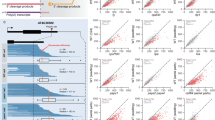
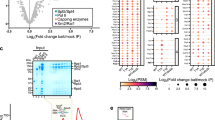
Abstract
Although small nucleolar (sno)RNAs, which guide ribosomal (r)RNA modification, are synthesized and function in the nucleus, some of them can be detected in the cytoplasm. Here, we identify Mex67 and Xpo1 as snoRNP export receptors, and Mtr10 and Cse1 as their re-import factors. Interestingly, only a fraction of snoRNAs shuttle, and we reveal that the mode of transcription determines whether or not the snoRNA is exported. In Saccharomyces cerevisiae, RNA polymerase II-transcribed RNAs are terminated either via the Nrd1-Nab3-Sen1 (NNS) complex or the cleavage and polyadenylation factor (CPF-CF) complex. NNS termination, which mostly occurs for snoRNAs, leads to nuclear retention. Conversely, fail-safe CPF-CF termination results in transcript polyadenylation and subsequent association of the guard proteins Hrp1 and Nab2, which in turn mediate Mex67–Mtr2 dependent export. Importantly, re-imported CPF-CF-terminated snoRNAs form functional snoRNPs. Together, we identified that transcription termination mode determines snoRNA export through a guard protein-controlled mechanism.
Similar content being viewed by others
Introduction
snoRNAs are small non-coding (nc)RNAs that primarily guide chemical modifications of other RNAs, mainly ribosomal (r)RNAs, small nuclear (sn)RNAs, and transfer (t)RNAs1. In Saccharomyces cerevisiae (baker’s yeast), there are 76 different snoRNAs, classified as box C/D or box H/ACA snoRNAs, which mostly guide co-transcriptional rRNA 2’-O-methylation or pseudouridylation, respectively2. To fulfil their functions, snoRNAs associate with proteins, including the 2’-O-methyltransferase Nop1 (box C/D) or pseudouridine synthetase Cbf5 (H/ACA), to form functional snoRNP complexes3. In yeast, snoRNAs are transcribed by RNA polymerase (RNAP)II, except snR52, which is synthesized by RNAPIII. They are either encoded independently or within intron sequences and can either be encoded individually or within clusters. Only the individually and independently transcribed snoRNA genes contain their own transcription termination sequences, which applies to 50 of the 76 snoRNAs in yeast4,5.
As snoRNAs are important for the modification of, e.g., pre-rRNAs, their biogenesis into functional snoRNPs is important for cell survival1,6. SnoRNAs are short ncRNAs, so their transcription is mostly terminated via the Nrd1-Nab3-Sen1 (NNS) complex7. NNS sites are found in all snoRNAs, but some also contain downstream-located polyadenylation signal (PAS) binding sites to which the cleavage and polyadenylation factor (CPF-CF) complex can bind and mediate termination. PAS sites can contribute to NNS-mediated termination and/or act as a fail-safe mechanism in case NNS termination is bypassed. However, within the short distance between the transcription start site and the end of the transcript (typically less than 150 nucleotides), NNS-mediated termination mostly occurs. At this stage of transcription, the Ser5 phosphorylation status of the RNAPII C-terminal domain (CTD) favors binding of Nrd1 and Nab38,9. Additionally, the early Tyr1-phosphorylation is believed to support Sen1-dependent termination via pausing10.
CPF-CF-mediated termination usually occurs on messenger (m)RNAs and long non-coding (lnc)RNAs that receive a poly(A) tail for their protection, whereas NNS-mediated termination is typical for ncRNAs that are either completely degraded in the case of pervasive transcripts and cryptic unstable transcripts (CUTs) or, if protected, immediately trimmed by the nuclear exosome, which includes most snoRNAs7,11,12,13. The terminator region of the snoRNA snR13, as well as its subsequent rapid processing were well studied, and its complete degradation upon NNS-termination is thought to be prevented by secondary structures and/or the co-transcriptional loading of protecting proteins14,15. NNS-mediated termination leads to binding of the TRAMP complex that marks the RNA with an oligo(A) tail for its subsequent recognition and partial degradation by the nuclear exosome. The TRAMP complex is composed of a zinc-finger RNA-binding protein Air1 or Air2, a poly(A) polymerase Trf4 or Trf5, and the RNA helicase Mtr4. For the snoRNA snR13, it was suggested that the oligoadenylated RNA is not degraded, as e.g., CUTs are, but that its tail is rather elongated by the poly(A) polymerase Pap114. Subsequent trimming of the 3’ end is then thought to occur via the nuclear exosome16,17.
All snoRNA transcripts in S. cerevisiae, with one exception, are synthesized by RNAPII and, therefore, their 5’ ends become m7G capped. However, unlike mRNAs, the pre-snoRNA cap is either removed and the RNA subsequently trimmed in the 5’ to 3’ direction, or the m7G cap is converted into a 2,2,7-trimethylguanosine (TMG) cap. TMG-capping usually occurs mostly on H/ACA snoRNAs, in a process involving Tgs1 that is very similar to that occurring on the spliceosomal snRNAs18,19. C/D snoRNAs are usually synthesized with a (150 to 200 nucleotides) long 5’ extension, including a stem-loop structure that is recognized by the endonuclease Rnt120. After cleavage, the unprotected extremity is subsequently processed by the 5’−3’ exonuclease Rat1 in the nucleus. As well as 5’ and 3’ trimming of independently transcribed snoRNAs, snoRNAs synthesized within clusters or encoded within introns require separation, which is achieved through several cleavage and processing steps performed by Rnt1, Rat1, and the exosome. Finally, all snoRNAs assemble with the core snoRNP proteins in the nucleus, facilitated by several assembly factors, to become fully complexed and capable of, e.g., rRNA modification, which takes place in the nucleolar sub-compartment of the nucleus. Therefore, it is assumed that most snoRNAs do not leave the nucleus during their lifetime.
It has been shown that yeast Lhp1 and Lsm2-8 (like-Sm) are involved in the maturation of ncRNAs such as tRNAs, U6 snRNA, and U3 snoRNA by binding to an oligo(U) tract at their 3’ ends21. Interestingly, Lhp1 functions as a guard protein that controls the nuclear loading of the Lsm-ring22). Both Sm and Lsm proteins form a ring-like structure that protects the bound RNA from exonucleolytic degradation23,24,25. In contrast to the Sm-ring, which is loaded onto RNA in the cytoplasm18,26, the Lsm2-8 ring is assembled in the nucleus. It is distinct from another complex, the Lsm1-7 ring, which associates with exoribonucleases and participates in cytoplasmic mRNA degradation27. Assembly of an Sm-ring on the snRNAs U2, U4, and U5, as well as the TLC1 RNA of the yeast telomerase, was shown to occur in the cytoplasm18,26. These RNAs are exported to the cytoplasm after transcription as pre-RNAs. This is necessary for certain maturation steps that occur within this compartment, but importantly, physical separation of the pre-RNA from the mature RNP prevents immature or partially matured but still unfunctional snRNPs or telomerase from impairing pre-mRNA splicing or telomere maintenance, respectively. Only upon successful assembly of the Est- and Pop-proteins with TLC1, for instance, the Sm-ring assembles, which in turn allows association of the nuclear re-import receptors Cse1 and Mtr10 and thus, nuclear re-import of the matured pre-RNP. In this way, cytoplasmic Sm-ring assembly represents a quality control checkpoint for maturation of the pre-RNPs18,26.
Interestingly, besides the core snoRNP proteins, Lhp1 and the Lsm-proteins, snoRNAs also interact with the guard proteins Gbp2, Npl3, Nab2 and Hrp128,29,30,31,32. This group of homologous proteins has been shown to surveil different aspects of pre-mRNA maturation, such as 5’-capping, intron splicing, 3’ cleavage and polyadenylation33,34,35,36,37,38,39. Upon successful completion of each step, a guard protein recruits the export receptor heterodimer Mex67-Mtr2, which increases the export competence of the mRNP. As more Mex67 is bound, the propensity for the mRNP to be exported into the cytoplasm increases38,39,40,41. Strikingly, Mex67 binding was also detected on snoRNAs32. Mex67-mediated RNA export is supported by Xpo1/Crm1, which contacts the 5’cap binding complex (CBC)18,37. As all snoRNA processing and snoRNP assembly steps seemed to occur within the nucleus, the binding of guard proteins and Mex67 to snoRNAs remained unexplained. It has been shown that some snoRNAs and their derivatives have been detected in the cytoplasm in human cells42,43,44,45, however, robust evidence of snoRNAs in the cytoplasm in yeast was lacking. Building on the hypothesis that under certain conditions some snoRNAs might leave the nucleus via Mex67-dependent export, this study aimed to uncover why some snoRNAs interact with Mex67, which features of these snoRNAs determine this interaction and what the role of snoRNA export to the cytoplasm is.
Here, we show that some snoRNA transcripts are exported into the cytoplasm during their life cycle. We present evidence that they do not require the cytoplasmic phase for their maturation, but that their nuclear export is rather a consequence of the way their transcription is terminated. When NNS-mediated termination takes place, the snoRNAs remain nuclear. However, when a snoRNA is terminated by the CPF-CF complex, it shuttles inevitably into the cytoplasm. Re-import occurs via its Lsm-ring, which contacts the import factors Cse1 and Mtr10. We suggest that exported snoRNAs are matured after nuclear re-import. Importantly, the mode of transcription termination and the export into the cytoplasm do not alter snoRNP functionality. Together, these findings rationalize why various snoRNAs are present in the yeast cytoplasm.
Results
Different types of snoRNAs are present in the cytoplasm
To explore whether the association of the export receptor Mex67 with snoRNAs leads to their nuclear export, we re-analyzed the transcriptome of nucleo-cytoplasmic fractionated yeast for snoRNAs present in the cytoplasm46. We compared the amount of sequencing reads derived from snoRNAs in wild type to those in the temperature-sensitive export mutant mex67-5 xpo1-1 and found a substantial amount of snoRNAs to be detectable in the cytoplasm in wild type cells (Fig. 1a, Supplementary Fig. 1a). Strikingly, the amount of snoRNAs present in the cytoplasm was reduced when the export pathway was blocked through shifting mex67-5 xpo1-1 mutant to 37 °C for 1 h (Fig. 1a). Notably, the amount of snoRNP core proteins such as Nop1 is not affected in export mutants in this time frame (Supplementary Fig. 1b, c). The remaining snoRNAs detected in the cytoplasm could either reside in the cytoplasm for a longer time, or the cytoplasmic fraction contained low levels of nuclear RNA contaminants, although the fractionation seemed successful by other RNA parameters and the detection of nuclear and cytoplasmic proteins46. These data suggest that at least half of the snoRNAs present in the cytoplasm are actively transported into this compartment. To investigate whether the export is related to the type of the snoRNA, we categorized these snoRNAs into seven groups based on their motifs (C/D or H/ACA), genomic organization (monocistronic, polycistronic, and intronic), and cap structure (capped and uncapped). Interestingly, the snoRNAs detected in the cytoplasm were diverse, and representatives of all different types of snoRNAs were present (Fig. 1b). Importantly, the levels of the snoRNAs in the cytoplasm were not particularly high and were comparable to those of most mRNAs (Supplementary Fig. 1d, e). For intronic snoRNAs, it remains possible that they are exported within their pre-mRNA hosts in situations where the Gbp2- and Hrb1-mediated quality control of splicing failed, and snoRNA-containing pre-mRNAs escape mRNA surveillance and reach the cytoplasm. Such RNAs are normally captured by a second quality control system in the cytoplasm, nonsense-mediated decay (NMD), which recognizes mRNAs with premature termination codons in initial rounds of translation and subsequently eliminates them47,48. However, as all types of snoRNAs were affected by the mutation of the RNA export factors, this suggests that snoRNAs can, at least under some circumstances, be exported into the cytoplasm.
