Abstract
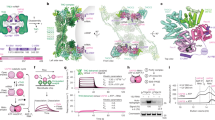
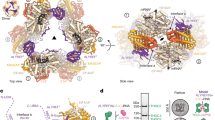
The nuclear export of mRNA represents a critical regulatory node in eukaryotic gene expression. This process is orchestrated by two conserved multi-subunit assemblies: the transcription-and-export complex (TREX) and TREX-2. While TREX facilitates mRNP packaging through multivalent RNA-protein interactions, the precise mechanism by which TREX-2 contributes to mRNA export has remained elusive. Here, we report a functional interaction between UAP56 and TREX-2 and resolve the structures of TREX-2 in both apo and UAP56-bound states. UAP56 engages TREX-2 via its N-terminal region, positioning its RecA domains on the V-shaped surface of the complex. A conserved loop from TREX-2 inserts between the RecA domains of UAP56, stabilizing an open conformation. Biochemical assays demonstrate that TREX-2 significantly stimulates the ATPase activity of UAP56, thereby promoting RNA release. These findings provide structural and mechanistic insights into TREX-2-mediated regulation of mRNA export through UAP56 remodeling.
Similar content being viewed by others
Data availability
The atomic coordinates for human TREX-2M and UAP56-TREX-2M have been deposited in the Protein Data Bank (https://www.rcsb.org/search) under the accession codes 9UPC and 9UPB, respectively. The cryo-EM maps for TREX-2M and UAP56-TREX-2M have been deposited in the EMDB with accession codes EMD-64391 and EMD-64390, respectively. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (https://proteomecentral.proteomexchange.org) via the iProX partner repository74 with the dataset identifier PXD068408. Source data are provided with this paper.
References
Köhler, A. & Hurt, E. Exporting RNA from the nucleus to the cytoplasm. Nat. Rev. Mol. Cell Biol. 8, 761–773 (2007).
Wickramasinghe, V. O. & Laskey, R. A. Control of mammalian gene expression by selective mRNA export. Nat. Rev. Mol. Cell Biol. 16, 431–442 (2015).
Singh, G., Pratt, G., Yeo, G. W. & Moore, M. J. The clothes make the mRNA: past and present trends in mrnp fashion. Annu. Rev. Biochem. 84, 325–354 (2015).
Khong, A. & Parker, R. The landscape of eukaryotic mRNPs. RNA 26, 229–239 (2020).
Chen, S., Jiang, Q., Fan, J. & Cheng, H. Nuclear mRNA export. Acta Biochim Biophys Sin (Shanghai), (2024).
Muller-McNicoll, M. & Neugebauer, K. M. How cells get the message: dynamic assembly and function of mRNA-protein complexes. Nat. Rev. Genet 14, 275–287 (2013).
Cheng, H. et al. Human mRNA export machinery recruited to the 5’ end of mRNA. Cell 127, 1389–1400 (2006).
Oeffinger, M. & Montpetit, B. Emerging properties of nuclear RNP biogenesis and export. Curr. Opin. Cell Biol. 34, 46–53 (2015).
Xie, Y. & Ren, Y. Mechanisms of nuclear mRNA export: A structural perspective. Traffic 20, 829–840 (2019).
Luo, M. L. et al. Pre-mRNA splicing and mRNA export linked by direct interactions between UAP56 and Aly. Nature 413, 644–647 (2001).
Le Hir, H., Gatfield, D., Izaurralde, E. & Moore, M. J. The exon-exon junction complex provides a binding platform for factors involved in mRNA export and nonsense-mediated mRNA decay. EMBO J. 20, 4987–4997 (2001).
Strasser, K. & Hurt, E. Splicing factor Sub2p is required for nuclear mRNA export through its interaction with Yra1p. Nature 413, 648–652 (2001).
Knockenhauer, K. E. & Schwartz, T. U. The nuclear pore complex as a flexible and dynamic gate. Cell 164, 1162–1171 (2016).
