Abstract
Tau accumulates in a group of neurodegenerative diseases known as tauopathies. A prevailing hypothesis has been that Tau degradation is impaired due to an age-related imbalance in the autophagy-lysosome pathway, but whether these defects are a cause or consequence of Tau accumulation remains unclear. Here we show that a disease-causing mutation in the MAPT gene, which encodes Tau, p.R406W, is sufficient to disrupt multiple steps of the autophagy-lysosome pathway in human neurons. Using Airyscan super-resolution imaging, we find that mutant Tau neurons accumulate Tau and phosphorylated Tau in dysfunctional lysosomes, exhibit reduced lysosome motility, impaired fusion of autophagosomes and lysosomes, and increased undegraded cellular cargo. Pharmacological enhancement of autophagy improves cargo clearance and lowers Tau levels, without restoring defects in lysosomal motility. Together, these findings demonstrate that mutant Tau directly perturbs cellular clearance pathways and suggest that boosting autophagy may help restore Tau homeostasis in tauopathies.
Similar content being viewed by others
Introduction
The dysregulation of protein degradation pathways has been implicated across neurodegenerative disorders1,2,3. In the context of frontotemporal lobar dementia with Tau inclusions (FTLD-Tau), pathological protein aggregates are composed of hyperphosphorylated microtubule-associated protein Tau4,5. The Tau protein, encoded by the MAPT gene, serves to stabilize and organize microtubules within neurons6,7. Mutations within the MAPT gene cause familial forms of FTLD-Tau, leading to altered Tau protein structure8,9,10 and Tau aggregation5,10. However, the mechanism by which these events occur remains uncertain.
As polarized, terminally differentiated cells, neurons encounter distinctive challenges in maintaining the quality and composition of their proteome11. Unlike proliferating cells, neurons cannot dilute proteotoxins through cell division12. Consequently, neurons tightly regulate proteostasis through parallel mechanisms, including the proteasome, macroautophagy, chaperone-mediated autophagy, and endo-microautophagy1,13,14,15. Defects in neuronal autophagy have been implicated in the pathogenesis of numerous neurodegenerative disorders16. However, whether disease-causing mutations are sufficient to disrupt these pathways in the absence of robust protein aggregates remains to be understood.
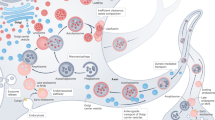
In the autophagy-lysosome pathway, autophagosomes engage with dynein for retrograde transport to the soma, where they undergo maturation into degradative compartments through fusion with LAMP1-positive lysosomes, ultimately leading to cargo degradation1,15. Numerous motor-associated proteins have been identified to regulate the dynein-driven retrograde transport of autophagic vesicles along neurites17,18,19,20,21. Alterations in these motor-associated proteins can disrupt the transport of autophagic vesicles and impede their maturation into degradative lysosomes22. Consequently, there exists a link between lysosome localization and motility with its degradative function18,20,23. Therefore, defects in lysosomal trafficking pathways can perturb neuronal clearance mechanisms, thereby promoting Tau aggregation and contributing to neurodegeneration3.
Here, we sought to determine whether alterations in the autophagy-lysosome pathway are early features in neurons from MAPT mutation carriers and whether restoring these pathways is sufficient to rescue Tau dysfunction. This report focuses on MAPT p.R406W, a pathogenic, missense mutation in exon 13 of the MAPT gene that occurs outside the microtubule binding domain. MAPT p.R406W carriers typically exhibit amnestic dementia, similar to Alzheimer’s disease (AD)24. While MAPT p.R406W brains are free of amyloid plaques, cryo-EM studies demonstrate that Tau filaments adopt an AD-like Tau fold 25. Thus, understanding Tau accumulation in MAPT p.R406W may inform FTLD-Tau as well as AD. We found that MAPT p.R406W is sufficient to alter multiple steps of the autophagy-lysosome pathway and impair Tau degradation. We discovered that mutant Tau neurons have Tau and phosphorylated Tau (pTau)-laden lysosomes, reduced lysosome motility, reduced autophagosome-lysosome fusion, and impaired cargo degradation. Treatment of mutant Tau neurons with an autophagy-enhancing drug promotes autophagy and cargo degradation, reduces Tau and pTau levels, and reduces Tau accumulation in lysosomes without restoring defects in lysosomal motility. Together, our findings point to effects of mutant Tau that can be rescued by targeting autophagy.
Results
Post-translational modifications impact Tau localization in lysosomes in human neurons
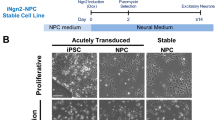
Post-translational modifications (PTMs) on Tau, particularly phosphorylation, play an important role in the normal function of Tau to bind to and stabilize microtubules26,27. Hyperphosphorylation of Tau alters its conformation, promoting Tau aggregation28,29,30. Phosphorylated Tau may be sequestered into autophagic vacuoles and targeted for degradation through the autophagy-lysosome pathway13,31. The intricate interplay between Tau phosphorylation and degradation pathways highlights the significance of PTMs in regulating Tau homeostasis13. While Tau processing in degradative vesicles has been described, these studies have relied on overexpression systems. Little is known of how human neurons process endogenous Tau and how endogenous Tau PTMs impact these degradative pathways. To understand the impact of Tau phosphorylation on Tau degradation in lysosomes, we used patient-derived iPSC-derived neurons harboring a doxycycline-inducible neurogenin-2 (NGN2) cassette stably integrated in the AAVS1 locus (Supplementary Figs. 1 and 2). NGN2 expression in iPSCs results in a homogeneous population of excitatory neurons within seven days32. These neurons express the neuronal markers Microtubule-Associated Protein 2 (MAP2) and β-tubulin III (Tuj1) by DIV7 and synapsin by DIV14 (Supplementary Fig. 3).
To understand where endogenous Tau localizes within the endolysosomal compartments of human neurons, we visualized total Tau (Tau5) and phosphorylated Tau (AT180; pThr231; pTau) in LAMP1-positive (LAMP1 + ) vesicles of DIV14 neurons using immunocytochemistry and Airyscan super-resolution microscopy (Fig. 1A and Supplementary Fig. 4A). LAMP1+ vesicles appeared as donut-shaped structures (Supplementary Fig. 4A). LAMP1+ vesicles were Cathepsin D positive (74.2%; Supplementary Fig. 4B) and lysosomal membrane protein 2 (LIMP2) positive (95.2%; Supplementary Fig. 4C)33,34,35,36. Thus, the majority of LAMP1+ structures we detected were lysosomes. Super-resolution images of LAMP1+ vesicles showed that total Tau and pTau localize to different compartments in the lysosome (Fig. 1A, B). Signal intensity measurements illustrate that total Tau localization was enriched within the lysosomal lumen (Fig. 1B, C and Supplementary Fig. 4D), while pTau localization was enriched at the lysosomal membrane (Figs. 1B, D and Supplementary Fig. 4D). We discovered a difference in the localization of total Tau and pTau in lysosomes, which may point to a difference in degradative efficiency: 70% of lysosomes were free of Tau (Tau- or empty), while only 40.7% of lysosomes were free of pTau (Fig. 1B, E). Applying immuno-electron microscopy, pTau (anti-AT180; pTau-Thr231) was immunogold labeled to detect pTau localization in human neurons. Phosphorylated Tau was predominantly observed on vesicles containing electron-dense material, consistent with the morphology of a lysosome (Fig. 1F). We observed significantly more pTau on the lysosomal membrane compared with the lumen (Fig. 1F, G). Thus, our findings point to differences in the way wild-type (WT) neurons handle Tau and pTau in the lysosome, which may ultimately lead to differences in degradation efficiency (Fig. 1H).
A iPSC-derived NGN2 neurons (control) were cultured until DIV14 and immuno-stained for LAMP1+ vesicles (green), MAP2 (magenta). Scale bar 5 µm. Right inset: LAMP1+ vesicles (Red) co-stained for total Tau (Tau5; green) and pTau (AT180; green). Imaging by Airyscan super-resolution microscopy. Scale bar 0.5 µm. B Tau and pTau localization within LAMP1+ vesicles differ. Dashed line used as a reference for subsequent quantification. LAMP1+ vesicles (red) co-stained for total Tau (Tau5; green) and pTau (AT180; green). Scale bar 0.5 µm. C, D Quantification of fluorescent intensity of Tau (Tau5; C) and pTau (AT180; D) in LAMP1+ vesicles across the dashed line in (B). The data displayed is a representation of the mean localization calculated across 3 independent experiments. See Supplementary Fig. 4D for individual traces. C Tau, n = 35 LAMP1+ vesicles. D pTau, n = 27 LAMP1+ vesicles. E Quantification of the percentage of LAMP1+ vesicles per cell that contain total Tau or pTau (Tau+ or free of Tau, Tau-). Graph represents mean ± SEM. 186 LAMP1+ vesicles were quantified from three independent experiments. Two-tailed Mann–Whitney U-test. ***p = 0.0043. F Immunogold electron microscopy of pTau in iPSC-derived NGN2 neurons. Immunogold labeling with the anti-pTau antibody (AT180) is associated with the single-membrane vesicles containing electron-dense material (defined as lysosomes). Mitochondria (Mito), the endoplasmic reticulum (ER), and the nucleus (Nuc) are visible but remain unlabeled. pTau predominantly localized to the membranes of electron-dense structures with single-limiting membranes (lysosomes, right inset). Additionally, some structural labeling was observed along microtubules. Scale bar 1 µm (left) and 50 nm (right, inset). G Quantification of gold particle-labeled pTau localization within lysosomal structures. The graph represents mean ± SEM. 26 lysosomes were quantified. Two-tailed Mann–Whitney U-test. ****p < 0.0001. H Diagram of Tau and pTau localization in lysosomes based on our observations. Source data are provided as a Source Data file. Created in BioRender. Mirfakhar, F. (2026) https://BioRender.com/8bzn5l2.
Tau accumulates in MAPT p.R406W neurons
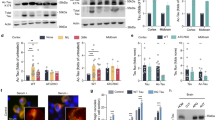
Given our finding that total Tau and pTau differ in their lysosomal localization in WT neurons, we sought to understand the impact of a FTD-causing MAPT mutation on Tau phosphorylation and Tau degradation in lysosomes. Studying Tau degradation in neurons has been challenging due to the lack of proper human cell models, resulting in the overreliance on Tau overexpression in immortalized cell lines37,38. NGN2 was engineered into the AAVS locus of iPSC lines from patients carrying the FTD-causing MAPT p.R406W mutation and its isogenic, WT pair (Fig. 2A; Supplementary Figs. 1 and 2). MAPT p.R406W neurons (DIV14) produced significantly more total Tau (Tau5), Tau phosphorylated at pThr231 (AT180; Fig. 2B-D), and Tau phosphorylated at pSer202/pThr205 (AT8; Supplementary Fig. 5A, B) compared to isogenic control neurons. The ratio of pTau/total Tau was also significantly higher in mutant neurons using immunofluorescence (Fig. 2E) and immunoblotting (Supplementary Fig. 5C–F).
A Diagram of the stem cell model used in this study. Human-derived iPSCs with NGN2 integrated into the AAVS locus were differentiated into neurons (NPC: neuronal progenitor cells). MAPT mutation carrier (p.R406W) and CRISPR/Cas9-corrected isogenic control (wild-type; WT) were differentiated into cortical neurons upon doxycycline induction. B MAPT p.R406W neurons display more total Tau and pTau. Neurons were defined with MAP2 staining (green). MAPT WT and p.R406W neurons were stained for total Tau (Tau5; Magenta) and pTau (AT180; cyan). Scale bar 20 µm. C–E Quantification of the relative mean fluorescence intensity for total Tau (C), pTau (D), and pTau/Tau ratio (E). Data are represented as mean ± SEM. Data on the y-axis are normalized to WT. 240 cells were quantified from 3 independent experiments. Two-tailed Mann–Whitney U-test; **p = 0.0077; ***p = 0.0002; ****p < 0.0001. F Schematic illustrating the in vitro Tau SILK protocol to monitor Tau production. iPSC-derived neurons were cultured for 4 weeks, followed by labeling with 50 mol % (100% TTR) 13C6-leucine for 15 days from week 4 to 6. Cells were collected every 3 days. G Kinetic profiles of mid-domain Tau peptides (TPSLPTTPR) in the cells. Production of the Tau peptides in the cell lysates was similar in iPSC-neurons with and without MAPT p.R406W mutation. Two-tailed Wilcoxon test, p = 0.437. H Neurons were treated with Bafilomycin A1 (Baf,100 nM) and MG-132 (5 µM) to transiently block all protein degradation. Neurons were defined with MAP2 staining (green). MAPT WT and p.R406W neurons were stained for Tau (Tau5; Magenta) and pTau (AT180; cyan). Scale bar 20 µm. I, J Quantification of relative mean fluorescence intensity for total Tau (I) and pTau (J). Data are represented as mean ± SEM. Data on the y-axis are normalized to WT-DMSO. 167 cells were quantified across 3 independent experiments. Kruskal-Wallis test followed by Dunn’s test; *p = 0.024; **p = 0.0085; ***p = 0.0005; ****p < 0.0001; ns, not significant. Source data are provided as a Source Data file. Created in BioRender. Mirfakhar, F. (2026) https://BioRender.com/8bzn5l2.
