nearly isoenergetic and separated by a unique intermediate M0 and a transition state TS. M0 is an intermediate formed after proton transfer from MC, with a free-energy difference of less than 2 kcal/mol between them, indicating that proton transfer occurs relatively easily. Subsequently, the hydride ion is transferred to the C4’ atom of NADP+ via the transition state TS, forming the key 3-keto intermediate (see the Supplementary Movie 1). These results suggest that C3-epimerization of DON needs the oxidation of DON to form 3-keto-DON firstly, which proceeds through a concerted but asynchronous mechanism, where the activation energy required for hydride transfer determines the overall catalytic step (MC → TS) with a free-energy barrier of 11.8 kcal/mol.
a Collective variables used in QM/MM metadynamics simulations of the oxidation of DON catalyzed by DONepi. b Free-energy landscape (FEL) obtained from the simulation (isolines at 2 kcal/mol). c Representative structures of stationary states along the reaction coordinate. d Definition of the rotational dihedral angle collective variable (CV3) in QM/MM MD simulations of the rotation of the 3-keto intermediate in the enzyme active site. e Representative structures during the rotation process. f Free-energy profile based on the dihedral angle distribution. MC, initial DONepi-DON-NADP+ complex. M0, intermediate formed after proton transfer from MC. TS, transition state. I, the enzyme complex containing the 3-keto intermediate and NADPH. INT, transition state during the rotation process. IROT, the enzyme complex containing the rotated 3-keto intermediate and NADPH.
Actually, DONepi can utilize 3-keto-DON as a substrate to form 3-epi-DON (Supplementary Fig. 33). However, 3-keto-DON was not detected during the transformation of DON to 3-epi-DON by DONepi, suggesting that it is a transient intermediate. To test whether 3-keto-DON is released from the enzyme active site during catalysis, hydroxylamine hydrochloride (NH2OH·HCl)—a strong nucleophilic reagent—was used as a competitive probe to target the carbonyl group of 3-keto-DON. We found that in the DONepi-catalyzed reaction system using 3-keto-DON as the substrate, hydroxylamine can readily outcompete NADPH, leading to almost complete oximation of free 3-keto-DON without enzymatic epimerization. In contrast, when hydroxylamine was added to the DONepi-catalyzed reaction system using DON as the substrate, the product detected was still 3-epi-DON, which is consistent with the reaction system without hydroxylamine (Supplementary Fig. 34). These data demonstrate that the intermediate 3-keto-DON is not released from the active site during the DON epimerization process catalyzed by DONepi. Furthermore, QM/MM MD simulations based on post-oxidation state conformation (denoted as I) revealed a rigid-body rotation of 3-keto-DON in the active site (see the Supplementary Movie 2). To simplify the description of this rotation, we defined a dihedral angle collective variable (CV3), constructed using the phenolic hydroxyl hydrogen atom of Tyr56, the carbonyl C3/O3 atoms, and the adjacent carbon C2 (Fig. 4d). The QM region included the 3-keto intermediate, the nicotinamide group of NADPH, and protonated Tyr56 (a total of 83 atoms). Three independent 5 ps unbiased QM/MM MD simulations consistently demonstrated that the 3-keto intermediate could spontaneously complete the rotation (Fig. 4e). The free-energy profile constructed from the CV3 distribution revealed a rotational energy barrier of ≤2.5 kcal/mol (Fig. 4f), confirming that this process is highly feasible under reaction conditions. Furthermore, this low-energy rotation provides the necessary conformational pre-organization for the subsequent NADPH-mediated reduction and epimerization, ensuring precise spatial orientation of the reactants. Overall, the octameric-structured DONepi catalyzes C3-epimerization of DON by directing the rotation of the transient 3-keto intermediate to enable stereoinverting reduction of the keto group by NADPH in the active site.
Additionally, we generated transgenic Arabidopsis thaliana lines harboring the codon-optimized DONepi sequence and evaluated their DON response via seedling growth assays on medium supplemented with 5, 10, or 20 μM DON (Supplementary Figs. 35 and 36). Root growth was comparable between transgenic and non-transgenic (wild-type) plants in DON-free medium. However, under all tested DON concentrations, transgenic lines exhibited longer roots and reduced leaf chlorosis (Supplementary Fig. 36). Specifically, at 20 μM DON, root lengths of lines DONepi-4 and DONepi-5 increased by 26% and 24%, respectively, relative to non-transgenic plants (Supplementary Fig. 36). These results demonstrate that DONepi confers enhanced DON tolerance to transgenic A. thaliana.
Stereochemical C8-reduction of DON catalyzed by DONrd
DONrd catalyzed the reduction of DON at the C8 position to produce a pair of epimers, α- and β-8-OH-DON. These epimers could be caused by stereochemical hydride attacks from NADPH on the re-face and si-face of the prochiral C8 carbonyl (Fig. 5a). To examine the proposed mechanisms of DONrd catalysis, molecular docking analysis was conducted using the predicted structure by AlphaFold229 (Supplementary Fig. 37). DONrd also has a typical (α/β)8-barrel motif similar to DONepi but different C-terminal tail loop (Fig. 5b). Docking analysis revealed that DON acting as a ligand could bind into the predicted active pocket of DONrd via two binding orientations (Fig. 5c and Supplementary Table 6). One orientation is conducive to hydride attack at the si-face, in which DON C7-OH and C8 carbonyl group interacted with DONrd residues Y59 and H125 via hydrogen bonds, respectively (Fig. 5d). Another orientation is conducive to hydride attack at the re-face, in which C8 carbonyl group, C12,13-epoxy group, and C15-OH of DON interacted with DONrd residues Y59, K129 and Y229 via hydrogen bonds, respectively (Fig. 5e). Among these, Y59 and H125 were the conserved residues making up the typical catalytic tetrad of AKRs coupled with D54, and K84 (Fig. 5f). The mutation of residue K84 in DONrd caused significant decrease in the reduction activity towards DON, and mutations in other three residues of the catalytic tetrad in DONrd could produce the loss of activity of DON reduction (Fig. 5g). Additionally, residues K129 and Y229 are also crucial for the reduction activity of DONrd. Replacing residue K129 in DONrd with residue H, the most frequently occurring amino acid in the corresponding alignment position of residue K129, resulted in a significant reduction of DON-reducing activity (Fig. 5f, g). Interestingly, residue Y229 is conserved in these DONrd homologs, and a mutation could produce the loss of DON reduction activity (Fig. 5f, g). These data indicated that the reaction of reducing DON at the C8 position catalyzed by DONrd is still dependent on the catalytic tetrad of D54-Y59-K84-H125, but residues K129 and Y229 play the key role in allowing the C8 carboxyl group to approach the catalytic tetrad in the appropriate orientation.
