Abstract
Herbivory has evolved multiple times during mammalian diversification, playing a key role in the success of this globally distributed clade. A central metabolic challenge for herbivores is the detoxification of glyoxylate. The enzyme alanine:glyoxylate aminotransferase (AGT) converts glyoxylate to glycine, preventing the formation of harmful calcium oxalate crystals. AGT localizes to mitochondria and peroxisomes based on the mitochondrial targeting sequence (MTS) and the peroxisomal targeting signal (PTS1), respectively. While most studies focused on MTS, MTS variation alone does not fully explain AGT localization patterns. To assess the relative importance of the PTS1 motif, we combined comparative sequence analyses with functional assays. We find that multiple herbivorous lineages underwent independent mutations resulting in disrupted or truncated MTS regions, whereas the PTS1 motif remains functional. Immunofluorescence assays revealed more efficient peroxisomal localization of AGT in herbivores, with PTS1 often overriding mitochondrial signals even when the MTS is intact. Additionally, transcriptomic analyses show that several herbivorous lineages preferentially use downstream transcriptional start sites, producing AGT isoforms lacking the MTS. Together, our findings reveal a central role of AGT peroxisomal targeting in evolution of plant-based diets, and demonstrate how increased targeting efficiency has evolved convergently via the interplay of transcriptional regulation and targeting signals.
Similar content being viewed by others
Introduction
The independent evolution of plant-based diets has occurred multiple times throughout the diversification of mammals, and is a key factor contributing to the evolutionary and ecological success of this group1. Fossil evidence indicates that early mammals were primarily insectivorous, as inferred from their jaw and dental morphology, and that these formed the foundation for numerous subsequent shifts into novel dietary niches2,3. Herbivory in mammals likely originated ~260 million years ago in groups such as Eleutherodontidae, which fed on non-flowering plants4, with further diversification of herbivorous lineages coinciding with angiosperm evolution in the Cretaceous5.
For successful transitions to plant-based diets, mammals evolved multiple traits for finding, ingesting, breaking down, and metabolizing plant tissues6. Adaptations for hosting symbiotic bacteria for the breakdown of cellulose7,8, as well as processing simple and complex carbohydrates9,10,11, have evolved convergently in multiple groups. One of the major challenges for mammals that subsist on plants is overcoming toxins and secondary compounds, many of which have evolved as defences against herbivory12,13.
The enzyme alanine:glyoxylate aminotransferase (AGT) is primarily located in the mitochondria of liver cells, where it converts glyoxylate to glycine14. Glyoxylate is a byproduct of metabolism and is otherwise converted into oxalate, which can accumulate and lead to the formation of toxic calcium oxalate crystals in the body14. In carnivores, the precursor of glyoxylate is hydroxyproline, which is broken down in the mitochondria15, whereas in herbivores, it is glycolate, which is produced in peroxisomes16. As a consequence, in some species, AGT has also evolved to target peroxisomes via distinct protein domains and post-translational modifications17. The tendency for AGT to localize in mitochondria in carnivores, and peroxisomes in herbivores, is supported by immunoelectron microscopy studies across over 70 mammalian species18,19.
AGT’s subcellular localization is directed by a mitochondrial targeting sequence (MTS) of ~22 amino acids at the N-terminus, and a peroxisomal targeting signal (PTS1), a C-terminal tripeptide within the mature protein14,20,21,22. To date, most studies relating AGT to diet have focused on the dominant role of the MTS in mitochondrial targeting18,19,23. However, independent MTS losses have been reported in several lineages, including bats and primates23,24. Many such losses arise from mutations affecting the 5′ start site of the AGXT, which disrupt MTS function and impair mitochondrial import25,26,27. Yet despite these results, MTS integrity alone cannot fully explain the AGT distribution across mammals. In many species, AGT localization exists along a continuum between mitochondria and peroxisomes. Moreover, in some species with intact MTS—such as herbivorous domestic horse (Equus ferus) and omnivorous olive baboon (Papio anubis)—AGT predominantly targets peroxisomes18.
Compared to the MTS region, the role of AGT’s PTS1 motif is less well understood. Some have suggested that PTS1 might only function when the MTS is weak or inactive21; however, the finding that enhancing AGT peroxisomal targeting leads to reduced mitochondrial localization implies that AGT distribution is controlled by a balance between mitochondrial and peroxisomal import systems28. Studies of other peroxisome targeting proteins have identified a so-called consensus PTS1 motif, [S/A/C]-[K/R/H]-[L/M] that confers high peroxisomal targeting efficiency28. However, while comparisons of AGT across vertebrate groups have recorded this consensus motif in some taxa, its relationship to diet in mammals is unclear. Indeed, the human AGT PTS1 sequence, KKL, is unique among human peroxisomal proteins and is necessary but insufficient for efficient peroxisomal import27,29,30,31.
In addition to amino acid variation in targeting regions, subcellular localization of AGT across mammals can also reflect differential use or loss of alternative transcription start sites. The complete AGXT has two transcription start sites: one upstream of the first translation start codon, and another located between two translation start codons14. In cats, mitochondrial AGT derives from the longer transcript, whereas in rabbits, peroxisomal AGT originates from the shorter one21. More omnivorous species, such as marmosets and rats have been shown to express both transcript variants21, suggesting a link between transcriptional regulation and dietary strategy.
In this work, we conduct the large-scale comparative study of AGT across mammals. By combining cell-based immunofluorescence assays of natural and mutant proteins with analyses of transcripts and regulatory regions, we demonstrate that variation in PTS1 sequences and alternative transcription initiation contribute to differences in AGT localization and dietary adaptation across species.
Results
Using tBLASTn32, we searched publicly available genome assemblies from 685 mammal species and retrieved 505 hits, with the remaining 180 genomes showing no hit due to low quality. Of the 505 hits, we recorded 472 AGT orthologues with an MTS region. In 33 species, the sequence was incomplete due to gaps in either the mature region and/or PTS1 targeting region.
Nonfunctional MTS and high-conserved PTS1 occur mostly in herbivores
We examined the 472 MTS regions and found that 429 were complete and thus considered putatively functional. MTS sequences showed significant variation between Eutheria (22 amino acids) and Prototheria/Metatheria (33 amino acids), indicating a lack of sequence homology.
In 43 species (Supplementary Fig. 1), the MTS was inferred to be non-functional due to the presence of a single non-canonical start codon (n = 21), premature stop codon (n = 6), or frameshift mutation (n = 3), or a combination of these (n = 13). Of these species, 35 (81.40%) are herbivores including 28 strict herbivores, four are omnivores (9.30%), and four are carnivores (9.30%) including three hypercarnivores (species with a diet comprising >70% animal tissue; e.g., Iberian mole, Talpa occidentalis) (Supplementary Fig. 2). Examples of herbivores with functional losses caused by frameshift mutations include the common wombat (Vombatus ursinus) from the order Diprotodontia, representing the first such case from the Metatheria. Losses arising from nonsense mutations were seen in lagomorphs, primates, bats, and several rodents. For example, we identified a shared premature stop codon (TGA) at nucleotide positions 43–45 in the MTS of five squirrel species (alpine marmot, Himalayan marmot, Vancouver Island marmot, woodchuck, and yellow-bellied marmot) and the mountain beaver (Aplodontia rufa), and a different stop (TAA) at the same site in the dormouse (Glis glis), Similarly, among the primates, the MTS of the white-faced saki (Pithecia pithecia) contained a premature stop codon at nucleotide positions 55–57 which can also be found in nutrias (Myocastor coypus). Other cases of loss of function involved missense mutations in the upstream translation start site of the AGXT, seen in diverse groups such as bats (e.g., Rousettus aegyptiacus) and primates (e.g., Cercopithecus mona) (Supplementary Fig. 1).
In contrast to the MTS, we recorded no cases of loss-of-function mutations in the PTS1 motif. A total of 498 complete orthologues contained a PTS1 motif, although the residues varied among species. We examined in detail the PTS1 motif composition and found that the consensus motif ([S/A/C]-[K/R/H]-[L/M]) was present in 106 species, comprising 75 (70.75%) herbivores and 69 (65.09%) strict herbivores (Figs. 1 and S2 and Supplementary Data 1). Additionally, the PTS1 in 330 species (66.27%) contained the [N/K]-K-L motif, which partially matches the consensus sequence (Supplementary Data 1). Interestingly, we recorded an overrepresentation of the motifs SRL (n = 17), SKL (n = 66), and SKM (n = 23), with most examples of the former two (94.12% and 86.36%, respectively) seen in herbivores, whereas cases of the latter motif were more common (91.30%) in carnivores (Supplementary Data 1).
