Abstract
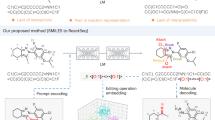
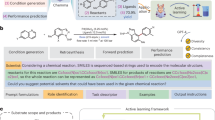
The application of artificial intelligence (AI) to chemical discovery is critically hindered by the inaccessibility of data locked within unstructured scientific literature. Existing data acquisition methods are often manual, limited in scope, or require extensive custom software development, impeding progress in leveraging AI for chemical discovery. Here, we introduce ReactionSeek, a framework that synergistically combines large language models (LLMs) with established cheminformatics tools to automate multi-modal data mining from organic synthesis literature. Through sophisticated prompt engineering with minimal custom code, ReactionSeek extracts and standardizes complex textual, graphical, and semantic chemical information. We validate this framework on the century-spanning Organic Syntheses collection, achieving over 95% precision and recall for key reaction parameters. This enables three applications: the generation of a large, AI-ready dataset; an interactive Synthetic Chatbot (SynChat) for natural language querying of chemical data; and an autonomous analysis that revealed decades-long trends in catalysis. ReactionSeek thus provides a general solution to the data curation bottleneck, representing a step forward in for AI-driven archive mining and knowledge discovery across the chemical sciences.
Similar content being viewed by others
Data availability
The mined data, code, and evaluation datasets generated in this study have been deposited in the Zenodo database under accession code (https://doi.org/10.5281/zenodo. 18523416).
The raw textual data utilized in this study are available under restricted access for copyright and licensing restrictions; access can be obtained by subscription to the original publishers. The processed evaluation data are available at the GitHub repository (https://github.com/DeepSynthesis/ReactionSeek.git) under the file path examples/OrganicSyntheses/reaction_extract/evaluate_raw.json. The literature list used in this study is available in the GitHub repository under the file path examples/OrganicSyntheses/OrgS_cite.csv. The Source Data underlying the main text figures are provided in the cited Zenodo repository54.
Code availability
The source code developed for this study, including all scripts and algorithms, has been deposited in a public GitHub repository https://github.com/DeepSynthesis/ReactionSeek.git.54 All code is available under the MIT License. The primary results reported in this work were generated using the GLM-4 model. The vision language model for image recognition tasks was based on GLM-4V. A comprehensive list of all large language models used for the multi-model evaluation is provided in the main text. Our framework is designed to be model-agnostic. The choice of specific models for testing was to benchmark performance and guide users in model selection. The provided codebase includes documentation to assist users in configuring and integrating their models, including proprietary models requiring user-provided API keys or other services compatible with the OpenAI API format. For the chemical structure recognition task, the InDraw34 OCSR tool was employed. Users may substitute this with any alternative chemical structure recognition tool suitable for their requirements.
References
Jiang, X. et al. Artificial intelligence and automation to power the future of chemistry. Cell Rep. Phys. Sci. 5, 102049 (2024).
Hong, X. et al. AI for organic and polymer synthesis. Sci. China Chem. 67, 2461–2496 (2024).
Tan, Z., Yang, Q. & Luo, S. AI molecular catalysis: Where are we now?. Org. Chem. Front. 12, 2759–2776 (2025).
Skoraczyński, G. et al. Predicting the outcomes of organic reactions via machine learning: are current descriptors sufficient?. Sci. Rep. 7, 3582 (2017).
Mercado, R., Kearnes, S. M. & Coley, C. W. Data sharing in chemistry: lessons learned and a case for mandating structured reaction data. J. Chem. Inf. Model. 63, 4253–4265 (2023).
Strieth-Kalthoff, F. et al. Artificial intelligence for retrosynthetic planning needs both data and expert knowledge. J. Am. Chem. Soc. 146, 11005–11017 (2024).
Ahneman, D. T., Estrada, J. G., Lin, S., Dreher, S. D. & Doyle, A. G. Predicting reaction performance in C–N cross-coupling using machine learning. Science 360, 186–190 (2018).
Kearnes, S. M. et al. The open reaction database. J. Am. Chem. Soc. 143, 18820–18826 (2021).
Su, Y. et al. Automation and machine learning augmented by large language models in a catalysis study. Chem. Sci. 15, 12200–12233 (2024).