a The proposed catalytic mechanism of reducing DON at the C8 position to form a pair of epimers. Hydride attacks from NADPH at the si-face and re-face of the prochiral C8 carbonyl generate α-8-OH-DON and β-8-OH-DON, respectively. b The modeled three-dimensional structure of DONrd in complex with DON and NADPH. c The predicted active pocket of DONrd. The binding models of DON within DONrd at binding orientation for hydride attack at the si-face (d) and re-face (e). Dashed yellow lines indicate the hydrogen bonding interactions between DONrd and DON. f Variation and conservation in prospective activity sites among DONrd homologs. The phylogenetic tree was constructed using the maximum likelihood method with MEGA X. The numbers on top of the sequence alignment represent the positions of amino acids in DONrd. Sequence logos denote the relative frequency of each amino acid at that position, which was calculated using the top 1000 BLASTp hits. The red stars below indicate DON-binding residues in DONrd and conserved catalytic tetrad active sites. g DON-reducing activity of wild-type DONrd and DONrd mutants of prospective activity sites. Amino acid residues in the catalytic tetrad and Y229 were mutated into Alanine. K129 was mutated to Histidine based on (f). n = 3 biological replicates per group. Data are presented as mean ± SD, and analyzed by two-tailed t-test. ** indicate the significant difference at P < 0.01 compared with DONrd.
Moreover, using the C8-reduction products of DON as substrate for strain S5-5, we identified an intermediate metabolite designated M4 (Supplementary Fig. 38). The mass spectrum of TMSI-derivatized M4 showed a molecular weight of 674.5, an 88 Da increase compared to TMSI-derivatized 8-OH-DON (586.3). Since the displacement of a hydroxyl hydrogen with a trimethylsilyl (TMSI) group increases molecular mass by 72 Da, this indicates an additional oxygen atom (16 Da) was incorporated into 8-OH-DON, introducing an extra hydroxyl group in M4. A previous study found that a cytochrome P450 enzyme mediates a similar catalytic process, hydroxylating DON at C16 to form 16-hydroxy-DON17. Notably, M4 was completely consumed with extended cultivation (Supplementary Fig. 38). These observations suggest that a potential oxidase (e.g., cytochrome P450 enzyme), along with other putative enzymes, contributes to the further degradation of the 8-hydroxylated intermediate, facilitating the complete degradation of DON (Fig. 6).
This process is initiated by two distinct metabolic pathways, C3-epimerization and C8-reduction, which are catalyzed by two AKRs, DONepi and DONrd, respectively. The 8-hydroxyl intermediate produced by DONrd catalysis can be further degraded by a potential oxidase (e.g., cytochrome P450 enzyme) and other putative enzymes, until complete degradation occurs.
Discussion
Mycotoxin contaminants are impactful due to their effects on food and feed safety, as well as international trade. In this study, the successful isolation of the DON-completely degrading bacterial strain, Nocardioides sp. S5-5, applying a screening strategy based on toxicity assessment, underscores the advantage of this process for screening the toxin-degrading microbes. Currently, nearly all microbe screening with toxin degradation relies on the decrease of the toxin levels at the initial stage, resulting in a potential risk of toxin-derived metabolites. Because some transformation products of DON retain toxicological potential and sometimes form more toxic metabolites10. For most DON-degrading bacterial strains, 3-epi-DON is the end product, using 3-keto-DON as the intermediate following DON epimerization25,26. In contrast, 3-epi-DON acts as an intermediate during DON degradation by the strain S5-5 and can be further degraded by this strain, leading to the complete elimination of toxicity. Nocardioides strains appear to retain the capacity to destroy the skeletal structure of DON initiated by C3-epimerization17. Alternatively, the strain S5-5 simultaneously adopts two metabolic pathways to initiate the complete degradation of DON, including C3-epimerization and C8-reduction (Fig. 6).
The enzymatic epimerization of DON at C3 in most previous studies consists of two independent reactions catalyzed by two different enzymes25,26,30,31. Notably, we demonstrated that an AKR member, DONepi, could alone catalyze the C3-epimerization of DON to produce 3-epi-DON. Known DON-associated AKRs primarily perform two types of reactions: either reducing the intermediate 3-keto-DON at the C3 ketone group to form 3-epi-DON25,30,31, or oxidizing the C3-OH of DON to generate 3-keto-DON32. Separately, a recent study reported an AKR protein (AKR11A2) from Bacillus velezensis strain Vel-HNGD-F2 that exhibits DON epimerization activity33. Nevertheless, DONepi shares low amino acid sequence identity (less than 32%) with AKR11A2 (Supplementary Fig. 39). Cryo-EM indicates that DONepi has the typical (α/β)8-barrel motif of AKRs and adopts an octameric quaternary structure, but has a significantly different C-terminal tail loop. This difference may lead to distinct substrate specificities or catalytic efficiencies, as the orientation of the C-terminal tail loops can affect the accessibility and conformation of the active sites in AKRs27. Moreover, the genes responsible for two-step epimerization of DON have been proven to improve the DON tolerance and resistance to Fusarium head blight (FHB) in wheat34. Our results indicated that the single gene DONepi could significantly increase the DON tolerance of transgenic Arabidopsis thaliana, suggesting its potential in resistance improvement against FHB. Considering that the reaction catalyzed by DONepi produces an equilibrium mixture of DON and 3-epi-DON, the synergistic action of DONepi and other subsequent enzymes is indispensable to achieve the complete detoxification of DON. Our current results demonstrated a catalytic reaction on DON, C8-reduction, which was also performed by an AKR, DONrd. More importantly, DONrd can catalyze the further degradation of 3-epi-DON, which serves as a final product in most DON-degrading bacterial strains. We found that the 8-hydroxyl intermediate produced by DONrd catalysis can be further degraded by a potential oxidase (e.g., cytochrome P450 enzyme) and other putative enzymes, facilitating the complete degradation of DON. As a feasible strategy, all enzymes involved in the complete degradation pathway of DON can be co-expressed in engineered strains (e.g., yeast, E. coli), which will facilitate the development of an efficient enzyme reagent for the complete degradation of DON in future applications.
