Abstract
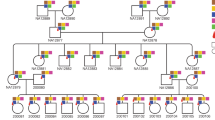
Long-read sequencing improves sensitivity to discover variation in complex repetitive regions, assign parent-of-origin, and distinguish germline from postzygotic mutations. We applied Illumina, Oxford Nanopore Technologies, and PacBio sequencing to discover and validate de novo mutations in 73 children from 42 autism families (157 individuals). We assay 2.77 Gbp of the human genome, yielding on average 95 de novo mutations per transmission (87.5 single-nucleotide substitutions, 7.8 indels), with no significant difference in mutation rate or profile between probands and their unaffected siblings. Long reads increase de novo mutation discovery by 20-40% and double the mutations classified as early embryonic. The germline mutation rate is 1.30×10−8 substitutions/base pair/generation; the postzygotic rate is 0.23×10−8. These rates are significantly increased in repetitive DNA, where segmental duplication mutability is dependent on length and percent identity. Here, we show that enrichment in repeats occurs predominantly postzygotically, likely resulting from faulty DNA repair and interlocus gene conversion.
Similar content being viewed by others
Data availability
The data used for analysis, including underlying sequence data, assemblies, and alignment files, are available to approved researchers in the SFARI Base under the accession number SFARI_DS0000104 (https://base.sfari.org/dataset/DS0000104) and through the National Institute of Mental Health Data Archive (NDA) under Collection ID 3780. Source data are provided with this paper.
Code availability
Code and scripts used for the analyses presented in this manuscript are available in GitHub at https://github.com/mdnoyes/denovo_calling69. Any additional information required to reanalyze the data reported in this work paper is available from the lead contact upon request.
References
Sasani, T. A. et al. Large, three-generation human families reveal post-zygotic mosaicism and variability in germline mutation accumulation. eLife 8, e46922 (2019).
Jónsson, H. et al. Parental influence on human germline de novo mutations in 1,548 trios from Iceland. Nature 549, 519–522 (2017).
Goldmann, J. M. et al. Parent-of-origin-specific signatures of de novo mutations. Nat. Genet. 48, 935–939 (2016).
Kong, A. et al. Rate of de novo mutations and the importance of father’s age to disease risk. Nature 488, 471–475 (2012).
Kessler, M. D. et al. De novo mutations across 1,465 diverse genomes reveal mutational insights and reductions in the Amish founder population. Proc. Natl. Acad. Sci. USA 117, 2560–2569 (2020).
Gao, Z. et al. Overlooked roles of DNA damage and maternal age in generating human germline mutations. Proc. Natl. Acad. Sci. USA 116, 9491–9500 (2019).
de Manuel, M., Wu, F. L. & Przeworski, M. A paternal bias in germline mutation is widespread in amniotes and can arise independently of cell division numbers. eLife 11, e80008 (2022).
Hahn, M. W., Peña-Garcia, Y. & Wang, R. J. The ‘faulty male’ hypothesis for sex-biased mutation and disease. Curr. Biol. 33, R1166–R1172 (2023).
Ju, Y. S. et al. Somatic mutations reveal asymmetric cellular dynamics in the early human embryo. Nature 543, 714–718 (2017).
Lindsay, S. J., Rahbari, R., Kaplanis, J., Keane, T. & Hurles, M. E. Similarities and differences in patterns of germline mutation between mice and humans. Nat. Commun. 10, 4053 (2019).
Porubsky, D. et al. Human de novo mutation rates from a four-generation pedigree reference. Nature 643, 427–436 (2025).
Jonsson, H. et al. Differences between germline genomes of monozygotic twins. Nat. Genet. 53, 27–34 (2021).
Huang, A. Y. et al. Postzygotic single-nucleotide mosaicisms in whole-genome sequences of clinically unremarkable individuals. Cell Res. 24, 1311–1327 (2014).
Acuna-Hidalgo, R. et al. Post-zygotic point mutations are an underrecognized source of de novo genomic variation. Am. J. Hum. Genet. 97, 67–74 (2015).
Wright, C. F. et al. Clinically-relevant postzygotic mosaicism in parents and children with developmental disorders in trio exome sequencing data. Nat. Commun. 10, 2985 (2019).
Dou, Y. et al. Postzygotic single-nucleotide mosaicisms contribute to the etiology of autism spectrum disorder and autistic traits and the origin of mutations. Hum. Mutat. 38, 1002–1013 (2017).
Lim, E. T. et al. Rates, distribution and implications of postzygotic mosaic mutations in autism spectrum disorder. Nat. Neurosci. 20, 1217–1224 (2017).
Telenti, A. et al. Deep sequencing of 10,000 human genomes. Proc. Natl. Acad. Sci. USA 113, 11901–11906 (2016).
