Abstract
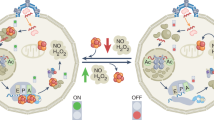
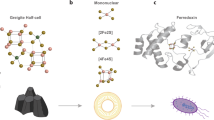
Pathogenic bacteria rely on the stringent response to adapt to hostile environments encountered within the host. However, the mechanisms by which host-induced stress activates this response remain poorly understood. Here, we identify iron-sulfur (Fe-S) cluster damage as a conserved trigger of the stringent response in major Gram-negative pathogens, including Salmonella enterica, Enterobacter cloacae, and Klebsiella pneumoniae. We demonstrate that Fe-S cluster disruption—triggered by oxidative stress or metal imbalance—limits intracellular pools of sulfur-containing and branched-chain amino acids, thereby activating the (p)ppGpp synthetase RelA. We further show that during Fe-S cluster stress, (p)ppGpp plays a dual role: enhancing bacterial fitness and promoting virulence by upregulating the Salmonella SPI-2 type III secretion system. These findings reveal a conserved mechanism by which pathogenic bacteria integrate host-associated stresses into an adaptive transcriptional response that promotes fitness and virulence, highlighting Fe-S cluster integrity as a central hub for environmental sensing during infection.
Similar content being viewed by others
Data availability
Raw and processed RNA-seq data have been deposited in the Gene Expression Omnibus (GEO) repository at the National Center for Biotechnology Information (NCBI) under accession number GSE305564. Data generated in this study are provided in the source data file. Biological materials are available upon request to the corresponding author. Source data are provided with this paper.
References
Fang, F. C., Frawley, E. R., Tapscott, T. & Vázquez-Torres, A. Bacterial stress responses during host infection. Cell Host Microbe 20, 133–143 (2016).
Liu, K., Bittner, A. N. & Wang, J. D. Diversity in (p)ppGpp metabolism and effectors. Curr. Opin. Microbiol. 24, 72–79 (2015).
Irving, S. E., Choudhury, N. R. & Corrigan, R. M. The stringent response and physiological roles of (pp)pGpp in bacteria. Nat. Rev. Microbiol. 19, 256–271 (2021).
Potrykus, K. & Cashel, M. p)ppGpp: still magical? Annu. Rev. Microbiol. 62, 35–51 (2008).
W. Steinchen, V. Zegarra, G. Bange, (p)ppGpp: Magic Modulators of Bacterial Physiology and Metabolism. Front. Microbiol. 11, 2072 (2020).
Dalebroux, Z. D. & Swanson, M. S. PpGpp: magic beyond RNA polymerase. Nat. Rev. Microbiol. 10, 203–212 (2012).
Ronneau, S. & Hallez, R. Make and break the alarmone: Regulation of (p)ppGpp synthetase/hydrolase enzymes in bacteria. FEMS Microbiol. Rev. 43, 389–400 (2019).
Irving, S. E. & Corrigan, R. M. Triggering the stringent response: Signals responsible for activating (p)ppGpp synthesis in bacteria. Microbiol. (U. Kingd.) 164, 268–276 (2018).
Battesti, A. & Bouveret, E. Acyl carrier protein/SpoT interaction, the switch linking SpoT-dependent stress response to fatty acid metabolism. Mol. Microbiol. 62, 1048–1063 (2006).
Battesti, A. & Bouveret, E. Bacteria possessing two RelA/SpoT-like proteins have evolved a specific stringent response involving The acyl carrier protein-SpoT interaction. J. Bacteriol. 191, 616–624 (2009).
Lee, J. W., Park, Y. H. & Seok, Y. J. Rsd balances (p)ppGpp level by stimulating the hydrolase activity of SpoT during carbon source downshift in Escherichia coli. Proc. Natl. Acad. Sci. USA. 115, E6845–E6854 (2018).
Haseltine, W. A. & Block, R. Synthesis of guanosine tetra and pentaphosphate requires the presence of a codon specific, uncharged transfer ribonucleic acid in the acceptor site of ribosomes. Proc. Natl. Acad. Sci. USA. 70, 1564–1568 (1973).
Vinella, D., Albrecht, C., Cashel, M. & D’Ari, R. Iron limitation induces SpoT-dependent accumulation of ppGpp in Escherichia coli. Mol. Microbiol. 56, 958–970 (2005).
L. F. Fitzsimmons, et al. SpoT induces intracellular Salmonella virulence programs in the phagosome. MBio 11, e03397-19 (2020).
Miethke, M., Westers, H., Blom, E. J., Kuipers, O. P. & Marahiel, M. A. Iron starvation triggers the stringent response and induces amino acid biosynthesis for bacillibactin production in Bacillus subtilis. J. Bacteriol. 188, 8655–8657 (2006).
C. Colomer-Winter, A. O. Gaca, J. A. Lemos, Association of metal homeostasis and (p)ppGpp regulation in the pathophysiology of Enterococcus faecalis. Infect. Immun. 85, e00260-17 (2017).