AKRs represent a class of critical oxidoreductases involved in the reversible reduction catalysis of ketone- and/or aldehyde-containing compounds to the corresponding alcohols, playing an essential role in metabolizing endogenous and exogenous toxicants35. However, little is known about the epimerization activity of these superfamily members. Our multiscale simulation results revealed the catalytic mechanism underlying this unique activity of DONepi, which involves the formation of a transient keto intermediate and its precise molecular rotation within the active site. This finding extends insights into the functional diversification of AKRs beyond their canonical redox roles. Notably, a conserved “transient keto intermediate” mechanism has been well characterized in the short-chain dehydrogenase/reductase (SDR)-type epimerases36,37,38. In these enzymes, the epimerization reaction initiates with hydride transfer to NAD(P)+, generating a transient keto intermediate. This intermediate then undergoes rotation within the active site, followed by hydride back-transfer from NAD(P)H to the carbonyl carbon, thereby completing the epimerization. The reduction of the generated keto intermediate is considerably faster than its formation, which effectively prevents its accumulation or release from the enzyme active site28,39,40. Our experimental and computational data show strong similarity to this conserved mechanism. Interestingly, significant amino acid sequence similarity is absent between DONepi and SDR-type epimerases; however, they exhibit convergent evolution in active site conformation. Specifically, the catalytically critical residues—Asp-Tyr-Lys-His in DONepi and Ser/Thr-Tyr-Lys in SDR-type epimerases—can be nearly superimposed in their 3D structures (Supplementary Fig. 40). This thus represents a striking example of convergent evolution between AKR and SDR-type epimerase families.
Horizontal gene transfer (HGT) is the transfer of genetic material between organisms without a parent-offspring relationship. Bacteria frequently obtain novel DNA via HGT, allowing them to adapt to changing environmental conditions and providing a competitive edge to persist in specific niches41,42. HGT is often mediated by various mobile genetic elements, such as genomic islands (GIs)43. GIs are commonly defined as clusters of consecutive genes of probable horizontal origin with lengths between 10 kb and 200 kb44. The functionalities conferred by GIs can be diverse, including antibiotic resistance, metal resistance, virulence, digestion of carbohydrates, and catabolism for bioremediation45,46,47,48,49. However, the roles of HGT regulated by GIs in mycotoxin degradation have not been explored. Current evidence indicates that the two specialized DON-degrading enzymes, DONepi and DONrd, were acquired by GIs-mediated HGT events. The homologous protein with the highest identity to DONepi was found in the genus Rhodococcus, and the homologs of DONepi in the genomes of two Rhodococcus strains were located in the predicted GIs, suggesting a probable HGT event involving gene transmission from the Rhodococcus strain to Nocardioides sp. S5-5. Interestingly, the population structure of the S5 consortium, from which strain S5-5 was purified, is composed of species from Rhodococcus, Nocardioides, and others. The severe mycotoxin contamination, resulting from the frequent outbreaks of Fusarium head blight (FHB) in the wheat fields where the consortium S5 was isolated, is likely to act as a selective pressure. This pressure drives the acquisition of toxin-degradation-related genes via HGT, thereby accelerating the adaptive evolution of microorganisms. Previous study revealed that a Fusarium-toxin detoxification gene associated with resistance to FHB, FHB7, encoding a glutathione S-transferase, was acquired via an HGT event from an endophytic fungus Epichloë to the host plant Thinopyrum elongatum6. Accordingly, these results underscore the roles that GI-mediated HGT has exerted in shaping microbial genomes, which advances the knowledge of mycotoxin-degrading gene evolution and offers a strategy for identifying mycotoxin-degrading genes.
In summary, by employing a combination of biochemical, molecular, Cryo-EM structural analyses and QM/MM metadynamics simulations, we have elucidated a pathway for the complete degradation of DON that is initiated by C3-epimerization and C8-reduction, and uncovered the catalytic mechanisms of two responsible enzymes, DONepi and DONrd, which were acquired via GIs-mediated horizontal gene transfer. The results not only deepen our understanding of the evolution of mycotoxin-degrading genes but also offer a strategy for identifying these genes. This, in turn, paves the way for the potential application of DON-degrading enzymes as a promising approach to control mycotoxin contamination both pre- and post-harvest, thereby ensuring food safety and security by effectively detoxifying DON.
Methods
Plants, soil, and chemicals
The aquatic plant Lemna minor was used for the toxicity assessment, which was grown in a half-strength Murashige and Skoog (1/2 MS) medium at 22 °C under a light regime of 16 h:8 h (light/dark). Arabidopsis thaliana (Col-0) was used for genetic transformation. Soil samples used for isolating DON-degrading bacteria were collected from wheat fields where Fusarium head blight is prevalent. Standard DON was purchased from Sigma-Aldrich (St. Louis, MO, USA). Stable carbon isotope (13C)-labelled DON was purchased from Alta Scientific Co., Ltd. (Tianjin, China) with catalog number 1ST7213C15-25A. 3-keto-DON and 3-epi-DON were prepared via biotransformation of DON, followed by isolation and purification using preparative high-performance liquid chromatography (preparative HPLC)32.
Isolation of DON-degrading bacteria based on toxicity assessment
Soil samples were suspended in the sterile deionized water (w/v, 1:10), and supernatants after sedimentation were inoculated to MM medium (v/v, 1:10) with DON (100 μg mL−1). Following incubation at 28 °C with shaking at 220 rpm for 7 days, these cultures were filtered through a 0.22 μm filter membrane and used for toxicity tests with L. minor bioassay in a 48-well plate following the methods previously described with a few modifications50. Sterilized culture filtrates starting from 100 μg mL−1 DON were diluted 10 times, resulting in an estimated final concentration of 10 μg mL−1 based on the amended DON-concentration. Three fronds of actively growing L. minor were placed in each well containing 0.8 mL diluted culture filtrates. The plants were incubated in triplicate for 6 days at 22 °C under a light regime of 16 h: 8 h (light/dark). The toxicity of DON-derived metabolites was analyzed based on the frond growth of L. minor. Culture samples with no significant inhibitory effects on L. minor growth were selected for isolating DON-degrading strains. The samples were treated with antibiotics to reduce microbial population diversity while maintaining community-level DON-degrading activity, and 16S rRNA gene sequencing was used to analyze the antibiotic-treated consortium for characterizing microbial population dynamics11,32.
Analysis of DON and its metabolites
High-performance liquid chromatography (HPLC) analysis of the levels of DON and its metabolized products in culture samples was performed using an Agilent 1260 Infinity Series LC (Agilent, Santa Clara, CA, USA). Chromatographic separation was achieved using an Agilent Zorbax Eclipse XDB-C18 column (4.6 × 150 mm, 5 μm) at 30 °C with methanol–water as mobile phases under gradient elution: initiation at 25% methanol; ramp to 75% over 15 min and then ramp to 80% over 5 min, hold for 3 min, then decrease back to 25% over 3 min, and hold for 3 min. The detection wavelength was 218 nm32.
