Abstract
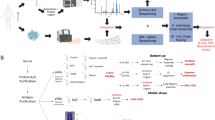
Elucidating antibody sequences by mass spectrometry-based de novo sequencing is essential but remains technically challenging. Here we present XA-Novo, an accurate and high-throughput de novo sequencing solution that integrates a single-pot multi-enzymatic gradient digestion method with a beam search-based assembler (Fusion) to reconstruct full-length antibody sequences directly from bottom-up mass spectrometry data. Benchmarking across well-characterized antibodies from multiple species demonstrates that XA-Novo outperforms commercial solutions in identification sensitivity, sequence completeness, and reconstruction accuracy. Furthermore, XA-Novo successfully reconstructs six immunotherapeutic antibodies with unknown sequences, and in vitro/vivo assays validate that these generated antibodies exhibit functionality equivalent to their commercial counterparts. Moreover, XA-Novo achieves over 99.54% accurate sequence coverage in distinguishing mixed COVID-19 neutralizing antibodies, exceeding the performance of current assemblers reported for single-antibody sequencing. Overall, XA-Novo establishes a reliable, scalable, and broadly applicable workflow for routine antibody sequencing, thereby accelerating both fundamental antibody research and therapeutic antibody development.
Similar content being viewed by others
Data availability
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the partner repository iProX under the dataset identifier PXD060500. The weights of the Casanovo model utilized in XA-Novo and the split datasets for training Casanovo are both available on Zenodo [https://zenodo.org/records/17266057] and [https://zenodo.org/records/18627093], respectively. Additionally, the revised nine-species benchmark dataset by Wen et al. is available on Zenodo [https://zenodo.org/records/13653420]. Unless otherwise stated, all data supporting the results of this study can be found in the article, supplementary, and source data files. Source data are provided with this paper.
Code availability
Result files and code to reproduce the results in this study are available on GitHub [https://github.com/biocc/SP-MEGD_Fusion]. An executable version of the code and the computational environment used in this study are also available as a Code Ocean capsule [https://codeocean.com/capsule/4653442/tree].
References
Lu, L. L., Suscovich, T. J., Fortune, S. M. & Alter, G. Beyond binding: antibody effector functions in infectious diseases. Nat. Rev. Immunol. 18, 46–61 (2018).
Oostindie, S. C., Lazar, G. A., Schuurman, J. & Parren, P. Avidity in antibody effector functions and biotherapeutic drug design. Nat. Rev. Drug Discov. 21, 715–735 (2022).
Watson, C. T., Glanville, J. & Marasco, W. A. The Individual and Population Genetics of Antibody Immunity. Trends Immunol. 38, 459–470 (2017).
Ejazi, S. A., Ghosh, S. & Ali, N. Antibody detection assays for COVID-19 diagnosis: an early overview. Immunol. Cell Biol. 99, 21–33 (2021).
Ning, L., Abagna, H. B., Jiang, Q., Liu, S. & Huang, J. Development and application of therapeutic antibodies against COVID-19. Int J. Biol. Sci. 17, 1486–1496 (2021).
Lu, R. M. et al. Development of therapeutic antibodies for the treatment of diseases. J. Biomed. Sci. 27, (2020).
Mason, D. M. et al. Optimization of therapeutic antibodies by predicting antigen specificity from antibody sequence via deep learning. Nat. Biomed. Eng. 5, 600–612 (2021).
Mattsson, J. et al. Sequence enrichment profiles enable target-agnostic antibody generation for a broad range of antigens. Cell Rep. Methods 3, 100475 (2023).
Parray, H. A. et al. Hybridoma technology a versatile method for isolation of monoclonal antibodies, its applicability across species, limitations, advancement and future perspectives. Int Immunopharmacol. 85, 106639 (2020).
Tomita, M. & Tsumoto, K. Hybridoma technologies for antibody production. Immunotherapy 3, 371–380 (2011).
Subas et al. NAb-seq: an accurate, rapid, and cost-effective method for antibody long-read sequencing in hybridoma cell lines and single B cells. MAbs 14, 2106621 (2022).
