Abstract
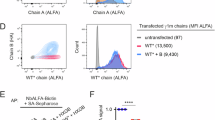
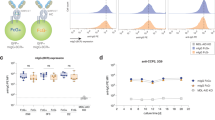
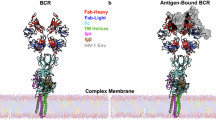
Although N-linked glycans play pivotal roles in the regulation of antibody effector functions, little is known about the composition and functional impact of glycans attached to human B cell receptors (BCR). Here, we describe the site-specific glycosylation profiles of all 4 N-linked glycosylation sites of human Igµ BCRs from primary naive and memory B cells. Our data indicate that the Igµ BCR glycans do not undergo structural changes during transition from naive to memory cells, as healthy donor-derived BCRs from both B cell subtypes carry similar glycans. Conversely, serum IgM antibodies display distinct glycosylation features to Igµ BCRs, which are in part explained by the different molecular form of Igµ BCRs and IgM antibodies. Moreover, using B cell lines expressing defined BCRs, we show that, although the absence of the Igµ BCR N209 glycans reduces antigen binding, the individual glycosylation sites are nonessential for cell surface expression and BCR internalization following BCR triggering. Collectively, we show high conservation of Igµ BCR glycosylation across B cell subsets and limited contribution to BCR expression and function, suggesting BCR glycans may have evolved to support IgM antibody functions.
Similar content being viewed by others
Data availability
The minimum dataset, including all experimental values and images are available in the source data file. Processed LC-MS tables and statistic tables are available in Supplementary Data 1–5. available in the source data file. The raw LC-MS proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE36 partner repository with the dataset identifier PXD062559 https://www.ebi.ac.uk/pride/archive/projects/PXD062559. The original Cryo-EM Igµ BCR structure file can be found on the protein data bank with code 7XQ8. https://doi.org/10.2210/pdb7XQ8/pdb Source data are provided in this paper.
References
Lu, L. L., Suscovich, T. J., Fortune, S. M. & Alter, G. Beyond binding: antibody effector functions in infectious diseases. Nat. Rev. Immunol. 18, 46–61 (2018).
Steffen, U. et al. IgA subclasses have different effector functions associated with distinct glycosylation profiles. Nat. Commun. 11, 120 (2020).
Beyer, H. et al. Functional studies with IgM and IgA immunoglobulins: binding to pIgR, FcalphamuR, FcmuR, and CDC activities. APMIS 132, 277–288 (2024).
Abes, R. & Teillaud, J. L. Impact of glycosylation on effector functions of therapeutic IgG. Pharmaceuticals 3, 146–157 (2010).
Nose, M. & Wigzell, H. Biological significance of carbohydrate chains on monoclonal antibodies. Proc. Natl. Acad. Sci. USA 80, 6632–6636 (1983).
Jefferis, R. Recombinant antibody therapeutics: the impact of glycosylation on mechanisms of action. Trends Pharm. Sci. 30, 356–362 (2009).
Larsen, M. D. et al. Afucosylated IgG characterizes enveloped viral responses and correlates with COVID-19 severity. Science 371, eabc8378 (2021).
Dekkers, G. et al. Decoding the human immunoglobulin G-glycan repertoire reveals a spectrum of Fc-receptor- and complement-mediated-effector activities. Front. Immunol. 8, 877 (2017).
Haslund-Gourley, B. S. et al. IgM N-glycosylation correlates with COVID-19 severity and rate of complement deposition. Nat. Commun. 15, 404 (2024).
Pabst, M. et al. A microarray-matrix-assisted laser desorption/ionization-mass spectrometry approach for site-specific protein N-glycosylation analysis, as demonstrated for human serum immunoglobulin M (IgM). Mol. Cell Proteom. 14, 1645–1656 (2015).
Kissel, T. et al. N-linked Fc glycosylation is not required for IgG-B-cell receptor function in a GC-derived B-cell line. Nat. Commun. 15, 393 (2024).
Rodríguez-Pinto, D. B cells as antigen presenting cells. Cell Immunol. 238, 67–75 (2005).
Oskam, N. et al. CD5L is a canonical component of circulatory IgM. Proc. Natl. Acad. Sci. USA 120, e2311265120 (2023).
Sitia, R. et al. Developmental regulation of IgM secretion: the role of the carboxy-terminal cysteine. Cell 60, 781–790 (1990).
Early, P. et al. 2 Messenger-Rnas can be produced from a single immunoglobulin-Mu gene by alternative RNA processing pathways. Cell 20, 313–319 (1980).
Su, Q. et al. Cryo-EM structure of the human IgM B cell receptor. Science 377, 875 (2022).
van Tol, B. D. M. et al. Comprehensive immunoglobulin G, A, and M glycopeptide profiling for large-scale biomedical research. Mol. Cell Proteom. 24, 100928 (2025).
Wortel, C. M. et al. Anti-myeloperoxidase IgM B cells in anti-neutrophil cytoplasmic antibody-associated vasculitis. Nat. Commun. 16, 1582 (2025).
He, X. C. et al. Continuous signaling of CD79b and CD19 is required for the fitness of Burkitt lymphoma B cells. Embo J. 37, e97980 (2018).