Krallinger, M., Rabal, O., Lourenço, A., Oyarzabal, J. & Valencia, A. Information retrieval and text mining technologies for chemistry. Chem. Rev. 117, 7673–7761 (2017).
Dagdelen, J. et al. Structured information extraction from scientific text with large language models. Nat. Commun. 15, 1418 (2024).
Kang, Y. et al. Harnessing large language models to collect and analyze metal–organic framework property data set. J. Am. Chem. Soc. 147, 3943–3958 (2025).
Leong, S. X., Pablo-García, S., Zhang, Z. & Aspuru-Guzik, A. Automated electrosynthesis reaction mining with multimodal large language models (MLLMs). Chem. Sci. 15, 17881–17891 (2024).
Wang, X., Huang, L., Xu, S. & Lu, K. How does a generative large language model perform on domain-specific information extraction?─A comparison between GPT-4 and a rule-based method on band gap extraction. J. Chem. Inf. Model. 64, 7895–7904 (2024).
Zhang, W. et al. Fine-tuning large language models for chemical text mining. Chem. Sci. 15, 10600–10611 (2024).
Zheng, Z. et al. Image and data mining in reticular chemistry powered by GPT-4V. Digit. Discov. 3, 491–501 (2024).
Zheng, Z., Zhang, O., Borgs, C., Chayes, J. T. & Yaghi, O. M. ChatGPT chemistry assistant for text mining and the prediction of MOF synthesis. J. Am. Chem. Soc. 145, 18048–18062 (2023).
Ruan, Y. et al. An automatic end-to-end chemical synthesis development platform powered by large language models. Nat. Commun. 15, 10160 (2024).
Ai, Q., Meng, F., Shi, J., Pelkie, B. & Coley, C. W. Extracting structured data from organic synthesis procedures using a fine-tuned large language model. Digit. Discov. 3, 1822–1831 (2024).
Leong, S. X., Pablo-García, S., Wong, B. & Aspuru-Guzik, A. MERMaid: universal multimodal mining of chemical reactions from PDFs using vision-language models. Matter 8, 102331 (2025).
Walker, N. et al. Extracting structured seed-mediated gold nanorod growth procedures from scientific text with LLMs. Digit. Discov. 2, 1768–1782 (2023).
Chen, Y. et al. A multi-agent system enables versatile information extraction from the chemical literature. Preprint at. https://ui.adsabs.harvard.edu/abs/2025arXiv250720230C (2025).
Fan, V. et al. OpenChemIE: an information extraction toolkit for chemistry literature. J. Chem. Inf. Model. 64, 5521–5534 (2024).
Rajan, K., Zielesny, A. & Steinbeck, C. DECIMER 1.0: deep learning for chemical image recognition using transformers. J. Cheminform. 13, 61 (2021).
Staker, J., Marshall, K., Abel, R. & McQuaw, C. M. Molecular structure extraction from documents using deep learning. J. Chem. Inf. Model. 59, 1017–1029 (2019).
Wilary, D. M. & Cole, J. M. ReactionDataExtractor: a tool for automated extraction of information from chemical reaction schemes. J. Chem. Inf. Model. 61, 4962–4974 (2021).
Wilary, D. M. & Cole, J. M. ReactionDataExtractor 2.0: a deep learning approach for data extraction from chemical reaction schemes. J. Chem. Inf. Model. 63, 6053–6067 (2023).
Guo, J. et al. Automated chemical reaction extraction from scientific literature. J. Chem. Inf. Model. 62, 2035–2045 (2022).
Vangala, S. R. et al. Suitability of large language models for extraction of high-quality chemical reaction dataset from patent literature. J. Cheminform. 16, 131 (2024).
Andrew, S., Kende & Jeremiah, P., Freeman. Organic Syntheses (John Wiley and Sons, 2003).
Clevert, D.-A., Le, T., Winter, R. & Montanari, F. Img2Mol – accurate SMILES recognition from molecular graphical depictions. Chem. Sci. 12, 14174–14181 (2021).
Oldenhof, M., Arany, A., Moreau, Y. & Simm, J. ChemGrapher: optical graph recognition of chemical compounds by deep learning. J. Chem. Inf. Model. 60, 4506–4517 (2020).