An increase in total Tau and pTau could reflect a defect in Tau production or clearance. To determine whether MAPT p.R406W neurons exhibit impaired Tau production, we performed stable isotope labeling kinetics (SILK)39. Briefly, iPSC-derived neurons from MAPT p.R406W and isogenic controls were cultured in media containing 13C6-leucine for 15 days (Fig. 2F). During this labeling period, a subset of any new proteins produced incorporated a 13C6 at leucine residues (rather than 12C6). We next isolated Tau using immunoprecipitation followed by mass spectrometry. When we monitored a major, mid-domain peptide (TPSL), we found that the production of Tau occurs at a similar rate in MAPT p.R406W and isogenic control neurons (Fig. 2G). These findings suggest that the mutation does not have a robust impact on Tau production. Using an orthogonal approach to evaluate the impact of MAPT p.R406W on Tau production, we treated neurons with Bafilomycin A1 (BAF) and MG132. By simultaneously inhibiting both lysosomal (BAF) and proteasomal (MG132) pathways, we were able to block all major Tau degradation routes and isolate Tau production without confounding clearance effects40. We then measured Tau and pTau (AT180) levels by immunofluorescence (Fig. 2H). Total Tau and pTau levels were similar in mutant and control neurons after blocking protein degradation, suggesting that changes in Total Tau and pTau levels are driven by clearance defects rather than production defects (Fig. 2H–J).
Tau degradation is impaired in MAPT p.R406W neurons
To determine whether MAPT p.R406W alters Tau localization in lysosomes, we immunostained MAPT p.R406W neurons and isogenic controls for total Tau (Tau5) and pTau (AT180) at DIV14. Total Tau was enriched in the lysosomal lumen in both MAPT p.R406W and isogenic control neurons (Fig. 3A, B); however, mutant neurons exhibited significantly more Tau in the lumen compared to isogenic controls (p.R406W: 69.6% vs. WT: 19.41%; p = 0.002; Fig. 3A, B, E and Supplementary Fig. 5G). We also observed a modest increase in membrane-bound Tau in the mutant neurons compared to the controls (p.R406W: 26.7% vs. WT: 11.8%; p = 0.017; Fig. 3A, B, E and Supplementary Fig. 5G). MAPT p.R406W neurons exhibited significantly more pTau in the lysosomal membrane than isogenic controls (p.R406W: 70.3% vs. WT: 26.2%; p = 0.016; Fig. 3C, D, F and Supplementary Fig. 5G). Strikingly, MAPT p.R406W neurons had fewer lysosomes free of total Tau (p.R406W: 3.6% vs. WT: 68.7%; p = 0.002) and pTau (p.R406W: 10.3% vs. WT: 51.2%; p = 0.015; Fig. 3C, D, F and Supplementary Fig. 5G) compared to isogenic controls. Immuno-electron microscopy using pTau (anti-AT180; pTau-Thr231) in MAPT p.R406W neurons revealed a significant increase in pTau on the lysosomal membrane (vesicles containing electron-dense material) compared with isogenic controls (Fig. 3G, H). Together, these findings suggest that Tau degradation in the lysosome is impaired in MAPT p.R406W neurons.
A, C MAPT WT and p.R406W neurons were stained for LAMP1 (red) and MAP2 (green). Scale bar 10 µm. B Magnified images of LAMP1+ vesicles (red) and total Tau (Tau5; green). Representative images of LAMP1+ vesicles free of total Tau (top panel), total Tau staining on the lysosomal membrane (middle panel), and total Tau staining in the lysosomal lumen (bottom panel). Scale bar 1 µm. D Magnified images of LAMP1+ vesicles (red) and pTau (AT180; green). Representative images of LAMP1+ vesicles free of pTau (top panel), pTau detected on the lysosomal membrane (middle panel), and pTau detected in the lysosomal lumen (bottom panel). Scale bar 1 µm. E, F Quantification of the percentage of LAMP1+ vesicles per cell. 186 LAMP1+ vesicles were quantified. Two-way ANOVA. E. LAMP1+ vesicles empty of total Tau or with total Tau in the membrane vs lumen. Total Tau; WT: Empty vs. p.R406W: Empty, **p = 0.002; WT: Membrane vs. p.R406W: Membrane, *p = 0.017; WT: Lumen vs. p.R406W: Lumen, **p = 0.002. % of lysosomes: Total Tau; WT: Empty = 68.75%; Membrane = 11.84%; Lumen = 19.41%. Total Tau; p.R406W: Empty = 3.65%; Membrane = 26.74%; Lumen = 69.61%. F. LAMP1+ vesicles empty of pTau or with pTau in the membrane vs lumen. pTau, WT: Empty vs. p.R406W: Empty, *p = 0.015; WT: Membrane vs. p.R406W: Membrane, *p = 0.016; WT: Lumen vs. p.R406W: Lumen p = 0.73. % of lysosomes: pTau; WT: Empty = 51.23%; Membrane = 26.27%; Lumen = 22.5%. pTau; p.R406W: Empty = 10.34%; Membrane = 70.3%; Lumen = 19.36%. G Immunogold labeling with the anti-pTau antibody (AT180) in MAPT WT and p.R406W neurons is associated with the single-membrane vesicles containing electron-dense material (defined as lysosomes). Scale bar 0.5 µm. H Quantification of gold particle-labeled pTau localization within lysosomal structures. 53 lysosomes were quantified (WT n = 26; p.R406W n = 27). Kruskal–Wallis test followed by Dunn’s test; **p = 0.0085; ***p = 0.0005; ****p < 0.0001; ns, not significant. Data represent mean ± SEM. Data representative of 3 independent experiments. Source data are provided as a Source Data file.
Autophagy-lysosome pathways are disrupted in MAPT p.R406W neurons
To assess whether MAPT p.R406W neurons display alterations in protein degradation pathways, we conducted a targeted reanalysis of previously published RNA sequencing data from MAPT p.R406W neurons and isogenic controls41. In the original study, the most significantly dysregulated pathways were related to endolysosomal function, synapse, and neuronal health41, providing a framework for the current analysis of protein degradation machinery. The significantly differentially expressed genes in MAPT p.R406W neurons41 were enriched in protein degradation pathways associated with phagosome maturation (ATP6V1B2, ATP6V1H, ATP6V1E1), regulation of autophagy (PLK2, ATP6V1B2, ATP6V1H, ATP6V1E1, EIF4G2), autophagosome (VMP1, HSPA8), and vesicle transport along the microtubule (RAB1A, KIF3B; Fig. 4A; Supplementary Fig. 6A). Based on this observation, we systematically evaluated the expression of genes involved in pathways associated with autophagy and lysosomal functions42,43. We also discovered that a number of lysosomal genes, including vATPases and autophagy genes, are significantly reduced in MAPT p.R406W neurons (Supplementary Fig. 6B–D; Supplementary Data 1). Thus, genes related to the autophagy-lysosome pathway are altered in mutant neurons.
A Differentially expressed genes in MAPT p.R406W neurons are enriched in pathways associated with the autophagy-lysosome pathway (FDR\(\le\)0.05). White bars, pathways with genes significantly upregulated in MAPT p.R406W neurons. Gray bars, pathways with genes significantly downregulated in MAPT p.R406W neurons. B Schematic. LSD, lysosomal storage disease. C Representative Tuj1+ (green) neurons at DIV14, stained for LAMP1 (red). Right, magnified images of LAMP1+ donut-shaped structures (red) distributed in neurons. Scale bar 5 µm. Inset, scale bar 0.5 µm. Images acquired from 3 independent experiments. D Tuj1+ (green) neurons at DIV14. MAPT p.R406W or isogenic controls (WT) were stained for LAMP1 (red). White arrows, lysosomes in neurites. Scale bar 5 µm. E–G Lysosomal morphology quantified in MAPT p.R406W neurons compared with isogenic controls (WT). Data are mean ± SEM from 3 independent experiments. Data normalized to WT. Two-tailed Mann-Whitney U test. E Quantification of LAMP1+ vesicle density defined as the total number of LAMP1+ puncta within Tuj1+ cells. 19 cells were quantified per genotype. **p = 0.0048. F LAMP1+ vesicle volume. Lysosomes quantified: MAPT WT n = 208; MAPT p.R406W n = 318. ****p < 0.0001. G Lysosome distance from the nucleus is defined as the shortest distance of LAMP1+ puncta to DAPI + . Lysosomes quantified: MAPT WT n = 164; MAPT p.R406W n = 214. ****p < 0.0001. H iPSC-derived NGN2 neurons (DIV14) were MAP2+ and co-stained with LAMP1 and Cathepsin (CatD). LAMP1+ vesicles (red) co-label with CatD (green) in MAPT p.R406W neurons and isogenic control cells (WT). Scale bar 10 µm. Inset, scale bar 0.5 µm. I Quantification of the percentage of LAMP1 + /CatD+ lysosomes in MAPT WT (n = 22) and MAPT p.R406W (n = 16) cells. J Quantification of the percentage of CatD+ vesicles in MAPT WT (n = 22) and MAPT p.R406W (n = 16) neurons expressed relative to WT control. p = 0.0373. K Quantification of LAMP1 + /CatD+ size (area). MAPT WT n = 219; MAPT p.R406W n = 242 lysosomes. ****p < 0.0001. Two-tailed Mann–Whitney U-test. 3D reconstruction of Airyscan microscopy images and analyses were performed by Imaris. L Diagram of hypothesized disrupted pathways derived from GO data. Source data are provided as a Source Data file. Created in BioRender. Mirfakhar, F. (2026) https://BioRender.com/8bzn5l2.
Autophagosome clearance is a multistep process involving vesicular trafficking, fusion, and degradation44. Autophagy impairment can lead to a disruption in normal fusion and degradation processes within the cell45,46,47. When autophagy is impaired, undegraded material may accumulate within the lysosomes, causing them to enlarge48,49. Lysosomal enlargement is an indication of undigested material reminiscent of the pathophysiology observed in lysosomal storage disorders (LSD; Fig. 4B)50,51. To evaluate the impact of MAPT p.R406W on lysosomal size, we used Airyscan super-resolution microscopy to visualize LAMP1 + , ring-shaped structures of different sizes along the neurites as well as in the perinuclear region (Fig. 4C). Quantification of LAMP1+ vesicle density and volume revealed increased lysosomal density (vesicle count) and volume in MAPT p.R406W neurons compared to the isogenic control neurons (Fig. 4D-F). LysoTracker+ vesicles were also significantly larger in mutant neurons (Supplementary Fig. 7A, B). Interestingly, the mean intensity of LysoTracker was significantly lower in MAPT p.R406W neurons compared with isogenic controls (Supplementary Fig. 7A,C). Together, the LysoTracker staining points to disruptions in lysosomal function. An increase in the volume of LAMP+ and LysoTracker+ vesicles, along with a decrease in Lysotracker intensity, is consistent with the observed accumulation of undigested material, including Tau and pTau, in MAPT p.R406W neurons (Fig. 3).
Our transcriptomic analyses identified defects in vesicle transport along the microtubule, which may influence lysosomal positioning within the neurons. Lysosomal distance from the nucleus has been shown to influence the degradative capacity of lysosomes (Fig. 4B, C)52. MAPT p.R406W neurons exhibited significantly more LAMP1+ vesicles (Fig. 4D, E) that were larger in size than LAMP1+ vesicles (Fig. 4D, F) measured in isogenic controls. To gain insight into the impact of MAPT p.R406W on lysosomal positioning along the neurites, we measured the LAMP1+ vesicle distance from the nucleus. Lysosomes in MAPT p.R406W neurons were located further from the nucleus compared to the lysosomes in isogenic control neurons (Fig. 4D, G). These LAMP1+ vesicles co-localized with Cathepsin D, consistent with a lysosomal vesicle (Fig. 4H, I). MAPT p.R406W and WT neurons displayed a similar percentage of LAMP1 + /Cathepsin D+ co-labeled lysosomes per cell (Fig. 4H, I). There were significantly more Cathepsin D+ vesicles in mutant neurons compared with isogenic controls (Fig. 4H, J). Dual LAMP1/Cathepsin D+ vesicles were significantly larger in MAPT p.R406W neurons compared to the isogenic controls (Fig. 4H, K). The peripheral localization of lysosomes in mutant neurons may reflect aberrant vesicle trafficking, particularly in retrograde trafficking (Fig. 4L)21,53. Taken together, molecular and morphologic evidence suggest that MAPT p.R406W induces defects in autophagy and lysosomal trafficking in human neurons, which may impact degradative capacity.
