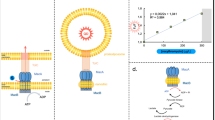
Abstract
Transport of proteins and small molecules across cellular membrane is crucial for bacterial interaction with the environment and survival against antibiotics. In Gram-negative bacteria that possess two layers of membranes, specialized macromolecular machines are required to transport substrates across the cell envelope, often via an indirect stepwise process. The major facilitator superfamily (MFS)-type tripartite efflux pumps use proton electrochemical gradient to extrude drugs in diverse bacterial species, but the architecture of the assembly and structural mechanisms remain elusive. A representative MFS-type tripartite efflux pump, EmrAB-TolC, mediates resistance to multiple antimicrobial drugs through proton-coupled EmrB, a member of the DHA2 transporter family. Here, we report the high-resolution (3.13 Å) structure of the EmrAB-TolC pump, revealing a distinct, asymmetric architecture emerging from the assembly of TolC:EmrA:EmrB with a ratio of 3:6:1 and contacts that are essential for the pump assembly. Key residues involved in drug transport are identified and corroborated by mutagenesis and antibiotic sensitivity assays. The structural and functional data support a model for one-step drug transport by the MFS pump across the entire envelope of Gram-negative bacteria.
Similar content being viewed by others
Data availability
Coordinates have been deposited in the Protein Data Bank (PDB) under PDB codes 8ZAL (EmrAB-TolC pump-EA) and 8ZAR (EmrAB-TolC pump-FA). Cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMD) under EMD codes EMD-39879 (EmrAB-TolC pump-EA) and EMD-39885 (EmrAB-TolC pump-FA). Molecular dynamics data are in the Figshare repository with the doi: 10.6084/m9.figshare.31240081 [doi.org/10.6084/m9.figshare.31240081]. Source data are provided with this paper.
Code availability
Molecular Dynamics input files, coordinates and simulation trajectories can be accessed at doi: 10.6084/m9.figshare.31240081 [doi.org/10.6084/m9.figshare.31240081].
References
Hodges, F. J., Torres, V. V. et al. Redefining the bacterial Type I protein secretion system. Adv. Micro. Physiol. 82, 155–204 (2023).
Costa, T. R. et al. Secretion systems in Gram-negative bacteria: structural and mechanistic insights. Nat. Rev. Microbiol 13, 343–359 (2015).
Du, D. et al. Multidrug efflux pumps: structure, function and regulation. Nat. Rev. Microbiol 16, 523–539 (2018).
Wang, Z. et al. An allosteric transport mechanism for the AcrAB-TolC multidrug efflux pump. Elife 6 https://doi.org/10.7554/eLife.24905 (2017).
Du, D. et al. Structure of the AcrAB-TolC multidrug efflux pump. Nature 509, 512–515 (2014).
Fitzpatrick, A. W. P. et al. Structure of the MacAB-TolC ABC-type tripartite multidrug efflux pump. Nat. Microbiol. 2, 17070 (2017).
Du, D., van Veen, H. W. & Luisi, B. F. Assembly and operation of bacterial tripartite multidrug efflux pumps. Trends Microbiol 23, 311–319 (2015).
Alav, I. et al. Structure, assembly, and function of tripartite efflux and type 1 secretion systems in gram-negative bacteria. Chem. Rev. 121, 5479–5596 (2021).
Varela, M. F., Stephen, J., Bharti, D., Lekshmi, M. & Kumar, S. Inhibition of multidrug efflux pumps belonging to the major facilitator superfamily in bacterial pathogens. Biomedicines 11 https://doi.org/10.3390/biomedicines11051448 (2023).
Yousefian, N. et al. Structural characterization of the EmrAB-TolC efflux complex from E. coli. Biochim. Biophys. Acta Biomembr. 1863, 183488 (2021).
