Abstract
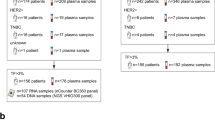
Endocrine therapy (ET) resistance in estrogen receptor positive (ER+) advanced breast cancer is often linked to ESR1 mutations, yet responses to oral selective ER degraders vary within mutant subgroups. Through a biomarker analysis of acelERA Breast Cancer (NCT04576455), we show that tumor ER transcriptional activity as well as circulating tumor DNA (ctDNA) genomics and dynamics effectively stratify response to ET, including giredestrant. We find that following first-line therapy, the ctDNA genomic landscape is diverse and influenced by CDK4/6 inhibitor exposure. Despite this complexity, ER activity in ESR1-mutant tumors remains comparable to early breast cancer but is reduced in most non-mutant cases. This maintained ER activity is associated with giredestrant benefit. Furthermore, early ctDNA clearance identifies responding patients, and the combination of low ER activity and high ctDNA burden predicts rapid clinical progression. These findings provide a framework for personalizing future breast cancer therapies by integrating liquid biopsies with tissue-based signatures.
Similar content being viewed by others
Data availability
The Source Data file contains derived, non-identifying data but does not include individual patient-level clinical records. Anonymized, publication-specific biomarker datasets, minimized to those necessary to reproduce the analyses reported here, may be made available upon request and subject to review and approval by Roche, including execution of a data sharing agreement. For up-to-date details on Roche’s Global Policy on the Sharing of Clinical Information and how to request access to related clinical study documents, see https://go.roche.com/data_sharing. The study protocol is available with the previously published primary analysis (DOI:10.1200/JCO.23.01500). For eligible studies, qualified researchers may request access to individual patient-level clinical trial data underlying the primary study endpoints through a data request platform. At the time of writing, this request platform is Vivli: https://vivli.org/ourmember/roche/. For up-to-date details on Roche’s Global Policy on the Sharing of Clinical Information and how to request access to related clinical study documents, see here: https://go.roche.com/data_sharing. Anonymized records for individual patients across more than one data source external to Roche cannot, and should not, be linked due to a potential increase in risk of patient reidentification. Source data are provided with this paper.
Code availability
Custom code or mathematical algorithms were not developed in this study. All analyses were completed using publicly available R packages (see Methods and Reporting Summary for versions) or using Prism (v10.1.1, see Statistical Methods).
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Giaquinto, A. N. et al. Breast cancer statistics, 2022. CA Cancer J. Clin. 72, 524–541 (2022).
Metcalfe, C., Friedman, L. S. & Hager, J. H. Hormone-targeted therapy and resistance. Annu. Rev. Cancer Biol. 2, 1–22 (2017).
Hanker, A. B., Sudhan, D. R. & Arteaga, C. L. Overcoming endocrine resistance in breast cancer. Cancer Cell 37, 496–513 (2020).
Patel, R., Klein, P., Tiersten, A. & Sparano, J. A. An emerging generation of endocrine therapies in breast cancer: a clinical perspective. npj Breast Cancer 9, 20 (2023).
Will, M., Liang, J., Metcalfe, C. & Chandarlapaty, S. Therapeutic resistance to anti-oestrogen therapy in breast cancer. Nat. Rev. Cancer 23, 673–685 (2023).
Fasching, P. A. et al. Neoadjuvant giredestrant (GDC-9545) plus palbociclib (P) versus anastrozole (A) plus P in postmenopausal women with estrogen receptor–positive, HER2-negative, untreated early breast cancer (ER+/HER2– eBC): Final analysis of the randomized, open-label, international phase 2 coopERA BC study. J. Clin. Oncol. 40, 589–589 (2022).
Sanchez, K. G., Nangia, J. R., Schiff, R. & Rimawi, M. F. Elacestrant and the promise of oral SERDs. J. Clin. Oncol. 40, 3227–3229 (2022).