MAPT p.R406W neurons exhibit defects in lysosomal transport
Many lysosomal functions, including degradative capacity, are influenced by their positioning and motility18,20. Although some lysosomes are relatively static, others move along microtubules between the soma and the periphery of the cell18. Our data suggest that lysosomes are farther away from the soma in MAPT p.R406W neurons (Fig. 4G). Given the relationship between Tau and microtubules, we hypothesized that MAPT p.R406W disrupts lysosomal trafficking. Lysosomal motility was monitored using time-lapse confocal live-cell imaging of LysoTracker+ vesicles (Fig. 5A). In MAPT p.R406W neurons, LysoTracker+ vesicles traveled significantly shorter distances compared to isogenic controls (Fig. 5A, B). LysoTracker+ vesicles also exhibited significantly reduced velocity in mutant neurons compared to isogenic controls (Fig. 5A, C). Together, these findings suggest that MAPT p.R406W impacts lysosomal motility.
MAPT p.R406W neurons compared with isogenic controls (WT) at DIV14. A Displacement and velocity of acidic vesicles were visualized using live imaging and analyzed by Imaris. Left panel: Lysotracker+ (red) acidic vesicles tracked for displacement length. Right, shorter distances (cyan) and longer distances (magenta). Scale bar 10 µm. Right panel: Lysotracker+ puncta (yellow). Right, slower velocity (cyan) and faster velocity (magenta). Scale bar 10 µm. B Quantification of relative distance traveled by Lysotracker+ vesicles in (A). MAPT WT n = 510 and p.R406W n = 856 vesicles. Two-tailed Mann–Whitney test. ***p = 0.0004. C Quantification of the relative mean velocity of Lysotracker+ vesicles in (A). MAPT WT n = 336 and p.R406W n = 415 vesicles. Two-tailed Mann–Whitney test. **p = 0.0095. D Dendritic complexity was evaluated using MAP2 (green). 3D reconstruction by Imaris (magenta) based on MAP2 signal (green). Scale bar 10 µm. E–G MAPT WT n = 30 and p.R406W n = 35 cells. E Quantification of arborization area. Two-tailed Mann-Whitney U test. p = 0.195. F Total dendrite length was quantified by measuring the lengths of all branches per dendritic compartment. Two-tailed Mann-Whitney U-test. p = 0.385. G Sholl analysis. Quantification of the number of intersections at concentric circles of increasing radii centered around the soma every 10μm. MAPT WT n = 9 and p.R406W n = 9 cells. Two-way ANOVA. p = 0.315. H Fluorescence recovery after photobleaching (FRAP) analysis of microtubules (Viafluor, green) in dendritic sections. Time-lapse sequences illustrate the recovery of fluorescence in the bleached ROI. I FRAP quantification of microtubule signals. Curves depict the average intensity normalized to the first frame. MAPT WT n = 9 and p.R406W n = 8 cells. Two-way ANOVA. p = 0.22. J Expression level of motor-associated genes detected by RNAseq. Transcripts per million (TPM). Two-tailed Mann-Whitney U-test. *p = 0.023. K MAPT p.R406W and WT neurons (MAP2; cyan) were immunostained for JIP3 (red). Scale bar 10 µm. L Quantification of JIP3 mean fluorescence intensity. MAPT WT n = 23 and p.R406W n = 25 cells. Two-tailed Mann–Whitney U-test. *p = 0.0358. M Schematic. All data are represented as mean ± SEM and were obtained from 3 independent experiments. Source data are provided as a Source Data file. Created in BioRender. Mirfakhar, F. (2026) https://BioRender.com/8bzn5l2.
To determine whether the observed alterations in lysosomal distribution and motility in MAPT p.R406W neurons could be attributed to mutation-induced alterations in cell morphology, we quantified dendritic arborization, dendritic length, and nodes in MAP2-immunostained neurons (Fig. 5D). Neuronal arborization, dendritic length, and the number of nodes (measured by Sholl analysis) were similar between MAPT p.R406W and isogenic control neurons (Fig. 5D–G). Thus, MAPT p.R406W likely does not impact lysosomal motility due to defects in neuronal morphology.
Vesicles move along microtubules, and the Tau protein promotes microtubule stability. Therefore, we tested whether lysosomal trafficking defects occur due to MAPT p.R406W-induced microtubule instability. Microtubule fluorescence recovery rate was measured in live cells by quantifying fluorescence recovery after photobleaching (FRAP) using microtubule probes. Monitoring the assembly of new fluorescent tubulins into the bleached area reflects microtubule polymerization rate and stability. The rate of recovery of the microtubule signal after photobleaching was similar in mutant and isogenic control neurons (Fig. 5H, I). Thus, it is unlikely that lysosomal trafficking defects are driven by microtubule alterations in MAPT p.R406W neurons. These findings are consistent with a role for Tau in neurodegeneration that is not primarily driven by its microtubule-stabilizing functions54,55.
The positioning and trafficking of lysosomes rely on a complex interplay of interactions between microtubule motors and adaptor proteins, such as JIP3, ARL8, and RILP56,57,58,59. To determine whether MAPT p.R406W impacts motor adaptor proteins, we evaluated expression of motor adaptor genes in our transcriptomic data (Fig. 5J). Among the candidate motoradaptor genes, only JIP3 (also known as MAPK8IP3) was significantly upregulated in MAPT p.R406W neurons compared with isogenic controls (Fig. 5J). JIP3 protein was also significantly elevated in mutant neurons (Fig. 5K, L). Motility of the most mature population of autophagic vesicles is regulated by the motor-interacting protein JIP322,56,58,59. As autophagosomes and lysosomes travel along axons, they acquire multiple motor-interacting proteins that sequentially regulate dynein activity and lysosome trafficking, depending on their maturation status and location within axons (Fig. 5M)22. Interestingly, lysosome distribution in neurites is sensitive to JIP3 levels: JIP3 knockout and JIP3 overexpression result in axonal swellings, suggesting that JIP3-mediated functions are tightly regulated by JIP3 expression58,59.
MAPT p.R406W neurons exhibit impairment in autophagy
Proper lysosome positioning is essential for autophagy60. This process starts with the engulfment of organelles into an autophagosome, which subsequently fuses with a lysosome (Fig. 6A). This process is pivotal to stymie neurodegenerative disease, as it facilitates the recycling of dysfunctional proteins44. Transcriptomic analyses pointed to the dysregulation of genes enriched in pathways associated with autophagosomes and regulation of autophagy (Fig. 4A). To determine whether MAPT p.R406W contributes to defective autophagy, we assessed the overall abundance of autophagosomes in MAPT p.R406W and isogenic controls using CYTO-ID (Fig. 6B and Supplementary Fig. 8A). Mutant neurons exhibited significantly more autophagosomes compared to isogenic controls (Fig. 6B–D). Dual treatment with rapamycin, an autophagy enhancer, and chloroquine, an inhibitor of autophagosome-lysosome fusion, (Rap/CQ) was used as a control to visualize autophagosome accumulation (Fig. 6B)61. Compared to DMSO-treated neurons, Rap/CQ treatment significantly enhanced CYTO-ID intensity and CYTO-ID positive vesicle density in both mutant and isogenic control neurons (Fig. 6B–D and Supplementary Fig. 8B). LC3B-II sits on the autophagosome membrane, where it serves as a marker of autophagosome initiation and maturation62. Consistent with our CYTO-ID findings, LC3B intensity was significantly elevated in mutant neurons (Fig. 6E, F and Supplementary Fig. 8C). Immunoblotting also revealed an increase in the ratio of LC3BII to LC3BI in MAPT p.R406W neurons (Supplementary Fig. 8D–F). Together, these results suggest that MAPT p.R406W increases autophagosomes.
MAPT p.R406W and isogenic control (WT) neurons at DIV14. A Diagram. B Live imaging using CYTO-ID (green) and microtubule marker (magenta). Rapamycin (Rap, 500 nM); Chloroquine (CQ, 20uM). Scale bar 10 µm. Lower panels, CYTO-ID (green). Scale bar 2 µm. C CYTO-ID mean fluorescence intensity per cell. Cells: WT (n = 26), WT-Rap+CQ (n = 29), p.R406W (n = 34), p.R406W-Rap+CQ (n = 29). Kruskal–Wallis test followed by Dunn’s test. **p = 0.0019; ****p < 0.0001. D CYTO-ID density (CYTO-ID+ puncta per cell). Cells: WT (n = 17), WT-Rap+CQ (n = 20), p.R406W (n = 17), p.R406W-Rap+CQ (n = 19). Kruskal–Wallis test followed by Dunn’s test. *p = 0.0366; *p = 0.0150; **p = 0.0021; ****p < 0.0001. E Neurons stained for MAP2 (magenta), LC3B (red), and p62 (green). Scale bar 10 µm. F LC3B mean intensity per cell. Cells: MAPT WT (n = 36) and p.R406W (n = 47). Two-tailed Mann–Whitney U-test. ****p < 0.0001. G p62 mean intensity per cell. Cells: MAPT WT (n = 33) and p.R406W (n = 36). Two-tailed Mann-Whitney U-test. ****p < 0.0001. H Live imaging with microtubule (magenta) and BODIPY (lipid droplets, green). Scale bar 20 µm. Lower panels, BODIPY. Scale bar 2 µm. I BODIPY mean intensity per cell. Cells: MAPT WT (n = 51) and p.R406W (n = 61). Mann–Whitney U-test. ****p < 0.0001. J Schematic. K Neurons expressing the pHluorin-mKate2-LC3 reporter. Scale bar 5 μm. Right panel: puncta/autophagic vesicles, pHLuorin (green), mKate2 (red), merged signals (yellow). Scale bars 1 µm. L Puncta size. Puncta: WT (n = 48), p.R406W (n = 87), WT-Rap+CQ (n = 181). Kruskal-Wallis test followed by Dunn’s test. ***p = 0.0002; ****p < 0.0001. M mKate2 to pHLuorin ratio. Puncta: WT (n = 64), p.R406W (n = 87), WT-Rap+CQ (n = 181). Kruskal–Wallis test followed by Dunn’s test. ***p = 0.0005, ****p < 0.0001. N Number of autophagosomes (mKate + / pHLuorin + ) and autolysosomes (mKate + /pHLuorin-) puncta. Total puncta: MAPT WT (n = 64), p.R406W (n = 87); WT-Rap+CQ (n = 111). Autolysosomes: MAPT WT (n = 53), p.R406W (n = 27), WT-Rap+CQ (n = 15). Autophagosomes: MAPT WT (n = 11), p.R406W (n = 60), WT-Rap+CQ (n = 70). Two-way ANOVA followed by Tukey multiple comparisons test, *p = 0.0205; *p = 0.0146; **p = 0.0059; ***p = 0.0006; ***p = 0.0008. WT (n = 12 cells), MAPT p.R406W (n = 11 cells), WT-Rap+CQ (n = 13 cells). All data are represented as mean ± SEM from 3 independent experiments. Source data are provided as a Source Data file. Created in BioRender. Mirfakhar, F. (2026) https://BioRender.com/8bzn5l2.
Autophagosomes undergo fusion with lysosomes, where the vesicle cargo goes on to be degraded by lysosomal hydrolases63. P62 (also known as sequestosome-1 or SQSTM1) binds to cargo, facilitating cargo degradation within autophagosomes64,65,66. When autophagic degradation is disrupted or inefficient, p62 levels increase66. MAPT p.R406W neurons exhibited significantly more p62 than isogenic controls (Fig. 6E, G). Thus, despite an increase in autophagosomes (CYTO-ID and LC3B), cargo was not degraded efficiently in mutant neurons.