Kazmierczak, K. M., Wayne, K. J., Rechtsteiner, A. & Winkler, M. E. Roles of relSpn in stringent response, global regulation and virulence of serotype 2 Streptococcus pneumoniae D39. Mol. Microbiol. 72, 590–611 (2009).
Fritsch, V. N. et al. The alarmone (p)ppGpp confers tolerance to oxidative stress during the stationary phase by maintenance of redox and iron homeostasis in Staphylococcus aureus. Free Radic. Biol. Med. 161, 351–364 (2020).
Kröger, C. et al. An infection-relevant transcriptomic compendium for Salmonella enterica serovar typhimurium. Cell Host Microbe 14, 683–695 (2013).
Fitzsimmons, L. et al. Zinc-dependent substrate-level phosphorylation powers Salmonella growth under nitrosative stress of the innate host response. PLoS Pathog. 14, 1–25 (2018).
Frangipani, E., Slaveykova, V. I., Reimmann, C. & Haas, D. Adaptation of aerobically growing Pseudomonas aeruginosa to copper starvation. J. Bacteriol. 190, 6706–6717 (2008).
Harinarayanan, R., Murphy, H. & Cashel, M. Synthetic growth phenotypes of Escherichia coli lacking ppGpp and transketolase A (tktA) are due to ppGpp-mediated transcriptional regulation of tktB. Mol. Microbiol. 69, 882–894 (2008).
Sy, J. Reversibility of the pyrophosphoryl transfer from ATP to GTP by Escherichia coli stringent factor. Proc. Natl. Acad. Sci. Usa. 71, 3470–3473 (1974).
Fitzsimmons, L. F., Liu, L., Kim, J. S., Jones-Carson, J. & Vázquez-Torres, A. Salmonella reprograms nucleotide metabolism in its adaptation to nitrosative stress. MBio 9, 1–15 (2018).
Martin, J. E., Waters, L. S., Storz, G. & Imlay, J. A. The Escherichia coli small protein MntS and exporter MntP optimize the intracellular concentration of manganese. PLoS Genet 11, 1–31 (2015).
Kaur, G. et al. Affected energy metabolism under manganese stress governs cellular toxicity. Sci. Rep. 7, 1–11 (2017).
Pudek, M. R. & Bragg, P. D. Inhibition by cyanide of the respiratory chain oxidases of Escherichia coli. Arch. Biochem. Biophys. 164, 682–693 (1974).
M. Unciuleac, et al. In vitro activation of apo-aconitase using a [4Fe-4S] cluster-loaded form of the IscU [Fe - S] cluster scaffolding protein. 46, 6812–6821 (2007).
Takahashi, Y. & Tokumoto, U. A third bacterial system for the assembly of iron-sulfur clusters with homologs in Archaea and plastids. J. Biol. Chem. 277, 28380–28383 (2002).
Hyduke, D. R., Jarboe, L. R., Tran, L. M., Chou, K. J. Y. & Liao, J. C. Integrated network analysis identifies nitric oxide response networks and dihydroxyacid dehydratase as a crucial target in Escherichia coli. Proc. Natl. Acad. Sci. USA. 104, 8484–8489 (2007).
Jang, S. & Imlay, J. A. Micromolar intracellular hydrogen peroxide disrupts metabolism by damaging iron-sulfur enzymes. J. Biol. Chem. 282, 929–937 (2007).
Richardson, A. R. et al. Multiple targets of nitric oxide in the tricarboxylic acid cycle of Salmonella enterica serovar Typhimurium. Cell Host Microbe 10, 33–43 (2011).
Py, B. & Barras, F. Building Fe-S proteins: Bacterial strategies. Nat. Rev. Microbiol. 8, 436–446 (2010).
Roche, B. et al. Iron/sulfur proteins biogenesis in prokaryotes: formation, regulation and diversity. Biochim. Biophys. Acta - Bioenerg. 1827, 455–469 (2013).
Ding, H., Harrison, K. & Lu, J. Thioredoxin reductase system mediates iron binding in IscA and iron delivery for the iron-sulfur cluster assembly in IscU. J. Biol. Chem. 280, 30432–30437 (2005).
Ding, H., Clark, R. J. & Ding, B. IscA mediates iron delivery for assembly of iron-sulfur clusters in IscU under the limited accessible free iron conditions. J. Biol. Chem. 279, 37499–37504 (2004).
Zorin, N. A., Zabelin, A. A., Shkuropatov, A. Y. & Tsygankov, A. A. Interaction of HydSL hydrogenase from Thiocapsa roseopersicina with cyanide leads to destruction of iron-sulfur clusters. J. Inorg. Biochem. 177, 190–197 (2017).
Tosa, T. & Pizer, L. I. Biochemical bases for the antimetabolite action of L-serine hydroxamate. J. Bacteriol. 106, 972–982 (1971).
Imlay, J. A. The molecular mechanisms and physiological consequences of oxidative stress: Lessons from a model bacterium. Nat. Rev. Microbiol. 11, 443–454 (2013).