Gas chromatography-mass spectrometry (GC-MS) analysis of trichothecene mycotoxins and their metabolites, derivatized by the mixture of trimethylsilyl imidazole/ trimethylchlorosilane (v/v, 100:1), was performed using an Agilent 7890A GC equipped with an Agilent 5975C MS detector, and a DB-5MS capillary column (30 m × 0.25 mm × 0.1 µm, J&W Scientific, Santa Clara, CA, USA). High-purity helium was used as the carrier gas with a flow rate of 1 mL/min. The GC oven temperature was initially held at 80 °C for 1 min, then increased linearly to 280 °C at a rate of 25 °C/min, and finally held at this temperature for 6 min. MS data were collected in full scan mode at mass range m/z 100–70025.
Ultrahigh-speed liquid phase tandem mass spectrometer (UPLC-HRMS/MS) analysis of putative DON metabolites was performed using a Thermo Scientific Accela 1250 UPLC (Thermo Fisher, Waltham, MA, USA) equipped with an AB SCIEX TripleTOF 6600 high-resolution mass spectrometer (SCIEX, Framingham, MA, USA). Chromatographic separation was achieved using a BEH Amide C18 column (2.1 × 100 mm, 1.7 μm) (Waters Co., Milford, MA, USA) at 30 °C with 5 mM ammonium acetate and methanol as mobile phases under gradient elution: initiation at 15% methanol, hold for 1 min; ramp to 90% methanol over 4.5 min, hold for 3 min, then decrease back to 15% over 1 min, hold for 1 min. The mass spectrometry acquisition signal mode was negative ion mode. The parameters of the mass spectrometry: apillary temperature, 320 °C; spray voltage, 3.00 kV; auxiliary gas heater temperature, 300 °C; and the full scan mass range was m/z 50–1000 Da51.
1H-NMR spectra of DON and its metabolite were obtained at 298 K using a Bruker Avance III 600 MHz spectrometer equipped with a Bruker inverse cryogenic probe (Bruker Biospin, Germany). The samples were dissolved in deuterated chloroform. 1H-NMR chemical shifts were measured with reference to the residual chloroform signal and referenced to sodium 3-trimethylsilyl [2,2,3,3-d4] propionate (TSP-d4)32.
Analysis of CO2 generation
Strain S5-5 was cultured in nutrient broth (NB). Cells were harvested when the OD600 value reached 0.6–0.8, washed three times with sterilized water, and then inoculated into 5 mL of MM medium in a 100 mL flask. The final cell concentration was adjusted to OD600 = 1, and the medium was supplemented with either 200 μM DON or a mixture of DON and 13C-labelled DON (molar ratio, 50:1). The flask was incubated at 28 °C. After 7 days of incubation, the residual DON in the medium was detected by HPLC as described above. Gas in the flask was collected. The concentration of CO₂ was analyzed using an Agilent 7890A GC equipped with a flame ionization detector (FID) and a HayeSep Q 80/100 UltiMetal column (12 ft L × 1/8″ OD × 2 mm ID, J&W Scientific, Santa Clara, CA, USA). High-purity nitrogen (N2) was used as the carrier gas. The FID temperature was set to 250 °C, with a hydrogen (H2) flow rate of 70 mL/min, a zero-grade air flow rate of 350 mL/min, and high-purity N2 as the makeup gas at 3.7 mL/min. The column oven temperature was maintained at 60 °C for 10 min. The δ13C value of 13CO2 was determined using a Trace Gas concentrator (Elementar Analysensysteme GmbH, Hanau, Germany) coupled to an isotopic ratio mass spectrometer (IRMS) (IsoPrime100, GV Instruments, Manchester, UK). The collected gas was manually injected into the Trace Gas concentrator, then transported to the IRMS via helium carrier gas at a flow rate of 200 mL min−1. Ionized CO2 species were quantified as ionic currents using a multicollector system equipped with three Faraday cups. Ionic current intensities corresponding to mass-to-charge ratios 46, 45, and 44 were utilized to calculate the isotopic ratio R(13C/12C). This ratio was calibrated against the VPDB scale using a CO2 reference gas and expressed in parts per thousand (‰): δ13C = {[R(13C/12C)sample / R(13C/12C)standard] - 1} × 1000.
Genome sequencing and BAC library construction
The genome of strain S5-5 was sequenced using a combination of Illumina and PacBio RS II single-molecule real-time (SMRT) sequencing platforms by Majorbio Biotech (Shanghai, China). The complete genome sequence was assembled using both the PacBio reads and Illumina reads, according to their standard procedures. The CDS prediction was achieved using Glimmer52. Functional annotation for the predicted protein-coding gene sequences was performed by alignment against public databases including COG, GO, KEGG, NR, and SwissProt, using BLASTp with an e-value of 10−5.
A bacterial artificial chromosome (BAC) library for strain S5-5 was constructed according to Luo and Wing53 with some modifications. The high-copy composite vector, pHZAUBACFXJ1, was used for the BAC library construction. The restriction enzyme, BamH I, was employed to linearize the vector and digest the genomic DNA isolated from strain S5-5. Individual BAC clones were arrayed in 384-well plates and stored at −80 °C. To screen the BAC clone with DON degradation activity, 1 mL overnight culture for each BAC clone was suspended in 0.2 mL MM medium containing DON (30 μg mL–1). After incubation for 5 days at 37 °C, metabolites were extracted and analyzed by HPLC and GC-MS as described above. The BAC plasmid DNA was isolated from positive clones and sequenced at both ends. BAC end sequences were used to search for the gene information in the insert fragment by alignment against the genome sequence of strain S5-5 with BLASTn.
Gene cloning, protein expression, and enzyme assays
The selected genes were amplified by PCR using the genomic DNA of strain S5-5 as a template. The PCR products were cloned into the expression vector pET28a (Novagen, Cat. No. 69337-3), and then transformed into Escherichia coli BL21 (DE3) for expression of the 6×His-tagged recombinant protein induced by isopropyl-β-D-thioga-lactopyranoside (IPTG). Recombinant proteins were purified by using Ni-NTA agarose (QIAGEN, Hilden, Germany) and analyzed by SDS-PAGE.