Reed, R. & Cheng, H. TREX, SR proteins and export of mRNA. Curr. Opin. Cell Biol. 17, 269–273 (2005).
Fischer, T. et al. The mRNA export machinery requires the novel Sac3p-Thp1p complex to dock at the nucleoplasmic entrance of the nuclear pores. EMBO J. 21, 5843–5852 (2002).
Strasser, K. et al. TREX is a conserved complex coupling transcription with messenger RNA export. Nature 417, 304–308 (2002).
Gonzalez-Aguilera, C. et al. The THP1-SAC3-SUS1-CDC31 complex works in transcription elongation-mRNA export preventing RNA-mediated genome instability. Mol. Biol. Cell 19, 4310–4318 (2008).
Masuda, S. et al. Recruitment of the human TREX complex to mRNA during splicing. Genes Dev. 19, 1512–1517 (2005).
Fischer, T. et al. Yeast centrin Cdc31 is linked to the nuclear mRNA export machinery. Nat. Cell Biol. 6, 840–848 (2004).
Jimeno, S., Rondon, A. G., Luna, R. & Aguilera, A. The yeast THO complex and mRNA export factors link RNA metabolism with transcription and genome instability. EMBO J. 21, 3526–3535 (2002).
Pacheco-Fiallos, B. et al. mRNA recognition and packaging by the human transcription-export complex. Nature 616, 828–835 (2023).
Bonneau, F. et al. Nuclear mRNPs are compact particles packaged with a network of proteins promoting RNA-RNA interactions. Genes Dev. 37, 505–517 (2023).
Jones, A. L. et al. SAC3 may link nuclear protein export to cell cycle progression. Proc. Natl. Acad. Sci. USA 97, 3224–3229 (2000).
Gallardo, M., Luna, R., Erdjument-Bromage, H., Tempst, P. & Aguilera, A. Nab2p and the Thp1p-Sac3p complex functionally interact at the interface between transcription and mRNA metabolism. J. Biol. Chem. 278, 24225–24232 (2003).
Lei, E. P. et al. Sac3 is an mRNA export factor that localizes to cytoplasmic fibrils of nuclear pore complex. Mol. Biol. Cell 14, 836–847 (2003).
Wickramasinghe, V. O. et al. mRNA export from mammalian cell nuclei is dependent on GANP. Curr. Biol. 20, 25–31 (2010).
Wickramasinghe, V. O., Stewart, M. & Laskey, R. A. GANP enhances the efficiency of mRNA nuclear export in mammalian cells. Nucleus 1, 393–396 (2010).
Umlauf, D. et al. The human TREX-2 complex is stably associated with the nuclear pore basket. J. Cell Sci. 126, 2656–2667 (2013).
Herold, A., Klymenko, T. & Izaurralde, E. NXF1/p15 heterodimers are essential for mRNA nuclear export in Drosophila. RNA 7, 1768–1780 (2001).
Moore, M. J. & Rosbash, M. Cell biology. TAPping into mRNA export. Science 294, 1841–1842 (2001).
Katahira, J. et al. The Mex67p-mediated nuclear mRNA export pathway is conserved from yeast to human. EMBO J. 18, 2593–2609 (1999).
Bachi, A. et al. The C-terminal domain of TAP interacts with the nuclear pore complex and promotes export of specific CTE-bearing RNA substrates. RNA 6, 136–158 (2000).
Hautbergue, G. M., Hung, M. L., Golovanov, A. P., Lian, L. Y. & Wilson, S. A. Mutually exclusive interactions drive handover of mRNA from export adaptors to TAP. Proc. Natl. Acad. Sci. USA 105, 5154–5159 (2008).
Huertas, P. & Aguilera, A. Cotranscriptionally formed DNA:RNA hybrids mediate transcription elongation impairment and transcription-associated recombination. Mol. Cell 12, 711–721 (2003).