Qian, Y. et al. MolScribe: robust molecular structure recognition with image-to-graph generation. J. Chem. Inf. Model. 63, 1925–1934 (2023).
Integle Inc. InDraw. https://indrawforweb.integle.com (2025).
Li, Y. et al. The ChEMU 2022 evaluation campaign: information extraction in chemical patents. in Advances in Information Retrieval, 400–407 (Springer, 2022).
He, J. et al. ChEMU 2020: natural language processing methods are effective for information extraction from chemical patents. Front. Res. Metr. Anal. 6, 654438 (2021).
Jang, Y. J. et al. Context aware named entity recognition and relation extraction with domain-specific language model. in Conference and Labs of the Evaluation Forum (CEUR-WS.org, 2022).
Li, Y. et al. Overview of ChEMU 2022 evaluation campaign: information extraction in chemical patents. in Experimental IR Meets Multilinguality, Multimodality, and Interaction, 521–540 (Springer-Verlag, 2022).
Jablonka, K. M. et al. 14 examples of how LLMs can transform materials science and chemistry: a reflection on a large language model hackathon. Digit. Discov. 2, 1233–1250 (2023).
Kim, S. et al. PubChem 2025 update. Nucleic Acids Res. 53, D1516–D1525 (2025).
Lowe, D. M., Corbett, P. T., Murray-Rust, P. & Glen, R. C. Chemical name to structure: OPSIN, an open source solution. J. Chem. Inf. Model. 51, 739–753 (2011).
Zeng, A. et al. GLM-130B: an open bilingual pre-trained model. In Proc. 11th Int. Conf. Learn. Represent. (2023).
Machlab, D. & Battle, R. LLM in-context recall is prompt dependent. Preprint at. https://arxiv.org/abs/2404.08865 (2024).
Tom, B. et al. Language models are few-shot learners. Adv. Neural Inf. Process. Syst. 33, 1877–1901 (2020).
OpenAI et al. GPT-4 Technical report. Preprint at. https://arxiv.org/abs/2303.08774 (2023).
Touvron, H. et al. Llama 2: open foundation and fine-tuned chat models. Preprint at. https://arxiv.org/abs/2307.09288 (2023).
Jiang, A. Q. et al. Mixtral of experts. Preprint at. https://arxiv.org/abs/2401.04088 (2024).
Yang, A. et al. Baichuan 2: open large-scale language models. Preprint at. https://arxiv.org/abs/2309.10305 (2023).
Yang, A. et al. Qwen2 technical report. Preprint at. https://arxiv.org/abs/2407.10671 (2024).
DeepSeek-AI et al. DeepSeek-V3 technical report. Preprint at. https://arxiv.org/abs/2412.19437 (2024).
Google DeepMind. Gemini 2.5 Pro. https://deepmind.google/models/gemini/pro/ (2025).
Lewis, P. et al. Retrieval-augmented generation for knowledge-intensive NLP tasks. Adv. Neural Inf. Process. Syst. 33, 9459–9474 (2020).
Chroma. Chroma, https://www.trychroma.com/ (2025).
Li, J., Li, M.,Yang, Q. & Luo, S. DeepSynthesis/ReactionSeek: source code and data for nature communications. Zenodo https://doi.org/10.5281/zenodo.18523416 (2026).
Acknowledgements
We thank the Fundamental and Interdisciplinary Disciplines Breakthrough Plan of the Ministry of Education of China (JYB2025XDXM414), Natural Science Foundation of China (22031006; 22271192; 22373056 and 22393891), the National Key R&D Program of China (2023YFA1506401; 2023YFA1506402) and Haihe Laboratory of Sustainable Chemical Transformations (25HHWCSS00032, 24HHWCSS00018) for financial support. This work is supported by High performance computing Center, Tsinghua University and the robotic AI-Scientist platform of Chinese Academy of Sciences.
Author information
Authors and Affiliations
Contributions
S.L. and Q.Y. conceived and directed the project. J.L. designed the research workflow and developed the programs and prompt engineering for the data mining framework. M.L. built and deployed SynChat. All authors contributed to the writing and revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Masaharu Yoshioka, Yanyan Xu, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, J., Li, M., Yang, Q. et al. ReactionSeek: LLM-powered literature data mining and knowledge discovery in organic synthesis. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70180-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70180-1