Abstract
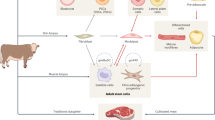
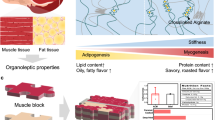
Cell-cultivated meat has emerged as an alternative approach for sustainable protein production, but replicating the texture and complex composition of conventional meat remains a critical challenge. Here, we developed an efficient approach to produce multitissue engineered meat by leveraging a serum- and animal-component-free differentiation system to direct porcine pregastrulation epiblast stem cells toward muscle, adipose, and endothelium with synergistic functions. These three types of cell progenitors demonstrated autonomous intercellular recognition and coculture compatibility in a scaffold-free 3D suspension system, enabling the spontaneous formation of tissue-mimetic spheroids with enhanced cellular multiplication efficiency. The products of multitissue cultivated meat recapitulated the textural properties of conventional pork and allowed for nutritional modulation. This platform overcomes key limitations in currently employed cultured meat technologies by integrating scalable 3D suspension culture with serum-free, species-specific stem cell multidirectional differentiation, providing an avenue for the development of multitissue cell-cultivated meat.
Similar content being viewed by others
Data availability
The raw RNA-seq data generated in this paper have been deposited in the Genome Sequence Archive (GSA) at the National Genomics Data Center, China National Center for Bioinformation (NGDC/CNCB), under accession number CRA026288, and are publicly accessible at https://ngdc.cncb.ac.cn/gsa/browse/CRA026288. The data generated in this study are provided in the Supplementary Information/Source Data file. Source data are provided with this paper.
References
Post, M. J. et al. Scientific, sustainability and regulatory challenges of cultured meat. Nat. Food 1, 403–415 (2020).
Post, M. J. Cultured meat from stem cells: challenges and prospects. Meat Sci. 92, 297–301 (2012).
Kupferschmidt, K. Food science. Lab burger adds sizzle to bid for research Funds. Science 341, 602–603 (2013).
Martins, B. et al. Advances and challenges in cell biology for cultured meat. Annu. Rev. Anim. Biosci. 12, 345–368 (2024).
Kawecki, N. S. et al. Scalable processes for culturing meat using edible scaffolds. Annu. Rev. Food Sci. Technol. 15, 241–264 (2024).
Samandari, M. et al. Repurposing biomedical muscle tissue engineering for cellular agriculture: challenges and opportunities. Trends Biotechnol. 41, 887–906 (2023).
Broucke, K., Van Pamel, E., Van Coillie, E., Herman, L. & Van Royen, G. Cultured meat and challenges ahead: a review on nutritional, technofunctional and sensorial properties, safety and legislation. Meat Sci. 195, 109006 (2023).
Messmer, T. et al. A serum-free media formulation for cultured meat production supports bovine satellite cell differentiation in the absence of serum starvation. Nat. Food 3, 74 (2022).
Ben-Arye, T. et al. Textured soy protein scaffolds enable the generation of three-dimensional bovine skeletal muscle tissue for cell-based meat. Nat. Food 1, 210–220 (2020).
Song, W., Liu, P., Li, H. & Ding, S. Large-scale expansion of porcine adipose-derived stem cells based on microcarriers system for cultured meat production. Foods 11, 3364 (2022).
Mitic, R., Cantoni, F., Borlin, C. S., Post, M. J. & Jackisch, L. A simplified and defined serum-free medium for cultivating fat across species. iScience 26, 105822 (2023).
Dohmen, R. G. J. et al. Muscle-derived fibro-adipogenic progenitor cells for production of cultured bovine adipose tissue. NPJ Sci. Food 6, 6 (2022).
Pasitka, L. et al. Spontaneous immortalization of chicken fibroblasts generates stable, high-yield cell lines for serum-free production of cultured meat. Nat. Food 4, 35–50 (2023).
Zhi, M. et al. Generation and characterization of stable pig pregastrulation epiblast stem cell lines. Cell Res. 32, 383–400 (2022).
Jara, T. C. et al. Stem cell-based strategies and challenges for production of cultivated meat. Nat. Food 4, 841–853 (2023).
Giordani, L. et al. High-dimensional single-cell cartography reveals novel skeletal muscle-resident cell populations. Mol. Cell 74, 609 (2019).
Li, C. H. et al. The production of fat-containing cultured meat by stacking aligned muscle layers and adipose layers formed from gelatin-soymilk scaffold. Front. Bioeng. Biotechnol. 10, 875069 (2022).