Phylogenetic tree of mammals and related ancestral nodes highlighting species with/without consensus PTS1 and nonfunctional MTS. The outer ring shows the proportion of plant tissue in the diet based on data obtained from the PHYLACINE 1.2.1 database. Silhouettes of mammals were taken from phylopic.org. Source data is provided as a Source Data file.
We performed ancestral reconstruction of PTS1 motifs across the mammal tree and identified numerous instances of convergent evolution of known consensus motifs (Fig. 1). Notably, we observed independent evolution of the GKL motif in all Old World fruit bats (Pteropodidae) and sloths (Folivora) examined, and the SKL motif in most members of Artiodactyla and the koala (Phascolarctos cinereus). Similarly, we observed convergent evolution of the NRL motif in several unrelated lineages, including the Hispaniolan solenodon (Solenodon paradoxus) and degu (Octodon degus), the SRL motif in the dama gazelle (Nanger dama), paca (Cuniculus paca), and mouse-deer (Tragulus javanicus), and the HKL motif in carnivorans such as the domestic dog (Canis lupus familiaris) and banded mongoose (Mungos mungo), along with non-carnivorans like the Sumatran rhinoceros (Dicerorhinus sumatrensis) and domestic pig (Sus scrofa). Less common convergent motifs included HRL seen in the South African springhare (Pedetes capensis) and Patagonian mara (Dolichotis patagonum), SKM seen in the kinkajou (Potos flavus) and European hedgehog (Erinaceus europaeus), and GRL in the Indochinese shrew (Crocidura indochinensis) and nectar bat (Macroglossus sobrinus). Interestingly, while the majority of these convergent motif changes occurred in herbivorous mammals, the presence of distinct motifs was also seen in some omnivorous and carnivorous species, and no consensus motifs were recorded in primates (Fig. 1).
AGT peroxisomal targeting efficiency correlates with degrees of herbivory
We conducted cell-based immunofluorescence assays to assess the mitochondrial and peroxisomal targeting efficiency of the mature AGT protein in 37 mammal species for which immunoelectron microscopy data were also available18. Based on co-localization data, we found that mitochondrial targeting efficiencies ranged from 21.43% in the rhesus monkey (Macaca mulatta) to 98.66% in the greater horseshoe bat (Rhinolophus ferrumequinum), while peroxisomal targeting efficiencies ranged from 5.66% in the greater horseshoe bat to 92.61% in the rhesus monkey (Fig. 2a/l and Supplementary Fig. 3). Notably, we observed generally lower peroxisomal targeting efficiencies in species from the Carnivora and Chiroptera orders. Within Carnivora, peroxisomal targeting efficiencies varied from 8.06% in the lion (Panthera leo) to 29.40% in the brown bear (Ursus arctos), and only three species—including the giant panda (Ailuropoda melanoleuca) and the North American raccoon (Procyon lotor)—showed peroxisomal targeting efficiencies exceeding 20%. Similarly, in Chiroptera, peroxisomal targeting efficiencies ranged from 5.66% in greater horseshoe bats to 30.74% in the black flying fox (Pteropus alecto), the only herbivorous bat species in our sample (Fig. 2a).
a Comparison of peroxisomal targeting efficiencies with and without the PTS1 region across 37 mammal species. b PGLS regression analysis of peroxisomal targeting efficiency versus the proportion of plant tissue in the diet. c PGLS regression analysis of difference of peroxisomal targeting efficiency between AGT with/without PTS1 versus the proportion of plant tissue in the diet. In (b and c), species left of cut-offs (brown vertical lines) at 0.1 and 0.3 correspond to hypercarnivores and carnivores, respectively, and species right of cut-offs at 0.7 and 0.95 correspond to herbivores and strict herbivores, respectively. P values were calculated using two-sided t-tests. d Synonymous (dS) and nonsynonymous (dN) substitution rates estimated using the free-ratio model. e Synonymous (dS) and nonsynonymous (dN) substitution rates estimated using the modified Nei-Gojobori method. In (d and e), black vertical lines represent dN/dS = 1. f, g, l, and m are confocal images showing contrasting patterns of AGT peroxisomal localization in the koala (P. cinereus) and greater horseshoe bat (R. ferrumequinum), respectively. For each species, localization is shown for AGT with and without the PTS1 (left and right, respectively), revealing a strong effect of PTS1 removal in the former species and negligible effect in the latter. AGT, mitochondria, and peroxisomes are labeled in magenta, cyan, and green, respectively. All experiments were repeated independently three times with similar results. h–k display plot profiles of fluorescence intensity along dashed lines indicated in (f, g, l, and m), with arrows marking regions of colocalization. Y-axis represents pixel intensity in arbitrary units (a.u.). Scale bars: 10 μm in merged images and 1 μm in magnified views. Dietary data were obtained from the PHYLACINE 1.2.1 database. Source data is provided as a Source Data file.
To investigate the relationship between the AGT subcellular targeting quantified from our cell-based assays with published data on diet, we ran phylogenetic Bayesian mixed models (MCMCglmm)33 and found contrasting effects for peroxisomal and mitochondrial localization. In a model of peroxisomal targeting efficiency as a function of dietary plant content, diet had a significant positive effect (posterior mean = 0.28, 95% CI = 0.110–0.430, pMCMC = 0.0016, Supplementary Table 1), consistent with enhanced peroxisomal localization in herbivorous species. An equivalent model assessing mitochondrial targeting revealed a negative, but non-significant, association with plant-based diets (posterior mean = –0.12, 95% CI = –0.280 to 0.037, pMCMC = 0.13, Supplementary Table 1). We also ran phylogenetic generalized least squares (PGLS) regressions34 and obtained consistent findings in which herbivory showed a significant positive association with AGT peroxisomal targeting efficiency (R² = 0.30, p = 4.09 × 10−4, Fig. 2b and Supplementary Table 2) but not with mitochondrial targeting efficiency (R² = 0.07, p = 0.11, Supplementary Table 2). In both models, the phylogenetic signal remained strong (Pagel’s λ = 1.00 and 0.97, respectively).
PTS1 is significant in determining the subcellular localization of full-length AGT
To investigate the impact of PTS1 on peroxisomal targeting ability in mammals, we also constructed AGT plasmids lacking PTS1 for our 37 focal species. These plasmids were then used in immunofluorescence assays to assess the subcellular localization of AGT when only the MTS and mature regions were present. The experimental procedure followed a similar protocol to that described earlier. By comparing the results with previously obtained peroxisomal targeting efficiencies, we observed that the peroxisomal targeting efficiency of AGT in the absence of PTS1 was reduced by over 50% in most species, with the most marked decreases seen in herbivorous mammals, ranging from 73.16% in the strict herbivorous koalas to 8.96% in omnivorous olive baboons. In contrast, carnivorous mammals maintained low peroxisomal targeting efficiencies, with the greater horseshoe bats showing almost no change (3.71%) in efficiency (Fig. 2a and Supplementary Fig. 4). Closer examination of these strongly contrasting species showed that, following removal of PTS1, the peak intensity of AGT in koalas no longer overlapped with that of peroxisomes (Fig. 2g/j), whereas AGT in the greater horseshoe bat retained a wildtype condition of mitochondrial localization (Fig. 2k/l).
We modeled the reduction in peroxisomal targeting efficiency following PTS1 removal as a function of plant tissue in the diet, and found a significant positive association after controlling for phylogeny (posterior mean = 0.20, 95% CI = 0.091–0.310, pMCMC = 0.0014, Supplementary Table 1). This result was further supported using a PGLS regression (R² = 0.41, p = 1.88 × 10−5, Fig. 2c and Supplementary Table 2) with strong phylogenetic signal (Pagel’s λ = 0.83). These results suggest herbivorous species experience a greater loss in targeting efficiency when PTS1 is disrupted.
Because variation in peroxisomal localization among species likely reflects a balance between the relative efficiencies of PTS1 and MTS targeting, we also tested whether the greater targeting in herbivores over carnivores was mainly attributable to their PTS1 sequences, or might instead also reflect adaptive decreases in MTS targeting efficiency, as previously suggested for anthropoid lineages24. We therefore generated AGT constructs lacking the MTS for all focal species and compared their subcellular localization via immunofluorescence assays with those of our first experiment based on vectors containing full AGT sequences. Our results revealed that deletion of the MTS led to substantial reductions of mitochondrial targeting efficiency—over 35% in most mammalian species—ranging from a 31.67% decrease in the crested porcupine (Hystrix cristata) to 71.47% in the African hunting dog (Lycaon pictus) (Supplementary Figs. 3, 5, and 6). Thus, the MTS contributes significantly to mitochondrial localization of AGT, with particularly pronounced effects in members of Laurasiatheria. However, this was not universally the case. For example, in the olive baboon and rhesus monkey, removal of the MTS resulted in only modest (~10%) reduction in mitochondrial localization, suggesting a reduced role for MTS in mitochondrial targeting without a compensatory enhancement of PTS1 function. Together, these findings indicate that the PTS1 motif retains the capacity to direct AGT to peroxisomes even in the presence of a functional MTS, so contradicting earlier suggestions that PTS1 becomes functionally relevant only when the MTS is absent or compromised21.