To determine whether the observed defects in autophagy impact cargo degradation more broadly, we evaluated neutral lipids. While these lipid droplets are not harmful by themselves, they can represent disrupted lipid processing and distribution67. To determine whether MAPT p.R406W impacts lipid droplet biogenesis, we used BODIPY 493/503. We observed significantly more lipid droplet content in mutant neurons relative to controls based on the overall BODIPY fluorescence per cell (Fig. 6H, I and Supplementary Fig. 8G). These data suggest that MAPT p.R406W induces defects in autophagy that have a broad impact on cargo, which ultimately promotes lipid dysregulation.
To further pinpoint the defects in autophagy, we next examined the impact of MAPT p.R406W on fusion between autophagosomes and lysosomes. MAPT p.R406W and isogenic control neurons were transduced with Lenti viral particles containing a tandem pH-sensing fluorescent pHLuorin-mKate2-tagged LC3 probe, which has been shown to distinguish pre-fusion autophagic compartments from mature, acidic autolysosomes based on the differential pH sensitivity of pHLuorin versus mKate2 (Fig. 6J)68. In autophagosomes, both green (pHLuorin) and red (mKate2) signals are present. When autophagosomes successfully fuse with lysosomes to form autolysosomes, the green (pHLuorin) is quenched, and the vesicles remain red (mKate2). We observed the more enlarged yellow puncta in transduced mutant neurons (Fig. 6K, L and Supplementary Fig. 8H), representing more autophagosomes and consistent with our CYTO-ID findings. Furthermore, the ratio of mKate2/pHLuorin was significantly decreased in MAPT p.R406W compared to isogenic control neurons (Fig. 6K, M). The reduced mKate2/pHLuorin ratio in MAPT p.R406W neurons was driven by significantly more autophagosomes (mKate2 + /pHLuorin+ puncta) and significantly fewer autolysosomes (mKate2 + /pHLuorin− puncta) compared to isogenic control neurons (Fig. 6N). Rap/CQ treatment was performed in WT neurons as a control for autophagosome accumulation. Together, our findings suggest that the MAPT p.R406W mutation is sufficient to disrupt the efficient fusion of autophagosomes with lysosomes.
Autophagy enhancer G2-567 rescues Tau defects in MAPT p.R406W neurons
Given our findings of Tau accumulation and autophagy defects in MAPT p.R406W neurons, we sought to determine whether pharmacological enhancement of autophagy could reverse Tau accumulation. G2-567 is a well characterized small molecule tool that enhances autophagy69,70,71. The 567 analog is more potent and stable than the 115 analog while retaining the same mechanism of action. G2 promotes autophagy via RCAN1-TFEB70, leading to increased autophagosome and autolysosome puncta per cell, reduced p62, and increased CytoID69,70,71. G2 analogs have been shown to reduce Huntingtin inclusions and misfolded α1-antitrypsin Z69,70,71. MAPT p.R406W neurons and isogenic controls were treated with G2-567 (0.5 µM) or DMSO vehicle control for 14 days beginning on DIV7. G2-567 significantly reduced pTau (Fig. 7A, B) and total Tau (Fig. 7C, D) levels in mutant neurons compared to DMSO-treated mutant neurons. Treatment of MAPT p.R406W neurons with G2-567 did not significantly alter the ratio of pTau/total Tau compared to DMSO-treated mutant neurons (Fig. 7E). Interestingly, total Tau was also significantly reduced in G2-567-treated WT neurons (Fig. 7C, D).
MAPT p.R406W and isogenic control neurons were treated with G2-567 (0.5 µM) or DMSO for 14 days beginning on DIV7 and were fixed on DIV21. A Neurons (MAP2, green) stained for pTau (AT180, red). Scale bar 20 µm. B Quantification of pTau mean intensity per cell (WT n = 52; WT-G2-567 n = 52; p.R406W n = 59; p.R406W-G2-567 n = 56). Data are represented as mean ± SEM. Data on the y-axis are normalized to WT-DMSO. Kruskal–Wallis test followed by Dunn’s test. ****p < 0.0001. C Neurons (MAP2, green) stained for total Tau (Tau5, magenta). Scale bar 20 µm. D Quantification of Tau mean intensity per cell (WT n = 43; WT-G2-567 n = 63; p.R406W n = 51; p.R406W-G2-567 n = 37). Data are represented as mean ± SEM. Data on the y-axis are normalized to WT-DMSO. Kruskal–Wallis test followed by Dunn’s test. *p = 0.0491 and p = 0.0506. E Quantification of ratio of pTau/total Tau (WT n = 43; WT-G2-567 n = 63; p.R406W n = 51; p.R406W-G2-567 n = 37). Data are represented as mean ± SEM. Data on the y-axis are normalized to WT-DMSO. Kruskal–Wallis test followed by Dunn’s test. **p = 0.0097. A–E Cells were quantified from 3 independent experiments. F LAMP1+ vesicles (red) co-stained with total Tau (Tau5, green) in MAPT p.R406W and isogenic control neurons after treatment with G2-567. Scale bar 1 µm. G Quantification of the percentage of lysosomes free of total Tau (empty, white) or co-localized with total Tau (Tau positive, gray). Data are represented as mean ± SEM. Tukey’s multiple comparison. Empty vs. Tau positive (+): *p = 0.0355, p = 0.0498, p = 0.0125; ***p = 0.0005; ****p < 0.0001. H LAMP1+ vesicles (red) in MAPT p.R406W and isogenic control neurons after treatment with G2-567. Scale bar 1 µm. I Quantification of lysosome volume (WT n = 135; WT-G2-567 n = 158; p.R406W n = 221; p.R406W-G2-567 n = 177). Data are represented as mean ± SEM. Kruskal-Wallis test followed by Dunn’s test. *p = 0.0109; **p = 0.0036; ****p < 0.0001. F–I Data are representative of 3 independent experiments. Source data are provided as a Source Data file.
To determine whether G2-567 impacts Tau accumulation in LAMP1+ vesicles, we imaged treated neurons using Airyscan super-resolution microscopy. In G2-567-treated mutant neurons, the percentage of lysosomes free of Tau was significantly higher than in DMSO-treated mutant neurons (p.R406W-DMSO: 32.8% vs. p.R406W-G2-567: 67.1%; Fig. 7F, G). G2-567 treatment also restored the proportion of pTau-free lysosomes in MAPT p.R406W neurons to control levels, without significantly altering the localization of pTau in the lumen (p.R406W-DMSO: 27.2% vs. p.R406W-G2-567: 40.3%; Supplementary Fig. 9). LAMP1+ vesicle size in MAPT p.R406W neurons was restored to sizes observed in controls upon G2-567 treatment (Fig. 7H, I and Supplementary Fig. 10). In fact, neurons treated with G2-567 displayed a significant reduction in lysosomal size in both mutant and isogenic control neurons (Fig. 7H, I). Our findings demonstrate that treating mutant neurons with an autophagy enhancer, G2-567, rescues total Tau and pTau levels, increases Tau-free lysosomes, and reduces lysosome size.
Autophagy enhancer G2-567 increases autophagy in MAPT p.R406W neurons without rescuing motility defects
Next, we sought to determine whether treatment with G2-567 rescues the motility and autophagic defects in MAPT p.R406W neurons. G2-567 treatment did not significantly alter the number of LAMP1+ vesicles compared to vehicle treatment in mutant or control neurons (Fig. 8A, B). G2-567-treated mutant neurons also maintained a significant increase in distally positioned lysosomes compared to DMSO-treated controls (Fig. 8A, C). To determine whether G2-567 restores the expression of the dysregulated adaptor protein JIP3, we evaluated JIP3 immunofluorescence signal intensity in MAPT p.R406W and isogenic control neurons. JIP3 levels remained significantly elevated in G2-567-treated mutant neurons compared with vehicle controls (Fig. 8D, E). Thus, G2-567 treatment does not restore lysosomal positioning, lysosomal number, or motor adaptors in mutant neurons.
MAPT p.R406W and isogenic control (WT) neurons were treated with G2-567 (0.5 µM) or DMSO for 14 days beginning on DIV7 and fixed on DIV21. Neurons were then immunostained for lysosomal positioning and autophagy markers. A Immunostaining for MAP2 (green) and LAMP1 (red). Lower panel, 3D reconstruction by Imaris. Scale bar 5 µm. B Quantification of lysosome density: WT (n = 16), WT-G2-567 (n = 14), p.R406W (n = 17), p.R406W-G2-567 (n = 19). Kruskal-Wallis test followed by Dunn’s test. *p = 0.0214, p = 0.0390; **p = 0.0017. C Quantification of lysosome distance from the nucleus as the shortest distance of LAMP1+ vesicles from DAPI. Quantification performed by Imaris 3D rendering. Data on the y-axis are normalized to WT-DMSO. Number of vesicles quantified: WT (n = 181), WT-G2-567 (n = 219), p.R406W (n = 319), p.R406W-G2-567 (n = 318). Kruskal–Wallis test followed by Dunn’s test. WT-DMSO vs. p.R406W-DMSO: **p = 0.0011. p.R406W-DMSO vs. p.R406W-G2-567: p = 0.3235. D Immunostaining with MAP2+ (magenta) and JIP3 (green). Scale bar 10 µm. E Quantification of JIP3 mean intensity per cell. WT (n = 43), WT-G2-567 (n = 55), p.R406W (n = 33), p.R406W-G2-567 (n = 54) cells. Kruskal–Wallis test followed by Dunn’s test; ****p < 0.0001. F Cells were probed for autophagosomes using CYTO-ID (green) and microtubule (magenta). Right panel, magnified CYTO-ID positive vesicles (green). Scale bar 5 µm. G Quantification of CYTO-ID mean density. Total number of cells quantified: WT (n = 71), WT-G2-567 (n = 79), p.R406W (n = 79), p.R406W-G2-567 (n = 78). Kruskal–Wallis test followed by Dunn’s test, **p = 0.0027; ****p < 0.0001. H Immunostaining for MAP2 (magenta), LC3B (red) and p62 (green). Scale bar 10 µm. I Quantification of LC3B mean intensity per cell. WT (n = 70), WT-G2-567 (n = 43), p.R406W (n = 68), p.R406W-G2-567 (n = 68) cells. Kruskal-Wallis test followed by Dunn’s test, *p = 0.0393; **p = 0.0062; ***p = 0.0005; ****p < 0.0001. J Quantification of p62 mean intensity. WT (n = 26), WT-G2-567 (n = 32), p.R406W (n = 32), p.R406W-G2-567 (n = 26) cells. Kruskal–Wallis test. *p = 0.0266, ****p < 0.0001. All data are mean ± SEM. All data are normalized to WT-DMSO. Data are representative of 3 independent experiments except in panels I and J which represent 4 independent experiments. Source data are provided as a Source Data file.
To understand the impact of G2-567 on autophagy, we quantified autophagic vacuoles in mutant and control neurons after G2-567 treatment. We observed a significant increase in the CYTO-ID signal with G2-567 treatment, representing more autophagic vacuoles, in both mutant and control neurons (Fig. 8F, G). This finding is consistent with the mechanism of action of G270. Autophagosome surface marker, LC3B, was also significantly elevated in G2-567 treatment compared with DMSO treatment in both mutant and isogenic control neurons (Fig. 8H, I). Thus, G2-567 enhances autophagy in mutant and control neurons. To determine whether the enhanced autophagic vacuoles lead to efficient cargo degradation, we measured p62 levels. Treatment of MAPT p.R406W neurons with G2-567 resulted in a significant reduction of p62 (Fig. 8G, J). However, G2-567 treatment did not alter p62 levels in isogenic control neurons (Fig. 8G, J). Thus, G2-567 restored productive autophagy in mutant neurons. We replicated the same phenotype in a second donor line, further confirming the robustness of our findings (Supplementary Fig. 11). Together, we show that G2-567 rescues pTau levels and Tau accumulation in lysosomes by enhancing autophagy without rescuing motility defects.
Discussion
Here, we provide a high-dimensional visualization of endogenous Tau degradation in human neurons. We discovered key differences in the way human neurons handle total Tau and pTau in the lysosome. These differences in total Tau and pTau are exaggerated in neurons expressing MAPT p.R406W, where there is an overall defect in protein degradation. Integrating super-resolution microscopy with biochemical manipulation, we reveal a spatially specific lysosomal abnormality not previously described in human MAPT mutant neurons. We find that MAPT p.R406W neurons exhibit numerous alterations in autophagy and lysosomal pathways, which contribute to the defects in degradative capacity. Beyond total Tau and pTau, the MAPT p.R406W mutation is sufficient to disrupt p62 and neutral lipid degradation. We were able to rescue mutant Tau-induced defects in total Tau and pTau degradation by restoring autophagy without rescuing lysosomal trafficking defects using an autophagy-enhancing compound, G2-567. Together, our findings create a more complete picture of the direct impact of a Tau mutation on the degradative capacity of human neurons (Fig. 9) and provide evidence that we can decouple lysosomal trafficking and degradative capacity to therapeutically target protein clearance.