a, b RNA-sequencing data from a nucleo-cytoplasmic fractionation experiment46 were analyzed for individual snoRNAs. Each snoRNA’s ratio of cytoplasmic to total reads in wild type and mex67-5 xpo1-1 was shown in (a). Box plot indicates median (centre), 25th-75th percentile (box), and minimum-maximum values (whiskers) as well as all single data points. snoRNAs are further divided in groups and the relative ratio of cytoplasmic to total reads in mex67-5 xpo1-1 related to wild type is shown in (b). n = 3. c Western blot of the RIP validates the pull-down of the GFP-tagged Mex67. The mitochondrial protein Hem15 served as a negative control. kDa=kilodaltons. d Binding of snoRNAs to the Mex67 was analyzed via qPCR, compared to lysate, and normalized to the mitochondrial 21S rRNA, which does not interact with Mex67. The mRNA ACT1 served as a positive control. The primer pair amplifying immature 3’-extended snoRNAs was indicated with an arrow. c, d n = 3. e qPCR analysis of total and immature 3’-extended snoRNAs in wild type and mex67-5 is shown after a temperature shift for 1 h to 37 °C using the indicated primer pairs. The fold change of the total and 3’-extended snoRNAs in mex67-5 was calculated compared to wild type and normalized to the 18S rRNA. n = 4. f FISH experiments using ~200 nucleotide long digoxygenin labelled probes that are complementary to the individual snoRNAs were used and detected via an FITC conjugated α-DIG antibody (green). Analysis was carried out in wild-type and mex67-5 after shifting the cells for 1 h to 37 °C. The DNA was stained with DAPI (blue), and the pre-rRNA fragment ETS1 served as negative control. n = 3. n indicates the number of biological replicates. Data in bar plots (d, e) are presented as mean +/- SD. A two-tailed Student’s t-test was used to calculate p-values. *p < 0.05; **p < 0.01; ***p < 0.001. Individual data points are represented by black circles.
Immature pre-snoRNAs are exported into the cytoplasm
To gain further insight into the maturation status of the snoRNAs associated with Mex67, we next investigated whether Mex67 interacts with pre-snoRNAs with extended 3’ ends. For this purpose, RNA co-immunoprecipitation (RIP) experiments were performed in which Mex67 was precipitated from cell lysates (Fig. 1c) and its interaction with the nascent 3’-extended pre-snoRNAs was investigated. We found clear interaction of Mex67 with 3’ extended versions of the chosen snoRNAs snR24, 42, 68, and 13, that represent different types of snoRNAs and in most cases, this enrichment was statistically significant (Fig. 1d). The mRNA ACT1 served as a positive control and mitochondrial 21S rRNA, which does not bind to Mex67, was used for normalization. This suggests that these immature snoRNAs are exported into the cytoplasm, which in turn means that these immature snoRNAs are likely to accumulate in mex67-5 mutants at the non-permissive temperature due to their nuclear retention. Indeed, qPCR experiments of the different example snoRNAs show that snoRNAs accumulate in the export mutant after a temperature shift to 37 °C for 1 h (Fig. 1e, Supplementary Fig. 1f). These are mainly derived from the pre-snoRNAs, as their amount increases as well in mex67-5. The strongest increase was seen for snR68, 42, and 13, while snR24 seemed to decrease. The nuclear accumulation of these pre-snoRNAs was also visualized by fluorescence in situ hybridization (FISH) experiments with a probe that detects both the mature and the extended immature pre-snoRNAs. To visualize the cytoplasmic snoRNAs preferentially, we used ~200 nt long digoxygenin labelled probes that likely encounter difficulties entering the nucleus18. These probes still show the strongest signal of the snoRNAs in the nucleolus, which is the place of their function49,50. However, this long probe enabled us to also detect signals in the cytoplasm of wild-type cells (Fig. 1f, Supplementary Fig. 1g). Importantly, the cytoplasmic signals were lost in the mex67-5 export mutant shifted to 37 °C for 1 h. Instead, this strain shows strong nuclear accumulations of the snoRNAs. Furthermore, the distribution of the snoRNAs also differed from the nucleolar signal observed in wild-type cells, as a dot-like staining along the nuclear rim was observed, which is in line with perturbed nuclear export. Although it cannot be excluded that this effect might arise as a secondary effect of the loss of mRNA export in mex67-5, importantly, the cytoplasmic signal for these snoRNAs completely vanished when nuclear export was blocked.
snoRNA re-import is mediated by the import receptors Cse1 and Mtr10
Upon successful completion of the cytoplasmic phases of their biogenesis, re-import of snRNAs and TLC1 is mediated by Cse1 and Mtr1018,26. It is, therefore, conceivable that snoRNAs use the same re-import pathway following their translocation to the cytoplasm. To visualize the differences in the localization of an import mutant, we used short ~30 to 50 nt long Cy3-labelled probes for the individual snoRNAs. Those short probes easily get into the nuclei of fixed cells18 and therefore reliably stain nucleolar snoRNAs in cells. In contrast to wild type, where the tested snoRNAs were predominantly nucleolar, snoRNA detection in the single import mutants and more strongly in the double mutant cse1-1 mtr10∆ showed cytoplasmic signals for all analyzed snoRNAs, except the intronic snR24, which does not seem to be re-imported into the nucleus (Fig. 2a, Supplementary Fig. 2). It might rather be degraded in the cytoplasm via NMD as part of an intron-containing pre-mRNA. This result was confirmed with qPCRs in which mostly the immature form of the snoRNAs accumulated in the import factor double mutant (Fig. 2b). We confirmed this result with a third method, a nucleo-cytoplasmic fractionation experiment, in which we isolated the cytoplasmic RNA content of wild type and the double import factor mutant and showed in qPCRs a significant increase of the immature forms of all tested snoRNAs (Fig. 2c, d). These findings suggest that exported snoRNAs are re-imported into the nucleus via the import receptor Mtr10 and Cse1.
a FISH experiments were carried out in wild type and cse1-1 mtr10∆ upon temperature shift for 1 h 15 min to 16 °C. ~30 to 50 nucleotide long Cy3 labelled probes (red) were used for hybridization with the indicated snoRNAs. The DNA was stained with DAPI. n = 3. b qPCR analysis of total and immature 3’-extended snoRNAs in wild type and cse1-1 mtr10∆ shifted for 1 h 15 min to 16 °C is shown. The indicated primer pairs are shown. The fold change was normalized to the 18S rRNA. n = 4. c The separation of nuclear and cytoplasmic fractions was verified by Western blot. Yra1 served as nuclear control, and Zwf1 served as cytoplasmic control. kDa = kilodaltons. d Immature 3’-extended snoRNAs in the cytoplasmic fraction were analyzed via qPCR, compared to wild type, and normalized to the 18S rRNA. c, d n = 6. n indicates the number of biological replicates. Data in bar plots (b, d) are presented as mean +/− SD. A two-tailed Student’s t-test was used to calculate p values. *p < 0.05; **p < 0.01; ***p < 0.001. Individual data points are represented by black circles.
Lhp1 and the Lsm-ring are loaded in the nucleus
snRNAs shuttle into the cytoplasm to receive their Sm-ring, except for U6, which receives its Lsm-ring in the nucleus in a process controlled by Lhp118,22. It is currently known that Lhp1 is initially loaded onto the snoRNA snR17a (U3a), but that it dissociates during maturation and is not part of the mature snoRNP51,52. Additionally, microarray studies indicate binding of Lhp1 to other snoRNAs as well53. To investigate in which compartment Lhp1 and the Lsm-ring are attached to the snoRNAs, we did the following RIP experiments. First, we analyzed the association of some exported snoRNAs with Lhp1 by reverse transcription coupled to quantitative PCR (RT-qPCR) using primers that amplify the immature pre-snoRNAs and detected binding (Fig. 3a, Supplementary Fig. 3a). We found that the binding of the snoRNAs to Lhp1 increased in the export and re-import mutants at their non-permissive temperature (Fig. 3b, Supplementary Fig. 3b, c). This suggests that Lhp1 is loaded onto the snoRNAs in the nucleus prior to export and re-import. The binding increased, in all cases, between 2- and almost 10-fold. Binding to the 18S rRNA served as a negative control53 and all RIP experiments were therefore normalized to 18S rRNA.
RIP experiments of GFP-tagged Lhp1 and Lsm8 were conducted in wild type (a, c), wild type and mex67-5 with a temperature shift for 1 h to 37 °C, wild type and cse1-1 mtr10∆ with a temperature shift for 1 h 15 min to 16 °C (b, d). Binding of immature 3’-extended snoRNAs to Lhp1 or Lsm8 was analyzed via qPCR with the indicated primer pairs and compared to lysate (a, c) or wild type (b, c) and normalized to the 18S rRNA. a, c n = 3, b n = 4 (mex67-5) and 6 (cse1-1 mtr10∆), d n = 5 (mex67-5) and 6 (cse1-1 mtr10∆). e RIP experiments pulling down GFP-tagged Lhp1 were carried out in wild type and lsm8-1 that was shifted for 2 h 30 min to 37 °C. qPCR with the indicated primer pairs is shown and the binding was compared to wild type and normalized to the 18S rRNA. n = 4. f RIP experiments pulling down GFP-tagged Lsm8 were carried out in wild type and lhp1∆. qPCR with the indicated primer pairs is shown and the binding was compared to wild type and normalized to the 18S rRNA. n = 3. g FISH experiments were carried out in wild type and lsm8-1 upon temperature shift for 2 h 30 min to 37 °C. ~30 to 50 nucleotide long Cy3 labelled probes were used for hybridization with the indicated snoRNAs. The DNA was stained with DAPI. n = 3. h Cytoplasmic fractionation experiments were conducted in wild type and lsm8-1 upon temperature shift for 2 h 30 min to 37 °C. Cytoplasmic immature 3’-extended snoRNAs were quantified by qPCR. n = 4. n indicates the number of biological replicates. Data in bar plots (a–g) are presented as mean +/− SD. Two-tailed student’s t-test was used to calculate p values. *p < 0.05; **p < 0.01; ***p < 0.001. Individual data points are represented by black circles.
To analyze if and in which compartment the Lsm-ring is loaded onto the snoRNAs, we repeated the RIP experiments with Lsm8. As Lsm8 is a unique protein of the Lsm2-8 ring, it excludes detection of other Lsm ring complexes, such as Lsm1-7, which plays a role in mRNA decay, and Lsm2-7, which associates with snR5 or the pre-RNase P RNA54. We detected binding of Lsm8 to all tested snoRNAs and observed a ~ 2-20-fold increased binding in the export and the re-import mutants (Fig. 3c, d, Supplementary Fig. 3d–f), suggesting that the Lsm-ring is loaded in the nucleus and stays bound during shuttling, as for Lhp1. Lhp1 was shown to act as a quality control factor for correct Lsm-ring loading for the snRNA U622. To investigate whether such an inter-dependency also occurs in the context of snoRNAs binding, we carried out RIP experiments with Lhp1 in the lsm8-1 mutant. Clearly, the binding increased at the non-permissive temperature (Fig. 3e, Supplementary Fig. 3g), indicating that Lhp1 binds independently of the Lsm-ring attachment and that its dissociation from the RNA is reduced in the absence of Lsm8. However, by contrast, the Lsm-ring loading is not altered in the absence of Lhp1 (Fig. 3f, Supplementary Fig. 3h), suggesting that Lsm-ring binding does not require Lhp1. Importantly, in the absence of a functional Lsm-ring in lsm8-1, the extended snoRNAs accumulate in the cytoplasm (Fig. 3g, Supplementary Fig. 3i), supporting that it is required for the re-import into the nucleus, as shown earlier22. The Lsm-ring serves as a contact for the import receptors Mtr10 and Cse122. In addition to the nucleo-cytoplasmic fractionation experiment, we could also visualize the cytoplasmic accumulation in the lsm8-1 mutant in FISH experiments. We detect an increased cytoplasmic signal for snoRNA 13, 42, and 68, and to a lesser extent for snoRNA U3a, but not for snoRNA 24 (Fig. 3h, Supplementary Fig. 3j). Taken together, these data show that Lhp1 and the Lsm-ring are loaded onto immature shuttling snoRNAs in the nucleus, prior to export, and that proper Lsm-ring assembly is important for the re-import of the cytoplasmic immature pre-snoRNAs into the nucleus via Mtr10 and Cse1.