Enzymatic activity assays were performed in a 50 μL solution containing purified protein (6 μg), DON or its intermediate metabolites (100 μM), and either NADP⁺ (1 mM) or NADPH (0.25 mM), at different temperatures. The effect of pH on enzymatic activity was assessed at a constant temperature of 37 °C using buffers (50 mM) with varying pH values: sodium acetate buffer (pH 4.0–5.8), sodium phosphate buffer (pH 5.8–7.5), Tris-HCl buffer (pH 7.5–8.6), and glycine-NaOH buffer (pH 8.6–10.0). The effects of temperature on enzymatic activity were tested for DONepi and DONrd in sodium phosphate buffer at pH 7.5 and pH 6.0, respectively. Unless otherwise specified, the standard enzyme activity assays for DONepi and DONrd were performed in sodium phosphate buffer (pH 7.5) at 45 °C and sodium phosphate buffer (pH 6.0) at 25 °C, respectively. Reactions were terminated by adding an equal volume of methanol. Substrates and their metabolites in the reaction solutions were detected by HPLC and GC-MS as described above. Reaction velocity was measured based on the reduction of the substrate. Origin 8 (Origin Software, Northampton, MA, USA) was used to calculate the Michaelis–Menten constants.
Construction of AKRs SSN
The protein sequences of about 370,000 AKRs in the InterPro database were downloaded. The SSN was then constructed via the EFI-EST web tool54 (http://efi.igb.illinois.edu/efi-est/) using 28297 UniRef50 cluster ID sequences from these AKRs as the input. Networks were generated with default parameters under an alignment score threshold of 45. The resulting representative node networks were visualized with Cytoscape v.3.755.
Cryo-EM data collection, processing, and model building
The large-scale purification of DONepi was performed using nickel-affinity chromatography (Qiagen). The protein was further purified successively by using ion exchange chromatography (Source 15Q 10/100, GE Healthcare) and size exclusion chromatography (Superdex 200 increase 10/300GL, GE Healthcare) in buffer (25 mM Tris, pH 7.5, 150 mM NaCl) on the AKTA pure system. For cryo-EM sample preparation, 3.5 μL of 1.0 mg mL−1 DONepi protein was dropped onto glow-discharged 300 mesh holey carbon grids (Quantifoil Au R1.2/1.3), which were then blotted using vitrobot Mark IV (Thermo Fisher Scientific) with a blotting time of 3.5 s at 100% humidity and 8 °C, and rapidly plunged into the liquid nitrogen pre-cooled liquid ethane. The cryo-EM data for DONepi was collected on a Titan Krios electron microscope operated at 300 kV, and equipped with a GatanK3 Summit detector and a GIF Quantum energy filter. Images for the DONepi protein were recorded in counting mode at 81,000× magnification, yielding a pixel size of 1.07 Å. Each image underwent dose fractionation into 32 frames, accumulating a total electron dose of 50 e− Å−2 during an exposure time of 3.5 s. A summit direct electron detector, featuring a 20-eV slit width on its energy filter, was utilized. The defocus was preset within the range of −1.2 to −2.2 µm. For fully automated data collection, EPU software was employed. All images were motion-corrected by MotionCor256. A binning factor of 2 was applied along with concurrent dose weighting during this process. Subsequently, Gctf was utilized to estimate the defocus values57.
The cryo-EM dataset was processed in cryoSPARC (3.2)58. A total of 1,441,738 particles were automatically picked from 3392 manually selected micrographs. After 2D classification, 1,127,418 good particles were selected and subjected to multiple cycles of 3D classification. The 404,379 particles belonging to the best-scoring class were selected, followed by non-uniform and local refinement. The DONepi achieved a particle density with an estimated resolution of 2.56 Å according to the gold-standard Fourier shell correlation (FSC)59.
For the structure building of DONepi, the initial template was generated using AlphaFold. The initially fitted model was further refined through iterative model adjustments in COOT60. The resultant model was then refined against the map in real-space using PHENIX, with secondary structure and geometry restraints applied respectively61. Model quality was assessed via Molprobity scores and Ramachandran plots.
Computational molecular docking and site-directed mutagenesis
For molecular modeling and substrate docking, the three-dimensional structure of DONrd was predicted by the AlphaFold2 online server with default parameters29. The coenzyme molecules, NADP+ or NADPH, were incorporated into the predicted models using AlphaFill online server (https://alphafill.eu/)62. Protein active pocket detection was achieved using Proteins Plus online server (https://proteins.plus/). Autodock 4.263 was used to perform the docking of modeled proteins and substrates with the genetic algorithm approach. The size of the grid box was set to 60 × 60 × 60 xyz points. Computational molecular docking was shown by Pymol software (Version 1.7.0.0, http://www.pymol.org/pymol).
Site-directed mutagenesis for the interacting amino acids was achieved using the Mut Express II Fast Mutagenesis (Vazyme, Nanjing, China) with recombinant plasmids pET28a-DONepi or pET28a-DONrd as the PCR template. A list of primers used for site-directed mutagenesis is shown in Supplementary Table 7. The sequences of mutant variants were confirmed by sequencing. The site-directed mutants of DONepi and DONrd were expressed, purified, and assayed for DON degradation activity following the same protocols described above.
Molecular dynamics (MD) simulations
The initial complex structure was derived from the Autodocking structure of DONepi enzyme in complex with DON and NADP⁺. To accurately assign the protonation states of titratable residues, we employed the Propka364, analyzing the local environment of each residue under the assumption of a system pH of 7.6. Based on the experimental results and prior studies on related enzymes, the hydroxyl group of the residue Y56 was set to a deprotonated state. Subsequently, the enzyme-substrate complex was solvated in a cubic unit cell (dimensions: 100.168 × 100.095 × 99.880 ų) using the solvate and genion modules of GROMACS 2023.5. The system contained 26,679 TIP3P water molecules and 10 Na⁺ ions to neutralize the overall charge. For classical molecular dynamics (MD) simulations, the following force field parameter sets were used: FF14SB for the protein, GAFF2 for the ligand DON and cofactor NADP⁺, and TIP3P for water molecules. To further enhance the accuracy of charge distribution, partial atomic charges for DON and NADP⁺ were calculated using Gaussian16 at the B3LYP/6-311++G** level of theory and fitted via the RESP method.
The MD simulations were performed using GROMACS 2023.5, following the protocol outlined below. First, the entire enzyme complex system underwent energy minimization. Next, weak positional restraints were applied to the protein, substrate, and cofactor in the NVT ensemble, and the system was simulated for 400 ps at 298 K to equilibrate the temperature. Subsequently, the system entered the density equilibration phase, where it was simulated for an additional 500 ps in the NPT ensemble. A time step of 2 fs was used, and all bonds involving hydrogen atoms were constrained using the LINCS algorithm. During the heating and density equilibration stages, weak restraints were applied to key reaction distances to maintain the substrate in its reactive position. The production phase simulations were carried out in the NPT ensemble, consisting of three independent 200 ns MD runs. Restraints were gradually released over the first 10 ns to allow the system to relax naturally. Supplementary Fig. 31 shows the changes in backbone RMSD during the simulation, indicating that the system achieved good stability.