Schematic of key findings. Created in BioRender. Mirfakhar, F. (2026) https://BioRender.com/8bzn5l2.
Our study suggests that Tau degradation is driven by an interplay between neuronal trafficking defects and faulty autophagy. In lysosomal storage disorders, cells respond to inefficient degradative machinery by upregulating macroautophagy components72. We reported increased autophagic vacuole accumulation coupled with increased cargo in MAPT p.R406W neurons, which is reminiscent of lysosomal storage disorders. Importantly, we discovered that the chemical enhancement of autophagy rescued total Tau and pTau levels and lysosomal phenotypes without restoring the trafficking defects, effectively decoupling motility from degradation. Together, the targeted and pathway-level analyses reveal that MAPT mutations disrupt lysosomal and autophagy-associated networks at the transcriptomic level. These transcriptomic signatures are consistent with the functional impairments we observe, such as reduced autophagic flux, impaired lysosomal proteolysis, and pTau mislocalization, and support the model in which MAPT mutations compromise degradative capacity in human neurons.
Our finding that phosphorylated Tau accumulates at the lysosomal limiting membrane in MAPT mutant neurons highlights a previously unrecognized defect in degradative organelle handling of pathological Tau species. Although the precise molecular events that mediate pTau recruitment to the lysosomal surface remain to be defined, several lines of evidence in the literature support the plausibility of this phenomenon. Prior studies have shown that misfolded or hyperphosphorylated Tau can associate with late endosomal and lysosomal membranes under conditions of impaired autophagy or proteostatic stress and may even impair lysosomal function upon mislocalization13,73,74,75,76. Tau has also been observed within autophagic vesicles and accumulated in autolysosomes when degradation is compromised77,78. These observations collectively suggest that Tau-lysosome interactions are part of a broader, stress-responsive degradative pathway that becomes pathologically engaged in tauopathy. However, prior to our study, it was unclear how early these defects occur.
We discovered that the presence of mutant Tau (p.R406W) was sufficient to induce several changes in the localization of Tau in the lysosome. Neurons expressing mutant Tau had fewer lysosomes free of total Tau or pTau protein, suggesting that degradation of Tau is impaired. These defects were restored upon correction of the mutant allele by CRISPR/Cas9. Additionally, we discovered that pTau accumulates at the lysosomal membrane in mutant neurons. Possible mechanisms by which Tau and pTau associate with the lysosomal membrane include interacting with membrane-associated chaperones, such as Hsc70, involved in chaperone-mediated autophagy; partial unfolding of Tau leading to transient membrane binding prior to lysosomal import; and post-translational modifications that may modulate membrane affinity. While phosphorylation of Tau is required for normal function, hyperphosphorylation is a key feature of Tau aggregates. Thus, as has been suggested for acetylation of Tau, pTau may directly impair autophagic machinery early in a manner that ultimately promotes Tau aggregation13,29.
Our data extend this conceptual framework by identifying a mutation-specific defect in lysosomal handling of pTau in human iPSC-derived neurons. The enrichment of pTau on the lysosomal surface suggests disruption at a specific step of the degradative pathway, such as impaired cargo translocation into the lumen, stalled autophagosome-lysosome fusion, altered lysosomal membrane remodeling, or compromised luminal proteolysis. Importantly, the rescue of this phenotype by enhancing autophagic-lysosomal capacity provides causal evidence that insufficient lysosomal degradation is a key upstream driver of pTau membrane recruitment. Thus, the combination of spatially resolved imaging, pathway-specific functional assays, and pharmacologic rescue supports a mechanistic model in which MAPT mutations impair degradative processing of pTau, leading to its accumulation at the lysosomal limiting membrane. Future studies will be required to identify the specific pathways that mediate pTau recruitment and retention at lysosomal membranes. These may include testing the roles of selective autophagy adaptors, endosomal-lysosomal fusion machinery, and proteolytic competence using targeted perturbations or organelle-resolved proteomics. Nonetheless, by defining a reproducible, mutation-associated defect in lysosomal pTau distribution and demonstrating its reversible nature, the present study establishes a foundational mechanistic framework that can be refined and expanded through deeper molecular investigation.
Precise regulation of lysosome localization and motility is essential for maximal degradative capacity in neurons18,20,23. Neurons show spatially restricted autophagosome biogenesis, which is mostly enriched at distal ends of axons in both growing and synaptically connected neurons79. Distal lysosomes are largely devoid of degradative enzymes and need to undergo retrograde transport to the neuronal soma for effective cargo degradation35,52,80. Late endosomes predominantly undergo long-range retrograde transport while simultaneously acidifying and maturing into lysosomes and ultimately fusing with mature lysosomes enriched in the cell body81. For efficient degradation, endolysosomal vesicles must reach the soma, where the vesicles fuse with lysosomes17,82. The expression of JIP3 has been shown to influence lysosomal distribution within neurons22,56,58,59. JIP3 silencing has been shown to reduce lysosome association with dynein motors, leading to failure in retrograde trafficking56,58,59. Loss-of-function mutants in JIP3 display lysosome accumulations in axonal terminal swellings and an increase in pTau protein58. Overexpression of GFP-JIP3 leads to JIP3 enrichment on lysosomes and the formation of lysosome-filled axonal swellings. This suggests that lysosome trafficking is highly sensitive to JIP3 levels. We report that MAPT p.R406W causes impairment in lysosomal transport that is mediated by the adaptor molecule JIP3. The increased JIP3 levels in mutant neurons may contribute to the slowing of lysosomal motility by disrupting the delicate balance required for proper lysosome transport. In addition to elevated mRNA and protein levels, JIP3 staining appears denser and more punctate in mutant neurons, which could mirror lysosome-filled axonal swellings observed in GFP-JIP3 neurons. Thus, the observed accumulation of pTau in MAPT p.R406W neurons may be driven by defects in lysosomal trafficking due to faulty motor-associated proteins, which present a potential therapeutic target. Rescuing JIP3 levels is challenging, asprior studies illustrate that JIP3 overexpression also induces lysosomal and axonal defects59. However, our finding that G2-567 treatment rescues Tau and pTau levels without restoring JIP3 levels suggests that JIP3 is upstream of autophagy defects or is not the primary contributor to Tau clearance defects in these cells.
Alterations in intracellular lipid content, such as in metabolic disorders, can significantly affect the fusion step of macroautophagy and influence the overall activity of this intracellular proteolytic pathway83. Membrane lipids are broken down into fatty acids that can serve as a source of energy production or be packaged into lipid droplets, acting as reservoirs for stored lipids destined for future utilization. It is common for patients with neurodegenerative disorders, including FTD, to have abnormal levels of lipid droplets in their nerve cells84. It has also been demonstrated that upregulated lipid synthesis and downregulated lipid turnover can lead to pathological lipid accumulation in both neurons and glia in multiple tauopathy model systems. Here, we showed that MAPT p.R406W neurons have a greater lipid droplet content compared to the control neurons, indicating a more general neuronal metabolic dysregulation.
Derivatives of glibenclamide, including a series of “G2” small molecules, have been reported to increase autophagy in an mTOR-independent manner without altering insulin secretion71. G2-567 is a derivative of G2-115, which has been well characterized69,70. Here, we used G2-567 as an experimental probe to test the responsiveness of an autophagy enhancer on Tau clearance. G2 is an autophagy enhancer that reduces the interaction between RCAN1 and calcineurin. This stimulates calcineurin activity and promotes the translocation of transcription factor EB (TFEB) into the nucleus, where it regulates expression of genes associated with lysosomal biogenesis69,70. G2-115 has been shown to increase autophagosomes, autolysosomes, and LC3; reduce p62; and reduce levels of α1-antitrypsin Z (ATZ) and Huntingtin proteins in C. elegans models, directly converted human neurons, and animal models70,71. Determining the precise molecular target and binding mechanism of G2-567 is an important question and will require additional biochemical, biophysical, and structural studies. Here, we found that enhancing autophagy with an analog of G2-115, G2-567, was sufficient to reduce total Tau and pTau levels and to increase the proportion of lysosomes free of Tau in mutant neurons. Calcineurin is a Ca²⁺/calmodulin-dependent phosphatase that dephosphorylates specific phospho-epitopes on Tau85,86. If calcineurin activity were the primary driver of the observed effect, we would expect a decrease in pTau levels without a corresponding change in total Tau, as calcineurin-mediated dephosphorylation alters phosphorylation state but does not affect Tau abundance. In contrast, our data show that G2-567 treatment leads to a reduction in both pTau and total Tau protein levels. This pattern is consistent with enhanced Tau clearance, rather than selective dephosphorylation. Therefore, the decrease in pTau observed upon G2-567 treatment cannot be explained solely by increased calcineurin activity and instead supports a model in which G2-567 promotes increased Tau degradation. In addition to reducing Tau levels, G2-567 treatment reduces lysosomal size and restores lysosomal morphology. G2-567 may act to promote Tau clearance by upstream or indirect signaling mechanisms that remain to be resolved. Together, we demonstrate that G2-567 and its derivatives promote broad protein degradation in vitro and in vivo.
A potential limitation of pharmacologic enhancement of autophagy is the possibility of off-target effects, including unintended modulation of the ubiquitin–proteasome system. However, multiple lines of evidence support an autophagy-lysosomal mechanism for pTau reduction in MAPT mutant neurons in this study. First, the pathological phenotype we identify, namely enrichment of pTau at the lysosomal limiting membrane, is intrinsically linked to lysosomal function and cannot be corrected by proteasomal activity, which does not act on membrane-associated or detergent-insoluble Tau species. Second, the biochemical properties of the affected Tau species are consistent with substrates known to rely preferentially on autophagy for clearance, rather than the proteasome77,87,88. Third, treatment with the autophagy activator produces changes in LC3B-II, p62, and lysosomal proteolysis that align closely with enhanced autophagic flux, and lysosomal pTau mislocalization is blocked by lysosomal inhibition in our system. These findings collectively support the conclusion that the restoration of lysosomal degradative capacity, rather than proteasomal activation, underlies the normalization of pTau localization.
Overall, our findings suggest that the MAPT p.R406W mutation impacts autophagy-lysosomal function in a manner that contributes to altered Tau homeostasis. In MAPT p.R406W neurons, we observed elevations in both total Tau and pTau, an increased pTau/total Tau ratio, as well as enlarged LAMP1+ lysosomes and impaired lysosomal trafficking dynamics. These results support a model in which lysosomal dysfunction precedes and contributes to Tau accumulation, rather than Tau accumulation being solely a downstream consequence of altered kinase or phosphatase activity. Treatment with G2-567 reduced both total Tau and pTau levels and restored lysosomal morphology, consistent with enhanced Tau degradation through improved autophagy-lysosomal processing. Given that the MAPT p.R406W mutation affects axonal transport and vesicle dynamics, it is plausible that altered trafficking contributes to both the autophagy phenotype and the increased pTau/total Tau ratio. Future studies disentangling these relationships will be valuable in defining the causal sequence linking MAPT p.R406W, autophagy dysfunction, and Tau accumulation. More than 50 mutations occur in the MAPT gene and cause FTLD-Tau89; therefore, the extent to which these phenotypes are shared across different Tau mutations must be investigated in the future.