CPF-CF-terminated snoRNAs are exported into the cytoplasm
From the above experiments, we concluded that some snoRNA transcripts shuttle into the cytoplasm. Importantly, this does not seem to include all snoRNAs, and of those snoRNAs that were shown to shuttle, only a small proportion of each seems to move into the cytoplasm, as judged by the very low cytoplasmic signal visible in the FISH experiments. Even in the import receptor mutants, a nuclear signal was still visible, which further supports that only a fraction of each snoRNA shuttles. This indicates that shuttling is not essential for snoRNA maturation and raises the question of what determines which snoRNA transcripts shuttle and how they differ from the non-shuttling snoRNA transcripts.
Transcription of some snoRNAs was already shown to be terminated by the NNS-complex7,55. However, the terminator sequences of some snoRNAs also contain CPF-CF sites, possibly as fail-safe termination sites as they are located downstream of the NNS sites (Fig. 4a and Supplementary Fig. 4). NNS-mediated termination is coupled with TRAMP/exosome-complex-mediated trimming of the snoRNA precursors17. This NNS-dependent trimming is defective when TRF4 of the TRAMP-complex is deleted. To analyze if, in the situation when NNS-mediated termination is generally impaired, more snoRNAs would be terminated by the CPF-CF fail-safe sites and that this subsequently might lead to nuclear export as for CPF-CF-terminated mRNAs, we carried out several experiments. First of all, we localized the snoRNAs in trf4∆ and found an increased cytoplasmic signal, which was slightly stronger in the trf4∆ mtr10∆ double mutant (Fig. 4b, c, Supplementary Fig. 5a). Moreover, we detected a genetic interaction between TRF4 and MTR10, as the double mutant was strongly impaired in its growth as shown in drop dilution assays, indicating a connected function of these gene products in the same pathway (Supplementary Fig. 5b). The cytoplasmic signal increased visibly for snR13, 42 and 68, but not for snR24, which is an intron-encoded snoRNA. We analyzed these snoRNAs via qPCRs in trf4∆ and showed with reverse primers hybridizing downstream of the NNS sites, that elongated transcripts accumulate in trf4∆ (Fig. 4d). Not only did the elongated forms of the snoRNAs increase, but also the total amount of the transcripts did, as the extended forms are part of the total snoRNAs. The cell might sense a lack of the mature form and upregulate transcription to obtain more mature snoRNAs. Importantly, this elongated form was significantly enriched in the cytoplasm, as shown in qPCRs after nucleo-cytoplasmic fractionation experiments (Fig. 4e, Supplementary Fig. 5c). These longer snoRNAs accumulated between ~13 and ~80-fold, suggesting that most likely all these longer snoRNAs shuttle through the cytoplasm.
a Simplified scheme for the transcription termination sites of the indicated snoRNAs. b FISH experiments were carried out in the indicated strains upon temperature shift for 1 h to 37 °C. ~30 to 50 nucleotide long Cy3 labelled probes were used for hybridization with the indicated snoRNAs. The DNA was stained with DAPI. Since trf4∆ shows much stronger signals, signal intensity in the other three strains was equally increased for better comparison of the nuclear and cytoplasmic localizations. n = 3. c snoRNA signal quantification from (b). 50 cells per sample were chosen. Box plot indicates median (centre), 25th–75th percentile (box) and minimum-maximum values (whiskers). An ordinary one-way ANOVA was used for multiple group comparisons. Total RNA isolation (d) and cytoplasmic fractionation (e) were conducted in wild type and trf4∆. Total snoRNA and CPF-CF terminated snoRNA (indicated as ‘over NNS’) were quantified by qPCR. n = 4. f Scheme of 3’-end PCR. An oligo d(T17) containing adapter primer was used with snoRNA forward primers of to amplify the polyadenylated pre-snoRNAs. g Total RNA from d was used for 3’-end PCR. The transcription termination site was indicated with pink arrows. bp=base pairs, n = 4. RIP experiments pulling down GFP-tagged Nab2 were conducted in wild type (h) or wild type and nrd1-102 (i) upon temperature shift for 1 h to 37 °C. qPCR amplifying the total snoRNA is shown, and the binding was compared to lysate (h) or wild type related to precipitated Nab2 amount (i) and normalized to the 18S rRNA. The mRNA ZWF1 served as positive (h) and negative (i) control. h n = 3, i n = 5. j The RNA from (i) was used for 3’-end PCR. The transcription termination site was indicated with pink arrows. n = 3. k Quantification of CPF-CF terminated snoRNAs from (j). n = 3. n indicates the number of biological replicates. Data in bar plots (d, e, h, i, k) are presented as mean values +/− SD. Two-tailed student’s t-test or one-way ANOVA was used to calculate p values. *p < 0.05; **p < 0.01; ***p < 0.001. Individual data points are represented by black circles.
These experiments suggest that impaired NNS termination might result in a shift to the downstream located CPF-CF sites. This would mean that those snoRNA transcripts would be polyadenylated, because CPF-CF-mediated termination results in the addition of a poly(A) tail56. To investigate if, in trf4∆ where the NNS-mediated termination is affected, snoRNA termination occurs downstream at CPF-CF sites, we carried out 3’ end PCRs to identify polyadenylated snoRNAs. We used an oligo d(T) reverse primer with gene-specific forward primers to amplify exclusively polyadenylated RNAs and separated them by length via agarose gel electrophoresis (Fig. 4f). In trf4∆, increased amounts of the CPF-CF-terminated polyadenylated transcripts were detected for snR13, 42, 65, and 128. The mRNA ACT1 served as a control for a CPF-CF-terminated transcript, which is not affected by the absence of Trf4. Notably, in wild type, low levels of polyadenylated transcripts were detected, indicating that some snoRNA transcripts might naturally receive a poly(A) tail. However, in trf4∆, the polyadenylated transcripts were longer and correspond to the location of the CPF-CF sites located downstream of the NNS sites (Fig. 4g). snR68 showed a similar phenotype in both the wild type and the trf4∆ strain, with a band that, based on its size, represents an amplification artifact. Within the termination sequence of snR68, there is a stretch of 11 adenosines, located before the NNS and the CPF-CF sites. Most likely, the oligo d(T) primer annealed to this sequence, resulting in the amplification of a <200 bp product. This artifact prevented the detection of the real termination site(s) of snR68 in this assay.
Poly(A) tail-containing mRNAs that were terminated by the CPF-CF complex bind the guard proteins Hrp1 and Nab2. Hrp1 controls the cleavage process and recruits, upon successful cleavage, the export receptor Mex67 to the RNA. On the RNA, Hrp1 binds to the efficiency element, interacts with Rna14 of the CPF-CF complex, and is the only CPF-CF complex component that moves with the polyadenylated RNAs into the cytoplasm36,39. Nab2 quality controls the successful addition of the poly(A) tail by binding several times to it and subsequently recruiting several molecules of Mex67 for nuclear export37,39,57,58,59. As both proteins are typical quality control factors for CPF-CF termination, we wanted to analyze whether they would bind to snoRNAs. In wild-type cells, RIP experiments with Nab2 revealed some binding for all analyzed snoRNAs, except snR24 and 128 (Fig. 4h, Supplementary Fig. 5d). The mRNA ZWF1 served as a positive control. Importantly, in a mutant of the NNS complex component NRD1, nrd1-102, the interaction of Nab2 with many snoRNAs significantly increased compared to wild type (Fig. 4i, Supplementary Fig. 5e), indicating a shift in the termination mode towards CPF-CF mediated termination. In this case, the mRNA ZWF1 served as a negative control as its binding to Nab2 was not affected. To further identify the termination site of Nab2-bound snoRNAs, the RNA recovered from nrd1-102 was used for cDNA synthesis and subsequent 3’ end PCR experiments. Indeed, CPF-CF-terminated transcripts are enriched in the eluate (Fig. 4j, k), indicating that the decreased NNS-mediated termination results in more Nab2-bound snoRNAs as a result of the CPF-CF-mediated termination. Like Nab2, Hrp1 also binds to some analyzed snoRNAs in wild-type cells (Supplementary Fig. 5f, g). Furthermore, we identified binding of the guard proteins Npl3 and Gbp2 to some snoRNAs (Supplementary Fig. 5g–k), which might support the nuclear export of the shuttling snoRNAs through Mex67 recruitment. The more Mex67 is bound to an RNA, the better it is exported39. These results suggest that CPF-CF-terminated transcripts are exported into the cytoplasm, assisted by guard proteins that recruit Mex67.
Mutation of the NNS sites in SNR13 force CPF-CF termination
To directly investigate whether a forced termination with the CPF-CF complex would lead to increased cytoplasmic accumulation of a snoRNA, we constructed an NNS mutant (pSNR13-NNS*), which was expressed from a plasmid and compared it to a wild type construct (pSNR13) of the snoRNA snR13, either of which were expressed in a SNR13 knock out strain (Fig. 5a, Supplementary Fig. 6a). Through 3’ end PCR, significantly more CPF-CF terminated elongated transcripts were detected in the NNS mutant compared to the wild type SNR13-containing plasmid. The artificial polyadenylation conducted by the Poly(A) polymerase (Pap) allowed the detection of the mature snR13. Importantly, the level of the mature snR13 remains unchanged upon the NNS binding site mutation (Fig. 5b, c), suggesting that processing is not disturbed. Similar results were obtained by qPCR analysis, and a strong increase of read-through transcripts was detected in the NNS mutant as well. Although most of the elongated transcripts end at the CPF-CF sites 1, 2 or 3, around 10% of the transcripts were read through these sites and might be terminated at the CPF-CF site of the neighboring gene TRS31 (Supplementary Fig. 6b)55. To see if these CPF-CF terminated transcripts shuttle between the nucleus and the cytoplasm, we carried out FISH experiments with a Cy3-labelled snoRNA-specific probe. Indeed, a weak cytoplasmic pool of the snoRNA was visible in the cytoplasm of wild-type cells (Fig. 5d, Supplementary Fig. 6c), and this was more than 10-fold increased in the NNS mutant, as judged by nucleo-cytoplasmic fractionation experiments (Fig. 5e, f). For the NNS mutant, the cytoplasmic FISH signal increased even further in the double import receptor mutant cse1-1 mtr10∆ at its non-permissive temperature (Fig. 5g, Supplementary Fig. 6d). Nucleo-cytoplasmic fractionation experiments revealed this increase to be more than 2-fold compared to cells expressing the wild type import receptors (Fig. 5h, i). These experiments show that mutation of the NNS site in SNR13 leads to elongated CPF-CF-terminated transcripts that increasingly shuttle into the cytoplasm.
a Partial map of pSNR13 and pSNR13-NNS*. Mutated nucleotides of terminator I are indicated in red. CPF-CF sites of terminator II determined from Fig. 4g, j are marked with pink arrows. b Untreated or Poly(A) polymerase (Pap) treated total RNA isolated in SNR13 and SNR13-NNS* was used for 3’ end PCR. The polyadenylated and mature snR13 was indicated with pink and green arrows, respectively. bp = base pairs, n = 3. c Quantification of snR13 transcripts from b. n = 3. FISH (d, g) and cytoplasmic fractionation (e, f, h, i) were performed in SNR13 and SNR13 NNS* expressed in a wild typic background (d–f)) or SNR13 NNS* expressed in the wild typic and a cse1-1 mtr10∆ background (g–i) upon temperature shift for 1 h 15 min to 16 °C. For FISH, a 50 nt long Cy3 labelled probe against snR13 was used for detection, and the DNA was stained with DAPI. For cytoplasmic fractionation, the correct separation of the fractions was verified by western blot for the cytoplasmic Zwf1 and the nuclear Nop1 (e) or Yra1 (h). kDa=kilodaltons. qPCR of cytoplasmic CPF-CF-terminated snR13 normalized to 18S rRNA is shown in (f) and (i). d–g n = 3, h, i n = 7. j–l RIP experiments precipitating GFP-tagged Nab2 and Hrp1 were carried out in SNR13 and SNR13-NNS*. Western blot of the RIP experiment validates the pull-down of the GFP-tagged Nab2 (j) and Hrp1 (k). Hem15 served as a negative control. kDa=kilodaltons. Nab2 and Hrp1 binding to CPF-CF terminated snR13 in the eluate is shown in (l) and the binding was compared to wild type and was normalized to the 18S rRNA. n = 3. (M) RNA from (l) was used for 3’-end PCR. The transcription termination site was indicated with pink arrows. n = 3. n indicates the number of biological replicates. Data in bar plots (c, f, i, l) are presented as mean +/− SD. A two-tailed Student’s t-test was used to calculate p-values. *p < 0.05; **p < 0.01; ***p < 0.001. Individual data points are represented by black circles.