Quantum mechanics/molecular mechanics (QM/MM) MD simulations
To explore the reaction mechanism of the oxidation of the substrate DON to the key 3-keto intermediate, we performed a series of QM/MM (quantum mechanics/molecular mechanics) dynamic simulations using the CP2K 2024.1 program65. In the QM region, the dynamics of atoms depend on the electronic density ρ(r) calculated via density functional theory (DFT), while the dynamics of atoms in the MM region are governed by empirical force fields. In the QM/MM simulations, the QM part includes the amino acid residue Y56, the ligand DON, and the nicotinamide moiety and phosphate groups of NADP+, comprising a total of 83 atoms, enclosed in a cubic box with an edge length of 18.156 Å. The QM/MM boundaries (Cα and Cβ of NADP+ and Cα and Cβ of Tyr56) were treated using the link atom scheme, and the QM/MM interactions were computed using the GEEP scheme66. The QM part was described using DFT with the Perdew-Burke-Ernzerhof (PBE) functional67,68. All atoms were modeled using the DZVP-MOLOPT-GTH basis set, with a cutoff of 450 Ry retained for projecting the electronic structure. The parameters for the MM part were consistent with those used in classical MD simulations. Periodic boundary conditions were applied to both the QM and MM systems. Points of interest were extracted from the MD simulations, and geometries and velocities were retained to initiate QM/MM MD simulations in the NVT ensemble for 5 ps, using a Nose-Hoover thermostat with a time step of 0.5 fs.
QM/MM metadynamics simulations
To explore the free-energy landscape (FEL) of the substrate oxidation reaction, we employed the well-tempered metadynamics (WT-MetaD)69 method at the same theoretical level as the QM/MM MD calculations. Two collective variables (CVs) were used, facilitated by the Plumed2 plugin (version 2.9.0). Each CV was defined as the difference between two distances that need to be broken or formed during the reaction. The first CV measures the proton transfer between Y56 and the substrate 3-OH [CV1 = (O3···H3) - (OY56···H3)], while the second CV measures the interaction between the C3-H hydrogen atom of the substrate DON and the nicotinamide moiety of NADP+ [CV2 = (C3···H) - (C4···H)]. Thus, the second CV describes the hydrogen atom transfer between DON and nicotinamide. To enhance sampling in the CV space, a Gaussian kernel was added to the system every 50 MD steps (10 fs), with an initial height of 1.0 kcal mol−1 and a Gaussian width of 0.5 Å (corresponding to oscillations of CVs in unbiased dynamics, with a deposition interval of 10 fs). A bias factor γ = 15 was set to reduce the height of the generated Gaussian kernels. The time step for the QM/MM WT-MetaD simulation was adjusted to 0.2 fs, and the simulation was stopped upon meeting convergence criteria, with a total simulation time of 5 ps. Convergence was monitored using reweighting (debiasing) methods and block analysis, and errors in the free-energy estimates were calculated.
Construction and characterization of the transgenic Arabidopsis thaliana
The DONepi gene was codon-optimized for expression in Arabidopsis thaliana and synthesized (Supplementary Table 8). Subsequently, it was integrated into the vector pRI101-AN (Takara, Code No. 3262). The recombinant plasmid carrying the DONepi gene was transformed into the Agrobacterium tumefaciens strain GV3101, which was used to transform Arabidopsis thaliana through the floral dip method. Positive transgenic plants were selected on half-strength Murashige and Skoog (MS) medium supplemented with 50 mg L−1 kanamycin and further verified by PCR.
For the DON tolerance assay, seeds of Arabidopsis thaliana were surface-sterilized by using 10% sodium hypochlorite, followed by incubation on the half-strength MS medium. seedlings with uniform shoot lengths (2–3 mm) were transferred to fresh half-strength MS medium supplemented with DON at final concentrations of 5, 10, and 20 μM, respectively. Seedlings were then cultured under a 16 h light/8 h dark photoperiod at 22–24 °C. For each experimental line, at least 15 independent seedlings were used, and their root lengths were measured 21 days post-transfer.
Statistical analysis
All assays were performed in triplicate. The Student’s t-test (unpaired, two-tailed) was used to evaluate the statistical differences between groups using the GraphPad Prism software. P < 0.05 was considered statistically significant. Asterisks indicate P-values (*P < 0.05, **P < 0.01).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The DNA sequencing data are available at NCBI GenBank under the accession numbers CP199913 (genome), PX396128 (DONepi), and PX396129 (DONrd). The cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-66036. The atomic coordinates have been deposited in the Protein Data Bank (PDB) under accession codes 9WK1. Previously published PDB structural data are referenced as 1UDA, 1ZSX [https://doi.org/10.2210/pdb6ZLA/pdb], 6KV9, 6ZLA, 7UTF [https://doi.org/10.2210/pdb6ZLA/pdb], 8JWN [https://doi.org/10.2210/pdb6ZLA/pdb]. All data supporting the findings described in this manuscript are available in the article and in the Supplementary Information. Source data are provided with this paper.
References
Johns, L. E., Bebber, D. P., Gurr, S. J. & Brown, N. A. Emerging health threat and cost of Fusarium mycotoxins in European wheat. Nat. Food 3, 1014–1019 (2022).
Lee, H. J. & Ryu, D. Worldwide occurrence of mycotoxins in cereals and cereal-derived food products: public health perspectives of their co-occurrence. J. Agric. Food Chem. 65, 7034–7051 (2017).
Guo, H., Ji, J., Wang, J. S. & Sun, X. Deoxynivalenol: masked forms, fate during food processing, and potential biological remedies. Compr. Rev. Food Sci. Food Saf. 19, 895–926 (2020).
Chen, Y., Kistler, H. C. & Ma, Z. H. Fusarium graminearum trichothecene mycotoxins: biosynthesis, regulation, and management. Annu. Rev. Phytopathol. 57, 15–39 (2019).
Savary, S. et al. The global burden of pathogens and pests on major food crops. Nat. Ecol. Evol. 3, 430–439 (2019).
Wang, H. W. et al. Horizontal gene transfer of Fhb7 from fungus underlies Fusarium head blight resistance in wheat. Science 368, eaba5435 (2020).
Mishra, S., Srivastava, S., Dewangan, J., Divakar, A. & Kumar Rath, S. Global occurrence of deoxynivalenol in food commodities and exposure risk assessment in humans in the last decade: a survey. Crit. Rev. Food Sci. Nutr. 60, 1346–1374 (2020).