Methods
The research described in this study complies with all relevant ethical regulations and was approved by the Washington University School of Medicine (IRB 201104178 and 201306108).
iPSC lines
iPSCs from a MAPT p.R406W carrier (F11362.1 or FA14530.1) and the CRISPR/Cas9 edited isogenic control (F11362.1Δ1B06 or FA14530.1Δ3 G12) were previously described90. To facilitate the generation of excitatory neuronal cultures, a Tet-ON 3G-controlled NGN2 transgene was integrated into the AAVS1 locus of the iPSC lines described above using TALENs32,91: MAPT p.R406W donor line (iPSC F11362.1.CRISPRiΔ5A01.NGN2Δ1B08 and iPSC FA14-530.1.NGN2Δ1C05) and isogenic control (iPSC F11362.1Δ1C11.CRISPRiΔ2D10.NGN2Δ1C07 and iPSC FA14-530.1Δ3G12.NGN2Δ1E04). Engineered cell lines were verified using Sanger sequencing for the MAPT mutation and the NGN2 cassette, karyotyping for chromosomal stability, and immunocytochemistry for pluripotency markers (Supplementary Figs. 1 and 2). Cell lines were maintained in mTesR medium (STEMCell Technologies, 85850) on Cultrex PathClear Basement Membrane Extract, Type II (R&D Systems, 3532-010-02). The cell lines were confirmed to be free of mycoplasma.
iPSC differentiation
MAPT p.R406W and isogenic WT control iPSCs with the integrated NGN2 transgene were differentiated into neural progenitor cells (NPCs)90. Briefly, iPSCs were plated at a density of 65,000 cells per well in Neural Induction Media (STEMdiff SMADi Neural Induction Kit, STEMCell Technologies, 08581) in a 96-well v-bottom plate to form neural aggregates. After 5 days, neural aggregates were passaged onto Poly-L-Ornithine (PLO, R&D Systems, 3436-100-01) and laminin-coated (R&D Systems, 3400-010-03) plates to form neural rosettes. Neural rosettes were isolated after 5 days in culture by selection using STEMdiff Neural Rosette Selection Reagent (STEMCell Technologies, 05832). The resulting NPCs were cultured on PLO and laminin-coated plates, and terminal differentiation was initiated with the addition of cortical maturation medium (BrainPhys Neuronal Medium, STEMCell Technologies, 05790) supplemented with B27 (Gibco, 17504001), BDNF (Peprotech, 450-02-100UG) (10 ng/mL), GDNF (Peprotech, 450-10-100UG) (10 ng/mL), cAMP (Sigma, D0627-1G), GlutaMax (Thermo Scientific, 35050061), N2 supplement (Gibco, 17502001), and doxycycline (Sigma, D5207-10G) (2 mg/mL). Neurons typically show neuronal identity 7 days after plating based on immunocytochemistry for β-tubulin III (Tuj1) and microtubule-associated protein 2 (MAP2), a marker of mature neuronal dendrite (Supplementary Fig. 3). Neurons were analyzed at 14 or 21 days after doxycycline induction. Experiments were performed in at least two independent differentiations (see Figure legends for experiment-specific details).
Antibodies
Primary antibodies used for immunostaining included: anti-MAP2 (Abcam, ab5392), anti-LAMP1 (D2D11; Cell signaling, 9091), anti-LAMP1 (D4O1S; Cell signaling, 15665), anti-Tau5 (total Tau; generously provided by Dr. Lester Binder), anti-AT180 (pTau-Thr23; Thermo Fisher Scientific, MN1040), anti-AT8 (pTau-Ser202, Thr 205; Thermo Scientific, MN1020B), anti-JIP3 (Thermo Fisher, PA5-59728), anti-LC3B (Cell Signaling, 2775), anti-SQSTM1 / p62 (Cell signaling, D5L7G 88588S), anti-Synapsin I (Millipore, AB1543), anti-Cathepsin D (C-5; SantaCruz, sc-377124), anti-Limp2 (Thermo Fisher Scientific, 712072), anti-Beta III Tubulin (Tuj1; Promega, G712A), and DAPI (Sigma, D-9542). The secondary antibodies included: Alexa-488 Goat anti-mouse IgG (H + L; Thermos Scientific, A-11001), Alexa-488 Goat anti-rabbit IgG (H + L; Thermos Scientific, A-11008), Alexa-568 Goat anti-mouse IgG (H + L; Thermos Scientific, A-11004), Alexa-568 Goat anti-rabbit IgG (H + L; Thermos Scientific, A-11011), Alexa-647 Goat anti-mouse IgG (H + L; Thermos Scientific A-32728), Alexa-647 Goat anti-rabbit IgG (H + L; Thermos Scientific A-32733), Alexa Fluor™ Plus 647 Goat anti-Chicken IgY (Thermos Scientific A-32933), and Alexa-405 Goat anti-chicken IgY (Abcam, ab175674).
Immunoblotting
iPSC-derived neurons were lysed in RIPA buffer (50 mM Tris, 150 mM NaCl, 1% SDS, 1% Triton X-100) supplemented with phosphatase and protease inhibitors. Lysates were mixed with 4× SDS sample buffer (Bio-Rad) containing 10% β-mercaptoethanol, heated at 95 °C for 5 min, and then separated on a 4–12% Bis-Tris gel (NuPAGE). Proteins were transferred onto PVDF membranes (Immobilon), and the membrane was subsequently blocked for 1 h at room temperature in 3% BSA in phosphate-buffered saline with 0.1% Tween-20 (PBST). Membranes were incubated overnight at 4 °C with primary antibodies. After washing, membranes were incubated with horseradish peroxidase-conjugated secondary antibodies for 1 h at room temperature. Signals were detected using SuperSignal™ West Pico PLUS Chemiluminescent Substrate (Thermo Scientific), and images were captured using a ChemiDoc Imaging System (Bio-Rad). Blots shown are representative of three independent experiments performed in duplicate, with actin used as a loading control.
Immunostaining
iPSC-derived neurons plated on coverslips were washed and fixed with 4% paraformaldehyde (Electron Microscopy Sciences, 15710) for 20 min at room temperature (RT) and permeabilized with 0.3% Triton X-100 in PBS for 5 min at room temperature (RT). Cells were blocked with 2% FBS, 1% BSA, and 0.1% saponin in PBS and incubated with primary antibodies at 4 °C overnight. After washing in PBS, neurons were incubated with secondary antibodies for 1 h at RT. Cells were incubated with DAPI (Sigma, D-9542) for 5 min, then washed with PBS. Coverslips were then mounted using Fluoromount-G (Southern Biotechnology, 00-4958-02). Confocal microscopy was performed with an LSM 980 confocal microscope (AiryScan, Zeiss). Each image reflects a Z-stack of 5–12 images, each taken at 0.2 μm depth intervals. Samples were imaged using identical acquisition parameters for direct comparison.
Monitoring acidic vesicle motility in live cells
To evaluate the motility of acidic vesicles, iPSC-derived neurons were cultured in live-imaging dishes pre-coated with poly-L-ornithine and laminin (35 mm glass-bottom dish #1.5H, Cellvis, D35-20-1.5H) at a density of 100,000-150,000 cells/dish. Neurons were probed for microtubules by ViaFluor® dye (Viafluor® 647, Biotium, 70063, 1× in media, 30-min incubation) to define cell boundaries. Cells were then incubated with 50 nM LysoTracker Red D99 (Invitrogen, L7528) for 10 min. Super-resolution confocal microscopy (Zeiss LSM 980 Multiphoton Airyscan 2; Carl Zeiss Ltd, Cambridge, UK) was used with a 40x or 63x objective lenses to image neurons or record and track LysoTracker-positive vesicles in time-lapse mode for up to 5 min. Each assay was repeated at least 3 times from independent cultures. Only modest bleaching was observed during the experiment. Cells showing signs of damage (pearling or blebbing) were discarded. Lysosomal motility/trafficking was measured in a semi-automatic manner using Imaris (Bitplane) by tracking LysoTracker-positive puncta over time, using the 3D time-lapse tracing module. The lysosomal tracks in neurites were visualized in order to quantify acidic vesicle velocity and vesicle displacement length. Acidic vesicles within the field were detected using the spot function, and tracks were semi-automatically created using the Brownian motion algorithm in Imaris.
Monitoring autophagy in live cells
For autophagy assays, iPSC-derived neurons were cultured in live-imaging dishes pre-coated with poly-L-ornithine and laminin. Rapamycin (Rap, 500 nM) and Chloroquine (CQ, 20uM) treatments were used as a positive control of autophagosome accumulation (treatment: 16 h). Neurons were incubated with CYTO-ID (1:1000; Enzo 175-0050) for 20 min based on the manufacturer’s recommendations, and live imaging was performed at 37 °C in a humidified and CO2-controlled chamber. Neurons were also probed for microtubules by ViaFluor® dye (Viafluor® 647, Biotium, 70063) to define cell boundaries. Images were acquired by a super-resolution confocal microscope (Zeiss LSM 980 Airyscan). CYTO-ID selectively probes the autophagosome membrane, and vesicles appear as round-shaped structures detectable under fluorescence excitation/emission of 495/519 nm (Supplementary Fig. 8A). Density and intensity of CYTO-ID-positive autophagic vacuoles were quantified within the neurons defined by microtubule signal using Fiji ImageJ.
Monitoring Lipids in Live Cells
Levels of intracellular neutral lipids were quantified by adding BODIPYTM FL C16 (Thermo Scientific D3821, final concentration 10 μg/ml in PBS) for 20 min at RT to iPSC-derived neurons. Live imaging was then immediately performed at 37 °C in a humidified and CO2-controlled chamber. Neurons were also probed for microtubules by ViaFluor® dye (Viafluor® 647, Biotium, 70063) to define cell boundaries. Images were acquired using a super-resolution confocal microscope (Zeiss LSM 980 Airyscan). BODIPYTM FL C16 probes neural lipid storage compartments, detectable under fluorescence excitation/emission of 505/512 nm. BODIPY intensity was measured within the microtubule signal using Fiji ImageJ.
Image quantification of total Tau/pTau localization in lysosomes
Total Tau (Tau5) and pTau (AT8) localization was quantified in LAMP1-positive vesicles using Fiji ImageJ. The signal intensity (Gray Value) of total Tau or pTau within each LAMP1-positive vesicle was measured. A single line spanning the LAMP1-positive vesicle was used to define the plane for quantification of total Tau or pTau signal intensity. Intensity curves were then generated from the selected pixels. Total Tau and pTau intensity measurements were performed using Fiji, as the total mean fluorescence intensity per cell using structural signals from MAP2 as the cell boundary. Measurements were corrected for background fluorescence, resulting in the Corrected Total Cell Fluorescence (CTCF) values.
Image quantification of lysosome characterization
Lysosome characterization analysis was performed using LAMP1 antibody (D2D11; Cell Signaling, 9091) on 60x confocal images acquired in Z-stacks. 3D reconstruction of lysosomes was performed in Imaris to capture lysosome count, volume, and distance from soma by first detecting the LAMP1+ puncta using the Imaris spot detection module. We then measured the spot density and volume, as well as the distance of the detected spot (lysosome) to the nucleus, as a measurement for lysosomal distribution.
Image quantification of neuronal morphology
Neuronal morphometric analysis was performed on 40x confocal images acquired in Z-stacks. 3D reconstruction was performed in Imaris software using the filament module and was semi-automatically analyzed for dendritic arborization and length as well as Sholl analysis (Imaris Sholl Analysis XTension, Sholl sphere resolution 1 μm).
Fluorescence recovery after photobleaching (FRAP)
FRAP experiments were performed using the FRAP module on the confocal microscope (Zen software). Neurons (DIV14) were incubated with the microtubule probe Viafluor® 488 (Biotium, final concentration: 1μM) for 15 min at RT. Neurons were then imaged for 20 seconds (pre-bleach phase). A circular region of interest (ROI) was selected for photobleaching along the neurites. Photobleaching was performed at 50% laser power and 50 iterations. Images were acquired at 2 s intervals over a period of 5 min following the recovery of the fluorescence intensity in the selected ROI. FRAP recovery curves were generated using Fiji with normalization to the initial pre-bleach value of fluorescence intensity.
pHluorin-mKate2-LC3 transduction and quantification
pHLuorin-mkate3-LC3 FUGW-PK-hLC3 was acquired from Addgene (Addgene plasmid #61460)68. Neurons were transduced on DIV7 with lentiviral particles at a final titer of 4.2 × 10⁷ IU/mL. Following overnight incubation, cells were then imaged live using confocal microscopy (Zeiss LSM 980 Multiphoton Airyscan 2; Carl Zeiss Ltd, Cambridge, UK). For quantification of puncta size, the area covered by mKate2 and pHLuorin signal was measured. Fluorescence intensity ratios of mKate2 to pHluorin within defined ROIs were quantified and normalized. For autophagosome (that is, Red+Green+ puncta) vs autolysosome (that is, Red+ Green− puncta), compartments were quantified in both MAPT WT and p.R406W cells. Cells with no transduction were excluded from analysis.
G2 treatment
The small molecule G2 was identified by a phenotypic screen in a C.elegans model of misfolded α1-antitrypsin Z variant (ATZ)92 and was later shown to mediate autophagic degradation of ATZ in mammalian cell models71. This analog has been subjected to chemical modification to derive the G2-567 compound. MAPT p.R406W and isogenic control iPSC-neurons were treated with 0.5 µM G2-567 (resuspended in DMSO) on DIV7, a time point where neurons are positive for neuronal markers MAP2 and Tuj1 (Supplementary Fig. 3). Neurons were treated every 2 days for 14 days. Neurons were then fixed on DIV21 for immunofluorescent staining or live imaging as described above.
