Abstract
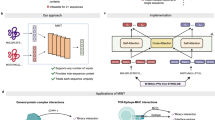
Understanding protein–protein interactions (PPIs) is crucial for deciphering cellular processes and guiding therapeutic discovery. While recent protein language models have advanced sequence-based protein representation, most are designed for individual chains and fail to capture inherent PPI patterns. Here, we introduce a Protein Pair Language Model (PPLM) that jointly encodes paired sequences, enabling direct learning of interaction-aware representations beyond what single-chain models can provide. Building on this foundation, we develop PPLM-PPI, PPLM-Affinity, and PPLM-Contact for binary interaction, binding affinity, and interface contact prediction. Large-scale experiments show that PPLM-PPI achieves state-of-the-art performance across different species on binary interaction prediction, while PPLM-Affinity outperforms both ESM2 and structure-based methods on binding affinity modeling, particularly on challenging cases including antibody–antigen and TCR–pMHC complexes. PPLM-Contact further surpasses existing contact predictors on inter-protein contact prediction and interface residue recognition, including those deduced from cutting-edge complex structure predictions. Together, these results highlight the potential of co-represented language models to advance computational modeling of PPIs.
Similar content being viewed by others
Data availability
The dataset used in this study is deposited in our GitHub repository: https://github.com/junliu621/PPLM/tree/main/data/. Protein pairs used to train PPLM are collected from the Protein Data Bank (PDB, https://www.rcsb.org/) and the STRING database (https://string-db.org/). The interaction dataset for PPLM-PPI is sourced from D-SCRIPT (https://github.com/samsledje/D-SCRIPT/), while the binding affinity dataset for PPLM-Affinity is taken from PPB-Affinity (https://github.com/ChenPy00/PPB-Affinity). The Uniref30_2021_03 database is available at https://www.user.gwdguser.de/~compbiol/uniclust/. Source data are provided with this paper.
Code availability
The PPLM webserver and source codes, including the language model, interaction prediction, binding affinity prediction, and inter-protein contact prediction, are freely accessible at https://zhanggroup.org/PPLM/. The source codes are also available on GitHub at https://github.com/junliu621/PPLM under the MIT License. The publication release is deposited on Zenodo at https://zenodo.org/records/1825639258.
References
Barabasi, A.-L. & Oltvai, Z. N. Network biology: understanding the cell’s functional organization. Nat. Rev. Genet. 5, 101–113 (2004).
Keskin, O., Gursoy, A., Ma, B. & Nussinov, R. Principles of protein-protein interactions: what are the preferred ways for proteins to interact? Chem. Rev. 108, 1225–1244 (2008).
Siebenmorgen, T. & Zacharias, M. Computational prediction of protein–protein binding affinities. Adv. Rev. 10, e1448 (2019).
Vangone, A. & Bonvin, A. M. Contacts-based prediction of binding affinity in protein–protein complexes. elife 4, e07454 (2015).
Rives, A. et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proc. Natl. Acad. Sci. 118, e2016239118 (2021).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Hayes, T. et al. Simulating 500 million years of evolution with a language model. bioRxiv, 2024.2007. 2001.600583 (2024).
Elnaggar, A. et al. Prottrans: Toward understanding the language of life through self-supervised learning. IEEE Trans. Pattern Anal. Mach. Intell. 44, 7112–7127 (2021).
Chowdhury, R. et al. Single-sequence protein structure prediction using a language model and deep learning. Nat. Biotechnol. 40, 1617–1623 (2022).
Fang, X. et al. A method for multiple-sequence-alignment-free protein structure prediction using a protein language model. Nat. Mach. Intell. 5, 1087–1096 (2023).
Wang, S., You, R., Liu, Y., Xiong, Y. & Zhu, S. NetGO 3.0: protein language model improves large-scale functional annotations. Genomics Proteom. Bioinforma. 21, 349–358 (2023).
Ferruz, N., Schmidt, S. & Höcker, B. ProtGPT2 is a deep unsupervised language model for protein design. Nat. Commun. 13, 4348 (2022).
Meier, J. et al. Language models enable zero-shot prediction of the effects of mutations on protein function. Adv. Neural Inf. Process. Syst. 34, 29287–29303 (2021).