The co-transcriptional loading of guard proteins determines whether a snoRNA transcript is exported
CPF-CF-terminated mRNAs are controlled for proper cleavage by the guard protein Hrp1 and for the correct addition of the poly(A) tail by Nab2 binding39. These proteins act as nuclear retention factors until cleavage and poly(A) tail addition are successfully completed, and only then do they recruit the export receptor Mex67 to allow passage of the mRNP through the nuclear pore complex (NPC). To investigate whether the CPF-CF termination-controlling guard proteins, Hrp1 and Nab2, bind to CPF-CF-terminated snR13, we carried out RIP experiments in the SNR13 knock-out strain, expressing with plasmid-derived wild-type or NNS mutant snR13. We detected some binding of wild-type snR13 but importantly, this was increased significantly for the NNS mutant (Fig. 5j–l). To monitor 3’ end status of the Nab2-bound snR13 transcripts, we carried out 3’ end PCRs using the RNA samples derived from RIP experiments. The Nab2-bound transcripts of wild type SNR13 and SNR13-NNS* were amplified with a reverse oligo d(T17) primer and a sequence-specific forward primer, which revealed that a) more poly(A)-tailed product was precipitated from the NNS mutant and b) that termination did not only occur on CPF-CF site 3, but rather downstream of this site, termed read-through (Fig. 5m). A similar outcome was observed for Hrp1 (Fig. 5m). Together these results show that the CPF-CF-mediated transcription termination of snoRNAs is a fail-safe mechanism for defects in the NNS termination. It furthermore shows that CPF-CF termination leads to the export of snoRNAs into the cytoplasm, via the guard protein-mediated recruitment of Mex67.
As described above, Lhp1 associates with pre-snoRNAs during maturation. It has been shown that Lhp1 facilitates the nuclear export of the snRNA U6 by recruiting Mex6722 and it seems possible that Lhp1 might have a similar function on snoRNAs as well. However, when snoRNAs are terminated by the NNS pathway, their precursors are initially also bound to Lhp1. Since NNS-terminated snoRNAs are not considered to shuttle, it is contradictory if Lhp1 recruits Mex67 for these transcripts. To verify whether the Mex67 recruitment by Lhp1 is dependent on the transcription termination mechanism, co-immunoprecipitation (co-IP) between Lhp1 and Mex67 was conducted in wild type and in cft2-1 after a temperature shift for 1 h to 37 °C. Cft2 is a subunit of the CPF complex and is crucial for proper CPF-CF termination60. GFP-tagged Lhp1 was precipitated and the associated Mex67 was detected in both strains (Fig. 6a). After quantification, the relative interaction between Lhp1 and Mex67 was significantly reduced to nearly 60% in cft2-1 (Fig. 6b), revealing that Lhp1 might recruit Mex67 only when snoRNAs are terminated via the CPF-CF pathway. A co-transcriptional assembly of box C/D snoRNPs has been suggested by several studies52,61. This assembly might finally replace Lhp1 and thus prevent export of NNS-terminated snoRNAs. However, when the CPF-CF-terminated snoRNAs are exported via the Lhp1-, Hrp1-, and Nab2-bound Mex67, they can return to the nucleus due to the interaction of the bound Lsm-ring with the import receptors Mtr10 and Cse1. As it remains unclear in which compartment the core proteins are assembled on the snoRNAs when they shuttle, we investigated the interaction between the CPF-CF-terminated snR13 in wild-type SNR13 or SNR13-NNS* with two core proteins, the early associating Snu13 and the later recruited Nop161,62. We found that in the double import mutant cse1-1 mtr10∆, the binding of Snu13 was increased, while the Nop1 binding slightly decreased (Fig. 6c, d, f, Supplementary Fig. 6e). Also, a slight mis-localization of Snu13 to the cytoplasm was observed in the double import mutant cse1-1 mtr10∆ expressing SNR13-NNS* (Supplementary Fig. 6f). Even though the pull down of Nop1 was lower in the double import mutant, which might be due to reduced protein amounts, visible already in the lysate, when equal amounts of precipitated RNA were compared, we found a reduced binding of the core protein to SNR13-NNS*. As CPF-CF terminated snR13 mislocalizes to the cytoplasm in cse1-1 mtr10∆, and Nop1 is a canonical nuclear protein, this outcome supports a model in which the association of Nop1 to CPF-CF terminated snR13 occurs probably after re-import, while Snu13 binds prior to the re-import of SNR13. To distinguish if Snu13 joins the complex before export or in the cytoplasm, we repeated the RIP experiment in the mex67-5 export mutant. We found that the binding of Snu13 and the snoRNAs increased, which indicates that this core protein binds prior to export (Fig. 6e, f). It appears that Snu13 binding is not able to retain the snoRNA in the nucleus, either because Mex67 association outweighs retention, or because Snu13 is masked, e.g. by some guard proteins. Nevertheless, although snoRNP assembly is interrupted through the cytoplasmic phase, it is not prevented. To investigate whether the 3’ end processing of CPF-CF-terminated snoRNA transcripts occurs already in the cytoplasm, 3’end PCR was carried out in wild type and a strain lacking Ski2, a cofactor of the cytoplasmic exosome. This demonstrated that the level of the polyadenylated snoRNA species was not affected, suggesting that the processing occurs after the re-import (Supplementary Fig. 6g). This fits well to the detection of the extended forms in the export and import mutants (Figs. 1e and 2b). Therefore, these finding provoke the question whether these shuttling snoRNAs will, despite their export and re-import, mature to functional snoRNAs that are assembled into snoRNPs capable of rRNA modification.
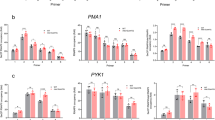
a Western blot of co-IP precipitating GFP-tagged Lhp1 and Mex67 is shown. Hem15 served as a negative control. n = 4. b Quantification of the interaction from (a). n = 4. RIP experiments precipitating GFP-tagged Snu13 (c) and Nop1 (d) in SNR13 and SNR13-NNS* with a wild type and cse1-1 mtr10∆ background upon temperature shift for 1 h 15 min to 16 °C is shown. For Snu13, RIP experiments precipitating GFP-tagged Snu13 (e) in SNR13 and SNR13-NNS* in the mex67-5 mutant either unshifted (25 °C) or shifted for 1 h to 37 °C is shown. Western blot validates the pull down. Aco1 served as a negative control. c n = 3, d, e n = 4. f The binding of core proteins to CPF-CF terminated snR13 in mex67-5 or cse1-1 mtr10∆ is shown and the binding was compared to wild type and normalized to a control RNA (for Snu13:ZWF1 mRNA, for Nop1: 21S rRNA). g Scheme for the primer extension method. The modification site on RNA was marked with a green circle. Under low dNTPs the reverse transcription stops at the modification site, while under high dNTPs the reverse transcriptase reads through the site. h 2’-O-methylation sites in the 25S rRNA guided by snR13 were detected in the indicated strains and are marked with black arrows. b = bases, n = 3. i cdc28-13 cells were synchronized and arrested at G1 by shifting them to 37 °C for 3 h. Subsequently, the cells were shifted back to 25 °C for release. Cells were harvested before the temperature shift, 0 min and 60 min after the arrest release. Total RNA was isolated and used for 3’ end PCR. The CPF-CF-terminated snR13 are indicated with a pink arrow. bp = base pairs, n = 4. j Quantification of the main CPF-CF terminated snR13 transcripts from i. n = 4. n indicates the number of biological replicates. Data in bar plots (b, f, j) are presented as mean +/− SD. A two-tailed Student’s t-test was used to calculate p-values. *p < 0.05; **p < 0.01; ***p < 0.001. Individual data points are represented by black circles.
To determine whether an snR13 transcript that was CPF-CF terminated and had to shuttle through the cytoplasm was able to guide rRNA modification, the 2’-O-methylation of 25S-A2280 and 25S-A2281 was monitored in cells lacking endogenous snR13 or expressing plasmid-derived wild-type or NNS mutant snR13. Total RNAs extracted from these strains were subjected to primer extension using a labelled oligonucleotide hybridizing downstream of the modification sites of interest. Reactions were performed in the presence of high and low amounts of dNTPs as decreasing the availability of nucleotides increases the probability of reverse transcriptase stalling at 2’-O-methylated sites (Fig. 6g). The modification of both nucleotides was detected in wild type cells, albeit it Am2281 more strongly than Am2280, and these modifications were lost in the absence of snR13 (Fig. 6h). Re-expression of either wild type or the NNS mutant snR13 restored the modifications to wild type levels (Fig. 6h), demonstrating that CPF-CF-mediated transcription termination of the snoRNA and its shuttling through the cytoplasm does not impact its ability to form a functional snoRNP complex.
An important reason for this CPF-CF-mediated fail-safe termination mechanism is probably to save the energy that was invested in the synthesis of the snoRNA when NNS termination does not take place. However, when does this occur in cells? The number of CPF-CF terminated transcripts is low, and the poly(A) tail containing transcripts are below 5% (Supplementary Fig. 6b). Interestingly, it was shown that the level of Sen1 from the NNS-complex fluctuates during the cell cycle; Sen1 levels decrease during G1, and this decrease was shown to result in reduced transcription termination of a reporter gene63,64. To investigate the preferences of the termination mode during the cell cycle, we synchronized cells of the cdc28-13 mutant by shifting it to 37 °C for 3 h in the G1 phase. Subsequently, we released the arrest by shifting the strain back to 25°C. Total RNA was isolated from the cells at 0 and 60 minutes upon the release and analyzed via 3’ end PCR. When compared to asynchronously growing cells, we clearly see a reduction in the CPF-CF terminated snR13 in the G1 phase and a strong increase after 60 minutes when cells have reached the G2 phase (Fig. 6i, j). This result supports that the availability of the NNS-termination complex is not consistent and shows that the termination mode of snR13 alters during the cell cycle. This highlights the requirement for an alternative termination strategy, namely CFP-CF, to maintain snoRNA production.
