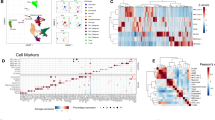
Abstract
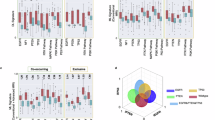
Glioblastoma invasion into brain parenchyma presents significant challenges for treatment but remains poorly understood. In this study, we combine single-cell RNA sequencing, spatial transcriptomics, and multiplexed imaging of orthotopic xenograft models to investigate glioblastoma invasion. We first screen 20 patient-derived gliomasphere models for their distal (i.e., extending to the contralateral hemisphere) and local invasive potential in mice. We show that models with distal invasion potential are enriched with oligodendrocyte progenitor-like cells, while models with only local invasion potential are enriched with mesenchymal-like cells. These patterns reflect predominantly peri-axonal vs peri-vascular invasion routes, respectively. Next, we analyze the transcriptomes of invading cells within models (compared to tumor core) and identify programs associated with distal and local invasion. Thus, we decouple transcriptional features associated with invasion potential from those associated with the process of invasion. We validate our findings by spatial transcriptomics and multiplexed imaging, further describing the spatial niche of invasive cells. Taken together, our results provide a blueprint for the invasive potential of glioblastoma cell states and of the programs associated with invasion across different scales.
Similar content being viewed by others
Data availability
The single-cell RNA sequencing and spatial transcriptomics data generated in this study have been deposited in the Gene Expression Omnibus (GEO) database under accession code GSE281796 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE281796) The published spatial transcriptomics data that was used in this study are available in Datadryad (https://doi.org/10.5061/dryad.wpzgmsbv6) The published bulk RNA-seq data of GBM PDXs that was used in this study are available in the NCBI BioProject database under accession code: PRJNA548556 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA548556) The source data are provided in the Source Data file. Source data are provided with this paper.
Code availability
The code used to create the figures in this manuscript was uploaded to (https://github.com/rchanoch/GBM_Invasion_script).
References
Ostrom, Q. T. et al. CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2012–2016. Neuro-Oncol. 21, v1–v100 (2019).
Horbinski, C., Berger, T., Packer, R. J. & Wen, P. Y. Clinical implications of the 2021 edition of the WHO classification of central nervous system tumours. Nat. Rev. Neurol. 18, 515–529 (2022).
Wen, P. Y. et al. Glioblastoma in adults: a Society for Neuro-Oncology (SNO) and European Society of Neuro-Oncology (EANO) consensus review on current management and future directions. Neuro-Oncol. 22, 1073–1113 (2020).
Stupp, R. et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N. Engl. J. Med. 352, 987–996 (2005).
Drumm, M. R. et al. Extensive brainstem infiltration, not mass effect, is a common feature of end-stage cerebral glioblastomas. Neuro-Oncol. 22, 470–479 (2020).
Holland, E. C. Glioblastoma multiforme: The terminator. Proc. Natl. Acad. Sci. 97, 6242–6244 (2000).
Brown, T. J. et al. Association of the extent of resection with survival in glioblastoma: a systematic review and meta-analysis. JAMA Oncol. 2, 1460 (2016).
Jelsma, R. & Bucy, P. C. The treatment of glioblastoma multiforme of the brain. J. Neurosurg. 27, 388–400 (1967).
Cuddapah, V. A., Robel, S., Watkins, S. & Sontheimer, H. A neurocentric perspective on glioma invasion. Nat. Rev. Neurosci. 15, 455–465 (2014).
Huang-Hobbs, E. et al. Remote neuronal activity drives glioma progression through SEMA4F. Nature 619, 844–850 (2023).
Peiffer, J. & Kleihues, P. Hans-Joachim Scherer (1906-1945), pioneer in glioma research. Brain Pathol. 9, 241–245 (1999).
Miyai, M., Iwama, T., Hara, A. & Tomita, H. Exploring the vital link between glioma, neuron, and neural activity in the context of invasion. Am. J. Pathol. 193, 669–679 (2023).
Scherer, H. J. Etude sur les gliomes. IV. Croissance des gliomes dans leurs rapports avec les substances blanches et grises du cerveau. Bull. Assoc. Franç Etude du Cancer 25, 451–469 (1936).
Scherer, H. J. Sur le développement des structures dans les gliomes. Deuxième Congrès. Int. de. Lutte Scientifique et. Soc. Contre le. Cancer 2, 250–254 (1937).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Neftel, C. et al. An integrative model of cellular states, plasticity, and genetics for glioblastoma. Cell 178, 835–849.e21 (2019).
Couturier, C. P. et al. Single-cell RNA-seq reveals that glioblastoma recapitulates a normal neurodevelopmental hierarchy. Nat. Commun. 11, 3406 (2020).
Chanoch-Myers, R., Wider, A., Suva, M. L. & Tirosh, I. Elucidating the diversity of malignant mesenchymal states in glioblastoma by integrative analysis. Genome Med. 14, 106 (2022).