Turner, T. N. et al. Genomic patterns of de novo mutation in simplex autism. Cell 171, 710–722.e12 (2017).
Wilfert, A. B. et al. Recent ultra-rare inherited variants implicate new autism candidate risk genes. Nat. Genet. 53, 1125–1134 (2021).
Noyes, M. D. et al. Familial long-read sequencing increases yield of de novo mutations. Am. J. Hum. Genet. 109, 631–646 (2022).
Ebbert, M. T. W. et al. Systematic analysis of dark and camouflaged genes reveals disease-relevant genes hiding in plain sight. Genome Biol. 20, 97 (2019).
Porubsky, D. & Eichler, E. E. A 25-year odyssey of genomic technology advances and structural variant discovery. Cell 187, 1024–1037 (2024).
Holt, G. S. et al. Phasing of de novo mutations using a scaled-up multiple amplicon long-read sequencing approach. Hum. Mutat. 43, 1545–1556 (2022).
Kucuk, E. et al. Comprehensive de novo mutation discovery with HiFi long-read sequencing. Genome Med. 15, 34 (2023).
Fu, J. M. et al. Rare coding variation provides insight into the genetic architecture and phenotypic context of autism. Nat. Genet. 54, 1320–1331 (2022).
An, J.-Y. et al. Genome-wide de novo risk score implicates promoter variation in autism spectrum disorder. Science 362, eaat6576 (2018).
Sui, Y. et al. Using the linear references from the pangenome to discover missing autism variants. Nat. Commun. 17, 1681 (2026).
McLaren, W. et al. The Ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
Seplyarskiy, V. et al. A mutation rate model at the basepair resolution identifies the mutagenic effect of polymerase III transcription. Nat. Genet. 55, 2235–2242 (2023).
Koch, Z., Li, A., Evans, D. S., Cummings, S. & Ideker, T. Somatic mutation as an explanation for epigenetic aging. Nat. Aging 5, 709–719 (2025).
Michaelson, J. J. et al. Whole-genome sequencing in autism identifies hot spots for de novo germline mutation. Cell 151, 1431–1442 (2012).
Besenbacher, S. et al. Multi-nucleotide de novo mutations in humans. PLOS Genet. 12, e1006315 (2016).
Ng, J. K. & Turner, T. N. HAT: de novo variant calling for highly accurate short-read and long-read sequencing data. Bioinformatics 40, btad775 (2024).
Mitchell, E. et al. Clonal dynamics of haematopoiesis across the human lifespan. Nature 606, 343–350 (2022).
Horebeek, L. V., Dubois, B. & Goris, A. Somatic variants: new kids on the block in human immunogenetics. Trends Genet. 35, 935–947 (2019).
Fabre, M. A. & Vassiliou, G. S. The lifelong natural history of clonal hematopoiesis and its links to myeloid neoplasia. Blood 143, 573–581 (2024).
Bernstein, N. et al. Analysis of somatic mutations in whole blood from 200,618 individuals identifies pervasive positive selection and novel drivers of clonal hematopoiesis. Nat. Genet. 56, 1147–1155 (2024).
Harland, C. et al. Frequency of mosaicism points towards mutation-prone early cleavage cell divisions in cattle. Preprint at https://doi.org/10.1101/079863 (2017).
Shoag, J. E. et al. Direct measurement of the male germline mutation rate in individuals using sequential sperm samples. Nat. Commun. 16, 2546 (2025).
Kunisaki, J. et al. Sperm from infertile, oligozoospermic men have elevated mutation rates. Preprint at https://doi.org/10.1101/2024.08.22.24312232 (2024).
Braude, P., Bolton, V. & Moore, S. Human gene expression first occurs between the four- and eight-cell stages of preimplantation development. Nature 332, 459–461 (1988).
Musson, R., Gąsior, Ł, Bisogno, S. & Ptak, G. E. DNA damage in preimplantation embryos and gametes: specification, clinical relevance and repair strategies. Hum. Reprod. Update 28, 376–399 (2022).
Park, S. et al. Clonal dynamics in early human embryogenesis inferred from somatic mutation. Nature 597, 393–397 (2021).
Rahbari, R. et al. Timing, rates and spectra of human germline mutation. Nat. Genet. 48, 126–133 (2016).
Kiktev, D. A., Sheng, Z., Lobachev, K. S. & Petes, T. D. GC content elevates mutation and recombination rates in the yeast Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 115, E7109–E7118 (2018).
Harpak, A., Lan, X., Gao, Z. & Pritchard, J. K. Frequent nonallelic gene conversion on the human lineage and its effect on the divergence of gene duplicates. Proc. Natl. Acad. Sci. USA 114, 12779–12784 (2017).