Tests of molecular adaptation
To investigate whether positive selection has acted on the AGXT—particularly within the MTS and PTS1 regions that govern subcellular localization—we analyzed separate datasets corresponding to the MTS, PTS1, and the mature region excluding PTS1 across all mammals as well within major superorders. For each dataset, we estimated rates of synonymous and nonsynonymous substitutions (dS and dN, respectively) and the dN/dS ratio ω for all branches on the tree under the free-ratio model.
Across all mammals, we recorded ω values > 1 in 642 out of 854 branches for PTS1 (Supplementary Fig. 7), compared to 465 branches for the MTS region, and 55 branches for the mature region without PTS1. Pairwise ω estimates using the modified Nei-Gojobori method35 further supported these results: 62.38% of pairwise comparisons in the PTS1 region showed dN > dS, compared to 12.90% in the MTS region, and none in the mature region without PTS1 (Supplementary Fig. 7). These findings suggest that the PTS1 region of AGXT is subject to more positive selection than both the MTS and the rest of the mature protein sequence. These trends were also seen when analyzing separate superorders and 37 species in experiments with the PTS1 region showing the strongest selection pressure compared to the MTS and the rest of the mature protein sequence in all groups except Monotremata (Fig. 2e/d and Supplementary Fig. 7).
Branch-site tests with herbivore lineages marked in Supplementary Fig. 8 as the foreground revealed no evidence for positive selection when applied across mammals as a whole (ω₂ = 1.0; LRT p = 0.92, model A in Supplementary Data 2), nor within Afrotheria or Metatheria (ω₂ ≈ 1.0–1.3; LRT p ≥ 0.38, model G/I in Supplementary Data 2). In contrast, analyses restricted to Laurasiatheria and Euarchontoglires detected a strongly supported class of codons with ω > 1 on herbivore branches (ω₂ ≈ 33 and 32, respectively; LRT p = 0.001 in both cases, model C/E in Supplementary Data 2). The inferred proportion of such sites was extremely small (p2a + p2b ≈ 0, model C/E in Supplementary Data 2), indicating that adaptive changes are highly localised. BEB analysis identified a handful of candidate codons in these two clades, including 215 I and 286 T in Laurasiatheria (PP ≥ 0.95, model C in Supplementary Data 2), and 20 V and 286 T in Euarchontoglires (PP > 0.8, model E in Supplementary Data 2), whereas no codons had supported ω > 1 in Afrotheria or Metatheria.
Dietary adaptation is associated with alternative AGT transcription start site usage
To investigate whether dietary strategies influence the transcriptional regulation of AGT in mammals, we analyzed RNA-sequencing (RNA-seq) data from normal liver tissue across 172 mammalian species. We assessed usage of upstream transcription start sites by quantifying the ratio of read coverage of the upstream versus the downstream translation start site (ATG1 and ATG2 in Fig. 3b, respectively) and found differences between species with plant- versus meat-based diets. Among the 148 species with a complete AGT sequence, 75 species exhibited a low upstream-to-downstream start site usage ratio (ATG1/ATG2 ≤ 0.5, Fig. 3a), including 14 carnivorous and 51 herbivorous species (including 12 hypercarnivores and 37 strict herbivores). In contrast, 73 species showed higher upstream usage (ATG1/ATG2 > 0.5, Fig. 3a), comprising 50 carnivorous and 18 herbivorous species (including 47 hypercarnivores and 9 strict herbivores). An additional 24 species lacked a complete MTS but retained the mature AGT region and the PTS1 motif. Of these, five herbivorous species had an ATG1/ATG2 ratio > 0.5, while the remaining 19 species (3 carnivorous and 15 herbivorous, including 1 hypercarnivore and 14 strict herbivores) had lower ratios (≤ 0.5, Fig. 3a).
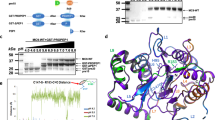
a ATG1/ATG2 ratios obtained from RNA-seq coverage at the AGT translation start sites across 172 mammal species. The average coverage of the upstream (ATG1) and downstream (ATG2) translation start sites was obtained from liver RNA-seq data of 148 mammals. The ATG1/ATG2 ratio reflects the relative usage of the upstream transcription start site, where a lower ratio indicates reduced usage of the upstream start site. Standard error bars are shown for species with multiple biological replicates. Individual data points are overlaid on the bars. The outer ring shows the proportion of plant tissue in the diet. b Schematic representation of the upstream (ATG1) and downstream (ATG2) translation start sites within the ATG encoding sequence. c PGLS regression of the ATG1/ATG2 expression ratio against the proportion of plant material in the diet, based on species with complete AGT sequences. P values were calculated using two-sided t-tests. d RNA-seq analysis, ChIP-seq analysis, and ATAC-seq analysis of liver tissue from 7 mammal species, showing peaks covering the region within ±250 bp around the translation start site of AGT gene. Dietary data were obtained from the PHYLACINE 1.2.1 database. Source data is provided as a Source Data file.
For species with an intact MTS, we again ran a Bayesian mixed model to examine upstream AGT transcription start site usage as a function of dietary plant content. A significant negative association (posterior mean = –0.29, 95% CI = −0.420 to −0.110, pMCMC = 0.0014, Supplementary Table 1) indicates that herbivorous species are more likely to initiate AGT transcription from the downstream transcription start site, bypassing the upstream start codon that includes the mitochondrial targeting signal. A significant negative relationship between the ATG1/ATG2 ratio and dietary plant content was also supported by the results of PGLS analysis (R² = 0.11, p = 3.53 × 10−5, Fig. 3c and Supplementary Table 2) with some remaining phylogenetic signal (Pagel’s λ = 0.60). These findings independently corroborate the results of the MCMCglmm model and further support the link between diet and alternative AGT transcriptional regulation, with plant-rich diets associated with reduced usage of the upstream transcript isoform.
Variation in promoter activity across species
We hypothesised that herbivory is associated with reduced usage of the upstream AGXT promoter (ATG1) and compensatory activation of a downstream start region, which should manifest as weaker upstream promoter marks/accessibility and lower upstream promoter activity. To probe the regulatory basis of the start-site shift, we examined liver ChIP-seq profiles for active promoter marks (H3K4me3, H3K27ac)36 across 13 species (Fig. 3d and Supplementary Fig. 9). In multiple herbivores, enrichment was weaker upstream of ATG1 and comparatively stronger near the downstream start region, whereas dog (Carnivora) showed extended upstream signal—patterns consistent with reduced upstream promoter activity in herbivores and greater upstream activity in some non-herbivores. Consistent with these profiles, RNA-seq coverage over the AGXT start-site region was higher in the dog than in the herbivores within this subset (Fig. 3d).
We then assessed chromatin accessibility using liver ATAC-seq (subset of species in Fig. 3d). Herbivores generally showed lower accessibility upstream of ATG1 with more open chromatin nearer the downstream start region, mirroring the ChIP-seq profiles, while the dog showed accessible chromatin extending upstream of ATG1. Taken together, concordant shifts in active promoter marks, chromatin accessibility, and subset RNA-seq coverage support a diet-associated reduction in upstream promoter activity in herbivores, providing a mechanistic link to the ATG1/ATG2 usage patterns described above.
To test whether upstream promoter sequence contributes to these differences, we edited the upstream start codon (ATG1) within the promoter fragment cloned upstream of luciferase (single base-triplet substitution ATG → GCG at ATG1; +1..+3 relative to ATG1). The effect of this mutation on AGT promoter activity was then quantified using dual-luciferase reporter assays across 19 mammalian species. The selection of these species was informed by prior immunofluorescence colocalization data, the availability of complete AGT sequences, and the presence of a functional MTS. When comparing across mammals, we observed no clear association between diet and the magnitude of difference in luciferase activity between ATG1-edited and corresponding wildtype binding sites (p = 0.16, Supplementary Fig. 10 and Supplementary Table 2). However, within bats, we recorded lower impacts of ATG1 substitution on promoter activity in the insectivorous species Rhinolophus ferrumequinum and Myotis myotis, and in the blood-feeding Desmodus rotundus, compared to in plant-feeding bats (Supplementary Fig. 10). Striking exceptions were also seen, notably in the donkey (Equus asinus) in which the mutant construct displayed a strong increase in promoter activity.