Hara, T. et al. Interactions between cancer cells and immune cells drive transitions to mesenchymal-like states in glioblastoma. Cancer Cell 39, 779–792.e11 (2021).
Shibue, T. & Weinberg, R. A. EMT, CSCs, and drug resistance: the mechanistic link and clinical implications. Nat. Rev. Clin. Oncol. 14, 611–629 (2017).
Puram, S. V. et al. Single-cell transcriptomic analysis of primary and metastatic tumor ecosystems in head and neck cancer. Cell 171, 1611–1624.e24 (2017).
Jin, X. et al. Frizzled 4 regulates stemness and invasiveness of migrating glioma cells established by serial intracranial transplantation. Cancer Res. 71, 3066–3075 (2011).
Yachi, K. et al. miR-23a promotes invasion of glioblastoma via HOXD10-regulated glial-mesenchymal transition. Signal Transduct. Target. Ther. 3, 33 (2018).
Kim, Y. et al. Perspective of mesenchymal transformation in glioblastoma. Acta Neuropathol. Commun. 9, 50 (2021).
Kahlert, U. D., Nikkhah, G. & Maciaczyk, J. Epithelial-to-mesenchymal(-like) transition as a relevant molecular event in malignant gliomas. Cancer Lett. 331, 131–138 (2013).
Jin, X. et al. Targeting glioma stem cells through combined BMI1 and EZH2 inhibition. Nat. Med. 23, 1352–1361 (2017).
Minata, M. et al. Phenotypic plasticity of invasive edge glioma stem-like cells in response to ionizing radiation. Cell Rep. 26, 1893–1905.e7 (2019).
Greenwald, A. C. et al. Integrative spatial analysis reveals a multi-layered organization of glioblastoma. Cell 187, 2485–2501.e26 (2024).
Venkataramani, V. et al. Glioblastoma hijacks neuronal mechanisms for brain invasion. Cell 185, 2899–2917.e31 (2022).
Yu, K. et al. Surveying brain tumor heterogeneity by single-cell RNA sequencing of multi-sector biopsies. Natl. Sci. Rev. 7, nwaa099 (2020).
Darmanis, S. et al. Single-Cell RNA-Seq Analysis of Infiltrating Neoplastic Cells at the Migrating Front of Human Glioblastoma. Cell Rep. 21, 1399–1410 (2017).
Vaubel, R. A. et al. Genomic and phenotypic characterization of a broad panel of patient-derived xenografts reflects the diversity of glioblastoma. Clin. Cancer Res. 26, 1094–1104 (2020).
Comba, A. et al. Spatiotemporal analysis of glioma heterogeneity reveals COL1A1 as an actionable target to disrupt tumor progression. Nat. Commun. 13, 3606 (2022).
Heaton, H. et al. Souporcell: robust clustering of single-cell RNA-seq data by genotype without reference genotypes. Nat. Methods 17, 615–620 (2020).
Neavin, D. et al. Demuxafy: improvement in droplet assignment by integrating multiple single-cell demultiplexing and doublet detection methods. Genome Biol. 25, 94 (2024).
Kinker, G. S. et al. Pan-cancer single-cell RNA-seq identifies recurring programs of cellular heterogeneity. Nat. Genet. 52, 1208–1218 (2020).
Mathewson, N. D. et al. Inhibitory CD161 receptor identified in glioma-infiltrating T cells by single-cell analysis. Cell 184, 1281–1298.e26 (2021).
Civita, P., Valerio, O., Naccarato, A. G., Gumbleton, M. & Pilkington, G. J. Satellitosis, a crosstalk between neurons, vascular structures and neoplastic cells in brain tumours; early manifestation of invasive behaviour. Cancers 12, 3720 (2020).
Tsai, H.-H. et al. Oligodendrocyte precursors migrate along vasculature in the developing nervous system. Science 351, 379–384 (2016).
Wu, Y. et al. Neurodevelopmental hijacking of oligodendrocyte lineage programs drives glioblastoma infiltration. Dev. Cell 60, 2420–2433 (2025).
Ratliff, M. et al. Individual glioblastoma cells harbor both proliferative and invasive capabilities during tumor progression. Neuro-Oncol. 25, 2150–2162 (2023).
Manoharan, V. T. et al. Spatiotemporal modeling reveals high-resolution invasion states in glioblastoma. Genome Biol. 25, 264 (2024).
Venkatesh, H. S. et al. Neuronal activity promotes glioma growth through neuroligin-3 secretion. Cell 161, 803–816 (2015).
Brooks, L. J. et al. The white matter is a pro-differentiative niche for glioblastoma. Nat. Commun. 12, 2184 (2021).
Kang, S. et al. Glioblastoma shift from bulk to infiltrative growth is guided by plexin-B2-mediated microglia alignment in invasive niches. Nat. Cancer 6, 1505–1523 (2025).
Ubellacker, J. M. et al. Lymph protects metastasizing melanoma cells from ferroptosis. Nature 585, 113–118 (2020).