Loubresse, N. G. D., Holtkamp, W., Rodnina, M. V., Yusupova, G. & Yusupov, M. Structural basis for the inhibition of the eukaryotic ribosome. Nature 513, 517–522 (2014).
Tu, Y., Liu, S., Cai, P. & Shan, T. Global distribution, toxicity to humans and animals, biodegradation, and nutritional mitigation of deoxynivalenol: a review. Compr. Rev. Food Sci. Food Saf. 22, 3951–3983 (2023).
Adegoke, T. V. et al. Microbial enzymes involved in the biotransformation of major mycotoxins. J. Agric. Food Chem. 71, 35–51 (2022).
He, W. J. et al. Novel soil bacterium strain Desulfitobacterium sp. PGC-3-9 detoxifies trichothecene mycotoxins in wheat via de-epoxidation under aerobic and anaerobic conditions. Toxins 12, 363 (2020).
Hou, B. Q. et al. A novel strategy for detoxification of deoxynivalenol via modification of both toxic groups. Food Chem. 456, 139886 (2024).
Yang, J. et al. Enzymatic degradation of deoxynivalenol with the engineered detoxification enzyme Fhb7. JACS Au 4, 619–634 (2024).
Yang, J. et al. Developing Fhb7-derived enzymes with high thermostability for detoxification of T-2 toxin through ancestral sequence reconstruction. ACS Catal. 15, 11664–11672 (2025).
Shima, J. et al. Novel detoxification of the trichothecene mycotoxin deoxynivalenol by a soil bacterium isolated by enrichment culture. Appl. Environ. Microbiol. 63, 3825–3830 (1997).
Poppenberger, B. et al. Detoxification of the Fusarium mycotoxin deoxynivalenol by a UDP-glucosyltransferase from Arabidopsis thaliana. J. Biol. Chem. 278, 47905–47914 (2003).
Ikunaga, Y. et al. Nocardioides sp. strain WSN05-2, isolated from a wheat field, degrades deoxynivalenol, producing the novel intermediate 3-epi-deoxynivalenol. Appl. Microbiol. Biotechnol. 89, 419–427 (2011).
Ito, M. et al. Bacterial cytochrome P450 system catabolizing the Fusarium toxin deoxynivalenol. Appl. Environ. Microbiol. 79, 1619–1628 (2013). 03227-12.
Huang, J. Q. et al. Aromatization of natural products by a specialized detoxification enzyme. Nat. Chem. Biol. 16, 250–256 (2020).
Hu, Y. M. et al. Crystal structure and biochemical analysis of the specialized deoxynivalenol–detoxifying glyoxalase SPG from Gossypium hirsutum. Int. J. Biol. Macromol. 200, 388–396 (2022).
Lee, S. H. et al. Discovery to engineering of mycotoxin deoxynivalenol degrading enzymes based on the specialized glyoxalase I. Adv. Sci. 12, e02914 (2025).
Berthiller, F. et al. Masked mycotoxins: a review. Mol. Nutr. Food Res. 57, 165–186 (2013).
He, J. W. et al. Toxicology of 3-epi-deoxynivalenol, a deoxynivalenol-transformation product by Devosia mutans 17-2-E-8. Food Chem. Toxicol. 84, 250–259 (2015).
Li, X. Z., Hassan, Y. I., Lepp, D., Zhu, Y. & Zhou, T. 3-keto-DON, but not 3-epi-DON, retains the in planta toxicological potential after the enzymatic biotransformation of deoxynivalenol. Int. J. Mol. Sci. 23, 7230 (2022).
He, W. J. et al. A quinone-dependent dehydrogenase and two NADPH-dependent aldo/keto reductases detoxify deoxynivalenol in wheat via epimerization in a Devosia strain. Food Chem. 321, 126703 (2020).
Carere, J., Hassan, Y. I., Lepp, D. & Zhou, T. The enzymatic detoxification of the mycotoxin deoxynivalenol: identification of DepA from the DON epimerization pathway. Microb. Biotechnol. 11, 1106–1111 (2017).
Xiao, Z. et al. A three-level regulatory mechanism of the aldo-keto reductase subfamily AKR12D. Nat. Commun. 15, 2128 (2024).
Borg, A. J. E., Esquivias, O., Coines, J., Rovira, C. & Nidetzky, B. Enzymatic C4-epimerization of UDP-glucuronic acid: precisely steered rotation of a transient 4-keto intermediate for an inverted reaction without decarboxylation. Angew. Chem. Int. Ed. Engl. 62, e202211937 (2023).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Carere, J., Hassan, Y. I., Lepp, D. & Zhou, T. The identification of DepB: an enzyme responsible for the final detoxification step in the deoxynivalenol epimerization pathway in Devosia mutans 17-2-E-8. Front. Microbiol. 9, 1573 (2018).
Abraham, N. et al. Structure-function characterization of an aldo-keto reductase involved in detoxification of the mycotoxin, deoxynivalenol. Sci. Rep. 12, 14737 (2022).
He, W. J. et al. An aldo-keto reductase is responsible for Fusarium toxin-degrading activity in a soil Sphingomonas strain. Sci. Rep. 7, 9549 (2017).
Liu, J. X. et al. Expression, characterization, and application of an aldo-keto reductase mined from Bacillus velezensis Vel-HNGD-F2 for deoxynivalenol biodegradation. Food Chem. Toxicol. 196, 115159 (2025).
He, W. J. et al. Detoxifying bacterial genes for deoxynivalenol epimerization confer durable resistance to Fusarium head blight in wheat. Plant Biotechnol. J. 22, 2395–2409 (2024).
Penning, T. M. The aldo-keto reductases (AKRs): overview. Chem. Biol. Interact. 234, 236–246 (2015).
Kavanagh, K. L., Jörnvall, H., Persson, B. & Oppermann, U. The SDR superfamily: functional and structural diversity within a family of metabolic and regulatory enzymes. Cell. Mol. Life Sci. 65, 3895–3906 (2008).
Van Overtveldt, S. et al. A structural classification of carbohydrate epimerases: From mechanistic insights to practical applications. Biotechnol. Adv. 33, 1814–1828 (2015).
Fushinobu, S. Molecular evolution and functional divergence of UDP-hexose 4-epimerases. Curr. Opin. Chem. Biol. 61, 53–62 (2021).
Iacovino, L. G. et al. Crystallographic snapshots of UDP-glucuronic acid 4-epimerase ligand binding, rotation, and reduction. J. Biol. Chem. 295, 12461–12473 (2020).
