Abstract
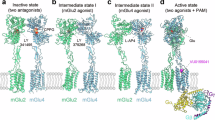
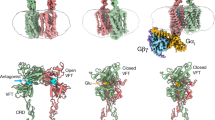
The metabotropic glutamate receptor 6 (mGlu6) is essential for synaptic communication of rod photoreceptors, and mutations in mGlu6 lead to a blinding disorder. However, its structural organization remains unknown. Here, we present the structure of agonist-bound mGlu6, revealing an asymmetric dimer arrangement in the absence of a G protein. This indicates that agonist binding alone can induce the homodimeric receptor asymmetry in metabotropic glutamate receptors and structurally prime mGlu6 for activation by pre-organizing the transmembrane domain dimer interface for G protein binding. The structure also identifies noncanonical interactions between the cysteine-rich domain and extracellular loop 2, forming a unique interface that likely stabilizes the activation state. Mutational analyses of this interface reveal its role in maintaining rapid Gαo activation and surface targeting. The structure also permits mechanistic investigation of congenital stationary night blindness and reveals diverse effects of pathogenic mutations on surface trafficking, Gαo coupling, and activation dynamics, including unexpected gain-of-function. These results provide critical insight into the intermediate asymmetric structure of mGlu6 and offer a molecular framework for understanding the pathogenesis of inherited retinal disorders.
Similar content being viewed by others
Data availability
The cryo-EM density maps and corresponding coordinates generated in this study have been deposited in the Electron Microscopy Data Bank (EMDB) and the Protein Data Bank (PDB) under the following accession codes: EMD-63210 and 9LLZ (L-SOP bound mGlu6, global), EMD-63211 and 9LM0 (L-SOP bound mGlu6, local), EMD-64620 and 9UYO (L-SOP bound mGlu6, class 1), EMD-64622 and 9UYQ (L-SOP bound mGlu6, class 2), and EMD-64621 and 9UYP (L-SOP bound mGlu6, class 3). The source data underlying Figs. 1c, d, 4g–j, and 5b, c, and Supplementary Figs. 4b–g and 7b–g are provided in the Source Data file. Uncropped scans of all gels and blots are provided in the Supplementary Information. Source data are provided with this paper.
References
Schiller, P. H., Sandell, J. H. & Maunsell, J. H. Functions of the ON and OFF channels of the visual system. Nature 322, 824–825 (1986).
Ray, T. A. et al. GPR179 is required for high sensitivity of the mGluR6 signaling cascade in depolarizing bipolar cells. J. Neurosci. 34, 6334–6343 (2014).
Werblin, F. S. & Dowling, J. E. Organization of the retina of the mudpuppy, Necturus maculosus. II. Intracellular recording. J. Neurophysiol. 32, 339–355 (1969).
Nelson, R. & Connaughton, V. Bipolar cell pathways in the vertebrate retina. In Webvision: the organization of the retina and visual system, eds. Kolb, H., Fernandez, E. & Nelson, R. (University of Utah Health Sciences Center, 1995).
Koike, C. et al. TRPM1 is a component of the retinal ON bipolar cell transduction channel in the mGluR6 cascade. Proc. Natl. Acad. Sci. USA 107, 332–337 (2010).
Morgans, C. W. et al. TRPM1 is required for the depolarizing light response in retinal ON-bipolar cells. Proc. Natl Acad. Sci. USA 106, 19174–19178 (2009).
Shen, Y. et al. A transient receptor potential-like channel mediates synaptic transmission in rod bipolar cells. J. Neurosci. 29, 6088–6093 (2009).
Orlandi, C. et al. GPR158/179 regulate G protein signaling by controlling localization and activity of the RGS7 complexes. J. Cell Biol. 197, 711–719 (2012).
Orlandi, C. et al. Transsynaptic binding of orphan receptor GPR179 to dystroglycan-pikachurin complex is essential for the synaptic organization of photoreceptors. Cell Rep 25, 130–145.e5 (2018).
Peachey, N. S. et al. GPR179 is required for depolarizing bipolar cell function and is mutated in autosomal-recessive complete congenital stationary night blindness. Am. J. Hum. Genet. 90, 331–339 (2012).
Huang, Y.-Y., Haug, M. F., Gesemann, M. & Neuhauss, S. C. Novel expression patterns of metabotropic glutamate receptor 6 in the zebrafish nervous system. PLoS ONE 7, e35256 (2012).
Pin, J.-P., Galvez, T. & Prézeau, L. Evolution, structure, and activation mechanism of family 3/C G-protein-coupled receptors. Pharmacol. Ther. 98, 325–354 (2003).
Chun, L., Zhang, W. -h & Liu, J. -f Structure and ligand recognition of class C GPCRs. Acta Pharmacol. Sin. 33, 312–323 (2012).
Koehl, A. et al. Structural insights into the activation of metabotropic glutamate receptors. Nature 566, 79–84 (2019).