Chen, Y. et al. Barcoded sequencing workflow for high throughput digitization of hybridoma antibody variable domain sequences. J. Immunol. Methods 455, 88–94 (2018).
Schardt, J. S., Sivaneri, N. S. & Tessier, P. M. Monoclonal antibody generation using single B-cell screening for treating infectious diseases. BioDrugs 38, 477–486 (2024).
de Graaf, S. C., Hoek, M., Tamara, S. & Heck, A. J. R. A perspective toward mass spectrometry-based de novo sequencing of endogenous antibodies. MAbs 14, 2079449 (2022).
Schulte, D., Peng, W. & Snijder, J. Template-based assembly of proteomic short reads for de novo antibody sequencing and repertoire profiling. Anal. Chem. 94, 10391–10399 (2022).
Le Bihan, T. et al. De novo protein sequencing of antibodies for identification of neutralizing antibodies in human plasma post SARS-CoV-2 vaccination. Nat. Commun. 15, 8790 (2024).
Sen, K. I. et al. Automated Antibody De Novo sequencing and its utility in biopharmaceutical discovery. J. AM Soc. Mass Spectr. 28, 803–810 (2017).
Gadush, M. V. et al. Template-Assisted De Novo Sequencing of SARS-CoV-2 and Influenza Monoclonal Antibodies By Mass Spectrometry. J. Proteome Res. 21, 1616–1627 (2022).
He, M.-T. et al. Do-It-Yourself De Novo Antibody Sequencing Workflow That Achieves Complete Accuracy Of The Variable Regions. J. Proteome Res. 24, 3062–3073 (2025).
Pinto, D. et al. Broad betacoronavirus neutralization by a stem helix-specific human antibody. Science 373, 1109–1116 (2021).
Ye, X. et al. Integrated proteomics sample preparation and fractionation: Method development and applications. Trends Anal. Chem. 120, 115667 (2019).
Tsiatsiani, L. & Heck, A. J. R. Proteomics beyond trypsin. FEBS J. 282, 2612–2626 (2015).
Morsa, D. et al. Multi-enzymatic limited digestion: the next-generation sequencing for proteomics? J. Proteome Res. 18, 2501–2513 (2019).
Yilmaz M., Fondrie W., Bittremieux W., Oh S., Noble W. S. De novo mass spectrometry peptide sequencing with a transformer model. Int. Conf. Mach. Learn. 162, 25514-25522 (2022).
Yilmaz, M. et al. Sequence-to-sequence translation from mass spectra to peptides with a transformer model. Nat. Commun. 15, 6427 (2024).
Cao, Y. et al. BA.2.12.1, BA.4 and BA.5 escape antibodies elicited by Omicron infection. Nature 608, 593–602 (2022).
Wu, Y. et al. Lineage-mosaic and mutation-patched spike proteins for broad-spectrum COVID-19 vaccine. Cell Host Microbe 30, 1732–1744.e1737 (2022).
Tran, N. H. et al. Complete De Novo assembly of monoclonal antibody sequences. Sci. Rep. 6, 31730 (2016).
Schulte, D. & Snijder, J. A Handle on Mass Coincidence Errors in De Novo sequencing of antibodies by bottom-up proteomics. J. Proteome Res. 23, 3552–3559 (2024).
Guthals, A. et al. De Novo MS/MS sequencing of native human antibodies. J. Proteome Res. 16, 45–54 (2017).
Guthals, A., Clauser, K. R., Frank, A. M. & Bandeira, N. Sequencing-Grade De novo Analysis of MS/MS Triplets (CID/HCD/ETD) From Overlapping Peptides. J. Proteome Res. 12, 2846–2857 (2013).
Peng, W. et al. Reverse-engineering the anti-MUC1 antibody 139H2 by mass spectrometry-based de novo sequencing. Life Sci. Alliance 7, e202302366 (2024).