Finally, we asked whether transcription factor (TF) binding architecture differs among lineages. Using motif prediction in the upstream region for the species with matched RNA-seq/ChIP-seq/ATAC-seq, canonical liver TF motifs (e.g., HNF1A, C/EBP) were conserved in position. By contrast, we observed positional shifts for ZNF454 and VAX2: in ungulates, the predicted ZNF454 motif overlaps the ATG1 codon, whereas in dog and in both rodents it lies downstream; VAX2 shows the opposite pattern (upstream in dog and rodents, downstream in ungulates) (Supplementary Fig. 11). Given that Carnivora and Euungulata are sister clades within Laurasiatheria, these shifts are consistent with lineage-specific regulatory remodeling on the branch leading to ungulates. Functional validation (e.g., TF ChIP or targeted reporter mutagenesis of the predicted sites) will be required to establish causality for liver regulation of AGXT.
Discussion
We combined immunofluorescence assays of intact and mutant proteins with phylogenetics models and analyses of transcriptome data to examine AGT targeting efficiency in relation to diet across mammals. Our results reveal a significant positive correlation between peroxisomal localization and plant-rich diets, consistent with findings from birds and select mammalian groups. While previous studies emphasized the dominant role of the MTS and suggested PTS1 is secondary or redundant, we show that PTS1 can drive peroxisomal targeting even when a functional MTS is present.
AGT evolution with PTS1 and MTS during mammal dietary diversification
Comparison of AGT sequences from 498 mammals revealed divergent evolutionary trajectories with some lineages having retained gene function, and others showing evidence of accruing loss-of-function mutations in either the mature AGXT or its mitochondrial targeting region. The finding that the functional MTS region spans 22 amino acids in all eutherian species examined, with the exception of the Middle East blind mole-rat Nannospalax galili (20 amino acids), is consistent with earlier observations from much smaller datasets18,37. At the same time, we found that the same region spans 33 amino acids in both prototherians examined (duck-billed platypus and short-beaked echidna) as well as in the 15 species of marsupial (Metatheria). Interestingly, although not our focus, we also examined MTS length in publicly-available AGT sequences of other vertebrate groups and find a similar 33-amino-acid MTS region in divergent reptiles such as the estuarine crocodile (Crocodylus porosus) and painted turtle (Chrysemys picta). We thus speculate that the 33-amino-acid MTS was present at the common ancestor of Amniotes, and was retained in non-eutherian mammals but evolved to be shorter in the ancestor of Eutheria, with subsequent lineage-specific modifications.
Our results add to the number of mammals known to have lost their MTS function, with losses arising via a range of mutations. In total, 21 species were seen to harbor a non-canonical start codon, six species exhibited premature stop codons, four had frameshift mutations, and 13 species showed combinations of these defects. Notably, of the 43 species lacking a functional MTS, five are carnivorous or omnivorous (e.g., Iberian mole, Talpa occidentalis; Desmarest’s hutia, Capromys pilorides), while the majority (35 species) are herbivorous. Herbivorous species include the first documented cases of MTS losses in Metatheria, with the tammar wallaby (Notamacropus eugenii), Leadbeater’s possum (Gymnobelideus leadbeateri), and common wombat (Vombatus ursinus), each found have undergone an independent frameshift mutation, two resulting in loss of the upstream translation site. Marsupials appear to have evolved from a putative insectivorous common ancestor38, thus further implying adaptive loss linked to diversification in diet. Additional cases of MTS loss arising from missense mutations at the upstream translation start site were seen in diverse species, including bats (e.g., Rousettus aegyptiacus) and primates (e.g., Cercopithecus mona). We also identified several instances of nonsense mutations causing MTS dysfunction in lagomorphs and rodents, including a shared premature stop codon (TGA) at positions 43–45 in the MTS of five squirrel species, and a different stop codon (TAA) at the same site in the dormouse (Glis glis). Similarly, a premature stop codon (TGA) was observed at positions 55–57 in the MTS of the white-faced saki (Pithecia pithecia), which can also be found in nutrias (Myocastor coypus).
Our reconstruction of PTS1 motifs across mammals revealed repeated, convergent evolution of canonical targeting sequences. Many cases—such as SKL in artiodactyls (Giraffa camelopardalis, Bos taurus, Ovis aries) and the koala (Phascolarctos cinereus), GKL in fruit bats (Noctilio leporinus) and sloths (Bradypus torquatus), and SRL in the dama gazelle (Nanger dama), paca (Cuniculus paca) and mouse-deer (Tragulus javanicus)—occurred in herbivores. However, convergence was not limited to a single dietary group. Carnivorans such as the dog (Canis lupus familiaris) and banded mongoose (Mungos mungo) independently evolved HKL motifs, and rarer variants such as GRL (e.g., Crocidura indochinensis, Macroglossus sobrinus), HRL (Pedetes capensis, Dolichotis patagonum), and SKM (Potos flavus, Erinaceus europaeus) appeared across diverse lineages. These patterns suggest that while herbivory may drive some convergence in peroxisomal targeting, broader lineage-specific constraints are also influential. In contrast, birds appear to have retained a consensus motif that supports efficient targeting22, consistent with the proposed dietary shift from carnivory to herbivory during the transition from non-avian dinosaurs to birds39,40. However, AGT in Afrotheria and Metatheria was observed to lack significant selection signals in our nested models (Supplementary Table 2), possibly because these herbivorous lineages originated much earlier, around 60–70 million years ago41,42. Over such a long period, the initial selection signals from their dietary transitions may have been masked, whereas stronger selection signals remain detectable in herbivores of Laurasiatheria and Euarchontoglires, reflecting their later dietary shifts.
Impacts of PTS1 and MTS on AGT targeting
Immunofluorescence assays conducted on AGT proteins of 37 species, all with complete MTS regions, revealed wide variation in subcellular localization. Peroxisomal targeting efficiency was significantly positively associated with the proportion of plant matter in the diet, whereas mitochondrial targeting was not significantly explained by diet. Together, these results support a shift in AGT localization away from mitochondria and toward peroxisomes in herbivorous mammals, reflecting a potential convergent adaptation to increased glyoxylate production from plant-based diets. Notably, several herbivores showed strong peroxisomal routing despite intact MTS, consistent with an active contribution of PTS1.
When comparing the degree of peroxisomal targeting in cells expressing AGT proteins with and without the PTS1 motif, we observed clear differences among species. Overall, species with more plant-rich diets exhibit significantly greater decreases in peroxisomal targeting efficiency when the PTS1 motif is removed, indicating a stronger dependence on peroxisomal localization in herbivores, and a dominant role of the PTS1 in spite of an intact MTS. Indeed, marked reductions in peroxisomal targeting were seen in the sheep, orangutan, degu, and, most dramatically, the koala, consistent with the latter species’ need for effective peroxisomal localisation given its specialized and chemically-challenging diet of eucalyptus leaves43,44. Such dietary specialization may have driven tighter metabolic regulation, reinforcing the need for efficient peroxisomal targeting and enhancing reliance on the PTS1 pathway. Interestingly, for two species—the rhesus monkey and olive baboon—we observed high peroxisomal localisation even after removal of the PTS1, suggesting that additional unknown mechanisms of targeting might also exist. Furthermore, several species with the same PTS1 motif (e.g., the horse and cat) showed different degrees of loss in targeting efficiency following removal of PTS1, suggesting possible epistatic interactions with other variable parts of the AGT protein. In contrast to most herbivores, we detected much smaller reductions in peroxisomal targeting following removal of the PTS1 for species with animal-based diets, with almost no change seen in divergent taxa such as insectivorous bats, meerkat, Sunda slow loris, and short-beaked echidna.
AGT peroxisomal targeting via transcriptional regulation
In addition to an effect of the PTS1 and MTS, we also found that AGXT in herbivores tends to be transcribed from downstream transcription start sites, facilitating more effective peroxisomal targeting by avoiding translation from upstream start codons associated with mitochondrial localization. This dietary association with alternative transcription start site usage was supported by phylogenetic models, and aligns with shifts in AGT localization observed in our functional assays.
Regulatory differences in transcription start site usage were further supported by histone modification patterns. Across the species subset with epigenomic data, active promoter marks (H3K27ac and H3K4me3) extended further upstream of the upstream start region (ATG1) in non-herbivores such as dog, whereas in herbivores enrichment was comparatively concentrated nearer the downstream start region. This pattern is consistent with the presence of an extended upstream promoter region in some non-herbivores, potentially reflecting divergent regulatory architecture associated with dietary specialization. Although functional validation via luciferase assays revealed no clear association between promoter activity and diet across mammals, we observed that editing the conserved sequence around the upstream start site significantly reduced AGT promoter activity in some carnivorous species, including in three divergent bats that appear to have evolved similar patterns of promoter regulation. Together, these observations indicate that shifts in transcriptional architecture can complement changes in targeting signals, with clade-specific solutions.