Doroszko, M. et al. The invasion phenotypes of glioblastoma depend on plastic and reprogrammable cell states. Nat. Commun. 16, 6662 (2025).
El-Botty, R. et al. Oxidative phosphorylation is a metabolic vulnerability of endocrine therapy and palbociclib resistant metastatic breast cancers. Nat. Commun. 14, 4221 (2023).
LeBleu, V. S. et al. PGC-1α mediates mitochondrial biogenesis and oxidative phosphorylation in cancer cells to promote metastasis. Nat. Cell Biol. 16, 992–1003 (2014).
Davis, R. T. et al. Transcriptional diversity and bioenergetic shift in human breast cancer metastasis revealed by single-cell RNA sequencing. Nat. Cell Biol. 22, 310–320 (2020).
Cañellas-Socias, A. et al. Metastatic recurrence in colorectal cancer arises from residual EMP1+ cells. Nature 611, 603–613 (2022).
Ridenour, D. A. et al. The neural crest cell cycle is related to phases of migration in the head. Development 141, 1095–1103 (2014).
Vega, S. et al. Snail blocks the cell cycle and confers resistance to cell death. Genes Dev. 18, 1131–1143 (2004).
Mazzoleni, S. et al. Epidermal Growth Factor Receptor Expression Identifies Functionally and Molecularly Distinct Tumor-Initiating Cells in Human Glioblastoma Multiforme and Is Required for Gliomagenesis. Cancer Res. 70, 7500–7513 (2010).
Wakimoto, H. et al. Maintenance of primary tumor phenotype and genotype in glioblastoma stem cells. Neuro-Oncol. 14, 132–144 (2011).
Hara, T. & Verma, I. M. Modeling gliomas using two recombinases. Cancer Res. 79, 3983–3991 (2019).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform 12, 323 (2011).
Tirosh, I. et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313 (2016).
Spitzer, A. et al. Deciphering the longitudinal trajectories of glioblastoma ecosystems by integrative single-cell genomics. Nat. Genet. 57, 1168–1178 (2025).
Nomura, M. et al. The multilayered transcriptional architecture of glioblastoma ecosystems. Nat. Genet. 57, 1155–1167 (2025).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Van den Berge, K. et al. Trajectory-based differential expression analysis for single-cell sequencing data. Nat. Commun. 11, 1201 (2020).
Ravi, V. M. et al. Spatially resolved multi-omics deciphers bidirectional tumor-host interdependence in glioblastoma. Cancer Cell 40, 639–655.e13 (2022).
Acknowledgements
This work was supported by Grant-in-Aid for JSPS Fellows from the Japan Society for the Promotion of Science (to T.H.), SENSHIN Medical Research Foundation (to T.H.), Kanae Foundation (to T.H.), Brain Research Foundation (to T.H.), American Brain Tumor Association (to T.H.), Seth M Boyer Foundation (to T.H.), N.I.H. R37CA245523 (to M.L.S.), N.I.H. R01CA258763 (to M.L.S.), Mark Foundation Emerging Leader Award (to M.L.S.), Sontag Foundation Distinguished Scientist Award (to M.L.S.), MGH Research Scholars (to M.L.S.), Broad Institute-Israel Science Foundation Collaborative Project Award (to I.T. and M.L.S.), European Research Council Consolidator Grant 101044318 (to I.T.), Zuckerman STEM Leadership Program (to I.T.), Mexican Friends New Generation (to I.T.),. I.T. is the incumbent of the Dr. Celia Zwillenberg-Fridman and Dr. Lutz Zwillenberg Career Development Chair. IMMediate Advanced Clinician Scientist-Program, Department of Medicine II, Medical Center–University of Freiburg and Faculty of Medicine, University of Freiburg, funded by the Bundesministerium für Bildung und Forschung (BMBF, Federal Ministry of Education and Research) − 01EO2103 (to R.H.).
Author information
Authors and Affiliations
Contributions
R.C.M., T.H., A.C.G., M.L.S., and I.T. conceived the project, designed the study, interpreted results, and wrote the manuscript. R.C.M. and A.C.G. performed computational analyses. T.H. performed GBM model experiments. A.C.G. and E.C.F. performed spatial transcriptomics. R.H. performed antibody-based multiplexed imaging (CODEX) experiments. L.B. and H.R.W. helped generate plate-based scRNA-seq data. E.N.B., J.G., Z.B., A.J., J.M.H.F., W.A., and S.C.P. helped perform animal and pathological studies. C.B., J.B., R.G., and H.W. helped GBM model development.
Corresponding authors
Ethics declarations
Competing interests
I.T. is an advisory board member of Immunitas Therapeutics, and a scientific co-founder, equity holder and advisory board member of Cellyrix Therapeutics. M.L.S. is an equity holder, scientific co-founder, and advisory board member of Immunitas Therapeutics. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chanoch-Myers, R., Hara, T., Greenwald, A.C. et al. A blueprint for local and distal invasion programs in glioblastoma. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70470-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70470-8