Nasrallah, C. et al. Agonists and allosteric modulators promote signaling from different metabotropic glutamate receptor 5 conformations. Cell Rep 36, 109648 (2021).
Du, J. et al. Structures of human mGlu2 and mGlu7 homo- and heterodimers. Nature 594, 589–593 (2021).
Lin, S. et al. Structures of G(i)-bound metabotropic glutamate receptors mGlu2 and mGlu4. Nature 594, 583–588 (2021).
Seven, A. B. et al. G-protein activation by a metabotropic glutamate receptor. Nature 595, 450–454 (2021).
Zhang, J. et al. Structural insights into the activation initiation of full-length mGlu1. Protein Cell 12, 662–667 (2021).
Fang, W. et al. Structural basis of the activation of metabotropic glutamate receptor 3. Cell Res. 32, 695–698 (2022).
Mao, C. et al. Cryo-EM structures of inactive and active GABAB receptor. Cell Res. 30, 564–573 (2020).
Kim, Y., Jeong, E., Jeong, J.-H., Kim, Y. & Cho, Y. Structural basis for activation of the heterodimeric GABAB receptor. J. Mol. Biol. 432, 5966–5984 (2020).
Shaye, H. et al. Structural basis of the activation of a metabotropic GABA receptor. Nature 584, 298–303 (2020).
Shen, C. et al. Structural basis of GABAB receptor–Gi protein coupling. Nature 594, 594–598 (2021).
Ling, S. et al. Structural mechanism of cooperative activation of the human calcium-sensing receptor by Ca2+ ions and L-tryptophan. Cell Res 31, 383–394 (2021).
Wen, T. et al. Structural basis for activation and allosteric modulation of full-length calcium-sensing receptor. Sci. Adv. 7, eabg1483 (2021).
Gao, Y. et al. Asymmetric activation of the calcium-sensing receptor homodimer. Nature 595, 455–459 (2021).
Park, J. et al. Symmetric activation and modulation of the human calcium-sensing receptor. Proc. Natl Acad. Sci. USA 118, e2115849118 (2021).
Chen, X. et al. Structural insights into the activation of human calcium-sensing receptor. eLife 10, e68578 (2021).
He, F. et al. Allosteric modulation and G-protein selectivity of the Ca2+-sensing receptor. Nature 626, 1141–1148 (2024).
Zuo, H. et al. Promiscuous G-protein activation by the calcium-sensing receptor. Nature 629, 481–488 (2024).
Wang, X. et al. Structural insights into dimerization and activation of the mGlu2–mGlu3 and mGlu2–mGlu4 heterodimers. Cell Res. 33, 762–774 (2023).
Strauss, A. et al. Structural basis of positive allosteric modulation of metabotropic glutamate receptor activation and internalization. Nat. Commun. 15, 6498 (2024).
Laurie, D. J., Schoeffter, P., Wiederhold, K. H. & Sommer, B. Cloning, distribution and functional expression of the human mGlu6 metabotropic glutamate receptor. Neuropharmacology 36, 145–152 (1997).
Kang, H. J. et al. Selectivity and evolutionary divergence of metabotropic glutamate receptors for endogenous ligands and G proteins coupled to phospholipase C or TRP channels. J. Biol. Chem. 289, 29961–29974 (2014).
Kunishima, N. et al. Structural basis of glutamate recognition by a dimeric metabotropic glutamate receptor. Nature 407, 971–977 (2000).
Vafabakhsh, R., Levitz, J. & Isacoff, E. Y. Conformational dynamics of a class C G-protein-coupled receptor. Nature 524, 497–501 (2015).
Krishna Kumar, K. et al. Stepwise activation of a metabotropic glutamate receptor. Nature 629, 951–956 (2024).
Huang, W. et al. Structural basis of orientated asymmetry in a mGlu heterodimer. Nat. Commun. 15, 10345 (2024).
Latorraca, N. R. et al. Domain coupling in activation of a family C GPCR. bioRxiv, (2024).
Patil, D. N. et al. Cryo-EM structure of human GPR158 receptor coupled to the RGS7-Gβ5 signaling complex. Science 375, 86–91 (2022).
Dore, A. S. et al. Structure of class C GPCR metabotropic glutamate receptor 5 transmembrane domain. Nature 511, 557–562 (2014).
Yun, Y., Jeong, H., Laboute, T., Martemyanov, K. A. & Lee, H. H. Cryo-EM structure of human class C orphan GPCR GPR179 involved in visual processing. Nat. Commun. 15, 8299 (2024).
Punjani, A. & Fleet, D. J. 3D variability analysis: resolving continuous flexibility and discrete heterogeneity from single particle cryo-EM. J. Struct. Biol. 213, 107702 (2021).
Olofsson, L. et al. Fine tuning of sub-millisecond conformational dynamics controls metabotropic glutamate receptors agonist efficacy. Nat. Commun. 5, 5206 (2014).