In yeast, most snoRNAs are encoded individually, while only a few are intronic. In human cells, it is the other way around; most snoRNAs arise from intronic regions and only a few are independently transcribed. The individually encoded snoRNAs were suggested to be terminated by the Integrator complex, which could be the functional analog to the yeast NNS-complex, as it facilitates transcription termination of rather short transcripts without addition of a poly(A) tail65,66. Interestingly, it was also proposed that components of the CPSF-CF complex (the homolog to the yeast CPF-CF-complex) influence their transcription termination67. To investigate whether CPSF-CF-mediated termination might also affect snoRNA termination, at least to some extent, we divided the snoRNAs into two groups: intronic and independent, and analyzed the mature transcript ends for the presence of PAS sites. For individually transcribed snoRNAs, we found an enrichment of the CPSF-CF positioning element (AATAAA) and the efficiency element (TATATA) downstream of the transcript end sites (TES) compared to randomized sequences and also compared to intronic snoRNAs (Fig. 7a, Supplementary Fig. 7a–c). To see whether these sites might serve as fail-safe termination sites, we analyzed available RNA-seq data generated upon a 4 h depletion of the largest subunit of the Integrator complex, INTS1, from HEK293T cells and analyzed the area 100 nt upstream and 150 nt downstream of the TES68. Upon INTS1 depletion, we detected reduced coverage for the individually transcribed snoRNAs upstream of the TES and increased coverage downstream, indicating the presence of longer transcripts; however, no change was observed for the intronic snoRNAs (Fig. 7c, d). This suggests that in the absence of a functional Integrator, individually transcribed snoRNAs might be terminated via the CPSF-CF complex. To confirm experimentally that the extended snoRNAs are polyadenylated, 3’-end PCR was carried out with the randomly-chosen, individually-encoded snoRNA, SNORA51L9. Indeed, we found polyadenylated transcripts extended beyond the mature form at putative PAS sites (Fig. 7d, Supplementary Fig. 7d). These data suggest that a similar mechanism as in yeast cells might exist for human individually-encoded snoRNAs; if not terminated by the NNS/Integrator complex, these short transcripts are terminated by the CPF-CF/CPSF-CF complex, resulting in polyadenylated transcripts.
A tail. a Percentual presence of AATAAA motifs is shown around the transcript end site (-20 and +150 nucleotide) for individual human snoRNAs compared to a set of random sequences. Percentual read coverage of untreated (grey) and INTS1 depleted (blue) HEK293T cells is shown for individual (b) and intronic (c) snoRNAs around the transcript end site. d Untreated or Poly(A) polymerase (Pap) treated total RNA isolated from HCT116 cells was used for 3’ end PCR. Polyadenylated immature and mature SNORA51L9 transcripts are indicated in the blue and red boxes, respectively. bp = base pairs, n = 3 biological replicates.
Taken together, we suggest alternative pathways for snoRNA maturation, which are distinguished by NNS- and CPF-CF-mediated transcription termination in yeast (Fig. 8). While NNS-terminated transcripts are not exported to the cytoplasm and associate with their protein components in the nucleus, the CPF-CF-terminated transcripts are exported into the cytoplasm. The CPF-CF sites are located downstream of the NNS sites and act as fail-safe sites for perturbing NNS-mediated termination. The binding of the guard proteins Hrp1 and Nab2 supports snoRNA export through the recruitment of the export receptor Mex67. Re-import is facilitated by the Lsm-ring, which contacts the import receptors Cse1 and Mtr10. Final maturation, including transcript trimming and snoRNP protein assembly, occurs in the nucleus to form a functional snoRNP competent for RNA modification.
After the transcription by RNAP II, pre-snoRNAs undergo two different maturation pathways depending on their transcription termination. While the majority of snoRNAs is terminated by the NNS complex, snoRNAs that escape the NNS termination sites are terminated further downstream by the CPF-CF machinery. The NNS-terminated snoRNAs are mostly oligoadenylated, whereas CPF-CF terminated snoRNAs own a poly(A) tail. Coupled to the NNS termination, core proteins are loaded co- or post-transcriptionally, assisted by various assembly factors. Lhp1, as well as the Lsm2-8 complex, binds to the oligo(U) tract at the 3’-end of pre-snoRNAs, and oligoadenylated snoRNAs are rapidly processed in the nucleus. Besides the core protein Snu13, several guard proteins such as Gbp2, Npl3, Hrp1, and Nab2 associate with CPF-CF-terminated snoRNAs and recruit the export receptor Mex67-Mtr2 heterodimer for the nuclear export. Lhp1 and Lsm2-8 complex also bind to the CPF-CF-terminated snoRNAs. Similar to the guard proteins, Lhp1 can also recruit Mex67-Mtr2. During the cytoplasmic phase, the Mex67-Mtr2 and guard proteins dissociate. Lhp1 and Lsm2-8 remain bound to the pre-snoRNAs to protect them and to allow the binding of the import receptors Cse1 and Mtr10 to the Lsm-ring, to mediate the re-import. Subsequently, the remaining core proteins, such as Nop1, assemble with the CPF-CF-terminated snoRNAs. The assembly factors, Lhp1 and Lsm2-8, leave the snoRNPs to allow the final 3’-end trimming by the nuclear, Rrp6-containing exosome. Finally, mature snoRNPs reside in their final destination, the nucleolus. RNAP II = RNA polymerase II.
Discussion
snoRNAs are not only synthesized in the nucleus but they also predominantly function in this compartment as they modify mainly rRNAs in the nucleolus69,70. Although cytoplasmic functions of some snoRNAs have been described in human cells45,71,72, this is not the case for yeast snoRNAs, which likely only act within the nuclear compartment. To fulfill their functions as RNA modification guides, snoRNAs associate with different proteins, Nop1, Nop56, Nop58, and Snu13 in the case of C/D-box snoRNAs and Cbf5, Nop10, Nhp2 and Gar1 for H/ACA box snoRNAs. All proteins of the snoRNPs are nuclear and are loaded co- or post-transcriptionally onto the pre-snoRNAs73. Therefore, the current literature suggests that yeast snoRNAs normally do not leave the nucleus. The observation that snoRNAs can interact with the nucleo-cytoplasmic shuttling mRNA guard proteins Gbp2, Hrp1, and Nab2, as well as the export receptor Mex67, therefore remained unexplained32 as, so far, only a handful of snoRNAs have been found in the cytoplasm under particular circumstances42,43,74. These findings raised the possibility that snoRNAs leaving the nucleus may be a more wide-spread phenomena than previously thought, motivating us to examine their presence in cytoplasmic fractionation experiments46. We confirmed that several snoRNAs are detectable in the cytoplasm of logarithmically growing wild-type cells (Fig. 1a). Remarkably, these snoRNAs shifted their localization towards the nucleus when the RNA export pathway was blocked in the mex67-5 xpo1-1 double mutant strain, suggesting that the portion of these snoRNAs that are usually exported into the cytoplasm are then retained in the nucleus. To identify what specifies the translocation of snoRNA transcripts to the cytoplasm, we systematically analyzed the exported snoRNAs and found a rather random distribution of C/D and H/ACA box snoRNAs that included capped, uncapped, monocistronic, polycistronic and intron-encoded snoRNAs (Fig. 1b). Moreover, the amount of the shuttling snoRNAs of each particular type seemed low, because in FISH experiments with short sequence specific Cy3-labelled probes (<50 nucleotides), individual snoRNAs were not detectable in the cytoplasm of wild type cells (Fig. 2a). Detection of the cytoplasmic pool was only possible when using a long, sequence-specific probes (between 75 and 200 nucleotides) that encounter difficulties in traversing the nuclear envelope and entering the nuclear compartment (Fig. 1f). However, also in this case most of the signal was still nuclear, indicating that either only a very low proportion of snoRNA transcripts move to the cytoplasm or they remain there only for a very short time in their life cycle. The reason why some of them would leave the nucleus only became clear when we analyzed how their transcription is terminated. Usually, short ncRNAs are terminated via the NNS-system, and pervasive transcripts and CUTs are subsequently degraded by an NNS-connected TRAMP/exosome-complex mediated degradation mechanism7,11,12,13. snoRNAs are mainly NNS-terminated as well. NNS-mediated termination elicits the binding of the TRAMP complex that marks the RNA with an oligo(A) tail for its subsequent recognition and processing by the nuclear exosome16,17. However, snoRNAs are protected from degradation by their immediate association with snoRNP core proteins and strong secondary structures14,15. For the snoRNA snR13, it was shown that the oligoadenylated RNA is not degraded as e.g., CUTs are, but that its tail is rather elongated by the poly(A) polymerase Pap114.
Interestingly, many snoRNAs contain not only one NNS-binding motif, but often several and in addition, also CPF-CF motifs (Fig. 4a), as has been reported previously for SNR13 and SNR6514. A usual arrangement seems to be that the NNS site(s) are located upstream of the CPF-CF site(s), suggesting that the CPF-CF sites are rather fail-safe termination sites in case transcription is not terminated at the upstream NNS sites. CPF-CF termination is typical for mRNAs and results in loading of the guard protein Hrp1 close to the cleavage site and the guard protein Nab2, which is loaded onto the poly(A) tail36,37,39. These guard proteins quality control the successful cleavage and addition of a poly(A)tail and signal this through recruitment of the export receptor Mex67. This mode of termination thus drives export of snoRNAs, because the recruitment of Mex67 to an RNA enables it to pass through the NPC38,39,41. Thus, CPF-CF terminated snoRNAs are more prone to be exported and indeed, mutation of the NNS sites in SNR13 resulted in increased nuclear export of this snoRNA (Fig. 5). Other guard proteins such as Gbp2, Lhp1 and Npl3 associate at least with some snoRNAs (Fig. 3 and Supplementary Fig. 5) and32 and might support Mex67-mediated nuclear export of some snoRNAs under certain circumstances. Interestingly, Npl3 interacts with the two Lsm-ring binding site-interacting proteins Lhp1 and Lsm8, indicating that the action of this guard protein is not limited to mRNA but also surveils proper maturation of ncRNAs22. Importantly, Lhp1 was suggested to fit well into the group of guard proteins, as it shows not only high structural similarities with them, but also functional resemblance in surveilling correct maturation of the RNP and recruiting Mex67. In the case of the snoRNA maturation, it supports nuclear export by recruitment of Mex67, when CPF-CF is terminated and Lsm-ring bound, like the snRNA U6 (Fig. 6a, b and ref. 22). This is necessary for re-import to the nucleus via Cse1 and Mtr10, which contact the Lsm-ring (Fig. 2)22. It seems likely that the complete association of the core proteins might take place, at least in part, after shuttling, because guard proteins are bound to recruit the export receptor Mex67 and we have shown that the binding of Nop1 to the CPF-CF terminated pre-snR13 (SNR13-NNS*) is reduced in the cse1-1 mtr10∆ import mutant (Fig. 6f). However, importantly CPF-CF terminated, exported snR13 becomes a functional RNP, after re-import, as it is still able to modify the rRNA (Fig. 6h). This mechanism of fail-safe termination via the CPF-CF complex does not seem to be restricted to yeast as human individually-encoded snoRNAs also have PAS sites and can be terminated via the CPSF-CF complex, resulting in a polyadenylated snoRNA (Fig. 7 and Supplementary Fig. 7).
Taken together, we suggest a model (Fig. 8) in which snoRNAs remain nuclear when they are terminated by the NNS-complex. However, in case this termination is impaired, potentially when NNS factors are limited or work inefficiently, many snoRNAs are instead terminated at CPF-CF sites and through the intrinsic mechanism of loading the guard proteins and, in turn, Mex67, commit the snoRNAs for nuclear export. Fail-safe termination is important for several reasons. First, it ensures that the energy spent in making the RNA is not wasted and functional RNAs can be produced from the extended forms (Fig. 6h). Secondly, it is an important mechanism for ensuring transcriptome integrity. Extending transcripts not only traps RNA polymerases, but ongoing transcription can also interfere with the transcription of neighboring genes and increase the danger of colliding polymerases and DNA damage. Generally, transcription and replication conflicts can result in DNA strand breaks, leading to diseases such as cancer in multicellular organisms75,76. Therefore, not only does transcription initiation have a regulatory importance, but termination can also impact the fate of RNAs. In fact, the availability of the NNS complex fluctuates within the cell cycle. The Sen1 level decreases during G1 phase, leading to a reduced efficiency of NNS termination63,64. The level of the CPF-CF-terminated snR13 differs during the cell cycle as well, and more polyadenylated transcripts can be detected in G2 (Fig. 6i, j), possibly due to the limited availability of Sen1. Moreover, besides the termination of short ncRNAs, Nrd1 and Nab3 have additional individual roles in the quality control of overlong transcripts77. If many such transcripts are generated, the availability of the proteins in the NNS complex might not be sufficient, so NNS termination fails. In this case, the fail-safe CPF-CF termination becomes relevant to prevent run-on transcription and produce sufficient snoRNAs.