Chen, Y.-C. et al. Latest generation estrogen receptor degraders for the treatment of hormone receptor-positive breast cancer. Expert Opin. Investig. Drugs 31, 515–529 (2022).
Jeselsohn, R., Buchwalter, G., Angelis, C. D., Brown, M. & Schiff, R. ESR1 mutations—a mechanism for acquired endocrine resistance in breast cancer. Nat. Rev. Clin. Oncol. 12, 573–583 (2015).
Chaudhary, N. et al. CDK4/6i-treated HR+/HER2- breast cancer tumors show higher ESR1 mutation prevalence and more altered genomic landscape. NPJ Breast Cancer 10, 15 (2024).
Tolaney, S. M. et al. 212MO AMEERA-3, a phase II study of amcenestrant (AMC) versus endocrine treatment of physician’s choice (TPC) in patients (pts) with endocrine-resistant ER + /HER2− advanced breast cancer (aBC). Ann. Oncol. 33, S634–S635 (2022).
Oliveira, M. et al. Abstract GS3-02: GS3-02 Camizestrant, a next generation oral SERD vs fulvestrant in post-menopausal women with advanced ER-positive HER2-negative breast cancer: results of the randomized, multi-dose Phase 2 SERENA-2 trial. Cancer Res. 83, GS3-02–GS3-02 (2023).
Bidard, F.-C. et al. Elacestrant (oral selective estrogen receptor degrader) Versus Standard Endocrine Therapy for Estrogen Receptor–Positive, Human Epidermal Growth Factor Receptor 2–Negative Advanced Breast Cancer: Results From the Randomized Phase III EMERALD Trial. J. Clin. Oncol. 40, 3246–3256 (2022).
Martín, M. et al. Giredestrant for estrogen receptor-positive, HER2-negative, previously treated advanced breast cancer: results from the randomized, phase II acelERA breast cancer study. J. Clin. Oncol. JCO2301500, https://doi.org/10.1200/jco.23.01500 (2024).
Liang, J. et al. GDC-9545 (giredestrant): a potent and orally bioavailable selective estrogen receptor antagonist and degrader with an exceptional preclinical profile for ER+ breast cancer. J. Med. Chem. 64, 11841–11856 (2021).
Jhaveri, K. L. et al. Phase Ia/b study of giredestrant ±palbociclib and ±luteinizing hormone-releasing hormone agonists in estrogen receptor-positive, HER2-negative, locally advanced/metastatic breast cancer. Clin. Cancer Res. 30, 754–766 (2023).
Guan, J. et al. Therapeutic ligands antagonize estrogen receptor function by impairing its mobility. Cell 178, 949–963.e18 (2019).
Jimenez, M. M. et al. 211MO Giredestrant (GDC-9545) vs physician choice of endocrine monotherapy (PCET) in patients (pts) with ER+, HER2– locally advanced/metastatic breast cancer (LA/mBC): primary analysis of the phase II, randomised, open-label acelERA BC study. Ann. Oncol. 33, S633–S634 (2022).
Desai, A. et al. Putting comprehensive genomic profiling of ctDNA to work: 10 proposed use cases. J. Liq. Biopsy 4, 100140 (2024).
Parseghian, C. M. et al. Novel potential mechanisms of acquired resistance to anti-EGFR monoclonal antibody (mAb) therapy detected in liquid biopsies (LBx) from patients (pts) with advanced colorectal cancer (CRC). J. Clin. Oncol. 42, 199–199 (2024).
Lim, Z.-F. & Ma, P. C. Emerging insights of tumor heterogeneity and drug resistance mechanisms in lung cancer targeted therapy. J. Hematol. Oncol. 12, 134 (2019).
Sweeney, C. et al. Evaluating early changes in circulating tumor DNA (ctDNA) tumor fraction (TF) as a value add to PSA in predicting early progression in metastatic castrate resistant prostate cancer (mCRPC). J. Clin. Oncol. 42, 196–196 (2024).