Lee, M. et al. Cultured meat with enriched organoleptic properties by regulating cell differentiation. Nat. Commun. 15, 77 (2024).
Liu, Y. et al. Engineered meatballs via scalable skeletal muscle cell expansion and modular micro-tissue assembly using porous gelatin micro-carriers. Biomaterials 287, 121615 (2022).
Yen, F. C. et al. Cultured meat platform developed through the structuring of edible microcarrier-derived microtissues with oleogel-based fat substitute. Nat. Commun. 14, 2942 (2023).
Ghaben, A. L. & Scherer, P. E. Adipogenesis and metabolic health. Nat. Rev. Mol. Cell Biol. 20, 242–258 (2019).
Tan, J. Y., Sriram, G., Rufaihah, A. J., Neoh, K. G. & Cao, T. Efficient derivation of lateral plate and paraxial mesoderm subtypes from human embryonic stem cells through GSKi-mediated differentiation. Stem Cells Dev. 22, 1893–1906 (2013).
Lian, Q. et al. Functional mesenchymal stem cells derived from human induced pluripotent stem cells attenuate limb ischemia in mice. Circulation 121, 1113–1123 (2010).
Lamouille, S., Xu, J. & Derynck, R. Molecular mechanisms of epithelial-mesenchymal transition. Nat. Rev. Mol. Cell Biol. 15, 178–196 (2014).
Hudak, C. S. et al. Pref-1 marks very early mesenchymal precursors required for adipose tissue development and expansion. Cell Rep. 8, 678–687 (2014).
Deng, P., Zhou, C., Alvarez, R., Hong, C. & Wang, C. Y. Inhibition of IKK/NF-κB signaling enhances differentiation of mesenchymal stromal cells from human embryonic stem cells. Stem Cell Rep. 6, 456–465 (2016).
Lefterova, M. I. et al. PPARγ and C/EBP factors orchestrate adipocyte biology via adjacent binding on a genome-wide scale. Gene Dev. 22, 2941–2952 (2008).
Siersbaek, R., Nielsen, R. & Mandrup, S. Transcriptional networks and chromatin remodeling controlling adipogenesis. Trends Endocrinol. Metab. 23, 56–64 (2012).
Ahfeldt, T. et al. Programming human pluripotent stem cells into white and brown adipocytes. Nat. Cell Biol. 14, 209–219 (2012).
Lazarescu, O. et al. Human subcutaneous and visceral adipocyte atlases uncover classical and nonclassical adipocytes and depot-specific patterns. Nat. Genet. 57, 413–426 (2025).
Orlova, V. V. et al. Generation, expansion and functional analysis of endothelial cells and pericytes derived from human pluripotent stem cells. Nat. Protoc. 9, 1514–1531 (2014).
Wimmer, R. A. et al. Human blood vessel organoids as a model of diabetic vasculopathy. Nature 565, 505–510 (2019).
Patsch, C. et al. Generation of vascular endothelial and smooth muscle cells from human pluripotent stem cells. Nat. Cell Biol. 17, 994–1003 (2015).
Sahara, M. et al. Manipulation of a VEGF-Notch signaling circuit drives formation of functional vascular endothelial progenitors from human pluripotent stem cells. Cell Res. 25, 148 (2015). 24, 820 (2014).
Hamad, S. et al. High-efficient serum-free differentiation of endothelial cells from human iPS cells. Stem Cell Res. Ther. 13, 251 (2022).
Zhu, G. et al. Generation of three-dimensional meat-like tissue from stable pig epiblast stem cells. Nat. Commun. 14, 8163 (2023).
Zhang, S. et al. Implantation initiation of self-assembled embryo-like structures generated using three types of mouse blastocyst-derived stem cells. Nat. Commun. 10, 496 (2019).
Bonnans, C., Chou, J. & Werb, Z. Remodelling the extracellular matrix in development and disease. Nat. Rev. Mol. Cell Biol. 15, 786–801 (2014).
Gonzalez-Soto, M. et al. Comparing soy and milk protein regulation of hepatic omega-3 fatty acid biosynthesis. FASEB J. 39, e70805 (2025).
Li, M. M. et al. Effects of dietary n-6:n-3 PUFA ratios on lipid levels and fatty acid profile of Cherry Valley ducks at 15-42 days of age. J. Agr. Food Chem. 65, 9995–10002 (2017).
Hua, T. et al. Studies of human 2,4-dienoyl CoA Reductase Shed New Light On Peroxisomal β-Oxidation Of Unsaturated Fatty Acids. J. Biol. Chem. 287, 28956–28965 (2012).