Puertolas-Balint, F., Warsi, O., Linkevicius, M., Tang, P. C. & Andersson, D. I. Mutations that increase expression of the EmrAB-TolC efflux pump confer increased resistance to nitroxoline in Escherichia coli. J. Antimicrob. Chemother. 75, 300–308 (2020).
Lomovskaya, O. L., K. Emr, an Escherichia coli locus for multidrug resistance. Cell Biol. 89, 8938–42 (1992).
Hinchliffe, P. et al. Structure of the periplasmic adaptor protein from a major facilitator superfamily (MFS) multidrug efflux pump. FEBS Lett. 588, 3147–3153 (2014).
Jiang, D. et al. Structure of the YajR transporter suggests a transport mechanism based on the conserved motif A. Proc. Natl. Acad. Sci. USA 110, 14664–14669 (2013).
Yin, Y., He, X., Szewczyk, P., Nguyen, T. & Chang, G. Structure of the multidrug transporter EmrD from Escherichia coli. Science 312, 741–744 (2006).
Nagarathinam, K. et al. Outward open conformation of a Major Facilitator Superfamily multidrug/H+ antiporter provides insights into switching mechanism. Nat. Commun. 9 https://doi.org/10.1038/s41467-018-06306-x (2018).
Heng, J. et al. Substrate-bound structure of the E. coli multidrug resistance transporter MdfA. Cell Res. 25, 1060–1073 (2015).
Majumder, P. et al. Dissection of protonation sites for antibacterial recognition and transport in QacA, a multi-drug efflux transporter. J. Mol. Biol. 431, 2163–2179 (2019).
Remm, S. et al. Structural basis for triacylglyceride extraction from mycobacterial inner membrane by MFS transporter Rv1410. Nat. Commun. 14 https://doi.org/10.1038/s41467-023-42073-0 (2023).
Tanabe, M. et al. The multidrug resistance efflux complex, EmrAB from Escherichia coli forms a dimer in vitro. Biochem. Biophys. Res. Commun. 380, 338–342 (2009).
Symmons, M. F., Marshall, R. L. & Bavro, V. N. Architecture and roles of periplasmic adaptor proteins in tripartite efflux assemblies. Front. Microbiol. 6 https://doi.org/10.3389/fmicb.2015.00513 (2015).
Chun, E. et al. Fusion partner toolchest for the stabilization and crystallization of G protein-coupled receptors. Structure 20, 967–976 (2012).
Gotzke, H. et al. The ALFA-tag is a highly versatile tool for nanobody-based bioscience applications. Nat. Commun. 10, 4403 (2019).
Xie, P. et al. A fiducial-assisted strategy compatible with resolving small MFS transporter structures in multiple conformations using cryo-EM. Nat. Commun. 16, 7 (2025).
Higgins, M. K., Bokma, E., Koronakis, E., Hughes, C. & Koronakis, V. Structure of the periplasmic component of a bacterial drug efflux pump. Proc. Natl. Acad. Sci. USA 101, 9994–9999 (2004).
Daury, L. et al. Tripartite assembly of RND multidrug efflux pumps. Nat. Commun. 7, 10731 (2016).
Sulavik, M. C. et al. Antibiotic susceptibility profiles of Escherichia coli strains lacking multidrug efflux pump genes. Antimicrob. Agents Chemother. 45, 1126–1136 (2001).
Su, C. C. et al. Crystal structure of the CusBA heavy-metal efflux complex of Escherichia coli. Nature 470, 558–562 (2011).
Zhao, Y. et al. Crystal structure of the E. coli peptide transporter YbgH. Structure 22, 1152–1160 (2014).
Newstead, S. et al. Crystal structure of a prokaryotic homologue of the mammalian oligopeptide-proton symporters, PepT1 and PepT2. EMBO J. 30, 417–426 (2011).
Fowler, P. W. et al. Gating topology of the proton-coupled oligopeptide symporters. Structure 23, 290–301 (2015).