Jin, M. et al. ProLLM: protein chain-of-thoughts enhanced LLM for protein-protein interaction prediction. bioRxiv, 2024.2004. 2018.590025 (2024).
Sledzieski, S., Singh, R., Cowen, L. & Berger, B. D-SCRIPT translates genome to phenome with sequence-based, structure-aware, genome-scale predictions of protein-protein interactions. Cell Syst. 12, 969–982.e966 (2021).
Liu, J., Liu, D., He, G. & Zhang, G. Estimating protein complex model accuracy based on ultrafast shape recognition and deep learning in CASP15. Proteins: Struct. Funct. Bioinforma. 91, 1861–1870 (2023).
Zhou, Z. et al. ProAffinity-GNN: a novel approach to structure-based protein–protein binding affinity prediction via a curated data set and graph neural networks. J. Chem. Inf. Model. 64, 8796–8808 (2024).
Romero-Molina, S. et al. PPI-affinity: A web tool for the prediction and optimization of protein–peptide and protein–protein binding affinity. J. Proteome Res. 21, 1829–1841 (2022).
Guo, Z., Liu, J., Skolnick, J. & Cheng, J. Prediction of inter-chain distance maps of protein complexes with 2D attention-based deep neural networks. Nat. Commun. 13, 6963 (2022).
Lin, P., Tao, H., Li, H. & Huang, S.-Y. Protein–protein contact prediction by geometric triangle-aware protein language models. Nat. Mach. Intell. 5, 1275–1284 (2023).
Bernett, J., Blumenthal, D. B. & List, M. Cracking the black box of deep sequence-based protein–protein interaction prediction. Brief. Bioinforma. 25, bbae076 (2024).
Singh, R., Devkota, K., Sledzieski, S., Berger, B. & Cowen, L. Topsy-Turvy: integrating a global view into sequence-based PPI prediction. Bioinformatics 38, i264–i272 (2022).
Li, Y., Wang, C., Gu, H., Feng, H. & Ruan, Y. ESMDNN-PPI: a new protein–protein interaction prediction model developed with protein language model of ESM2 and deep neural network. Meas. Sci. Technol. 35, 125701 (2024).
Meda, R. S. & Farimani, A. B. BAPULM: Binding Affinity Prediction using Language Models. arXiv preprint arXiv:2411.04150 (2024).
Gorantla, R. et al. Learning Binding Affinities via Fine-tuning of Protein and Ligand Language Models. bioRxiv, 2024.2011. 2001.621495 (2024).
Siebenmorgen, T. & Zacharias, M. Computational prediction of protein–protein binding affinities. Wiley Interdiscip. Rev.: Comput. Mol. Sci. 10, e1448 (2020).
Guo, Z. & Yamaguchi, R. Machine learning methods for protein-protein binding affinity prediction in protein design. Front. Bioinforma. 2, 1065703 (2022).
Liu, H. et al. PPB-Affinity: Protein-Protein Binding Affinity dataset for AI-based protein drug discovery. Sci. Data 11, 1–11 (2024).
Xue, L. C., Rodrigues, J. P., Kastritis, P. L., Bonvin, A. M. & Vangone, A. PRODIGY: a web server for predicting the binding affinity of protein–protein complexes. Bioinformatics 32, 3676–3678 (2016).
Wang, M., Cang, Z. & Wei, G.-W. A topology-based network tree for the prediction of protein–protein binding affinity changes following mutation. Nat. Mach. Intell. 2, 116–123 (2020).
Zeng, H. et al. ComplexContact: a web server for inter-protein contact prediction using deep learning. Nucleic Acids Res. 46, W432–W437 (2018).
Lin, P., Yan, Y. & Huang, S.-Y. DeepHomo2.0: improved protein–protein contact prediction of homodimers by transformer-enhanced deep learning. Brief. Bioinforma. 24, bbac499 (2023).
Hu, L., Wang, X., Huang, Y.-A., Hu, P. & You, Z.-H. A survey on computational models for predicting protein–protein interactions. Brief. Bioinforma. 22, bbab036 (2021).
Si, Y. & Yan, C. Protein language model-embedded geometric graphs power inter-protein contact prediction. Elife 12, RP92184 (2024).
Xie, Z. & Xu, J. Deep graph learning of inter-protein contacts. Bioinformatics 38, 947–953 (2022).
Rao, R. M. et al. in International Conference on Machine Learning 8844–8856 (PMLR, 2021).