Bhatia, V. et al. BRCA2 prevents R-loop accumulation and associates with TREX-2 mRNA export factor PCID2. Nature 511, 362–365 (2014).
Zhang, K. et al. Structural basis for influenza virus NS1 protein block of mRNA nuclear export. Nat. Microbiol 4, 1671–1679 (2019).
Ren, Y., Schmiege, P. & Blobel, G. Structural and biochemical analyses of the DEAD-box ATPase Sub2 in association with THO or Yra1. Elife 6, 10.7554/eLife.20070 (2017).
Puhringer, T. et al. Structure of the human core transcription-export complex reveals a hub for multivalent interactions. Elife 9, 10.7554/eLife.61503 (2020).
Schuller, S. K. et al. Structural insights into the nucleic acid remodeling mechanisms of the yeast THO-Sub2 complex. Elife 9, 10.7554/eLife.61467 (2020).
Xie, Y. et al. Cryo-EM structure of the yeast TREX complex and coordination with the SR-like protein Gbp2. Elife 10, 10.7554/eLife.61467 (2021).
Faraway, R., Zenklusen, D. & Plaschka, C. Mechanisms of messenger RNA packaging and export. Annu. Rev. Cell Developmental Biol. 41, 479–504 (2025).
Stewart, M. From transcription to export: mRNA’s winding path to the cytoplasm. Trends Biochemical Sci. 50, 748–765 (2025).
Dimitrova, L. et al. Structural characterization of the chaetomium thermophilum trex-2 complex and its interaction with the mrna nuclear export factor mex67:mtr2. Structure 23, 1246–1257 (2015).
Vdovina, Y. A., Georgieva, S. G. & Kopytova, D. V. Interaction of mRNA with the C-Terminal Domain of PCID2, a Subunit of the TREX-2 complex, is required for its export from the nucleus to the cytoplasm in Drosophila melanogaster. Dokl. Biochem Biophys. 513, 328–331 (2023).
Valkov, E., Dean, J. C., Jani, D., Kuhlmann, S. I. & Stewart, M. Structural basis for the assembly and disassembly of mRNA nuclear export complexes. Biochim Biophys. Acta 1819, 578–592 (2012).
Garcia-Oliver, E., Garcia-Molinero, V. & Rodriguez-Navarro, S. mRNA export and gene expression: the SAGA-TREX-2 connection. Biochim Biophys. Acta 1819, 555–565 (2012).
Jani, D. et al. Functional and structural characterization of the mammalian TREX-2 complex that links transcription with nuclear messenger RNA export. Nucleic Acids Res 40, 4562–4573 (2012).
Ellisdon, A. M., Dimitrova, L., Hurt, E. & Stewart, M. Structural basis for the assembly and nucleic acid binding of the TREX-2 transcription-export complex. Nat. Struct. Mol. Biol. 19, 328–336 (2012).
Kuang, Z., Ke, J., Hong, J., Zhu, Z. & Niu, L. Structural assembly of the nucleic-acid-binding Thp3-Csn12-Sem1 complex functioning in mRNA splicing. Nucleic Acids Res 50, 8882–8897 (2022).
Xie, Y. et al. Structures and mRNP remodeling mechanism of the TREX-2 complex. Structure 33, 566–582.e6 (2025).
Schneider, M. et al. The Nuclear Pore-Associated TREX-2 Complex Employs Mediator to Regulate Gene Expression. Cell 162, 1016–1028 (2015).
Aibara, S., Bai, X. C. & Stewart, M. The Sac3 TPR-like region in the Saccharomyces cerevisiae TREX-2 complex is more extensive but independent of the CID region. J. Struct. Biol. 195, 316–324 (2016).
Gordon, J. M. B., Aibara, S. & Stewart, M. Structure of the Sac3 RNA-binding M-region in the Saccharomyces cerevisiae TREX-2 complex. Nucleic Acids Res 45, 5577–5585 (2017).