Jin, J., Guffanti, A. A., Beck, C. & Krulwich, T. A. Twelve-transmembrane-segment (TMS) Version (ΔTMS VII-VIII) of the 14-TMS Tet(L) antibiotic resistance protein retains monovalent cation transport modes but lacks tetracycline efflux capacity. J. Bacteriol. 183, 2667–2671 (2001).
Majumder, P. et al. Cryo-EM structure of antibacterial efflux transporter QacA from Staphylococcus aureus reveals a novel extracellular loop with allosteric role. EMBO J. 42, e113418 (2023).
Henderson, P. J. F. et al. Physiological functions of bacterial “multidrug” efflux pumps. Chem. Rev. 121, 5417–5478 (2021).
Vogele, M. et al. Systematic analysis of biomolecular conformational ensembles with PENSA. J. Chem. Phys. 162 https://doi.org/10.1063/5.0235544 (2025).
Drew, D., North, R. A., Nagarathinam, K. & Tanabe, M. Structures and general transport mechanisms by the major facilitator superfamily (MFS). Chem. Rev. 121, 5289–5335 (2021).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature https://doi.org/10.1038/s41586-024-07487-w (2024).
Wayment-Steele, H. K. et al. Predicting multiple conformations via sequence clustering and AlphaFold2. Nature 625, 832–839 (2024).
Sauve, S., Williamson, J., Polasa, A. & Moradi, M. Ins and outs of rocker switch mechanism in major facilitator superfamily of transporters. Membranes (Basel) 13 https://doi.org/10.3390/membranes13050462 (2023).
Glavier, M. et al. Antibiotic export by MexB multidrug efflux transporter is allosterically controlled by a MexA-OprM chaperone-like complex. Nat. Commun. 11, 4948 (2020).
Wu, H.-H., Symersky, J. & Lu, M. Structure and mechanism of a redesigned multidrug transporter from the Major Facilitator Superfamily. Sci. Rep. 10 https://doi.org/10.1038/s41598-020-60332-8 (2020).
Kim, J.-S. et al. Crystal structure of a soluble fragment of the membrane fusion protein HlyD in a type I secretion system of gram-negative bacteria. Structure 24, 477–485 (2016).
Teelucksingh, T. et al. A genetic platform to investigate the functions of bacterial drug efflux pumps. Nat. Chem. Biol. 18, 1399–1409 (2022).
Chovancova, E. et al. CAVER 3.0: a tool for the analysis of transport pathways in dynamic protein structures. PLoS Comput Biol. 8, e1002708 (2012).
Carlson, M. L. et al. The Peptidisc, a simple method for stabilizing membrane proteins in detergent-free solution. Elife 7 https://doi.org/10.7554/eLife.34085 (2018).
Angiulli, G. et al. New approach for membrane protein reconstitution into peptidiscs and basis for their adaptability to different proteins. Elife 9 https://doi.org/10.7554/eLife.53530 (2020).
Fluman, N., Ryan, C. M., Whitelegge, J. P. & Bibi, E. Dissection of mechanistic principles of a secondary multidrug efflux protein. Mol. Cell 47, 777–787 (2012).
Bogdanov, M. Preparation of uniformly oriented inverted inner (cytoplasmic) membrane vesicles from gram-negative bacterial cells. Methods Mol. Biol. 2715, 159–180 (2024).
Verkhovskaya, M. Preparation of everted membrane vesicles from Escherichia coli cells. Bio Protoc. 7, e2254 (2017).
Schorb, M., Haberbosch, I., Hagen, W. J. H., Schwab, Y. & Mastronarde, D. N. Software tools for automated transmission electron microscopy. Nat. Methods 16, 471–477 (2019).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7 https://doi.org/10.7554/eLife.42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Wright, N. J. et al. Methotrexate recognition by the human reduced folate carrier SLC19A1. Nature 609, 1056–1062 (2022).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Wang, R. Y. et al. Automated structure refinement of macromolecular assemblies from cryo-EM maps using Rosetta. Elife 5 https://doi.org/10.7554/eLife.17219 (2016).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D. Struct. Biol. 74, 519–530 (2018).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D. Struct. Biol. 75, 861–877 (2019).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput Chem. 29, 1859–1865 (2008).