Su, J. et al. Roformer: Enhanced transformer with rotary position embedding. Neurocomputing 568, 127063 (2024).
Evans, R. et al. Protein complex prediction with AlphaFold-Multimer. biorxiv, 2021.2010. 2004.463034 (2021).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature, 1–3 (2024).
Zheng, W. et al. Improving deep learning protein monomer and complex structure prediction using DeepMSA2 with huge metagenomics data. Nat. Methods 21, 279–289 (2024).
Ko, Y. S., Parkinson, J., Liu, C. & Wang, W. TUnA: an uncertainty-aware transformer model for sequence-based protein–protein interaction prediction. Brief. Bioinforma. 25, bbae359 (2024).
Chatterjee, A. et al. Improving the generalizability of protein-ligand binding predictions with AI-Bind. Nat. Commun. 14, 1989 (2023).
Wang, Y. et al. ZeroBind: a protein-specific zero-shot predictor with subgraph matching for drug-target interactions. Nat. Commun. 14, 7861 (2023).
Brekke, O. H. & Sandlie, I. Therapeutic antibodies for human diseases at the dawn of the twenty-first century. Nat. Rev. Drug Discov. 2, 52–62 (2003).
Szeto, C., Lobos, C. A., Nguyen, A. T. & Gras, S. TCR recognition of peptide–MHC-I: Rule makers and breakers. Int. J. Mol. Sci. 22, 68 (2020).
Swapna, L. S., Bhaskara, R. M., Sharma, J. & Srinivasan, N. Roles of residues in the interface of transient protein-protein complexes before complexation. Sci. Rep. 2, 334 (2012).
Burley, S. K. et al. Protein Data Bank (PDB): the single global macromolecular structure archive. Protein crystallography: methods and protocols, 627–641 (2017).
Szklarczyk, D. et al. The STRING database in 2023: protein–protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Zhang, C., Shine, M., Pyle, A. M. & Zhang, Y. US-align: universal structure alignments of proteins, nucleic acids, and macromolecular complexes. Nat. Methods 19, 1109–1115 (2022).
Mirdita, M. et al. Uniclust databases of clustered and deeply annotated protein sequences and alignments. Nucleic Acids Res. 45, D170–D176 (2017).
Steinegger, M. et al. HH-suite3 for fast remote homology detection and deep protein annotation. BMC Bioinforma. 20, 1–15 (2019).
Gribskov, M., McLachlan, A. D. & Eisenberg, D. Profile analysis: detection of distantly related proteins. Proc. Natl. Acad. Sci. USA 84, 4355–4358 (1987).
Seemayer, S., Gruber, M. & Söding, J. CCMpred—fast and precise prediction of protein residue–residue contacts from correlated mutations. Bioinformatics 30, 3128–3130 (2014).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Studer, G., Tauriello, G. & Schwede, T. Assessment of the assessment—All about complexes. Proteins: Struct., Funct., Bioinforma. 91, 1850–1860 (2023).
Lin, T. et al. Focal Loss for Dense Object Detection. IEEE Transactions on Pattern Analysis and Machine Intelligence. Vol. 42, 318–327 (2020).
Liu, J., Chen, H. & Zhang, Y. A paired sequence language model for protein-protein interaction modeling. junliu621/PPLM: Publication release. URL https://zenodo.org/records/18256392 (2026).
Acknowledgements
We thank Dr. Yang Li for the discussions. This work was supported in part by the Ministry of Education, Singapore (T1251RES2309 and T2EP20125-0039 to Y.Z.), the Agency for Science, Technology and Research (A*STAR), Singapore (IAF-PP H25J6a0034 to Y.Z.), the National Research Foundation, Singapore (NRF-CRP33-2025-0048 to Y.Z.) and the National Medical Research Council Open Fund – Young Individual Research Grant, Singapore (MOH-OFYIRG25jan-0011 to J.L.). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Y.Z. conceived and designed the project and supervised the work. J.L. developed the methods, trained the models, and analyzed the results. J.L and H.C. prepared the initial data and performed the benchmarks. J.L and Y.Z wrote the manuscript. All authors reviewed and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Xiaoyong Pan and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, J., Chen, H. & Zhang, Y. A paired sequence language model for protein-protein interaction modeling. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70457-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70457-5