Borg, A. J. E., Dennig, A., Weber, H. & Nidetzky, B. Mechanistic characterization of UDP-glucuronic acid 4-epimerase. FEBS J. 288, 1163–1178 (2021).
Soucy, S. M., Huang, J. & Gogarten, J. P. Horizontal gene transfer: building the web of life. Nat. Rev. Genet. 16, 472–482 (2015).
Brito, I. L. Examining horizontal gene transfer in microbial communities. Nat. Rev. Microbiol. 19, 442–453 (2021).
Bertelli, C. et al. IslandViewer 4: expanded prediction of genomic islands for larger-scale datasets. Nucleic Acids Res. 45, W30–W35 (2017).
Hacker, J. & Kaper, J. B. Pathogenicity islands and the evolution of microbes. Ann. Rev. Microbiol. 54, 641–679 (2000).
Hehemann, J. H. et al. Transfer of carbohydrate-active enzymes from marine bacteria to Japanese gut microbiota. Nature 464, 908–912 (2010).
Ikuma, K. & Gunsch, C. K. Genetic bioaugmentation as an effective method for in situ bioremediation: functionality of catabolic plasmids following conjugal transfers. Bioengineered 3, 236–241 (2012).
Navarro, C. A., Von Bernath, D. & Jerez, C. A. Heavy metal resistance strategies of ccidophilic bacteria and their acquisition: importance for biomining and bioremediation. Biol. Res. 46, 363–371 (2013).
Bohm, M. E., Huptas, C., Krey, V. M. & Scherer, S. Massive horizontal gene transfer, strictly vertical inheritance and ancient duplications differentially shape the evolution of Bacillus cereus enterotoxin operons hbl, cytK and nhe. BMC Evol. Biol. 15, 246 (2015).
Hemme, C. L. et al. Lateral gene transfer in a heavy metal-contaminated-groundwater microbial community. mBio 7, e02234–02215 (2016).
Vanhoutte, I. et al. Microbial detoxification of deoxynivalenol (DON), assessed via a Lemna minor L. bioassay, through biotransformation to 3-epi-DON and 3-epi-DOM-1. Toxins 9, 63 (2017).
Tian, Y. et al. Confrontation assays and mycotoxin treatment reveal antagonistic activities of Trichoderma and the fate of Fusarium mycotoxins in microbial interaction. Environ. Pollut. 267, 115559 (2020).
Delcher, A. L., Harmon, D., Kasif, S., White, O. & Salzberg, S. L. Improved microbial gene identification with GLIMMER. Nucleic Acids Res. 27, 4636–4641 (1999).
Luo, M. & Wing, R. A. An improved method for plant BAC library construction. Methods Mol. Biol. 236, 3 (2003).
Zallot, R., Oberg, N. & Gerlt, J. A. The EFI web resource for genomic enzymology tools: leveraging protein, genome, and metagenome databases to discover novel enzymes and metabolic pathways. Biochemistry 58, 4169–4182 (2019).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular Interaction networks. Genome Res. 13, 2498–2504 (2003).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Hekkelman, M. L., de Vries, I., Joosten, R. P. & Perrakis, A. AlphaFill: enriching AlphaFold models with ligands and cofactors. Nat. Methods 20, 205–213 (2023).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J. Comput. Chem. 30, 2785–2791 (2010).
Olsson, M. H. M., Søndergaard, C. R., Rostkowski, M. & Jensen, J. H. PROPKA3: consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory. Comput. 7, 525–537 (2011).
VandeVondele, J. et al. Quickstep: fast and accurate density functional calculations using a mixed Gaussian and plane waves approach. Comput. Phys. Commun. 167, 103–128 (2005).
Laino, T., Mohamed, F., Laio, A. & Parrinello, M. An efficient linear-scaling electrostatic coupling for treating periodic boundary conditions in QM/MM simulations. J. Chem. Theory Comput. 2, 1370 (2006).
Jerves, C., Neves, R. P. P., Ramos, M. J., da Silva, S. & Fernandes, P. A. Reaction mechanism of the PET degrading enzyme PETase studied with DFT/MM molecular dynamics simulations. ACS Catal. 11, 11626–11638 (2021).
Makurat, S., Neves, R. P. P., Ramos, M. J. & Rak, J. QM/MM MD study on the reaction mechanism of thymidine phosphorylation catalyzed by the enzyme Thermotoga Maritima thymidine kinase 1. ACS Catal. 14, 9840–9849 (2024).
Laio, A. & Parrinello, M. Escaping free-energy minima. Proc. Natl. Acad. Sci. USA 99, 12562–12566 (2002).
Acknowledgements
We thank Miss Danyang Li from the Core Facility of Wuhan University for her assistance with cryo-EM analysis. We thank Dr. Jianbo Cao and Dr. Zeyuan Guan from Huazhong Agricultural University for their assistance with TEM experiments and cryo-EM structure refinement. We also thank Huan He, Tingyan Zhang, and Shiwen Cao from the Public Laboratory Platform of the College of Resources and Environment, Huazhong Agricultural University, for their help with the experimental detection of CO2 and 13CO2. We acknowledge the technicians at Shenzhen HUASUAN Technology Co., Ltd. for their assistance with theoretical calculations (https://huasuankeji.com). This work was supported by the National Science Fund for Distinguished Young Scholars of China (32025030, A.W.), National Natural Science Foundation of China (32572412, W.H., 32272170, Y.K.L.), the Key Program of Action Plan to Revitalize Inner Mongolia through Science and Technology (2022EEDSKJXM011-3, J.Z.), and China Postdoctoral Science Foundation (2021M701348, W.H.).
Author information
Authors and Affiliations
Contributions
W.H., A.W., P.Y., J.Z., and Y.C.L. contributed to the conception and design of the work. W.H., M.Z., T.Z., Y.Z., C.Z., K.T., X.M., P.L., G.F., X.B., and Q.L. performed the microbial, molecular, and biochemical experiments. Q.W., R.X., W.H., and P.Y. contributed to the cryo-EM data collection and structure elucidation. W.H. and Y.T. performed the chromatographic and mass spectrometric analysis. W.H., T.Z., and C.Z. contributed to the plant study. W.H. performed bioinformatic analysis. T.H., Y.K.L., and W.Y. discussed results and provided advice. W.H. drafted the manuscript. W.H., A.W., P.Y., and J.Z. contributed to the revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
He, W., Xiong, R., Zheng, M. et al. Specialized aldo-keto reductases trigger complete degradation of mycotoxin deoxynivalenol. Nat Commun 17, 3240 (2026). https://doi.org/10.1038/s41467-026-70007-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70007-z