Imlay, J. A. Pathways of oxidative damage. Annu. Rev. Microbiol. 57, 395–418 (2003).
Kim, J. S. et al. DksA–DnaJ redox interactions provide a signal for the activation of bacterial RNA polymerase. Proc. Natl. Acad. Sci. USA. 115, E11780–E11789 (2018).
Burton, N. A. et al. Disparate impact of oxidative host defenses determines the fate of Salmonella during systemic infection in mice. Cell Host Microbe 15, 72–83 (2014).
Waters, L. S., Sandoval, M. & Storz, G. The Escherichia coli MntR miniregulon includes genes encoding a small protein and an efflux pump required for manganese homeostasis. J. Bacteriol. 193, 5887–5897 (2011).
Mettert, E. L. & Kiley, P. J. Fe-S cluster homeostasis and beyond: the multifaceted roles of IscR. Biochim. Biophys. Acta - Mol. Cell Res 1871, 119749 (2024).
Rice, C. J., Ramachandran, V. K., Shearer, N. & Thompson, A. Transcriptional and post-transcriptional modulation of SPI1 and SPI2 expression by ppGpp, RpoS and DksA in Salmonella enterica sv Typhimurium. PLoS One 10, e0127523 (2015).
Tapscott, T. et al. Guanosine tetraphosphate relieves the negative regulation of Salmonella pathogenicity island-2 gene transcription exerted by the AT-rich ssrA discriminator region. Sci. Rep. 8, 1–12 (2018).
Thompson, A. et al. The bacterial signal molecule, ppGpp, mediates the environmental regulation of both the invasion and intracellular virulence gene programs of Salmonella. J. Biol. Chem. 281, 30112–30121 (2006).
Jennings, E., Thurston, T. L. M. & Holden, D. W. Salmonella SPI-2 type iii secretion system effectors: molecular mechanisms and physiological consequences. Cell Host Microbe 22, 217–231 (2017).
Haraga, A., Ohlson, M. B. & Miller, S. I. Salmonellae interplay with host cells. Nat. Rev. Microbiol. 6, 53–66 (2008).
Henard, C. A., Bourret, T. J., Song, M. & Vázquez-Torres, A. Control of redox balance by the stringent response regulatory protein promotes antioxidant defenses of Salmonella. J. Biol. Chem. 285, 36785–36793 (2010).
Kim, H. Y., Go, J., Lee, K. M., Oh, Y. T. & Yoon, S. S. Guanosine tetra- and pentaphosphate increase antibiotic tolerance by reducing reactive oxygen species production in Vibrio cholerae. J. Biol. Chem. 293, 5679–5694 (2018).
Keasling, J. D., Bertsch, L. & Kornberg, A. Guanosine pentaphosphate phosphohydrolase of Escherichia coli is a long-chain exopolyphosphatase. Proc. Natl. Acad. Sci. USA. 90, 7029–7033 (1993).
Sinha, A. K., Winther, K. S., Roghanian, M. & Gerdes, K. Fatty acid starvation activates RelA by depleting lysine precursor pyruvate. Mol. Microbiol. 112, 1339–1349 (2019).
Velayudhan, J., Castor, M., Richardson, A., Main-hester, K. L. & Fang, F. C. The role of ferritins in the physiology of Salmonella enterica sv. Typhimurium: a unique role for ferritin B in iron-sulphur cluster repair and virulence. Mol Microbiol. 63, 1495–1507 (2007).
Yang, S. et al. Salmonella effector SpvB interferes with intracellular iron homeostasis via regulation of transcription factor NRF2. FASEB J. 33, 13450–13464 (2019).
Lazarus, J. E. et al. A new suite of allelic-exchange vectors for the scarless modification of proteobacterial genomes. Appl. Environ. Microbiol. 85, e00990–19 (2019).
Acknowledgements
We thank all members of the Hallez lab, the BRM unit and Benjamin Ezraty for fruitful scientific discussions. We are also grateful to Jean-François Collet, Hélène Andrews-Polymenis, Michael McClelland, and Athanasios Typas for providing the Salmonella Single Gene Deletion library. We also thank Sophie Helaine for her critical reading of the manuscript. This work was supported by the Marie Sklodowska Curie COFUND action (No 101034383) and the ATIP-Avenir program to S.R., the CARE ANR grant (22-PAMR-0002) to C.M., and the Welbio Starting Grant (WELBIO-CR-2019S-05) to R.H. R.H. is a senior research associates of the F.R.S. – FNRS.
Author information
Authors and Affiliations
Contributions
S.R. conceived and designed the experiments. E.M., L.R., C.M., and S.R. performed the experiments. C.L., L.B. and C.M. analyzed the RNA-seq data. C.M., V.C. and R.H. provided intellectual input and contributed reagents, materials, and analysis tools. S.R. wrote the manuscript with input from all the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Michaud, E., Ricci, L., Lallement, C. et al. Stress-induced iron-sulfur cluster damage as a conserved trigger of the bacterial stringent response. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70079-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70079-x