Previous studies linking AGT subcellular targeting to alternative transcription or translation initiation have largely focused on cases involving loss of canonical start sites25 or human mutations that result in mistargeting45. Reports of differential AGT localization arising from alternative start site usage in otherwise intact proteins with a functional mitochondrial targeting signal (MTS) have been limited to just a few species26,37. Our results, however, suggest that transcriptional regulation of AGT targeting via changes in the promoter is likely to be much more widespread across mammals than previously recognized, and might represent a common and evolutionarily-flexible mechanism facilitating subcellular relocalization in response to ecological shifts in diet.
Interplay between multiple mechanisms
Insights into the potential interplay between transcriptional regulation and AGT targeting signals come from bats and carnivorans, both of which include species that have evolved plant-feeding. Based on our cell-based assays, we observed that plant-feeding members of both groups exhibit low peroxisomal targeting efficiency, suggesting that their MTS regions have retained strong mitochondrial targeting capability, possibly reflecting an evolutionary legacy. Yet, at the same time, in bats we observed differential usage of alternative transcription start sites, with animal-feeding bats tending to initiate transcription upstream, but plant-feeding bats favoring downstream initiation, thus improving peroxisomal targeting consistent with their need to metabolize glyoxylate. Interestingly, a similar change in start site usage was not seen in plant-feeding or omnivorous carnivorans, such as the giant and red pandas, possibly reflecting their more recent switches to plant-feeding. For example, fossil data suggest giant pandas fully switched to a bamboo diet just 2 million years ago46, with stable isotope analyses implying this switch could be as recent as a few thousand years ago47. Both scenarios imply giant pandas might retain metabolic adaptations for meat eating, including strong mitochondrial targeting with insufficient time for selection to act on sites underlying transcriptional regulation.
In some species, transcription start site usage appears to complement targeting signals. For example, in two primate species, the rhesus monkey and olive baboon, we recorded a strong preference for using downstream transcription start sites in AGXT, alongside weak MTS-directed mitochondrial targeting and strong PTS1-directed peroxisome targeting. Moreover, comparisons of proteins with and without the non-consensus PTS1 motif in these taxa revealed little change, suggesting other unidentified residues may also contribute to peroxisomal targeting, as previously reported for humans31. Together, these results highlight how subcellular targeting can remain robust through alternative mechanisms, even when classical targeting signals are lost.
A key finding of our study is that strong peroxisomal targeting of AGT mediated by PTS1 can often override mitochondrial signals even when the MTS is intact, and that this pattern is seen in several herbivorous species. To explain this, we propose three potential mechanisms. First, AGT may directly interact with the receptor PEX5 in the cytoplasm via its structural domains, allowing peroxisomal import, as previously reported in humans29. Second, AGT may transiently enter mitochondria before being released and redirected to peroxisomes. Third, AGT could be processed within mitochondria—where the N-terminal sequence is cleaved—before being imported into peroxisomes through PEX5-mediated recognition. Similar dual-targeting mechanisms have been observed in other metabolic enzymes48,49. These possibilities highlight the flexibility of subcellular targeting and open new directions for investigating AGT’s transport dynamics in diverse species.
Methods
Genome data and gene identification
Genome assemblies for 685 mammal species were obtained from the National Center for Biotechnology Information (NCBI; https://www.ncbi.nlm.nih.gov/, accessed February 2, 2023). Details of the genome assemblies used in this study can be found in the supplementary table (Supplementary Data 1). To identify the gene encoding AGT, we performed tBLASTn searches using the mouse (Mus musculus) AGT protein sequence (NCBI accession number: NC_000067.7) as a query. Coding sequences for each mammal species were determined using GeneWise50 and were manually verified. Mutations in the 5′ translation start site, stop codons, or frame shifts in the mitochondrial targeting signal (MTS) region were considered indicative of a nonfunctional MTS. The genetic identification results are provided in the supplementary table (Supplementary Data 1). The phylogenetic tree topology used in this study was derived from previous research51,52,53,54.
Assessment of diet composition
For all mammals in our study, we obtained published dietary information from the PHYLACINE 1.2.1 database55,56. For each species, we calculated the proportion of animal tissue (vertebrate prey and/or invertebrate prey) in the diet. We classified species with diets of ≥70% animal tissue as carnivores, 70%–30% as omnivores, and ≤30% as herbivores57. Within these categories, we also defined species with diets of ≥90% animal tissue as hypercarnivores, and ≤5% as strict herbivores (see Supplementary Data 1).
Plasmid construction for assessing AGT mitochondrial and peroxisomal targeting efficiency
To assess AGT mitochondrial and peroxisome targeting efficiency, we performed cell-based immunofluorescence assays on natural AGT orthologues as well as corresponding mutants that lacked either the MTS or the PTS1. For these assays, we selected 37 mammal species across 11 orders, representing major branches of all three mammalian subclasses. All focal species were selected on the basis of having an available complete AGXT sequence containing an intact MTS, as well as available published immunoelectron microscopy data18.
To establish normal targeting, complete orthologues containing both the MTS and PTS1 were synthesized and inserted into the multiple cloning site of the pMSCV-puro vector. Next, to assess MTS- and PTS-mediated targeting, we generated constructs containing either the corresponding non-MTS AGT fused to a N-terminal 3 × FLAG-tag peptide, or the non-PTS1 AGT fused to a C-terminal 3 × FLAG-tag peptide. Oligonucleotide sequences for these experiments are provided in Supplementary Data 3. The 3 × FLAG-tag peptide enables confirmation of the subcellular localization of AGT in subsequent experiments using specific antibodies, and was chosen because its small size and hydrophilic properties allow for normal protein folding and biological activity58,59. Both constructs were produced with Phusion Plus DNA Polymerase (Thermo Scientific, USA, Cat#F630S) and inserted into the pMSCV-puro vector.
We also constructed plasmids for establishing AGT co-localization with mitochondria and peroxisomes. Here we transfected our cells with a retroviral vector containing either the mito-DsRed (a mitochondrial marker), so providing stable expression of red fluorescence, or the consensus tripeptide sequence of the PTS1 (SKL) fused to a green fluorescent protein (GFP), so providing stable expression of green fluorescence. Finally, because we could not record from both stable fluorescent markers in the same cells, for establishing co-localization of intact AGT with peroxisomes, we also labelled background peroxisomes in the cell by using a pEGFP-N1 vector (MiaoLingPlasmid, China, Cat#P0133) in which we fused the consensus tripeptide sequence of the PTS1 (SKL) to the GFP. Oligonucleotide sequences for these experiments are provided in Supplementary Data 3.
In comparisons of peroxisomal targeting efficiency across taxa, the results from the AGT of human and African clawed toad (Xenopus laevis) served as positive and negative controls, respectively. The human AGT protein lacks a functional MTS start codon and has been shown to exclusively target the peroxisome27,60, whereas the Xenopus protein exclusively targets the mitochondria61.
In vitro cell culture and transfection
We transiently transfected HeLa cells (CCTCC, China, Cat#GDC0009), which do not naturally express AGT, with plasmids containing full-length AGT sequences. HeLa cells were cultured in Dulbecco’s Modified Eagle’s Medium (Gibco, USA, Cat#C11995500BT) supplemented with 10% fetal bovine serum (FBS; Excell Bio, China, Cat#FSP500) and 1% penicillin/streptomycin (Gibco, USA, Cat#15140122) at 37 °C with 5% CO2. To facilitate subsequent experiments, circular slides were pretreated with Hanks’ balanced salt solution (Monad, China, Cat#14025-092) and used for cell culture. For transient co-transfection of AGT expression constructs and EGFP-SKL plasmid, we used Lipofectamine 2000 (Thermo Fisher Scientific, USA, Cat#11668019) and Opti-MEM medium (Invitrogen, Carlsbad, CA, USA, Cat#51985-034), following the manufacturer’s instructions.
Immunofluorescence staining for assessing MTS- and PTS1-mediated targeting efficiency
At 24 h post-transfection, the culture medium was removed, and cells were fixed with 4% paraformaldehyde (Aladdin, China, Cat#C104188) for 20 min. To enhance antibody permeability, cells were then incubated with 0.1% Triton ×-100 (BioFroxx, Germany, Cat#1139) in phosphate-buffered saline (PBS; Cytiva, USA, Cat#SH3025601) for 20 min. After permeabilization, cells were blocked with 10% FBS in PBS at room temperature for 1 h. For the subcellular localization of AGT, the cells were incubated with a primary antibody against AGT (polyclonal, Proteintech, China, Cat#22394-1-AP, 1:200) diluted in PBS containing 10% FBS for 1 h at room temperature. Subsequently, the cells were incubated with a goat anti-rabbit IgG (H + L) cross-adsorbed secondary antibody conjugated with Alexa Fluor® 647 (Invitrogen, USA, Cat#A-21244, 1:800) for 1 h at room temperature. Fluorescence visualization was then performed to determine the subcellular localization of AGT. Localization data for both positive and negative controls were consistent with previous immunoelectron microscopy findings61,62, validating our approach.