Hohmann, U. et al. An ATP-gated molecular switch orchestrates human mRNA export. Nature, https://doi.org/10.1038/s41586-025-09832-z (2025).
Clarke, B. P. et al. Structural mechanism of DDX39B regulation by human TREX-2 and a related complex in mRNP remodeling. Nat. Commun. 16, https://doi.org/10.1038/s41467-025-60547-1 (2025).
Wickramasinghe, V. O. et al. Selective nuclear export of specific classes of mRNA from mammalian nuclei is promoted by GANP. Nucleic Acids Res 42, 5059–5071 (2014).
Fujita, K. I. et al. Structural differences between the closely related RNA helicases, UAP56 and URH49, fashion distinct functional apo-complexes. Nat. Commun. 15, 455 (2024).
Saguez, C. et al. Mutational analysis of the yeast RNA helicase Sub2p reveals conserved domains required for growth, mRNA export, and genomic stability. RNA 19, 1363–1371 (2013).
Wilmes, G. M. et al. A genetic interaction map of RNA-processing factors reveals links between Sem1/Dss1-containing complexes and mRNA export and splicing. Mol. Cell 32, 735–746 (2008).
Xie, Y. et al. Structural basis for high-order complex of SARNP and DDX39B to facilitate mRNP assembly. Cell Rep. 42, 112988 (2023).
Stewart, M. Structure and Function of the TREX-2 Complex. Subcell. Biochem 93, 461–470 (2019).
Dufu, K. et al. ATP is required for interactions between UAP56 and two conserved mRNA export proteins, Aly and CIP29, to assemble the TREX complex. Genes Dev. 24, 2043–2053 (2010).
Lusong, et al. LENG8 mediates RNA nuclear retention and degradation in eukaryotes. bioRxiv https://doi.org/10.1101/2025.08.14.670437 (2025).
Bugai, A. et al. Molecular basis of polyadenylated RNA fate determination in the nucleus. bioRxiv https://doi.org/10.4161/cc.9.17.12792 (2025).
Yang, F. et al. Mechanisms of the RNA helicases DDX42 and DDX46 in human U2 snRNP assembly. Nat. Commun. 14, 897 (2023).
Zhang, X. et al. Structural insights into branch site proofreading by human spliceosome. Nat. Struct. Mol. Biol. 31, 835–845 (2024).
Zhang, X. et al. Structural basis of pre-tRNA intron removal by human tRNA splicing endonuclease. Mol. Cell, https://doi.org/10.1016/j.molcel.2023.03.015 (2023).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Chen, S. X. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290 (2017).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. Sect. D.-Struct. Biol. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. Sect. D.-Struct. Biol. 66, 213–221 (2010).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Chen, T. et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522–D1527 (2022).
Acknowledgements
We thank the Cryo-EM Center at the University of Science and Technology of China for facility access and technical assistance. We also acknowledge the Core Facility Center for Life Sciences for facility access. This work was supported by funds from the National Natural Science Foundation of China (32471302 to X.Z.), the Natural Science Foundation of Anhui Province (2408085MC068 to X.Z.), the Research Funds of the Center for Advanced Interdisciplinary Science and Biomedicine of IHM (QYPY20230023 to X.Z.), the USTC Research Funds of the Double First-Class Initiative (YD9100002059 to X.Z.), and the Major Program of Shenzhen Bay Laboratory (C1012523007).
Author information
Authors and Affiliations
Contributions
X.G., R.T., H.Z., M.L., and Y.C. designed and performed all the experiments. X.G. and Y.G. prepared cryo-EM samples and collected the cryo-EM data. Xiaofei G., X.Z., and J.H. processed the cryo-EM data, calculated the cryo-EM map, and built the atomic model. All authors contributed to data analysis. X.Z. supervised the project and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Eugene Valkov and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gong, X., Tao, R., Ge, X. et al. Molecular insights into mRNA export regulation by the human TREX-2 complex. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70088-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70088-w