Immuno-electron microscopy
Neurons were fixed in 4% paraformaldehyde (PFA) in PHEM buffer (60 mM PIPES, 25 mM HEPES, 10 mM EGTA, 2 mM MgCl₂, pH 7.4) for 1 h at room temperature and then quenched in 20 mM glycine/PHEM. Cells were permeabilized with 0.05% Triton X-100 in PHEM buffer for 5 min on ice, followed by blocking in block buffer (1% BSA/0.1% Ac-BSA in PHEM) for 45 min at room temperature. Cells were incubated with primary antibody (AT180, Thermo Fisher Scientific, MN1040) in blocking buffer for 1 h, followed by incubation with 1.4 nm Nanogold-conjugated secondary antibody (1:200) (#2001-0.5 ML, Nanoprobes, Inc.) for 45 min at room temperature. After washing, cells were fixed in a mixture of 2% PFA and 2% glutaraldehyde in 0.1 M cacodylate buffer for 30 min at room temperature. Cells were then washed and quenched in 20 mM glycine/PHEM and washed again in distilled water. The gold-enhance mixture was prepared following Nanoprobes directions (GoldEnhance EM kit, #2113-8 ml) and incubated in the gold-enhance solution for 7 min. The reaction was stopped by washing in DI water (5× 1 min), followed by 0.1 M cacodylate buffer (3× 5 min), and stained in 1% osmium tetroxide/1.5% potassium ferricyanide for 1 h in the dark on a shaker. Cells were rinsed in DI water (4× 10 min), incubated overnight in 2% aqueous uranyl acetate at 4 °C, washed in DI water (3 ×10 min), dehydrated in a graded ethanol series, embedded in epon resin (LX 112, EMS), and cured at 60 °C for 48 h. Sections 70 nm thick were cut using a Leica UC7 ultramicrotome, transferred to carbon-coated grids, and imaged with a JEOL JEM-1400PLUS TEM operated at 120 kV.
Tau SILK in iPSC-derived neurons
iPSC-derived neurons were cultured for four weeks prior to labeling as described above39. After four weeks in culture, 13C6-labeled leucine was added to the cortical maturation media at 105 mg/L to achieve 50% mol (100% TTR) labeled media. The iPSC-derived neurons were cultured in labeled media for 15 days. Cell pellets were collected every three days and solubilized in 1% NP-40 and centrifuged to remove the insoluble fraction.
Tau proteoforms were analyzed as previously described with minor modifications39. Samples were spiked with full-length 13C/15N labeled 2N4R and 15N labeled 0N3R Tau internal standards for absolute quantitation and were immunoprecipitated with a cocktail of Tau1 (mid-domain) and HJ8.5/HJ8.7 (N-terminus) Tau antibodies. After washing, samples were digested with trypsin (Promega), desalted using C18 Toptip (Glygen), and analyzed on an Eclipse Orbitrap mass spectrometer (Thermo Scientific). Tryptic peptides were normalized with 13C/15N labeled 2N4R Tau internal standards for absolute quantitation. 13C6-Leucine labeled and unlabeled tryptic peptides were quantified, and the ratio of labeled to unlabeled peptide was calculated as tracer-to-tracee ratio (TTR) for SILK analyses.
RNAseq
To evaluate the impact of MAPT p.R406W on molecular pathways, we analyzed publicly available RNAseq data from MAPT p.R406W and isogenic control iPSC-derived neurons41. Briefly, DNA libraries of individual samples were constructed using the TruSeq Stranded Total RNA Sample Prep with Ribo-Zero Gold kit (Illumina) and then sequenced by Illumina HiSeq 4000 Systems Technology with a read length of 1x150bp, and an average library size (mapped reads) of 43.2 ± 9.5 million reads per sample. The average percentage of mapped reads to GRCh38, 94.0% ± 2.0. STAR (v.2.6.0)93,94, was used to align the newly generated RNA sequences to the human reference genome, GRCh38.p13 (hg38). Salmon (v. 0.11.3)95 was used to quantify the expression of the genes annotated within the human reference genome used in this project (GRCh38.p13). Protein-coding genes were selected for further analyses. Differential gene expression analysis between samples carrying the mutation and controls was performed using the DESeq2 (v.1.22.2) package96. Pathway enrichment was performed using ToppGene. For single gene analysis, genes involved in autophagy-lysosomal pathways were selected from Bordi et al.97, which defined a gene toolbox for monitoring autophagy transcription.
Statistics and reproducibility
Experiments were conducted using NGN2-induced neurons generated from two to five independent differentiations to ensure biological reproducibility. Although NGN2-induced neuronal differentiation may introduce some variability across differentiation batches, our protocol consistently generates highly pure and homogeneous neuronal cultures within three days. The reproducibility and uniformity of differentiation are demonstrated in Supplementary Fig. 3. Each biological replicate (n) corresponds to neurons derived from a distinct differentiation batch. Within a batch, cells were plated in multiple wells (technical replicates), and data from these wells were pooled and treated as a single n for statistical analysis. Cases where n represents the number of vesicles or lysosomes analyzed are explicitly stated in the corresponding figure legend. Statistical analyses were performed using GraphPad Prism. For normally distributed data, parametric tests, such as unpaired t-tests or ANOVA with post hoc Tukey’s correction, were used based on the comparisons being made. If data did not pass normality tests, we used non-parametric statistical analyses (e.g., Mann-Whitney U test for two-group comparisons and Kruskal-Wallis test with post hoc Dunn’s correction for multiple comparisons). A significance threshold of p < 0.05 was considered statistically significant. Data are presented as mean ± SEM. Data acquisition and analyses were performed in a blinded and automated or semi-automated manner using Imaris and ImageJ software.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files. Source data are provided with this paper.
References
Jiang, S. & Bhaskar, K. Degradation and transmission of Tau by autophagic-endolysosomal networks and potential therapeutic targets for tauopathy. Front. Mol. Neurosci. 13, 586731 (2020).
Papanikolopoulou, K. & Skoulakis, E. M. C. Altered proteostasis in neurodegenerative tauopathies. Adv. Exp. Med. Biol. 1233, 177–194 (2020).
Tang, M., Harrison, J., Deaton, C. A. & Johnson, G. V. W. Tau clearance mechanisms. Adv. Exp. Med. Biol. 1184, 57–68 (2019).
Bevan-Jones, W. R. et al. Neuroinflammation and protein aggregation co-localize across the frontotemporal dementia spectrum. Brain 143, 1010–1026 (2020).
Goedert, M. et al. Tau protein and frontotemporal dementias. Adv. Exp. Med. Biol. 1281, 177–199 (2021).
Barbier, P. et al. Role of Tau as a microtubule-associated protein: structural and functional aspects. Front. Aging Neurosci. 11, 204 (2019).
Cabrales Fontela, Y. et al. Multivalent cross-linking of actin filaments and microtubules through the microtubule-associated protein Tau. Nat. Commun. 8, 1981 (2017).
Ando, K. et al. de novo MAPT mutation G335A causes severe brain atrophy, 3R and 4R PHF-tau pathology and early onset frontotemporal dementia. Acta Neuropathol. Commun. 8, 94 (2020).
Shafei, R. et al. Two pathologically confirmed cases of novel mutations in the MAPT gene causing frontotemporal dementia. Neurobiol. Aging 87, 141 e115–141.e120 (2020).
Strang, K. H., Golde, T. E. & Giasson, B. I. MAPT mutations, tauopathy, and mechanisms of neurodegeneration. Lab Invest. 99, 912–928 (2019).
Nam, K. H. & Ordureau, A. How does the neuronal proteostasis network react to cellular cues? Biochem. Soc. Trans. 52, 581–592 (2024).
Bourdenx, M., Gavathiotis, E. & Cuervo, A. M. Chaperone-mediated autophagy: a gatekeeper of neuronal proteostasis. Autophagy 17, 2040–2042 (2021).
Caballero, B. et al. Interplay of pathogenic forms of human tau with different autophagic pathways. Aging Cell 17, e12692 (2018).
Gorantla, N. V. & Chinnathambi, S. Autophagic pathways to clear the tau aggregates in Alzheimer’s disease. Cell Mol. Neurobiol. 41, 1175–1181 (2021).
Hamano, T. et al. Autophagy and Tau protein. Int. J. Mol. Sci. 22, 7475 (2021).
Nixon, R. A. & Rubinsztein D. C. Mechanisms of autophagy-lysosome dysfunction in neurodegenerative diseases. Nat. Rev. Mol. Cell Biol. 25, 926–946 (2024).
Bejarano, E. et al. Defective recruitment of motor proteins to autophagic compartments contributes to autophagic failure in aging. Aging Cell 17, e12777 (2018).
Cabukusta, B. & Neefjes, J. Mechanisms of lysosomal positioning and movement. Traffic 19, 761–769 (2018).
Ferguson, S. M. Neuronal lysosomes. Neurosci. Lett. 697, 1–9 (2019).
Oyarzun, J. E. et al. Lysosome motility and distribution: Relevance in health and disease. Biochim. Biophys. Acta Mol. Basis Dis. 1865, 1076–1087 (2019).
Roney, J. C., Cheng, X. T. & Sheng, Z. H. Neuronal endolysosomal transport and lysosomal functionality in maintaining axonostasis. J. Cell Biol. 221, e202111077 (2022).
Cason, S. E. et al. Sequential dynein effectors regulate axonal autophagosome motility in a maturation-dependent pathway. J. Cell Biol. 220, e202010179 (2021).
Lie, P. P. Y. & Nixon, R. A. Lysosome trafficking and signaling in health and neurodegenerative diseases. Neurobiol. Dis. 122, 94–105 (2019).
Reed, L. A. et al. Autosomal dominant dementia with widespread neurofibrillary tangles. Ann. Neurol. 42, 564–572 (1997).
Qi, C. et al. Tau filaments with the Alzheimer fold in human MAPT mutants V337M and R406W. Nat. Struct. Mol. Biol. 32, 1297–1304 (2025).
Alquezar, C., Arya, S. & Kao, A. W. Tau post-translational modifications: dynamic transformers of tau function, degradation, and aggregation. Front. Neurol. 11, 595532 (2020).
Ye, H., Han, Y., Li, P., Su, Z. & Huang, Y. The role of post-translational modifications on the structure and function of Tau protein. J. Mol. Neurosci. 72, 1557–1571 (2022).
Alonso, A. D. et al. Hyperphosphorylation of Tau associates with changes in its function beyond microtubule stability. Front. Cell Neurosci. 12, 338 (2018).
Liu, M. et al. Hyperphosphorylation renders Tau prone to aggregate and to cause cell death. Mol. Neurobiol. 57, 4704–4719 (2020).
Naseri, N. N., Wang, H., Guo, J., Sharma, M. & Luo, W. The complexity of tau in Alzheimer’s disease. Neurosci. Lett. 705, 183–194 (2019).
Silva, M. C. et al. Prolonged tau clearance and stress vulnerability rescue by pharmacological activation of autophagy in tauopathy neurons. Nat. Commun. 11, 3258 (2020).
Wang, C. et al. Scalable production of iPSC-derived human neurons to identify Tau-lowering compounds by high-content screening. Stem Cell Rep. 9, 1221–1233 (2017).
Barral, D. C. et al. Current methods to analyze lysosome morphology, positioning, motility and function. Traffic 23, 238–269 (2022).
Heybrock, S. et al. Lysosomal integral membrane protein-2 (LIMP-2/SCARB2) is involved in lysosomal cholesterol export. Nat. Commun. 10, 3521 (2019).
Cheng, X. T. et al. Characterization of LAMP1-labeled nondegradative lysosomal and endocytic compartments in neurons. J. Cell Biol. 217, 3127–3139 (2018).
Lie, P. P. Y. et al. Post-Golgi carriers, not lysosomes, confer lysosomal properties to pre-degradative organelles in normal and dystrophic axons. Cell Rep. 35, 109034 (2021).
Kuhn, R., Mahajan, A., Canoll, P. & Hargus, G. Human induced pluripotent stem cell models of frontotemporal dementia with Tau pathology. Front. Cell Dev. Biol. 9, 766773 (2021).
Motoi, Y., Hanger, D. P. & Hasegawa, M. Editorial: Tau propagation mechanisms: cell models, animal models, and beyond. Front Neurosci. 14, 456 (2020).