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013).
Coughtrie, D. J. & Tew, D. P. The Nose-Hoover looped chain thermostat for low temperature thawed Gaussian wave-packet dynamics. J. Chem. Phys. 140, 194106 (2014).
Parrinello, M., Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52 https://doi.org/10.1063/1.328693 (1981).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Jorgensen, W. L. & Tirado-Rives, J. Potential energy functions for atomic-level simulations of water and organic and biomolecular systems. Proc. Natl. Acad. Sci. USA 102, 6665–6670 (2005).
Michaud-Agrawal, N., Denning, E. J., Woolf, T. B. & Beckstein, O. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput Chem. 32, 2319–2327 (2011).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput Chem. 31, 455–461 (2010).
Valdes-Tresanco, M. S., Valdes-Tresanco, M. E., Valiente, P. A. & Moreno, E. AMDock: a versatile graphical tool for assisting molecular docking with Autodock Vina and Autodock4. Biol. Direct 15, 12 (2020).
Acknowledgements
This work was supported by the National Key R&D Program of China (2022YFC2303200); National Natural Science Foundation of China (31971133 to D.D.; 32270064 and 92478118 to Y.C.), the Science and Technology Commission of Shanghai Municipality (19PJ1407900, 19JC1414000 and 22WZ2504100 to D.D.; 24ZR1493200 to Y.C.), and the Chinese Academy of Sciences (XDB0570000 and 176002GJHZ2022022MI to Y.C.). BFL was supported by ERC Advanced grant (742210) and a Wellcome Trust Investigator award (200873/Z/16/Z). MLJ is supported by a UK Medical Research Council Intramural Programme Award MC_UU_000254/4 (RG94521).. Cryo-EM data were collected at the Bio-Electron Microscopy Facility of ShanghaiTech University with the assistance of Q. Sun, D. Liu, Z. Zhang, L. Wang and Y. Yang. We thank the Molecular Imaging Core Facility, the Molecular and Cell Biology Core Facility, and the Multi-Omics Core Facility at the School of Life Science and Technology for providing technical support. We are also grateful for the support of Lajos Kalmar of the MRC Toxicology Unit in the use of high-performance computing used in this study. We thank Sofiya Mason for help with molecular docking.
Author information
Authors and Affiliations
Contributions
Z.Z. performed cloning and overexpression of the EmrAB-TolC complex; Z.Z. and T.M. purified the EmrAB-TolC complex, prepared cryo-EM samples, collected cryo-EM data, determined structures, performed model building and refinement and prepared figures for the manuscript; J.G. and X.G. carried out drug-proton antiport assay; W.W., X.G., W.S., Q.W., J.G., S.L., H.L. and Q.O. optimized in-column peptide-disc methods, prepared homemade graphene monolayer grids, and assisted the collection of cryo-EM data; R.D., H.J., Z.Z., T.M., X.G., S.Z. and W.S. performed antibiotics sensitivity assays; T.M. and H.L. carried out Western blot assay; Y.C. supervised antibiotics sensitivity assays and discussed project design; M.L.J. performed protein structure predictions and modeling; U.Z. performed and analysed the molecular dynamics simulations; D.D. and B.L. conceived the project; D.D. designed and supervised all experiments; D.D., Y.C. and B.L. wrote the manuscript. All the authors contributed to the data interpretation and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Konstantinos Beis, Gabriel Da Hora and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhong, Z., Maimaiti, T., Jackson, M.L. et al. A model for drug transport across two membranes of Gram-negative bacteria by an MFS tripartite assembly. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70500-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70500-5