To assess MTS-mediated mitochondrial targeting efficiency and PTS1-mediated peroxisomal targeting efficiency, cells were first incubated with anti-FLAG rabbit polyclonal antibody (Proteintech, China, Cat#20543-1-AP, 1:400) for 1 h at room temperature. This was followed by incubation with Cy3-conjugated Affinipure Goat Anti-Rabbit IgG (H + L) (Proteintech, China, Cat#SA00009-2, 1:800) and Alexa Fluor® 647-conjugated goat anti-rabbit IgG (H + L) cross-adsorbed secondary antibody (ThermoFisher, USA, Cat#A-21244, 1:800). Each incubation step was followed by three washes with PBS. Finally, slices were mounted with anti-fading fluorescent mounting medium (YEASEN, China, Cat#36307ES08) and examined under a confocal microscope (Leica, TCS SP8).
Confocal microscopy and image processing
Confocal microscopy was performed using a Leica TCS SP8 confocal laser scanning microscope equipped with a high-resolution 63× oil immersion objective. Image acquisition and processing were carried out using LAS AF Lite 2.6 software (Leica Microsystems, Wetzlar, Germany). To quantify colocalization in confocal fluorescence images, we employed the EzColocalization plug-in for ImageJ63. For each cell, the perimeter was manually outlined using the area selection tools, and colocalization measurements were performed for individual cells. For each cell, we calculated the mitochondrial and peroxisomal targeting efficiency of AGT, calculated as the percentage of total pixel intensity for AGT that colocalized with, respectively mitochondrial and peroxisomal fluorescence. Colocalization analysis was performed using Mander’s Colocalization Coefficients, applying an automatic threshold to ensure consistency64,65. To ensure reliable mitochondrial or peroxisomal targeting results, this was repeated for 20–25 cells per species and an average taken. A dashed line was drawn across regions of interest using the Straight Line tool in ImageJ, traversing cellular compartments with distinct fluorescence signals. Fluorescence intensity values along the dashed line were extracted using the Plot Profile function in ImageJ.
Molecular evolutionary analysis
We aligned 429 mammal AGXT orthologues in MUSCLE66, after removing pseudogenes and incomplete gene sequences. To characterize differences in selection pressure across the AGT gene, we built and analyzed separate datasets based on the full AGXT, MTS motif, Mature Region without PTS1 motif, and PTS1 motif. For each dataset, we generated Nei-Gojobori estimates of ω (dN/dS ratio) for each pairwise species comparison using the modified method implemented in MEGA735. An ω value < 1 indicates purifying selection, ω = 1 suggests neutral evolution, and ω > 1 implies positive selection. Next, we also obtained independent ω estimates for each branch in the tree using the free-ratio model in PAML 4.967 based on the accepted species topology51,52,53,54.
To test for positive selection in lineages that feed on plant tissue, we also conducted branch-site models in PAML 4.967. We used Model A (model = 2, NSsites = 2, fix_omega = 0) with herbivore lineages as the foreground and compared it to the corresponding null model in which the ω > 1 class is constrained to ω₂ = 1 (fix_omega = 1, omega = 1). Positive selection on herbivore branches was inferred only when this one-sided likelihood ratio test was significant (p < 0.05) and ω₂ > 1 in the alternative model; BEB posteriors for sites in classes 2a/2b were interpreted only under these conditions. All analyses were repeated for Laurasiatheria (239 species), Euarchontoglires (159 species), Afrotheria (10 species), and Metatheria (12 species). Xenarthra was not analyzed due to insufficient data.
To gain insights into the evolutionary history of herbivory in mammals, we also reconstructed ancestral protein sequences of the PTS1 region using the full dataset. Ancestral state inference was conducted using maximum likelihood under the JTT substitution model in PAML 4.9, based on a species tree derived from recent phylogenomic studies51,52,53,54. For downstream analyses, we used the inferred ancestral sequences with the highest posterior probabilities. Notably, over 95% of sites in the reconstructed sequences had high confidence (posterior probability > 0.95).
Subcellular targeting and diet
We modelled subcellular targeting efficiency (peroxisomal and mitochondrial) of natural and mutant proteins, as quantified from our cell-based immunofluorescence assays, as a function of the proportion of plant tissue in the diet. We applied two approaches that account for statistical non-independence arising from phylogenetic inertia68. First, we ran phylogenetically-controlled Markov chain Monte Carlo generalized linear mixed models33 using the package MCMCglmm33 in Rstudio (v4.4.1). Phylogeny was fitted as a random effect, with the covariance matrix generated in the package ape69. The tree topology and branch lengths were generated using the TimeTree database (http://www.timetree.org/; last accessed April 15, 2025) with manual editing to add missing taxa51,52,53,54. The proportion of plant tissue in the diet was fitted as a fixed effect using data obtained from the PHYLACINE 1.2.1 database, as previously described. Our priors took the form of: prior <-list (G = list (G1 = list (V = 1, nu = 0.002)), R = list (V = 1, nu = 0.002)). The prior distribution for the variance components was assumed to be an inverse-Gamma density, with a small degree of belief (i.e., nu = 0.002). The Heidelberg was used to corroborate convergence of the MCMC runs. MCMCglmm was run with 130,000 iterations, after a burn-in of 30,000 iterations. Values from every 10 iterations were saved.
As a second test approach we used Phylogenetic Generalized Least Squares (PGLS) regressions, which test for phylogenetic autocorrelation in the residuals. PGLS allows for more direct modeling of the evolutionary relationships between species by incorporating a phylogenetic tree and estimating Pagel’s lambda (λ), which quantifies phylogenetic signal in the residuals. We ran PGLS in the R package Caper70, and the significance of each predictor was calculated using two-sided t-tests.
RNA-seq analysis for transcription start site usage
To determine variation in transcription start site usage, we analysed transcripts RNA-seq assemblies of liver tissue obtained for 172 mammal species from the NCBI (https://www.ncbi.nlm.nih.gov/, accessed December 16, 2024). Quality control of the assemblies and trimming of low-quality reads were performed using fastp (v0.23.2) and FastQC (v0.12.1, https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) with default settings to generate high-quality cleaned reads (Chen et al., 2018). Reads from each species were aligned to the respective reference genomes using HISAT2 (v2.2.1)71. Coverage of the AGT region was calculated using Samtools (v1.19)72. For species lacking a reference genome, de novo transcriptomes were assembled using Trinity (v2.9.1)73. These Trinity-assembled transcripts were then used to create databases with makeblastdb in BLAST+ v2.12.0, and the assemblies were compared to these databases using tBLASTn. Transcripts covering the upstream translation start site were considered to originate from the upstream transcription start site, while those covering the downstream translation start site were derived from both upstream and downstream transcription start sites. To quantify the usage of the upstream transcription start site, we calculated the ratio of coverage (ATG1/ATG2), where ATG1 refers to the coverage at the upstream translation start site and ATG2 refers to the coverage at the downstream translation start site. A higher ratio indicates a greater presence of longer AGT transcripts originating from the upstream transcription start site. We assessed ATG1/ATG2 usage as a function of diet using the same phylogenetic modelling approaches as described earlier.
ChIP-seq, ATAC-seq, and regulatory motif analysis
To investigate the regulatory basis of alternative AGT promoter usage, we analyzed publicly-available ChIP-sequencing (ChIP-seq) data from normal liver tissue across 13 mammalian species. These datasets were retrieved from the NCBI (https://www.ncbi.nlm.nih.gov/, accessed January 18, 2025). Data processing followed our RNA-seq pipeline, except that sequencing reads were aligned to each species’ reference genome using Bowtie2 (v2.5.4). Identification of enriched regions (peaks) was performed on two biological replicates per species using MACS2 (v2.2.9.1) with default settings and a p-value cutoff of 0.05 (one-sided Poisson test) for narrow peaks74.
We focused on two histone modifications associated with active promoter activity: H3K27ac and H3K4me336. Species were included if they had ChIP-seq data for two of these histone marks, a complete AGXT model, and a functional mitochondrial targeting signal (MTS). Visualisation of the identified peaks in relation to gene annotations and the underlying ChIP-seq signal, was conducted using the Integrated Genomics Viewer (IGV, v2.8.6)75.
To further assess regulatory architecture, we analyzed publicly available assay for transposase-accessible chromatin sequencing (ATAC-seq) data from normal liver tissue across seven mammalian species (NCBI; accessed 8 September 2025). Data were processed following the ChIP-seq pipeline, and chromatin accessibility patterns near AGXT promoters were visualized using IGV.