Vollger, M. R. et al. Increased mutation and gene conversion within human segmental duplications. Nature 617, 325–334 (2023).
Palsson, G. et al. Complete human recombination maps. Nature 639, 700–707 (2025).
Harris, K. & Nielsen, R. Error-prone polymerase activity causes multinucleotide mutations in humans. Genome Res. 24, 1445–1454 (2014).
Nagylaki, T. & Petes, T. D. Intrachromosomal gene conversion and the maintenance of sequence homogeneity among repeated genes. Genetics 100, 315–337 (1982).
Hallast, P. et al. Assembly of 43 human Y chromosomes reveals extensive complexity and variation. Nature 621, 355–364 (2023).
Rhie, A. et al. The complete sequence of a human Y chromosome. Nature 621, 344–354 (2023).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://doi.org/10.48550/arXiv.1303.3997 (2013).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. Preprint at https://doi.org/10.1101/201178 (2018).
Poplin, R. et al. A universal SNP and small-indel variant caller using deep neural networks. Nat. Biotechnol. 36, 983–987 (2018).
Smit, A. F. A., Hubley, R. & Green, P. RepeatMasker Open-3.0. https://www.repeatmasker.org/ (1996).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Hinrichs, A. S. et al. The UCSC genome browser database: update 2006. Nucleic Acids Res. 34, D590–D598 (2006).
Wang, T. et al. Large-scale targeted sequencing identifies risk genes for neurodevelopmental disorders. Nat. Commun. 11, 4932 (2020).
Satterstrom, F. K. et al. Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell 180, 568–584.e23 (2020).
Liu, X., Li, C., Mou, C., Dong, Y. & Tu, Y. dbNSFP v4: a comprehensive database of transcript-specific functional predictions and annotations for human nonsynonymous and splice-site SNVs. Genome Med. 12, 103 (2020).
Orenbuch, R. et al. Proteome-wide model for human disease genetics. Nat. Genet. 57, 3165–3174 (2025).
Zhao, Y. et al. A probabilistic graphical model for estimating selection coefficients of nonsynonymous variants from human population sequence data. Nat. Commun. 16, 4670 (2025).
Chen, S. et al. A genomic mutational constraint map using variation in 76,156 human genomes. Nature 625, 92–100 (2024).
Hennick, K. et al. Sex differences in the developing human cortex intersect with genetic risk of neurodevelopmental disorders. Preprint at https://doi.org/10.1101/2025.09.04.674293 (2025).
Noyes, Michelle D. Long-read sequencing of trios reveals increased germline and postzygotic mutation rates in repetitive DNA. mdnoyes/denovo_calling https://doi.org/10.5281/zenodo.18736932 (2026).
Helgason, A. et al. The Y-chromosome point mutation rate in humans. Nat. Genet. 47, 453–457 (2015).
Xue, Y. et al. Human Y Chromosome Base-Substitution Mutation Rate Measured by Direct Sequencing in a Deep-Rooting Pedigree. Curr. Biol. 19, 1453–1457 (2009).
Kuroki, Y. et al. Comparative analysis of chimpanzee and human Y chromosomes unveils complex evolutionary pathway. Nat. Genet. 38, 158–167 (2006).
Acknowledgements
This work was supported, in part, by the US National Institutes of Health (NIH R01MH101221 to E.E.E.) and the Simons Foundation (SFARI #810018EE to E.E.E.). E.E.E. is an investigator of the Howard Hughes Medical Institute. We thank Tonia Brown for assistance in editing this manuscript. This article is subject to HHMI’s Open Access to Publications policy. HHMI lab heads have previously granted a nonexclusive CC BY 4.0 license to the public and a sublicensable license to HHMI in their research articles. Pursuant to those licenses, the author-accepted manuscript of this article can be made freely available under a CC BY 4.0 license immediately upon publication.
Author information
Authors and Affiliations
Contributions
E.E.E. and M.D.N. conceptualized the study. K.M.M., K.H., J. Kordosky, G.H.G., J. Knuth, and A.P.L. generated the data. Y.S., Y.K., I.W., and N.K. performed data quality control. M.D.N. and Y.S. conducted the formal analysis. M.D.N. created the visualizations. M.D.N. developed the methodology. M.D.N. wrote the original draft. M.D.N., Y.S., and E.E.E. reviewed and edited the manuscript. E.E.E. supervised the study.
Corresponding author
Ethics declarations
Competing interests
E.E.E. is a scientific advisory board (SAB) member of Variant Bio, Inc. All other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ryan Yuen, who co-reviewed with Mahreen Khan; Jeremy Guez and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Noyes, M.D., Sui, Y., Kwon, Y. et al. Long-read sequencing of families reveals increased germline and postzygotic mutation rates in repetitive DNA. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70342-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70342-1