Chen, D. S. & Mellman, I. Oncology meets immunology: the cancer-immunity cycle. Immunity 39, 1–10 (2013).
Zhao, Y. L. et al. Comparison of the characteristics of macrophages derived from murine spleen, peritoneal cavity, and bone marrow. J. Zhejiang Univ. Sci. B 18, 1055–1063 (2017).
Bronte, V. & Pittet, M. ikaelJ. The spleen in local and systemic regulation of immunity. Immunity 39, 806–818 (2013).
Khoury, D. S. et al. Neutralizing antibody levels are highly predictive of immune protection from symptomatic SARS-CoV-2 infection. Nat. Med. 27, 1205–1211 (2021).
Cameroni, E. et al. Broadly neutralizing antibodies overcome SARS-CoV-2 Omicron antigenic shift. Nature 602, 664–670 (2022).
Tortorici, M. A. et al. Broad sarbecovirus neutralization by a human monoclonal antibody. Nature 597, 103–108 (2021).
Copin, R. et al. The monoclonal antibody combination REGEN-COV protects against SARS-CoV-2 mutational escape in preclinical and human studies. Cell 184, 3949–3961.e3911 (2021).
Cao, Y. et al. Rational identification of potent and broad sarbecovirus-neutralizing antibody cocktails from SARS convalescents. Cell Rep. 41, 111845 (2022).
Wang, M. et al. Assembling the Community-Scale Discoverable Human Proteome. Cell Syst. 7, 412–421.e415 (2018).
Beslic, D., Tscheuschner, G., Renard, B. Y., Weller, M. G. & Muth, T. Comprehensive evaluation of peptide de novo sequencing tools for monoclonal antibody assembly. Brief. Bioinform. 24, bbac542 (2022).
Wen, B. & Noble, W. S. A multi-species benchmark for training and validating mass spectrometry proteomics machine learning models. Sci. Data 11, 1207 (2024).
Tran, N. H., Zhang, X., Xin, L., Shan, B. & Li, M. De novo peptide sequencing by deep learning. Proc. Natl. Acad. Sci. USA 114, 8247–8252 (2017).
Kessner, D., Chambers, M., Burke, R., Agus, D. & Mallick, P. ProteoWizard: open source software for rapid proteomics tools development. Bioinformatics 24, 2534–2536 (2008).
Needleman, S. B. & Wunsch, C. D. A general method applicable to the search for similarities in the amino acid sequence of two proteins. J. Mol. Biol. 48, 443–453 (1970).
Acknowledgements
This work was supported by the National Science and Technology Major Project for Innovative Drug Research and Development (2025ZD1803703 to Q.Y.); the National Natural Science Foundation of China (32401237 to Y.X. and 92369110 to Q.Y.); the Fujian Provincial Natural Science Foundation of China (2024J08358 to Y.X.); the Natural Science Foundation of Xiamen, China (3502Z202371039 to Y.X.); the State Key Laboratory of Vaccines for Infectious Diseases, Xiang An Biomedicine Laboratory (2025XAKJ0200001 to R.Y.); and the Scientific Research Foundation of the State Key Laboratory of Vaccines for Infectious Diseases (2024SKLVDzy06 to Y.X.).
Author information
Authors and Affiliations
Contributions
Y.X., W.J. and J.X. contributed equally to this work. W.J., Y.X., R.Y., N.X. and Q.Y. conceived and designed the study. J.X. and J.W. conducted mass spectrometry experiments. Q.B., X.C. and Y.W. prepared recombinant antibodies and performed animal experiments. W.J. developed the algorithms. Z.J., L.L., Y.Q. and F.L. performed data analysis. Y.X. and W.J. wrote the draft of the manuscript. R.Y., N.X. and Q.Y. revised and edited the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.Y. is a shareholder of Aginome Scientific. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Albert Heck, Samantha Sarrett, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xiong, Y., Jiang, W., Xiao, J. et al. XA-Novo: high-throughput mass spectrometry-based de novo sequencing technology for monoclonal antibodies and antibody mixtures. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70496-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70496-y