Finally, we predicted transcription factor binding motifs across the ~2 kb upstream region (including exon 1) in seven focal species with available RNA-seq, ChIP-seq, and ATAC-seq data. Scans were performed using JASPAR 202476 with a relative score threshold of 80%. Motif presence, orientation, and position relative to the AGXT translation start site (ATG) were compared among species.
Luciferase reporter assay
To evaluate the functional consequences of a specific upstream promoter region—identified via ChIP-seq analysis as exhibiting interspecies variation in activity—we employed a dual-luciferase reporter assay in HeGp2 cells (Procell, China, Cat#CL-0103). We selected 19 species that represented a range of subcellular localisation patterns, prioritising those with available ChIP-seq data.
For each species, a ~ 2 kb DNA fragment upstream of the primary translation start site, including exon 1, was cloned into the pGL3-Basic vector (MiaoLingPlasmid, China, Cat#P0193) to drive firefly luciferase expression. The AGT coding region was not included in the reporter; thus, any AGT triplets in the insert lie in the reporter 5′ UTR and cannot initiate luciferase translation. Reporter plasmids (250 ng) were co-transfected with the pRL-TK Renilla luciferase control plasmid (10 ng, MiaoLingPlasmid, China, Cat#P0372) using Lipofectamine 2000 in Opti-MEM medium, following the manufacturer’s protocol. After 36 h of incubation in medium supplemented with 31.6 mM sodium glyoxylate (Macklin, China, Cat#S923532), luciferase activity was measured using the Dual-Luciferase® Firefly & Renilla Assay Kit (UElandy, China, Cat#F6075M) on a GloMax® 20–20 Luminometer (Promega, USA). Firefly luciferase activity was normalized to Renilla luciferase activity to calculate relative promoter activity77. Each assay was conducted in triplicate and averaged across replicates.
To test the contribution of the upstream start-site region, we introduced a single defined nucleotide substitution at the AGT ATG1 triplet within the promoter insert (overlap-extension PCR): ATG → GCG (coordinates +1..+3 relative to ATG1; A of ATG1 = +1). This triplet lies within a predicted motif overlapping the start-site region. Oligonucleotide sequences used for this experiment are provided in Supplementary Data 3. Wild-type and mutant constructs were transfected and assayed under identical conditions, with luciferase activity measured in triplicate for each construct.
To test whether the effect of the mutation on promoter activity varied with diet, we fitted a phylogenetic generalized least squares (PGLS) model. The response variable was the log-transformed difference in mean luciferase activity between the mutant and wild-type constructs. The predictor was the proportion of plant material in the species’ diet, treated as a continuous variable.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available within the article and its supplementary files. The genomic and transcriptomic data used in our manuscript were obtained from National Center for Biotechnology Information (NCBI) database, and the corresponding accession codes have been provided in the Supplementary Data 1. Plasmids and primers used in this study are available from the corresponding authors upon reasonable request. Any additional requests for information can be directed to and will be fulfilled by the corresponding authors. Source data are provided with this paper.
Code availability
R scripts that were used to analyze datasets and make figures are available in the GitHub repository https://github.com/Hcheni0131/Analysis-in-R.git. and archived on Zenodo78.
References
Price, S. A., Hopkins, S. S. B., Smith, K. K. & Roth, V. L. Tempo of trophic evolution and its impact on mammalian diversification. Proc. Natl. Acad. Sci. USA 109, 7008–7012 (2012).
Gill, P. G. et al. Dietary specializations and diversity in feeding ecology of the earliest stem mammals. Nature 512, 303–305 (2014).
Conith, A. J., Imburgia, M. J., Crosby, A. J. & Dumont, E. R. The functional significance of morphological changes in the dentitions of early mammals. J. R. Soc. Interface 13, 20160713 (2016).
Luo, Z. X. et al. New evidence for mammalia form ear evolution and feeding adaptation in a Jurassic ecosystem. Nature 548, 326–329 (2017).
Grossnickle, D. M., Smith, S. M. & Wilson, G. P. Untangling the multiple ecological radiations of early mammals. Trends. Ecol. Evol. 34, 936–949 (2019).
Freeland, W. J. & Janzen, D. H. Strategies in herbivory by mammals: the role of plant secondary compounds. Am. Nat. 108, 269–289 (1974).
Muegge, B. D. et al. Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science 332, 970–974 (2011).
Weimer, P. J. Degradation of cellulose and hemicellulose by ruminal microorganisms. Microorganisms 10, 2345 (2022).
Pope, P. B. et al. Adaptation to herbivory by the Tammar wallaby includes bacterial and glycoside hydrolase profiles different from other herbivores. Proc. Natl. Acad. Sci. USA 107, 14793–14798 (2010).
Hecker, N., Sharma, V. & Hiller, M. Convergent gene losses illuminate metabolic and physiological changes in herbivores and carnivores. Proc. Natl. Acad. Sci. USA 116, 3036–3041 (2019).
Potter, J. H. T. et al. Nectar-feeding bats and birds show parallel molecular adaptations in sugar metabolism enzymes. Curr. Biol. 31, 4667–4674 (2021).
Iason, G. The role of plant secondary metabolites in mammalian herbivory: ecological perspectives. Proc. Nutr. Soc. 64, 123–131 (2005).
Dearing, M. D. & Weinstein, S. B. Metabolic enabling and detoxification by mammalian gut microbes. Annu. Rev. Microbiol. 76, 579–596 (2022).
Danpure, C. J. Variable peroxisomal and mitochondrial targeting of alanine: glyoxylate aminotransferase in mammalian evolution and disease. Bioessays 19, 317–326 (1997).
Takayama, T. et al. Control of oxalate formation from L-hydroxyproline in liver mitochondria. J. Am. Soc. Nephrol. 14, 939–946 (2003).
Noguchi, T. Amino acid metabolism in animal peroxisomes. In Peroxisomes in Biology and Medicine (eds Fahimi, H. D. & Sies, H.) (Springer, 1987).
Danpure, C. J. & Jennings, P. R. Peroxisomal alanine:glyoxylate aminotransferase deficiency in primary hyperoxaluria type I. FEBS. Lett. 201, 20–24 (1986).
Birdsey, G. M. et al. A comparative analysis of the evolutionary relationship between diet and enzyme targeting in bats, marsupials and other mammals. Proc. Biol. Sci. 272, 833–840 (2005).
Birdsey, G. M. et al. Differential enzyme targeting as an evolutionary adaptation to herbivory in Carnivora. Mol. Biol. Evol. 21, 632–646 (2004).
Lumb, M. J., Purdue, P. E. & Danpure, C. J. Molecular evolution of alanine/glyoxylate aminotransferase 1 intracellular targeting - Analysis of the feline gene. Eur. J. Biochem. 221, 53–62 (1994).
Oatey, P. B., Lumb, M. J. & Danpure, C. J. Molecular basis of the variable peroxisomal and mitochondrial targeting of alanine:glyoxylate aminotransferase. Mol. Biol. Cell. 7, 2878–2878 (1996).
Wang, B. J. et al. Diet and adaptive evolution of alanine-glyoxylate aminotransferase mitochondrial targeting in birds. Mol. Biol. Evol. 37, 786–798 (2020).
Liu, Y. et al. Multiple adaptive losses of alanine-glyoxylate aminotransferase mitochondrial targeting in fruit-eating bats. Mol. Biol. Evol. 29, 1507–1511 (2012).
Holbrook, J. D. et al. Molecular adaptation of alanine: glyoxylate aminotransferase targeting in primates. Mol. Biol. Evol. 17, 387–400 (2000).
Birdsey, G. M. & Danpure, C. J. Evolution of alanine: glyoxylate aminotransferase intracellular targeting: structural and functional analysis of the guinea pig gene. Biochem. J. 331, 49–60 (1998).
Purdue, P. E., Lumb, M. J. & Danpure, C. J. Molecular evolution of alanine/glyoxylate aminotransferase 1 intracellular targeting - Analysis of the marmoset and rabbit genes. Eur. J. Biochem. 207, 757–766 (1992).
Takada, Y. et al. Human peroxisomal l-alanine: glyoxylate aminotransferase. Evolutionary loss of a mitochondrial targeting signal by point mutation of the initiation codon. Biochem. J. 268, 517–520 (1990).
Mesa-Torres, N. et al. Molecular recognition of PTS-1 cargo proteins by Pex5p: implications for protein mistargeting in primary hyperoxaluria. Biomolecules 5, 121–141 (2015).
Huber, P. A. J. et al. Peroxisomal import of human alanine: glyoxylate aminotransferase requires ancillary targeting information remote from its C terminus. J. Biol. Chem. 280, 27111–27120 (2005).