Assaf, Z. J. F. et al. A longitudinal circulating tumor DNA-based model associated with survival in metastatic non-small-cell lung cancer. Nat. Med. 29, 859–868 (2023).
Wyatt, A. W. et al. Plasma ctDNA as a treatment response biomarker in metastatic cancers: evaluation by the RECIST working group. Clin. Cancer Res. OF1–OF8, https://doi.org/10.1158/1078-0432.ccr-24-1883 (2024).
O’Leary, B. PADA-1 trial: ESR1 mutations in plasma ctDNA guide treatment switching. Nat. Rev. Clin. Oncol. 20, 67–68 (2023).
Pellini, B. et al. Circulating tumor DNA monitoring on chemo-immunotherapy for risk stratification in advanced non–small cell lung cancer. Clin. Cancer Res. 29, 4596–4605 (2023).
Wander, S. A. et al. The genomic landscape of intrinsic and acquired resistance to cyclin-dependent kinase 4/6 inhibitors in patients with hormone receptor–positive metastatic breast cancer. Cancer Discov. 10, 1174–1193 (2020).
Liang, J. et al. ERα dysfunction caused by ESR1 mutations and therapeutic pressure promotes lineage plasticity in ER+ breast cancer. Nat. Cancer 6, 357–371 (2025).
Parker, J. S. et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J. Clin. Oncol. 41, 4192–4199 (2023).
Caan, B. J. et al. Intrinsic subtypes from the PAM50 gene expression assay in a population-based breast cancer survivor cohort: prognostication of short- and long-term outcomes. Cancer Epidemiol. Prev. Biomark. 23, 725–734 (2014).
Ali, H. R. et al. Genome-driven integrated classification of breast cancer validated in over 7,500 samples. Genome Biol. 15, 431 (2014).
Toy, W. et al. Activating ESR1 mutations differentially affect the efficacy of ER antagonists. Cancer Discov. 7, 277–287 (2017).
Fanning, S. W. et al. Estrogen receptor alpha somatic mutations Y537S and D538G confer breast cancer endocrine resistance by stabilizing the activating function-2 binding conformation. eLife 5, e12792 (2016).
Tolaney, S. M. et al. Clinical significance of PIK3CA and ESR1 mutations in circulating tumor DNA: analysis from the MONARCH 2 study of abemaciclib plus fulvestrant. Clin. Cancer Res. 28, 1500–1506 (2022).
Bardia, A. et al. Abstract GS1-10: Giredestrant vs standard-of-care endocrine therapy as adjuvant treatment for patients with estrogen receptor-positive, HER2-negative early breast cancer: Results from the global Phase III lidERA Breast Cancer trial. Clin. Cancer Res. 32, GS1-10-GS1-10 (2026).
Sadeh, R. et al. ChIP-seq of plasma cell-free nucleosomes identifies gene expression programs of the cells-of-origin. Nat. Biotechnol. 39, 586–598 (2021).
Baca, S. C. et al. Liquid biopsy epigenomic profiling for cancer subtyping. Nat. Med. 29, 2737–2741 (2023).
Doebley, A.-L. et al. A framework for clinical cancer subtyping from nucleosome profiling of cell-free DNA. Nat. Commun. 13, 7475 (2022).
Esfahani, M. S. et al. Inferring gene expression from cell-free DNA fragmentation profiles. Nat. Biotechnol. 40, 585–597 (2022).
Stanley, K. E. et al. Cell type signatures in cell-free DNA fragmentation profiles reveal disease biology. Nat. Commun. 15, 2220 (2024).
Mathios, D. et al. Detection and characterization of lung cancer using cell-free DNA fragmentomes. Nat. Commun. 12, 5060 (2021).
Sjöström, M. et al. The 5-hydroxymethylcytosine landscape of prostate cancer. Cancer Res. 82, OF1–OF15 (2022).
Heeke, S. et al. Tumor- and circulating-free DNA methylation identifies clinically relevant small cell lung cancer subtypes. Cancer Cell 42, 225–237.e5 (2024).