Our findings furthermore suggest that the NNS-termination on snoRNAs retains these transcripts in the nucleus, while the CPF-CF termination results in the export of transcripts. For mRNAs, the CPF-CF mediated termination mode generally occurs, and coding transcripts are always exported as translation occurs in the cytoplasm. By contrast, no NNS-terminated transcript has so far been suggested to leave the nucleus, including snoRNAs. Our study resolves why some exceptions can be found for snoRNAs and suggests that the mode of termination determines whether an RNA is exported, which can be generalized to all independently transcribed RNAs.
Methods
Yeast strains, plasmids, and oligonucleotides
All yeast strains used in this study are listed in the Supplementary Table 1, plasmids in Supplementary Table 2 and oligonucleotides in Supplemental Tables 3–8. Yeast cells were cultivated either in YPD full medium or in selective media containing a plasmid. Cells were incubated at 25°C until the log phase (1–3 × 107 cells/ml) and subsequently harvested. For temperature-sensitive mutants, cells were shifted to the appropriate temperature for the indicated time. Plasmids were cloned either via Gibson assembly or restriction-free cloning78 and yeast strains were transformed with plasmids by standard methods79. Oligonucleotides were generated from Sigma-Aldrich.
Fluorescent in situ hybridization (FISH)
The experiments were essentially carried out as described earlier18. The probes were either Cy3-labeled oligonucleotides (Sigma-Aldrich) or digoxygenin (DIG)-labeled PCR products that were detected with a FITC-labelled anti-digoxygenin antibody. Cells were grown to the log phase (1–3 × 107 cells/ml) and shifted to the indicated temperatures if necessary. Cells were fixed by adding formaldehyde to a final concentration of 3,7% for 15 min at the relevant temperature and an additional 45 min at room temperature. Subsequently, cells were washed with P-solution (0.1 M phosphate buffer pH 6.5, 1.2 M sorbitol) and spheroplasted by adding 10 mM DTT and zymoylase (10 mg/ml). The cells were subsequently applied to 0.3% poly-L-lysine coated slides for 30 min at 4°C and permeabilized in P solution with 0.5% Triton X-100 for 10 min at room temperature. In the case of DIG-labeled probes, cells were further equilibrated by 0.1 M triethanolamine, pH 8 (TEA) for 2 min at room temperature, and the polar groups were blocked by 0.25% acetic anhydride, 0.1 M TEA for 10 min at room temperature. Afterwards, cells were pre-hybridized with Hybmix (50 % deionized formamide, 5× SSC (0.3 mM NaCl, 30 mM sodium citrate, pH 7), 1x Denhardts, 500 μg/ml tRNA, 500 μg/ml salmon sperm DNA, 50 μg/ml heparin, 2.5 mM EDTA pH 8.0, 0.1% Tween-20) for 1 h at 37 °C and hybridized in Hybmix with the specific probe over night at 37 °C. Cells were washed with 2x SSC and 1x SSC at room temperature for 1 h, respectively, followed by 0.5x SSC at 37 °C and room temperature for 30 min, respectively. For snR13, 4xSSC, 3x SSC, and 2x SSC were used in the first three washing steps instead. For DIG-labeled probes, cells were further blocked in antibody blocking buffer (ABB) (1x PBS, 5% heat-inactivated FCS, 0.3% Tween-20) for 1 h at room temperature, stained in ABB with anti-DIG-Fab-Cy3 (1:200) over night at 4°C and washed with ABB twice for 15 min, one time for 30 min and with PBS/0.1% Tween-20 for 30 min. The nuclei were stained with DAPI in PBS (1:10,000) and washed with PBS/0.1% Tween-20 for 5 min and PBS two times for 5 min. The slide was dried, coated with mounting medium (2% n-propyl gallate, 80% glycerol, 20% PBS pH 8), and sealed. Microscopy studies were carried out with a Leica AF6000 microscope, and pictures were obtained by using the LEICA DFC360FX camera and the LAS AF 2.7.3.9 software (Leica).
Total RNA Isolation
Cells were grown to the log phase (1–3 × 107 cells/ml) and shifted to the appropriate temperature for the indicated times. Subsequently, cells were harvested and lysed in RIP buffer (25 mM Tris HCl pH 7.5, 150 mM NaCl, 2 mM MgCl2, 0.5% (v/v) Triton X-100, 0.2 mM PMSF, 500 µM DTT, 1U/ml RiboLock RNase Inhibitor (Thermo Fisher Scientific)) using the FastPrep®-24 machine (MP Biomedicals) three times for 30 s at 5.5 m/s. The total RNAs were extracted via TRIzol (Ambion® RNA by Life technologies™)-chloroform. The RNA was mixed with 0.1 volume of the 10x TURBO DNase Buffer and 1 µl of TURBO DNase (Invitrogen by Thermo Fisher Scientific). The mixture was incubated for 30 min at 37 °C, before the TURBO DNase was deactivated by adding 0.1 volume of the DNase Inactivation Reagent and incubation for 5 min at room temperature. The purified RNA was measured via Nanodrop, and equal amounts (200–400 ng) of RNA were reverse transcribed with FastGene Scriptase II (Nippon Genetics) using random hexamer primers (Thermo Fisher Scientific) according to the manufacturer’s description. The resulting cDNA was diluted to the same concentration (either 1 or 2 ng/µl) for subsequent qPCR analyses.
HCT116 cells were grown in RPMI1640 medium supplemented with 10% fetal calf serum and 1% penicillin/streptomycin (Sigma, Germany) and cultured at 37 °C and 5% CO2. Cells were collected using PBS supplemented with 0.5 mM EDTA. Subsequently, RNA was extracted by adding 1 ml TRIzol following the protocol described above.
Cytoplasmic fractionation
Cells were grown to the log phase (1–2 × 107 cells/ml). After harvesting, cells were washed with YPD/ 1 M sorbitol/ 2 mM DTT and spheroplasted in YPD/ 1 M sorbitol/ 1 mM DTT by adding zymolyase (100 mg/ml). Cells were subsequently transferred into 50 ml YPD/ 1 M sorbitol and incubated for 30 min at 25°C prior to the indicated temperature shifts. The cultures were split into 10 ml for the preparation of the ‘total cell lysate’ and 40 ml for the ‘cytoplasmic fraction’. Cells were lysed in PBSKMT buffer (1x PBS, pH 7.4, 3 mM KCl, 2.5 mM MgCl2, 0.5% (v/v) Triton X-100, protease inhibitor (Roche)) using the FastPrep®-24 machine (MP Biomedicals) three times for 30 s at 5.5 m/s. For the cytoplasmic fractionation, the cells were lysed in 500 µl lysis buffer (18 % Ficoll 400, 10 mM HEPES pH 6.0), 1 ml buffer A (50 mM NaCl, 1 mM MgCl2, 10 mM HEPES pH 6.0), and 1 µl Ribolock RNase Inhibitor (Thermo Fisher Scientific), followed by vortexing and centrifugation to obtain the supernatant as the cytoplasmic fraction. To confirm the successful separation of the cytoplasmic fraction, the total lysate and cytoplasmic fraction were analyzed by western blot using antibodies of the nucleolar protein Nop1, the nuclear protein Yra1 and the cytoplasmic protein Zwf1. The total and cytoplasmic RNAs were extracted by Trizol-chloroform (Ambion® RNA by Life technologies™) as described above. The purified RNA was measured via Nanodrop, and the same amount (200–400 ng) of RNA was reverse transcribed with FastGene Scriptase II (Nippon Genetics) by using random hexamer primer (Thermo Fisher Scientific) according to the manufacturer’s description. The resulting cDNA was diluted to the same concentration (either 1 or 2 ng/µl) for subsequent qPCR analyses.
RNA co-immunoprecipitation (RIP)
Cells were grown to the log phase (1–3 × 107 cells/ml) and shifted to the requested temperatures for the indicated times. After harvesting, cells were lysed in RIP buffer (25 mM Tris HCl pH 7.5, 150 mM NaCl, 2 mM MgCl2, 0.5% (v/v) Triton X-100, 0.2 mM PMSF, 500 µM DTT, 1U/ml RiboLock RNase Inhibitor (Thermo Fisher Scientific) and 25 µl/ml protease inhibitor (Roche) using the FastPrep®-24 machine (MP Biomedicals) three times for 30 s at 5.5 m/s. After centrifugation and taking 20 µl and 40 µl of the lysate to control for protein and RNA, the supernatant was incubated with GFP-Selector beads (NanoTag) (prewashed five times with RIP buffer) for 1 h at 4°C. When Protein G Sepharose beads (GE Healthcare) were used, the beads were prewashed two times with RIP buffer, blocked with BSA (10 mg/ml) and glycogen (200 µg/ml) for 30 min at 4°C, followed by three times washing with RIP buffer. The remaining lysate was first mixed with Nop1 antibody for 30 min at 4°C before incubating with beads for 3 h at 4°C. After incubation, the beads were washed five times with RIP buffer and split into two portions after the last washing step. One portion was used for pull-down verification. Proteins were analyzed by western blot using anti-GFP, anti-Hem15, anti-Aco1, and anti-Nop1 antibodies. The other portion was used for RNA extraction by Trizol-Chloroform (Ambion® RNA by Life technologies™). The RNA was mixed with 0.1 volume of the 10x TURBO DNase Buffer and 1 µl of TURBO DNAse (Invitrogen by Thermo Fisher Scientific). The mixture was incubated for 30 minutes at 37 °C, before the reaction was terminated by adding 0.1 volume of 3 M sodium acetate (pH 5.2), 2.5 volumes of 100 % pure ethanol and glycogen, followed by incubation overnight at –20°C. The purified RNA was measured via Nanodrop, and the same amount (100-400 ng) of RNA was reverse transcribed with FastGene Scriptase II (Nippon Genetics) by using random hexamer primer (Thermo Fisher Scientific) according to the manufacturer’s description. The resulting cDNA was diluted to the same concentration (either 1 or 2 ng/µl) for subsequent qPCR analyses.
3’-end PCR
The experiment was essentially carried out as described earlier77. Total or precipitated RNA was isolated as described above. The RNA was either untreated or artificially polyadenylated by E. coli Poly(A) polymerase (Thermo Fisher Scientific). Subsequently, the same amounts (100–200 ng) of RNA were reverse transcribed with FastGene Scriptase II (Nippon Genetics) by using an oligo d(T17) primer with an adapter sequence at its 5’-end and either a C, G, or A at its 3’-end. The generated cDNA was subsequently amplified using the qPCRBIO SyGreen Mix ROX (PCR Biosystems) and a gene-specific forward primer together with the reverse primer, which is complementary to the adapter sequence. PCR products were separated on 2% agarose and purified from the gel using the NucleoSpin Gel and PCR Clean-up Kit (Macherey-Nagel) following the manufacturer’s protocol and the DNA was subsequently sequenced (Microsynth Seqlab GmbH, Germany).