Sato, C. et al. Tau kinetics in neurons and the human central nervous system. Neuron 97, 1284–1298.e1287 (2018).
Guo, J. L. et al. Unique pathological tau conformers from Alzheimer’s brains transmit tau pathology in nontransgenic mice. J. Exp. Med. 213, 2635–2654 (2016).
Minaya, M. A. et al. Conserved gene signatures shared among MAPT mutations reveal defects in calcium signaling. Front. Mol. Biosci. 10, 1051494 (2023).
Piccio, L. et al. Identification of soluble TREM-2 in the cerebrospinal fluid and its association with multiple sclerosis and CNS inflammation. Brain 131, 3081–3091 (2008).
Bourdenx, M. et al. Chaperone-mediated autophagy prevents collapse of the neuronal metastable proteome. Cell 184, 2696–2714.e2625 (2021).
Klionsky, D. J. et al. Autophagy in major human diseases. EMBO J. 40, e108863 (2021).
Fleming, A. et al. The different autophagy degradation pathways and neurodegeneration. Neuron 110, 935–966 (2022).
Ichimiya, T. et al. Autophagy and autophagy-related diseases: a review. Int. J. Mol. Sci. 21, 8974 (2020).
Saha, S., Panigrahi, D. P., Patil, S. & Bhutia, S. K. Autophagy in health and disease: a comprehensive review. Biomed. Pharmacother. 104, 485–495 (2018).
Mahapatra, K. K. et al. The lysosome as an imperative regulator of autophagy and cell death. Cell Mol. Life Sci. 78, 7435–7449 (2021).
Ren, H. & Wang, G. Autophagy and lysosome storage disorders. Adv. Exp. Med Biol. 1207, 87–102 (2020).
de Araujo, M. E. G., Liebscher, G., Hess, M. W. & Huber, L. A. Lysosomal size matters. Traffic 21, 60–75 (2020).
Toledano-Zaragoza, A. & Ledesma, M. D. Addressing neurodegeneration in lysosomal storage disorders: Advances in Niemann Pick diseases. Neuropharmacology 171, 107851 (2020).
Farfel-Becker, T. et al. Neuronal soma-derived degradative lysosomes are continuously delivered to distal axons to maintain local degradation capacity. Cell Rep. 28, 51–64 e54 (2019).
Sasazawa, Y., Hattori, N. & Saiki, S. JNK-interacting protein 4 is a central molecule for lysosomal retrograde trafficking. Bioessays 45, e2300052 (2023).
Chang, C. W., Evans, M. D., Yu, X., Yu, G. Q. & Mucke, L. Tau reduction affects excitatory and inhibitory neurons differently, reduces excitation/inhibition ratios, and counteracts network hypersynchrony. Cell Rep. 37, 109855 (2021).
Chang, C. W., Shao, E. & Mucke, L. Tau: Enabler of diverse brain disorders and target of rapidly evolving therapeutic strategies. Science 371, eabb8255 (2021).
Cason, S. E. & Holzbaur, E. L. F. Axonal transport of autophagosomes is regulated by dynein activators JIP3/JIP4 and ARF/RAB GTPases. J. Cell Biol. 222, e202301084 (2023).
Willett, R. et al. TFEB regulates lysosomal positioning by modulating TMEM55B expression and JIP4 recruitment to lysosomes. Nat. Commun. 8, 1580 (2017).
Gowrishankar, S. et al. Overlapping roles of JIP3 and JIP4 in promoting axonal transport of lysosomes in human iPSC-derived neurons. Mol. Biol. Cell 32, 1094–1103 (2021).
Gowrishankar, S., Wu, Y. & Ferguson, S. M. Impaired JIP3-dependent axonal lysosome transport promotes amyloid plaque pathology. J. Cell Biol. 216, 3291–3305 (2017).
Yang, Z. & Klionsky, D. J. Mammalian autophagy: core molecular machinery and signaling regulation. Curr. Opin. Cell Biol. 22, 124–131 (2010).
Masaki, N. et al. Targeting AUtophagy with the Synergistic Combination of Chloroquine and Rapamycin as A Novel Effective Treatment for Well-differentiated Liposarcoma. Cancer Genom. Proteom. 20, 317–322 (2023).
Zhao, Y. G., Codogno, P. & Zhang, H. Machinery, regulation and pathophysiological implications of autophagosome maturation. Nat. Rev. Mol. Cell Biol. 22, 733–750 (2021).
Cason, S. E., Mogre, S. S., Holzbaur, E. L. F. & Koslover, E. F. Spatiotemporal analysis of axonal autophagosome-lysosome dynamics reveals limited fusion events and slow maturation. Mol. Biol. Cell 33, ar123 (2022).
Gotz, J. Tau and transgenic animal models. Brain Res. Brain Res. Rev. 35, 266–286 (2001).
Gotzl, J. K., Lang, C. M., Haass, C. & Capell, A. Impaired protein degradation in FTLD and related disorders. Ageing Res. Rev. 32, 122–139 (2016).
Ono, M. et al. Central role for p62/SQSTM1 in the elimination of toxic tau species in a mouse model of tauopathy. Aging Cell 21, e13615 (2022).
Koga, H., Kaushik, S. & Cuervo, A. M. Inhibitory effect of intracellular lipid load on macroautophagy. Autophagy 6, 825–827 (2010).
Tanida, I., Ueno, T. & Uchiyama, Y. A super-ecliptic, pHluorin-mKate2, tandem fluorescent protein-tagged human LC3 for the monitoring of mammalian autophagy. PLoS ONE 9, e110600 (2014).
Oh, Y. M. et al. Age-related Huntington’s disease progression modeled in directly reprogrammed patient-derived striatal neurons highlights impaired autophagy. Nat. Neurosci. 25, 1420–1433 (2022).
Lee, S. W. et al. Longitudinal modeling of human neuronal aging reveals the contribution of the RCAN1-TFEB pathway to Huntington’s disease neurodegeneration. Nat. Aging 4, 95–109 (2024).
Wang, Y. et al. An analog of glibenclamide selectively enhances autophagic degradation of misfolded alpha1-antitrypsin Z. PLoS ONE 14, e0209748 (2019).
Myerowitz, R., Puertollano, R. & Raben, N. Impaired autophagy: the collateral damage of lysosomal storage disorders. EBioMedicine 63, 103166 (2021).
Hu, Y. B., Dammer, E. B., Ren, R. J. & Wang, G. The endosomal-lysosomal system: from acidification and cargo sorting to neurodegeneration. Transl. Neurodegener. 4, 18 (2015).
Caballero, B. et al. Acetylated tau inhibits chaperone-mediated autophagy and promotes tau pathology propagation in mice. Nat. Commun. 12, 2238 (2021).
Siano, G., Falcicchia, C., Origlia, N., Cattaneo, A. & Di Primio, C. Non-canonical roles of tau and their contribution to synaptic dysfunction. Int. J. Mol. Sci. 22, 10145 (2021).
Wong, E. & Cuervo, A. M. Autophagy gone awry in neurodegenerative diseases. Nat. Neurosci. 13, 805–811 (2010).
Wang, Y. et al. Tau fragmentation, aggregation and clearance: the dual role of lysosomal processing. Hum. Mol. Genet. 18, 4153–4170 (2009).
Xu, Y. et al. TFEB regulates lysosomal exocytosis of tau and its loss of function exacerbates tau pathology and spreading. Mol. Psychiatry 26, 5925–5939 (2021).
Maday, S., Twelvetrees, A. E., Moughamian, A. J. & Holzbaur, E. L. Axonal transport: cargo-specific mechanisms of motility and regulation. Neuron 84, 292–309 (2014).
Maday, S. & Holzbaur, E. L. Compartment-specific regulation of autophagy in primary neurons. J. Neurosci. 36, 5933–5945 (2016).
Maday, S., Wallace, K. E. & Holzbaur, E. L. Autophagosomes initiate distally and mature during transport toward the cell soma in primary neurons. J. Cell Biol. 196, 407–417 (2012).
Root, J., Merino, P., Nuckols, A., Johnson, M. & Kukar, T. Lysosome dysfunction as a cause of neurodegenerative diseases: Lessons from frontotemporal dementia and amyotrophic lateral sclerosis. Neurobiol. Dis. 154, 105360 (2021).
Koga, H., Kaushik, S. & Cuervo, A. M. Altered lipid content inhibits autophagic vesicular fusion. FASEB J. 24, 3052–3065 (2010).
Glasauer, S. M. K. et al. Human tau mutations in cerebral organoids induce a progressive dyshomeostasis of cholesterol. Stem Cell Rep. 17, 2127–2140 (2022).
Garver, T. D., Kincaid, R. L., Conn, R. A. & Billingsley, M. L. Reduction of calcineurin activity in brain by antisense oligonucleotides leads to persistent phosphorylation of tau protein at Thr181 and Thr231. Mol. Pharmacol. 55, 632–641 (1999).
Karch, C. M., Jeng, A. T. & Goate, A. M. Calcium phosphatase calcineurin influences tau metabolism. Neurobiol. Aging 34, 374–386 (2013).
Ciechanover, A. & Kwon, Y. T. Degradation of misfolded proteins in neurodegenerative diseases: therapeutic targets and strategies. Exp. Mol. Med 47, e147 (2015).
Dikic, I. Proteasomal and autophagic degradation systems. Annu. Rev. Biochem 86, 193–224 (2017).
Parra Bravo, C., Naguib, S. A. & Gan, L. Cellular and pathological functions of tau. Nat. Rev. Mol. Cell. Biol. 25, 845–864 (2024).
Karch, C. M. et al. A Comprehensive resource for induced pluripotent stem cells from patients with primary tauopathies. Stem Cell Rep. 13, 939–955 (2019).
Tian, R. et al. CRISPR interference-based platform for multimodal genetic screens in human iPSC-derived neurons. Neuron 104, 239–255.e212 (2019).
Long, O. S. et al. A C. elegans model of human alpha1-antitrypsin deficiency links components of the RNAi pathway to misfolded protein turnover. Hum. Mol. Genet. 23, 5109–5122 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2012).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Bordi, M. et al. A gene toolbox for monitoring autophagy transcription. Cell Death Dis. 12, 1044 (2021).
Acknowledgements
We thank the research subjects and their families who generously participated in this study. We thank Dr. Kanaan and Dr. Binder, who generously provided the Tau5 antibodies. We thank David Kast and Torri Ball for thoughtful discussions. We thank the Genome Engineering & Stem Cell Center (GESC@MGI) at Washington University in St. Louis School of Medicine for iPSC reprogramming services and the Hope Center Viral Vector Core at Washington University in St Louis School of Medicine for packaging the Lenti virus particles. This work was supported by access to equipment made possible by the Hope Center for Neurological Disorders, the Neurogenomics and Informatics Center, and the Departments of Neurology and Psychiatry at Washington University School of Medicine. Confocal images were generated on a Zeiss LSM 980 Airyscan Confocal Microscope which was purchased with support from NIH grant number S10 MH126964 through the use of Washington University Center for Cellular Imaging (WUCCI)-Neuro supported by Washington University School of Medicine, the Children’s Discovery Institute of Washington University and St. Louis Children’s Hospital (CDI-CORE-2015-505 and CDI-CORE-2019-813), and the Foundation for Barnes-Jewish Hospital (3770 and 4642). Funding provided by the National Institutes of Health (P30 AG066444, R01 AG056293, RF1 NS110890, U54 NS123985), Hope Center for Neurological Disorders (CMK), Rainwater Charitable Organization (CMK), Farrell Family Fund for Alzheimer’s Disease (CMK), and UL1TR002345. Diagrams were generated using BioRender.com.
Author information
Authors and Affiliations
Contributions
Designed experiments: F.S.M., C.M.K. Performed and analyzed experiments: F.S.M., J.A.M., C.S., K.J.S., M.A.M., C.M.K. Provided essential tools: R.E.D., S.C.P., G.A.S., D.H.P., S.L.M. Provided funding: C.M.K. Wrote the manuscript: F.S.M., C.M.K. Revised and approved manuscript: F.A.M., J.A.M., C.H., K.J.S., M.A.M., R.E.D., S.C.P., G.A.S., D.H.P., S.L.M., C.M.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jonathan Benn and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mirfakhar, F.S., Marsh, J.A., Sato, C. et al. A pathogenic Tau mutation drives autophagy-lysosome dysfunction that limits Tau degradation in a model of frontotemporal dementia. Nat Commun 17, 2699 (2026). https://doi.org/10.1038/s41467-026-70473-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70473-5