Pellini, B. et al. Computational Variant Origin Prediction (VOP) to distinguish germline, tumor somatic, and Clonal Hematopoiesis (CH) variants in Liquid Biopsy (LBX). J. Liq. Biopsy 5, 100242 (2024).
Woodhouse, R. et al. Clinical and analytical validation of FoundationOne Liquid CDx, a novel 324-Gene cfDNA-based comprehensive genomic profiling assay for cancers of solid tumor origin. PLoS ONE 15, e0237802 (2020).
Sweeney, C. J. et al. Circulating tumor DNA assessment for treatment monitoring adds value to PSA in metastatic castration-resistant prostate cancer. Clin. Cancer Res. 30, 4115–4122 (2024).
Acknowledgements
We would like to thank Steven Gendreau, Marc Hafner, Jackson Liang, Ciara Metcalfe, and Matthew Wongchenko for their review and discussion. This research was funded by F. Hoffmann-La Roche Ltd. F. Hoffmann-La Roche Ltd was involved in the study design, data interpretation, and decision to submit for publication in conjunction with the authors.
Author information
Authors and Affiliations
Contributions
A.E.C. wrote the manuscript. A.E.C and S.H. constructed the figures. A.E.C., S.H., L.W.P., T.M.F., and H.M.M. designed, performed, and interpreted the analyses. A.M.C. and X.S. provided additional computational support and data integration. M.CM., A.B., M.M., E.L, J.S., and P.D.PM. conducted the clinical study. C.D., P.D.PM., and H.M.M. conceived and executed the biomarker sampling plan for the clinical study. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The study was sponsored by F. Hoffmann-La Roche Ltd. The authors declare the following competing interests: A.E.C., S.H., A.M.C., P.D.P.-M., and H.M.M. are employees of Genentech, Inc. and hold stock in F. Hoffmann-La Roche Ltd. C.D. is an employee of Roche (China) Holding Ltd. and holds stock in F. Hoffmann-La Roche Ltd. and Bristol Myers Squibb. L.W.P. is an employee of Foundation Medicine and holds stock in Roche Holdings AG. M.CM. reports consulting fees from Novartis, AstraZeneca, and Lilly; honoraria from Exact Sciences; travel support from Novartis and AstraZeneca; advisory board participation for AstraZeneca and Roche/Genentech; and board membership for Legacy Community Health, The Hope Foundation, and ASCO. A.B. reports grants and personal fees from Pfizer, Novartis, Genentech, Merck, Menarini, Gilead, Sanofi, Daiichi Pharma/AstraZeneca, Alyssum, and Eli Lilly. M.M. reports honoraria and advisory fees from Roche/Genentech, Lilly, Pfizer, Novartis, Pierre Fabre, Seagen, AstraZeneca, and Daiichi-Sankyo; and research funding from Novartis, Roche, and Puma Biotechnology. E.L. reports institutional advisory/steering committee roles and research funding from AstraZeneca, Gilead, Lilly, MSD, Novartis, Pfizer, and Roche; royalties from Walter and Eliza Hall Institute; and leadership roles at Breast Cancer Trials Australia, Garvan Institute, St Vincent’s Hospital, and UNSW. J.S. reports institutional research funding from Seagen, MSD, Roche, Pfizer, Novartis, AstraZeneca, Lilly, GSK, Boehringer Ingelheim, Sanofi, Daiichi Sankyo, Qurient, Dragonfly, Eikon, Gilead, Celcuity, BMS, HLB Life Science, Sermonix, Olema, Hanmi, Ildong, and Samyang; and family stock ownership in Daiichi Sankyo. X.S. declares no competing interests.
Peer review
Peer review information
Nature Communications thanks Olufunmilayo Olopade, who co-reviewed with Frederick Howard and Sulin Wu, Jonathan Dudley and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Collier, A.E., Hilz, S., Chibly, A.M. et al. ctDNA and tumor-based biomarkers of giredestrant response in acelERA breast cancer. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70335-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70335-0