Co-immunoprecipitation (Co-IP)
Cells were grown to the log phase (1–3 × 107 cells/ml) and shifted to the requested temperatures for the indicated times. After harvesting, cells were lysed in PBSKMT buffer (1 x PBS, 3 mM KCl, 2.5 mM MgCl2, 0.5% (v/v) Triton X-100) and 50 µl/ml protease inhibitor using the FastPrep®-24 machine three times for 30 s at 5.5 m/s. After centrifugation and taking 20 µl of the lysate to control for protein, the remaining lysate was diluted with the same volume of PBSKM buffer with 50 µl/ml protease inhibitor to reduce the detergent concentration. Meanwhile, GFP-Selector beads were prewashed two times with PBSKMT buffer, blocking with BSA (10 mg/ml) and glycogen (200 µg/ml) for 30 min at 4 °C, followed by three times washing with PBSKMT buffer. The lysate was incubated with beads for 1 h at 4°C, and subsequently, the beads were washed five times with PBSKMT buffer, and each sample was mixed with the same volume of 2 × sample buffer and incubated for 5 min at 95 °C. Proteins were analyzed by western blot using anti-GFP, anti-Mex67, anti-Hem15, and anti-Nop1 (Santa Cruz) antibodies.
GFP microscopy
The experiments were essentially carried out as described earlier18. Cells were grown to the log phase (1–3 × 107 cells/ml) and shifted to the indicated temperatures if necessary. Cells were fixed by adding formaldehyde to a final concentration of 3% and immediately centrifuged. Subsequently, cells were washed with 0.1 M phosphate buffer, pH 6.5, P-solution (0.1 M phosphate buffer, pH 6.5, 1.2 M sorbitol). The cells were subsequently applied to 0.3% poly-L-lysine coated slides for 30-45 min at 4°C and permeabilized in P solution with 0.5% Triton X-100 for 10 min at room temperature. Cells were washed once with P-solution and once with Aby wash 2 (0.1 M Tris pH 9.5, 0.1 M NaCl) for 5 min respectively. The nuclei were stained with DAPI in Aby wash 2 (1:1000) for 5 min and washed three times with Aby wash 2 for 5 min. The slide was dried, coated, sealed, and Microscopy studies were performed as described above.
Ribose 2’-O-methylation detection by primer extension
2 μg of the total RNA, extracted using TRI reagent according to the manufacturer’s instructions, were annealed to 0.125 pmol 5′-[32P]-radiolabeled DNA oligonucleotide (5’-CTAGATAGTAGATAGGGACAGTGG-3’, located ~100 nt downstream to the 2’-O-methylation sites of interest) in 1x M-MuLV RT buffer (NEB) by incubation at 95°C for 3 min followed by slow cooling to 42 °C. Each reaction was supplemented with 2 × extension master mix (20 U/μl M-MuLV Reverse Transcriptase (NEB), 2 U/μl Ribolock (Thermo Fisher Scientific) and either 0.008 mM dNTPs (low) or 2 mM dNTPs (high) in 2 × M-MuLV RT buffer) and reactions were incubated at 42 °C for 30 min. Reactions were stopped by adding 2× formamide loading dye (95 % formamide, 0.5 mM EDTA, 0.025 % bromophenol blue, 0.025 % xylene cyanol, 0.025 % SDS). To generate a sequencing ladder, the USB® Thermo Sequenase Cycle Sequencing kit was used according to the manufacturer’s instructions. Genomic DNA was first isolated by incubation of wild-type cells at 70°C for 5 min in 200 mM LiAc with 1 % SDS, followed by ethanol precipitation and resuspension in water. Subsequently, genomic DNA served as a template to amplify the partial 25S rDNA containing the 2’-O-methylation sites using the following primers: 5’-CGCGCGTGACGCAATGTGATTTCTGCCC-3’ and 5’-CGCGCGTTCTTTCCCCGCTGATTCTGCC-3’. After purification of the PCR product using the NucleoSpin Gel and PCR Clean-up kit (Macherey-Nagel), the sequencing ladder was generated using the “Radiolabeled primer cycle sequencing” protocol with 50 ng of DNA for each ddNTP. Primer extension samples and the sequencing ladder reactions were heated at 85 °C for 5 min and resolved in 15 % denaturing (7 M urea) polyacrylamide gels. After migration, gels were dried and exposed to a phosphorimager screen (Cytiva). Radiolabelled cDNA fragments were detected using a phosphorimager Typhoon FLA 9500 (Cytiva).
Growth analysis
Cells were grown to the log phase (1-3 ×107 cells/ml), and 10-fold serial dilutions were prepared from 107 to 103 cells/ml. 5 µl of each dilution was spotted onto YPD full medium or selective media agar plates that were subsequently incubated for 2-4 days at 16 °C, 25 °C, and 37 °C. Pictures were taken after 2 and 3 days.
Motif distribution analysis
Annotations of human snoRNAs were taken from the snoDB 2.0 (https://doi.org/10.1093/nar/gkac835). snoRNAs were divided into individually expressed or intronic snoRNAs by using the “intersect” tool of the “bedtools” package80. Using the “getfasta” tool of the same package, the genomic sequence was extracted starting 20 nt upstream and ending 150 nt downstream of the transcript end site (TES). Using the “Biostrings” package80 the poly(A) signals TATATA (efficiency element) and AATAAA (positioning element) were quantified for each snoRNA around the TES, and every position of the motif was taken as a hit. Percentual abundance was averaged for each group and displayed around the TES. Motif presence was also determined for a set of 10,000 random sequences of the same length as a background control.
snoRNA level change upon INTS1 depletion
RNA sequencing data of HEK293T cells without and upon INTS1 depletion for 4 h via the AID system were analyzed for individual and included snoRNAs68. Reads were analyzed using the “computeMatrix” tool from the “deepTools2” package81. Percentual coverage per nucleotide was calculated for each snoRNA to avoid overrepresentation of highly expressed snoRNAs. The average was calculated for included and individual snoRNAs and displayed 100 nt upstream and 150 nt downstream of the transcript end site.
Statistics and Reproducibility
All experiments shown in this work were carried out at least three times in biologically independent repetitions as indicated in the figure legends. Quantification of fluorescence, western blot, and agarose gel signals was conducted by Bio-1D version 11.0 for western blot signals or Fiji (ImageJ version 1.54p) for fluorescence and agarose gel signals. qPCR results were normalized to 18S rRNA, the mitochondrial 21S rRNA (in case of Mex67, Npl3, and Nop1) or mRNA ZWF1 (in case of Snu13) as their expression was not affected in any mutant strain, and these RNAs do not interact with the precipitated target proteins. Statistical analyses were performed in Microsoft Excel version 16.77 or Prism version 9.0.2. Error bars represent the standard deviation (SD). p-values were calculated using either a two-tailed Student’s t-test or ordinary one-way analysis of variance (ANOVA) for multiple group comparison. *p < 0.05; **p < 0.01; ***p < 0.001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
None Source data are provided with this paper.
References
Watkins, N. J. & Bohnsack, M. T. The box C/D and H/ACA snoRNPs: key players in the modification, processing and the dynamic folding of ribosomal RNA. Wiley Interdiscip. Rev. RNA 3, 397–414 (2012).
Kos, M. & Tollervey, D. Yeast pre-rRNA processing and modification occur cotranscriptionally. Mol. Cell 37, 809–820 (2010).
Dominique, C. et al. The dual life of disordered lysine-rich domains of snoRNPs in rRNA modification and nucleolar compaction. Nat. Commun. 15, 9415 (2024).
Lui, L. & Lowe, T. Small nucleolar RNAs and RNA-guided post-transcriptional modification. Essays Biochem 54, 53–77 (2013).
Dieci, G., Preti, M. & Montanini, B. Eukaryotic snoRNAs: a paradigm for gene expression flexibility. Genomics 94, 83–88 (2009).
Tschochner, H. & Hurt, E. Pre-ribosomes on the road from the nucleolus to the cytoplasm. Trends Cell Biol. 13, 255–263 (2003).
Porrua, O. & Libri, D. Transcription termination and the control of the transcriptome: why, where and how to stop. Nat. Rev. Mol. Cell Biol. 16, 190–202 (2015).
Kubicek, K. et al. Serine phosphorylation and proline isomerization in RNAP II CTD control recruitment of Nrd1. Genes Dev. 26, 1891–1896 (2012).
Vasiljeva, L., Kim, M., Mutschler, H., Buratowski, S. & Meinhart, A. The Nrd1-Nab3-Sen1 termination complex interacts with the Ser5-phosphorylated RNA polymerase II C-terminal domain. Nat. Struct. Mol. Biol. 15, 795–804 (2008).
Collin, P., Jeronimo, C., Poitras, C. & Robert, F. RNA Polymerase II CTD Tyrosine 1 Is Required for Efficient Termination by the Nrd1-Nab3-Sen1 Pathway. Mol. Cell 73, 655–669.e657 (2019).
Chapon, C., Cech, T. R. & Zaug, A. J. Polyadenylation of telomerase RNA in budding yeast. RNA 3, 1337–1351 (1997).
Jamonnak, N. et al. Yeast Nrd1, Nab3, and Sen1 transcriptome-wide binding maps suggest multiple roles in post-transcriptional RNA processing. RNA 17, 2011–2025 (2011).
Vasianovich Y., Bajon E., Wellinger R. J. Telomerase biogenesis requires a novel Mex67 function and a cytoplasmic association with the Sm(7) complex. Elife 9, (2020).
Grzechnik, P. & Kufel, J. Polyadenylation linked to transcription termination directs the processing of snoRNA precursors in yeast. Mol. Cell 32, 247–258 (2008).
Steinmetz, E. J. et al. Genome-wide distribution of yeast RNA polymerase II and its control by Sen1 helicase. Mol. Cell 24, 735–746 (2006).
Grzechnik, P., Gdula, M. R. & Proudfoot, N. J. Pcf11 orchestrates transcription termination pathways in yeast. Genes Dev. 29, 849–861 (2015).
Tudek, A. et al. Molecular basis for coordinating transcription termination with noncoding RNA degradation. Mol. Cell 55, 467–481 (2014).
Becker, D. et al. Nuclear Pre-snRNA Export Is an Essential Quality Assurance Mechanism for Functional Spliceosomes. Cell Rep. 27, 3199–3214.e3193 (2019).
Mouaikel, J., Verheggen, C., Bertrand, E., Tazi, J. & Bordonne, R. Hypermethylation of the cap structure of both yeast snRNAs and snoRNAs requires a conserved methyltransferase that is localized to the nucleolus. Mol. Cell 9, 891–901 (2002).
Grzechnik, P. et al. Nuclear fate of yeast snoRNA is determined by co-transcriptional Rnt1 cleavage. Nat. Commun. 9, 1783 (2018).
Kufel, J., Allmang, C., Verdone, L., Beggs, J. & Tollervey, D. A complex pathway for 3’ processing of the yeast U3 snoRNA. Nucleic Acids Res 31, 6788–6797 (2003).
Wang X., Guo J., Li J., Krebber H. Initial spliceosomal U4/U6 di-snRNA formation occurs in the cytoplasm of Saccharomyces cerevisiae and requires a guard protein-mediated quality control. Nucleic Acids Res. 54, (2026).
Kufel, J., Allmang, C., Verdone, L., Beggs, J. D. & Tollervey, D. Lsm proteins are required for normal processing of pre-tRNAs and their efficient association with La-homologous protein Lhp1p. Mol. Cell Biol. 22, 5248–5256 (2002).
Kufel, J., Allmang, C., Petfalski, E., Beggs, J. & Tollervey, D. Lsm Proteins are required for normal processing and stability of ribosomal RNAs. J. Biol. Chem. 278, 2147–2156 (2003).
Urlaub, H., Raker, V. A., Kostka, S. & Lührmann, R. Sm protein-Sm site RNA interactions within the inner ring of the spliceosomal snRNP core structure. EMBO J. 20, 187–196 (2001).