Kurosaki, T. et al. Syk activation by the Src-family tyrosine kinase in the B cell receptor signaling. J. Exp. Med. 179, 1725–1729 (1994).
Ives, C. M. et al. Restoring protein glycosylation with GlycoShape. Nat. Methods 21, 2117–2127 (2024).
Su, Q. et al. Cryo-EM structure of the human IgM B cell receptor. Science 377, 875–880 (2022).
Ma, X. et al. Cryo-EM structures of two human B cell receptor isotypes. Science 377, 880–885 (2022).
Thaysen-Andersen, M. & Packer, N. H. Site-specific glycoproteomics confirms that protein structure dictates formation of -glycan type, core fucosylation and branching. Glycobiology 22, 1440–1452 (2012).
Yang, W. H. et al. An intrinsic mechanism of secreted protein aging and turnover. Proc. Natl. Acad. Sci. USA 112, 13657–13662 (2015).
Ripperger, T. J. & Bhattacharya, D. Transcriptional and metabolic control of memory B cells and plasma cells. Annu. Rev. Immunol. 39, 345–368 (2021).
Morel, M. et al. Abnormal B cell glycosylation in autoimmunity: A new potential treatment strategy. Front. Immunol. 13, 975963 (2022).
Youd, M. E., Luus, L. & Corley, R. B. IgM monomers accelerate disease manifestations in autoimmune-prone -deficient mice. J. Autoimmun. 23, 333–343 (2004).
Moens, L., Kane, A. & Tangye, S. G. Naive and memory B cells exhibit distinct biochemical responses following BCR engagement. Immunol. Cell Biol. 94, 774–786 (2016).
Kissel, T. et al. Surface Ig variable domain glycosylation affects autoantigen binding and acts as threshold for human autoreactive B cell activation. Sci. Adv. 8, eabm1759 (2022).
Vergroesen, R. D. et al. B-cell receptor sequencing of anti-citrullinated protein antibody (ACPA) IgG-expressing B cells indicates a selective advantage for the introduction of N-glycosylation sites during somatic hypermutation. Ann. Rheum. Dis. 77, 955 (2018).
Ubelhart, R. et al. N-linked glycosylation selectively regulates autonomous precursor BCR function. Nat. Immunol. 11, 759–765 (2010).
Cao, A. et al. Galectin-9 binds IgM-BCR to regulate B cell signaling. Nat. Commun. 9, 3288 (2018).
Blenda, A. V. et al. Galectin-9 recognizes and exhibits antimicrobial activity toward microbes expressing blood group-like antigens. J. Biol. Chem. 298, 101704 (2022).
Moar, P. & Tandon, R. Galectin-9 as a biomarker of disease severity. Cell Immunol. 361, 104287 (2021).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–D552 (2022).
Bern, M. W. & Kil, Y. J. Two-dimensional target decoy strategy for shotgun proteomics. J. Proteome Res. 10, 5296–5301 (2011).
Schulte, D., Leuvenink, R. W., Jager, S., Heck, A. J. R. & Snijder, J. A Universal Spectrum Annotator for Complex Peptidoforms in Mass Spectrometry-Based Proteomics. Anal. Chem. 97, 23120–23130 (2025).
Kissel, T. et al. Antibodies and B cells recognising citrullinated proteins display a broad cross-reactivity towards other post-translational modifications. Ann. Rheum. Dis. 79, 472–480 (2020).
Kerkman, P. F. et al. Identification and characterisation of citrullinated antigen-specific B cells in peripheral blood of patients with rheumatoid arthritis. Ann. Rheum. Dis. 75, 1170–1176 (2016).
Jumper, J. et al. Applying and improving AlphaFold at CASP14. Proteins 89, 1711–1721 (2021).
Varadi, M. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2022).
Agirre, J. et al. Privateer: software for the conformational validation of carbohydrate structures. Nat. Struct. Mol. Biol. 22, 833–834 (2015).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph 14, 33–38 (1996). 27-8.
Acknowledgements
This work is funded by the ERC GlycanSwitch grant 101071386.
Author information
Authors and Affiliations
Contributions
MDHK conceived the project, designed experiments, performed a majority of the experiments and analyzed the data. LH critically assessed project progress, contributed to intellectual discussions, provided literature resources and ideas, analyzed and visualized the LC-MS data. S.K. and R.B. contributed with the generation of B cell lines and their subsequent sorting, and performed antigen-titration experiments. RvdW performed B-cell isolation and B-cell sorting experiments. RTNT, PAvV designed and performed all LC-MS experiments and analyzed the data. O.S. and E.F. performed and analyzed the 3D reconstruction of the N209-glycosylated Igµ BCR. D.F. and M.W. contributed to the analysis, interpretation and visualization of LC-MS data. REMT conceived the project, designed experiments and supervised the project. All authors contributed to the revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Paolo Casali, Shipei Xing and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Holborough-Kerkvliet, M.D., Hafkenscheid, L., Kroos, S. et al. Site-specific profiling of structure and function of Igµ B cell receptor glycans. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70121-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70121-y