Chen, L. et al. Large-scale cultured meat production: trends, challenges and promising biomanufacturing technologies. Biomaterials 280, 121274 (2022).
Thorrez, L. & Vandenburgh, H. Challenges in the quest for ‘clean meat’. Nat. Biotechnol. 37, 215–216 (2019).
Hocquette, J. F. Is meat the solution for the future? Meat Sci. 120, 167–176 (2016).
Cheng, Y. M. et al. An immortal porcine preadipocyte cell strain for efficient production of cell-cultured fat. Commun. Biol. 6, 1202 (2023).
Khan, M. I., Jo, C. & Tariq, M. R. Meat flavor precursors and factors influencing flavor precursors-a systematic review. Meat Sci. 110, 278–284 (2015).
Fraeye, I., Kratka, M., Vandenburgh, H. & Thorrez, L. Sensorial and Nutritional Aspects Of Cultured Meat In Comparison To Traditional Meat: Much To Be Inferred. Front. Nutr. 7, 35 (2020).
Wang, Y., Zhuang, D., Munawar, N., Zan, L. & Zhu, J. A rich-nutritious cultured meat via bovine myocytes and adipocytes co-culture: Novel Prospect for cultured meat production techniques. Food Chem. 460, 140696 (2024).
Zagury, Y., Ianovici, I., Landau, S., Lavon, N. & Levenberg, S. Engineered marble-like bovine fat tissue for cultured meat. Commun. Biol. 5, 927 (2022).
Louis, F., Furuhashi, M., Yoshinuma, H., Takeuchi, S. & Matsusaki, M. Mimicking Wagyu beef fat in cultured meat: Progress in edible bovine adipose tissue production with controllable fatty acid composition. Mater. Today Bio. 21, 100720 (2023).
Rouwkema, J. & Khademhosseini, A. Vascularization and angiogenesis in tissue engineering: beyond creating static networks. Trends Biotechnol. 34, 733–745 (2016).
Kang, D. H. et al. Engineered whole cut meat-like tissue by the assembly of cell fibers using tendon-gel integrated bioprinting. Nat. Commun. 12, 5059 (2021).
Kong, Y. & Huang, D. Pumpkin seed proteins rival animal gelatin in increasing the cytoaffinity of edible microbeads for cell-based meat culture. Food Res. Int. 168, 112750 (2023).
Ianovici, I., Zagury, Y., Redenski, I., Lavon, N. & Levenberg, S. 3D-printable plant protein-enriched scaffolds for cultivated meat development. Biomaterials 284, 121487 (2022).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Zhou, Y. Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Acknowledgements
This work was supported by the Biological Breeding-National Science and Technology Major Project (2023ZD0407503 to J.H.), the Natural Science Foundation of China (32370846 to J.H., 31970825 to J.H., 32402757 to M.Z.), the National Key R&D Program of China (2022YFD1302201 to J.H.), the fellowship of China Postdoctoral Science Foundation (2022M720168 to M.Z.), China National Postdoctoral Program for Innovative Talents (BX20220344 to M.Z.), Pinduoduo-China Agricultural University Research Fund (PC2024A01001 to J.H.), the 2115 Talent Development Program of China Agricultural University and High-performance Computing Platform of China Agricultural University.
Author information
Authors and Affiliations
Contributions
J.H. conceptualized this project. J.H., S.C., and A.M. supervised the overall experiments. Y.Y. and G.Z. performed pgEpiSCs culture in vitro and characteristics analysis of differentiated cells. J.H., Y.Y., R.L., and M.Z. coordinated and performed bioinformatics analysis of the RNA-seq sequencing data. Y.Y., G.Z., M.Z., and T.W. performed spheroid culture in vitro and characteristics analysis. J.H., Y.Y., A.M, Y.Z., and Y.C. performed preparation of cell-based meat, nutritional components, and texture analysis. Y.Y., G.Z., L.L., and X.F. completed the preparation, cell inoculation of 3D edible scaffolds, and SEM. M.Z. performed pgEpiSCs derivation. Y.Y., G.Z., T.W., and L.Q. performed flow cytometric analysis. Y.Y., G.Z., Y.W., S.W., and H.Z. performed qRT-PCR primer design, quantitative validation, and immunofluorescence image processing. Y.Y., X.C., and M.Z. performed the preparation of the experimental schematic. J.H. and Y.Y. performed manuscript writing, review, and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jinkee Hong, Mark J. Post, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yao, Y., Zhu, G., Zhi, M. et al. Generation of multitissue cell-cultivated meat via multidirectional differentiation of stable porcine epiblast stem cells. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70177-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70177-w