Hirsch A. G., Becker D., Lamping J.-P., Krebber H. Unraveling the stepwise maturation of the yeast telomerase. bioRxiv, 2021.2004.2030.442090 (2021).
Bouveret, E., Rigaut, G., Shevchenko, A., Wilm, M. & Séraphin, B. A Sm-like protein complex that participates in mRNA degradation. EMBO J. 19, 1661–1671 (2000).
Chen, X. et al. Transcriptomes of six mutants in the Sen1 pathway reveal combinatorial control of transcription termination across the Saccharomyces cerevisiae genome. PLoS Genet. 13, e1006863 (2017).
Holmes, R. K. et al. Loss of the Yeast SR Protein Npl3 Alters Gene Expression Due to Transcription Readthrough. PLoS Genet. 11, 1–29 (2015).
Kim, M. et al. Distinct pathways for snoRNA and mRNA termination. Mol. Cell 24, 723–734 (2006).
Reuter L. M., Meinel D. M., Sträßer K. The poly (A) -binding protein Nab2 functions in RNA polymerase III transcription. Genes Dev. 1565–1575 (2015).
Tuck, A. C. & Tollervey, D. A Transcriptome-wide Atlas of RNP Composition Reveals Diverse Classes of mRNAs and lncRNAs. Cell 154, 996–1009 (2013).
Klama, S. et al. A guard protein mediated quality control mechanism monitors 5’-capping of pre-mRNAs. Nucleic Acids Res. 50, 11301–11314 (2022).
Hackmann, A. et al. Quality control of spliced mRNAs requires the shuttling SR proteins Gbp2 and Hrb1. Nat. Commun. 5, 3123 (2014).
Fasken, M. B., Stewart, M. & Corbett, A. H. Functional significance of the interaction between the mRNA-binding protein, Nab2, and the nuclear pore-associated protein, Mlp1, in mRNA export. J. Biol. Chem. 283, 27130–27143 (2008).
Li, J., Querl, L., Coban, I., Salinas, G. & Krebber, H. Surveillance of 3’ mRNA cleavage during transcription termination requires CF IB/Hrp1. Nucleic Acids Res. 51, 8758–8773 (2023).
Zander, G. et al. mRNA quality control is bypassed for immediate export of stress-responsive transcripts. Nature 540, 593–596 (2016).
Zander, G. & Krebber, H. Quick or quality? How mRNA escapes nuclear quality control during stress. RNA Biol. 14, 1642–1648 (2017).
Querl, L. & Krebber, H. Defenders of the Transcriptome: Guard Protein-Mediated mRNA Quality Control in Saccharomyces cerevisiae. Int J. Mol. Sci. 25, 10241 (2024).
Soheilypour M., Mofrad M. R. K. Regulation of the affinity between RNA-binding proteins and the export receptor enables nuclear basket proteins to distinguish and retain aberrant mRNAs. Scientific reports, 1–11 (2016).
Soheilypour, M. & Mofrad, M. R. K. Quality control of mRNAs at the entry of the nuclear pore: Cooperation in a complex molecular system. Nucleus 9, 202–211 (2018).
Holley, C. L. et al. Cytosolic accumulation of small nucleolar RNAs (snoRNAs) is dynamically regulated by NADPH oxidase. J. Biol. Chem. 290, 11741–11748 (2015).
Mleczko, A. M. et al. Levels of sdRNAs in cytoplasm and their association with ribosomes are dependent upon stress conditions but independent from snoRNA expression. Sci. Rep. 9, 18397 (2019).
Michel, C. I. et al. Small nucleolar RNAs U32a, U33, and U35a are critical mediators of metabolic stress. Cell Metab. 14, 33–44 (2011).
Ender, C. et al. A human snoRNA with microRNA-like functions. Mol. Cell 32, 519–528 (2008).
Coban, I. et al. dsRNA formation leads to preferential nuclear export and gene expression. Nature 631, 432–438 (2024).
Brogna, S., McLeod, T. & Petric, M. The Meaning of NMD: Translate or Perish. Trends Genet. 32, 395–407 (2016).
Lu Y. Y., Krebber H. Nuclear mRNA quality control and cytoplasmic NMD Are Linked by the Guard Proteins Gbp2 and Hrb1. Int. J. Mol. Sci. 22, (2021).
Qiu, H. et al. Identification of genes that function in the biogenesis and localization of small nucleolar RNAs in Saccharomyces cerevisiae. Mol. Cell Biol. 28, 3686–3699 (2008).
Narayanan, A. et al. Nuclear RanGTP is not required for targeting small nucleolar RNAs to the nucleolus. J. Cell Sci. 116, 177–186 (2003).
Kufel, J. et al. Precursors to the U3 small nucleolar RNA lack small nucleolar RNP proteins but are stabilized by La binding. Mol. Cell Biol. 20, 5415–5424 (2000).
Rothé, B. et al. Implication of the box C/D snoRNP assembly factor Rsa1p in U3 snoRNP assembly. Nucleic Acids Res 45, 7455–7473 (2017).
Inada, M. & Guthrie, C. Identification of Lhp1p-associated RNAs by microarray analysis in Saccharomyces cerevisiae reveals association with coding and noncoding RNAs. Proc. Natl. Acad. Sci. USA 101, 434–439 (2004).
Fernandez, C. F., Pannone, B. K., Chen, X., Fuchs, G. & Wolin, S. L. An Lsm2-Lsm7 complex in Saccharomyces cerevisiae associates with the small nucleolar RNA snR5. Mol. Biol. Cell 15, 2842–2852 (2004).
Steinmetz, E. J., Conrad, N. K., Brow, D. A. & Corden, J. L. RNA-binding protein Nrd1 directs poly(A)-independent 3’-end formation of RNA polymerase II transcripts. Nature 413, 327–331 (2001).
Boreikaitė, V. & Passmore, L. A. 3’-End processing of eukaryotic mRNA: machinery, regulation, and impact on gene expression. Annu Rev. Biochem 92, 199–225 (2023).
Green, D. M., Johnson, C. P., Hagan, H. & Corbett, A. H. The C-terminal domain of myosin-like protein 1 (Mlp1p) is a docking site for heterogeneous nuclear ribonucleoproteins that are required for mRNA export. Proc. Natl. Acad. Sci. USA 100, 1010–1015 (2003).
Green, D. M. et al. Nab2p Is Required for Poly(A) RNA Export in Saccharomyces cerevisiae and Is Regulated by Arginine Methylation via Hmt1p. J. Biol. Chem. 277, 7752–7760 (2002).
Hector, R. E. et al. Dual requirement for yeast hnRNP Nab2p in mRNA poly(A) tail length control and nuclear export. EMBO J. 21, 1800–1810 (2002).
Kyburz, A., Sadowski, M., Dichtl, B. & Keller, W. The role of the yeast cleavage and polyadenylation factor subunit Ydh1p/Cft2p in pre-mRNA 3’-end formation. Nucleic Acids Res. 31, 3936–3945 (2003).
Rothé, B. et al. Protein Hit1, a novel box C/D snoRNP assembly factor, controls cellular concentration of the scaffolding protein Rsa1 by direct interaction. Nucleic Acids Res. 42, 10731–10747 (2014).
Watkins, N. J., Dickmanns, A. & Lührmann, R. Conserved stem II of the box C/D motif is essential for nucleolar localization and is required, along with the 15.5K protein, for the hierarchical assembly of the box C/D snoRNP. Mol. Cell Biol. 22, 8342–8352 (2002).
Mischo, H. E. et al. Cell-cycle modulation of transcription termination Factor Sen1. Mol. Cell 70, 312–326 e317 (2018).
Schaughency, P., Merran, J. & Corden, J. L. Genome-wide mapping of yeast RNA polymerase II termination. PLoS Genet. 10, e1004632 (2014).
Matera, A. G. & Wang, Z. A day in the life of the spliceosome. Nat. Rev. Mol. Cell Biol. 15, 108–121 (2014).
Welsh, S. A. & Gardini, A. Genomic regulation of transcription and RNA processing by the multitasking Integrator complex. Nat. Rev. Mol. Cell Biol. 24, 204–220 (2023).
O’Reilly, D. et al. Human snRNA genes use polyadenylation factors to promote efficient transcription termination. Nucleic Acids Res. 42, 264–275 (2014).
Baluapuri, A. et al. Integrator loss leads to dsRNA formation that triggers the integrated stress response. Cell 188, 3184–3201.e3121 (2025).
Dupuis-Sandoval, F., Poirier, M. & Scott, M. S. The emerging landscape of small nucleolar RNAs in cell biology. Wiley Interdiscip. Rev. RNA 6, 381–397 (2015).
Kufel, J. & Grzechnik, P. Small nucleolar RNAs tell a different tale. Trends Genet. 35, 104–117 (2019).
Liu, B. et al. snoRNA-facilitated protein secretion revealed by transcriptome-wide snoRNA target identification. Cell 188, 465–483.e422 (2025).
Lemus-Diaz, N., Ferreira, R. R., Bohnsack, K. E., Gruber, J. & Bohnsack, M. T. The human box C/D snoRNA U3 is a miRNA source, and miR-U3 regulates expression of sortin nexin 27. Nucleic Acids Res. 48, 8074–8089 (2020).
Massenet, S., Bertrand, E. & Verheggen, C. Assembly and trafficking of box C/D and H/ACA snoRNPs. RNA Biol. 14, 680–692 (2017).
Li, M. W. et al. Nuclear export factor 3 regulates the localization of small nucleolar RNAs. J. Biol. Chem. 292, 20228–20239 (2017).
Hamperl, S., Bocek, M. J., Saldivar, J. C., Swigut, T. & Cimprich, K. A. Transcription-replication conflict orientation modulates R-loop levels and activates distinct DNA damage responses. Cell 170, 774–786.e719 (2017).
Lalonde M., Trauner M., Werner M., Hamperl S. Consequences and resolution of transcription-replication conflicts. Life 11, (2021).
Lamping J. P., Krebber H. The functionality of telomerase depends on CPF-CF induced 3’end processing of its RNA component, TLC1 and a novel Nrd1-Nab3 surveillance mechanism. Nucleic Acids Res. 53, (2025).
van den Ent, F. & Löwe, J. RF cloning: a restriction-free method for inserting target genes into plasmids. J. Biochem. Biophys. Methods 67, 67–74 (2006).
Gietz, D., St Jean, A., Woods, R. A. & Schiestl, R. H. Improved method for high efficiency transformation of intact yeast cells. Nucleic Acids Res. 20, 1425 (1992).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Acknowledgements
We are grateful to G. Braus, U. Mühlenhoff, E. Hurt, and R. Lill for providing plasmids, strains or antibodies. We thank M. Lange for technical assistance and the Krebber lab for fruitful discussions. This work was funded by the University of Göttingen and the Deutsche Forschungsgemeinschaft (DFG) via grant (1779/12-1) and SFB 1565 (Projektnummer 469281184) (to H.K and K.E.B) and by the China Scholarship Council (CSC) (to F.Y. and Y.D.)
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Experiments were designed and data interpreted by F.Y., G.Z., J.P.L., and H.K.; G.Z. conducted the experiments resulting in the Figs. 1f, 2a, 3h, S1d, e, g S2 and S3j. Y.D. contributed to the experiments resulting in Fig. 6a–d and S6f. J.P.L. conducted the experiments resulting in Fig. 7 and S7. S.T. and K.E.B. carried out the experiment resulting in Fig. 6h. All other experiments were performed by F.Y. The manuscript was written by H.K.; all authors discussed the results.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yu, F., Zaccagnini, G., Duan, Y. et al. CPF-CF-terminated snoRNAs shuttle through the cytoplasm via an mRNA guard protein-mediated surveillance mechanism. Nat Commun 17, 2328 (2026). https://doi.org/10.1038/s41467-026-70373-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70373-8