Motley, A., Hettema, E., Distel, B. & Tabak, H. Differential protein import deficiencies in human peroxisome assembly disorders. J. Cell. Biol. 125, 755–767 (1994).
Oatey, P. B., Lumb, M. J., Jennings, P. R. & Danpure, C. J. Context dependency of the PTS1 motif in human alanine:glyoxylate aminotransferase 1. Ann. N. Y. Acad. Sci. 804, 652–653 (1996).
Altschul, S. F. et al. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Hadfield, J. D. MCMC methods for multi-response generalized linear mixed models: the MCMCglmm R package. J. Stat. Softw. 33, 1–22 (2010).
Grafen, A. The phylogenetic regression. Philos. T. R.Sco. B. 326, 119–157 (1989).
Zhang, J. Z., Rosenberg, H. F. & Nei, M. Positive Darwinian selection after gene duplication in primate ribonuclease genes. Proc. Natl. Acad. Sci. USA 95, 3708–3713 (1998).
Shlyueva, D., Stampfel, G. & Stark, A. Transcriptional enhancers: from properties to genome-wide predictions. Nat. Rev. Genet. 15, 272–286 (2014).
Oda, T., Miyajima, H., Suzuki, Y. & Ichiyama, A. Nucleotide sequence of the cDNA encoding the precursor for mitochondrial serine: pyruvate aminotransferase of rat liver. Eur. J. Biochem. 168, 537–542 (1987).
Amador, L. I. & Giannini, N. P. Evolution of diet in extant marsupials: emergent patterns from a broad phylogenetic perspective. Mammal Rev. 51, 178–192 (2021).
Li, Z. H. et al. Ultramicrostructural reductions in teeth: implications for dietary transition from non-avian dinosaurs to birds. BMC Evol. Biol. 20, 46 (2020).
Larson, D. W., Brown, C. M. & Evans, D. C. Dental disparity and ecological stability in bird-like dinosaurs prior to the end-Cretaceous mass extinction. Curr. Biol. 26, 1325–1333 (2016).
Tian, R. et al. Sirenian genomes illuminate the evolution of fully aquatic species within the mammalian superorder Afrotheria. Nat. Commun. 15, 5568 (2024).
Meredith, R. W. et al. Impacts of the Cretaceous Terrestrial Revolution and KPg extinction on mammal diversification. Science 334, 521–524 (2011).
Alfano, N. et al. Variation in koala microbiomes within and between individuals: effect of body region and captivity status. Sci. Rep. 5, 1–12 (2015).
Cork, S. J., Hume, I. D. & Dawson, T. J. Digestion and metabolism of a natural foliar diet (Eucalyptus punctata) by an arboreal marsupial, the koala (Phascolarctos cinereus). J. Comp. Physiol. 153, 181–190 (1983).
Danpure, C. J. Primary hyperoxaluria type 1: AGT mistargeting highlights the fundamental differences between the peroxisomal and mitochondrial protein import pathways. Biochim. Biophys. Acta, Mol. Cell Res. 1763, 1776–1784 (2006).
Jin, C. Z. et al. The first skull of the earliest giant panda. Proc. Natl. Acad. Sci. USA 104, 10932–10937 (2007).
Han, H. et al. Diet evolution and habitat contraction of giant pandas via stable isotope analysis. Curr. Biol. 29, 664–669 (2019).
Stehlik, T. et al. Peroxisomal targeting of a protein phosphatase type 2C via mitochondrial transit. Nat. Commun. 11, 2355 (2020).
Stein, I., Peleg, Y., Evenram, S. & Pines, O. The single translation product of the FUM1 gene (Fumarase) is processed in mitochondria before being distributed between the cytosol and mitochondria in Saccharomyces cerevisiae. Mol. Cell. Biol. 14, 4770–4778 (1994).
Birney, E., Clamp, M. & Durbin, R. GeneWise and genomewise. Genome Res. 14, 988–995 (2004).
Alvarez-Carretero, S. et al. A species-level timeline of mammal evolution integrating phylogenomic data. Nature 602, 263–267 (2022).
Deakin, J. E. & O’Neill, R. J. Evolution of marsupial genomes. Ann. Rev. Anim. Biosci. 8, 25–45 (2020).
Foley, N. M. et al. A genomic timescale for placental mammal evolution. Science 380, eabl8189 (2023).
Hao, X. Y., Lu, Q. & Zhao, H. B. A molecular phylogeny for all 21 families within Chiroptera (bats). Integr. Zool. 19, 989–998 (2024).
Faurby, S. et al. PHYLACINE 1.2: the phylogenetic atlas of mammal macroecology. Ecology 99, 2626–2626 (2018).
Faurby, S. et al. PHYLACINE 1.2.1: an update to the phylogenetic atlas of mammal macroecology. https://doi.org/10.5281/zenodo.3690867 (2020).
Atwood, T. B. et al. Herbivores at the highest risk of extinction among mammals, birds, and reptiles. Sci. Adv. 6, eabb8458 (2020).
Delles, C. et al. Urinary proteomic diagnosis of coronary artery disease: identification and clinical validation in 623 individuals. J. Hypertens. 28, 2316–2322 (2010).
Hopp, T. P. et al. A short polypeptide marker sequence useful for recombinant protein identification and purification. Nat. Biotechnol. 6, 1204–1210 (1988).
Noguchi, T. & Takada, Y. Peroxisomal localization of serine:pyruvate aminotransferase in human liver. J. Biol. Chem. 253, 7598–7600 (1978).
Holbrook, J. D. & Danpure, C. J. Molecular basis for the dual mitochondrial and cytosolic localization of Alanine: glyoxylate aminotransferase in amphibian liver cells. J. Biol. Chem. 277, 2336–2344 (2002).
Blattner, J. et al. Glycosome assembly in trypanosomes: variations in the acceptable degeneracy of a COOH-terminal microbody targeting signal. J. Cell. Biol. 119, 1129–1136 (1992).
Stauffer, W., Sheng, H. J. & Lim, H. N. EzColocalization: an ImageJ plugin for visualizing and measuring colocalization in cells and organisms. Sci. Rep. 8, 15764 (2018).
Dunn, K. W., Kamocka, M. M. & McDonald, J. H. A practical guide to evaluating colocalization in biological microscopy. Am. J. Physiol. Cell Physiol. 300, C723–C742 (2011).
Manders, E. M. M., Verbeek, F. J. & Aten, J. A. Measurement of co-localization of objects in dual-colour confocal images. J. Microsc-Oxford. 169, 375–382 (1993).
Edgar, R. C. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinf. 5, 1–19 (2004).
Yang, Z. H. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Felsenstein, J. Phylogenies and the comparative method. Am. Nat. 125, 1–15 (1985).
Paradis, E., Claude, J. & Strimmer, K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004).
Orme, D. et al. The caper package: comparative analysis of phylogenetics and evolution in R. R. Package Version 5, 1–36 (2013).
Kim, D., Landmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods. 12, 357–360 (2015).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Rauluseviciute, I. et al. JASPAR 2024: 20th anniversary of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 52, D174–D182 (2023).
Xu, Y. R. et al. Tankyrases inhibit innate antiviral response by PARylating VISA/MAVS and priming it for RNF146-mediated ubiquitination and degradation. Proc. Natl. Acad. Sci. USA 119, e2122805119 (2022).
Huang, C. Convergent evolutionary shifts in AGT targeting between mitochondria and peroxisomes across mammal transitions to herbivory (Rscript). Zenodo. https://doi.org/10.5281/zenodo.18384691 (2026).
Acknowledgements
This study was supported in part by the Natural Science Foundation of Hubei Province (grant no. 2023AFA015 to H.Z.), National Natural Science Foundation of China (grant no. 32270436 to H.Z.), and Fundamental Research Funds for the Central Universities (grant no. 2042022dx0003 to H.Z.). C.H. was supported in part by the PhD Program for Outstanding and Innovative Talents in Basic Disciplines at Wuhan University to host an overseas collaborator and conduct a visit to the UK. We thank Dr. Yingcan Li and Dr. Yiran Xu for technical assistance with data analyses. We thank Yuanchun. Luo, a PhD student in Wuhan University for confocal microscopy.
Author information
Authors and Affiliations
Contributions
H.Z. conceived and designed the research. C.H. and B.W. designed and performed analyses. C.H. and J.Y. performed the functional experiments. C.H. analyzed data with input from S.J.R.; and C.H., S.J.R., and H.Z. discussed results and wrote the manuscript. All authors have read and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Inge Seim and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, C., Wang, B., Yu, J. et al. Convergent evolutionary shifts in AGT targeting between mitochondria and peroxisomes across mammal transitions to herbivory. Nat Commun 17, 2161 (2026). https://doi.org/10.1038/s41467-026-70246-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70246-0