Kang, H. J., Wilkins, A. D., Lichtarge, O. & Wensel, T. G. Determinants of endogenous ligand specificity divergence among metabotropic glutamate receptors. J. Biol. Chem. 290, 2870–2878 (2015).
Conn, P. J. & Pin, J. P. Pharmacology and functions of metabotropic glutamate receptors. Annu. Rev. Pharmacol. Toxicol. 37, 205–237 (1997).
Wright, R. A. & Schoepp, D. D. Differentiation of group 2 and group 3 metabotropic glutamate receptor cAMP responses in the rat hippocampus. Eur. J. Pharmacol. 297, 275–282 (1996).
Schkeryantz, J. M. et al. Determination of L-AP4-bound human mGlu8 receptor amino terminal domain structure and the molecular basis for L-AP4’s group III mGlu receptor functional potency and selectivity. Bioorg. Med. Chem. Lett. 28, 612–617 (2018).
Hack, Y. L. et al. Whole-genome sequencing identifies missense mutation in GRM6 as the likely cause of congenital stationary night blindness in a Tennessee walking horse. Equine Vet. J. 53, 316–323 (2021).
Huang, S. et al. Interdomain movements in metabotropic glutamate receptor activation. Proc. Natl Acad. Sci. USA 108, 15480–15485 (2011).
Dryja, T. P. et al. Night blindness and abnormal cone electroretinogram ON responses in patients with mutations in the GRM6 gene encoding mGluR6. Proc. Natl Acad. Sci. USA 102, 4884–4889 (2005).
O’Connor, E. et al. Congenital stationary night blindness associated with mutations in GRM6 encoding glutamate receptor MGluR6. Br. J. Ophthalmol. 90, 653–654 (2006).
Zeitz, C. et al. Mutations in GRM6 cause autosomal recessive congenital stationary night blindness with a distinctive scotopic 15-Hz flicker electroretinogram. Invest. Ophthalmol. Vis. Sci. 46, 4328–4335 (2005).
Wang, Q., Gao, Y., Li, S., Guo, X. & Zhang, Q. Mutation screening of TRPM1, GRM6, NYX and CACNA1F genes in patients with congenital stationary night blindness. Int. J. Mol. Med. 30, 521–526 (2012).
Xu, X. et al. Sequence variations of GRM6 in patients with high myopia. Mol. Vis. 15, 2094–2100 (2009).
Pindwarawala, M. et al. Defective glycosylation and ELFN1 binding of mGluR6 congenital stationary night blindness mutants. Life Sci. Alliance 8, e202403118 (2025).
Masu, M. et al. Specific deficit of the ON response in visual transmission by targeted disruption of the mGluR6 gene. Cell 80, 757–765 (1995).
Zeitz, C., Robson, A. G. & Audo, I. Congenital stationary night blindness: an analysis and update of genotype-phenotype correlations and pathogenic mechanisms. Prog. Retin. Eye Res. 45, 58–110 (2015).
Cannone, G. et al. Conformational diversity in class C GPCR positive allosteric modulation. Nat. Commun. 16, 619 (2025).
Zeitz, C. et al. Night blindness-associated mutations in the ligand-binding, cysteine-rich, and intracellular domains of the metabotropic glutamate receptor 6 abolish protein trafficking. Hum. Mutat. 28, 771–780 (2007).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Acknowledgements
We thank Dr. Bumhan Ryu at the Institute for Basic Science (IBS) for his assistance in cryo-EM data collection and Dr. Hyeongseop Jeong at the Korea Basic Science Institute (KBSI) for his assistance in cryo-EM data processing. We also thank Yeji Gil, Dong Kyu Chung, and Youngchae Joe for helpful discussions. Use of the cryo-EM facilities of the NEXUS Consortium was supported by a grant from the National Research Foundation of Korea (RS-2024-00440289). The authors thank the National Center for Inter-University Research Facilities (NCIRF) for assistance with cryo-EM. This work was supported by the Samsung Science and Technology Foundation (SSTF-BA2101-13 to H.H.L.) and NIH grants EY018139 and EY034339 (to K.A.M.).
Author information
Authors and Affiliations
Contributions
S.Y.L., C.-T.C., Y.Y., K.A.M., and H.H.L. conceived and designed the experiments. S.Y.L. solved the cryo-EM structures and performed the structural analyses. C.-T.C. performed biochemical assays. Y.Y. and J.S.J. helped with cryo-EM data collection and analysis. S.Y.L., C.-T.C., Y.Y., J.S.J., K.A.M., and H.H.L. analyzed the data and wrote the manuscript. K.A.M. and H.H.L. directed the work. All the authors have edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jean-Philippe Pin who co-reviewed with Anais Menny; and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lee, S.Y., Chang, CT., Yun, Y. et al. CryoEM structure of mGlu6 captures receptor activation prior to G protein coupling. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70436-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70436-w