Abstract
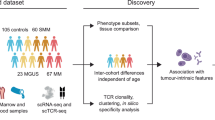
T cell states are prognostic in different cancer types. Recent technologies enable joint profiling of T cell RNA and T cell receptor (TCR) sequences at single-cell resolution. Here we present the TCR-RNA Integrating Model (TRIM), a multi-modal variational autoencoder framework that integrates RNA-TCR data and predicts T cell clonality and transcriptional states. TRIM learns a shared representation of the data conditioned on patient, tissue source, and treatment timepoint. We applied TRIM to three independent datasets that included T cells collected before and after checkpoint inhibitor treatment, sourced either from blood and tumor biopsies in patients with head and neck squamous cell carcinoma and colorectal cancer, or from tumor and adjacent tissue in a pan-cancer dataset. In all settings, TRIM accurately predicted intra-tumor T cell clonal expansion and transcriptional status based on T cells from blood or normal tissue before treatment, demonstrating its utility in modeling multimodal T cell data and predicting T cell response to treatment and disease progression.
Similar content being viewed by others
Data availability
All datasets used in this project have been previously published and are publicly available. The Head and Neck Squamous Cell Carcinoma (HNSCC) T cell dataset19 is available in the Gene Expression Omnibus (GEO) database under accession code GSE200996. The Colorectal Cancer (CRC) T cell dataset27 is available in the GEO database under accession code GSE236581. The pan-cancer T cell dataset11 is available in GEO database under accession code GSE156728. Source data are provided with this paper.
Code availability
All code for this project is available at https://github.com/uhlerlab/TRIMand on Zenodo https://doi.org/10.5281/zenodo.18421428. The repository also contains a list of the required open-source packages with version numbers, a tutorial for data pre-processing, the code for reproducing the figures in the manuscript.
References
Restifo, N. P., Dudley, M. E. & Rosenberg, S. A. Adoptive immunotherapy for cancer: harnessing the t cell response. Nat. Rev. Immunol. 12, 269–281 (2012).
Sharma, P. & Allison, J. P. Dissecting the mechanisms of immune checkpoint therapy. Nat. Rev. Immunol. 20, 75–76 (2020).
Alcover, A., Alarcón, B. & Di Bartolo, V. Cell biology of t cell receptor expression and regulation. Annu. Rev. Immunol. 36, 103–125 (2018).
Davis, M. M. & Bjorkman, P. J. T-cell antigen receptor genes and t-cell recognition. Nature 334, 395–402 (1988).
Mellman, I., Chen, D. S., Powles, T. & Turley, S. J. The cancer-immunity cycle: Indication, genotype, and immunotype. Immunity 56, 2188–2205 (2023).
Waldman, A. D., Fritz, J. M. & Lenardo, M. J. A guide to cancer immunotherapy: from t cell basic science to clinical practice. Nat. Rev. Immunol. 20, 651–668 (2020).
Galassi, C., Chan, T. A., Vitale, I. & Galluzzi, L. The hallmarks of cancer immune evasion. Cancer Cell 42, 1825–1863 (2024).
Oliveira, G. & Wu, C. J. Dynamics and specificities of t cells in cancer immunotherapy. Nat. Rev. Cancer 23, 295–316 (2023).
Pritykin, Y. et al. A unified atlas of cd8 t cell dysfunctional states in cancer and infection. Mol. cell 81, 2477–2493 (2021).
Andreatta, M. et al. Interpretation of t cell states from single-cell transcriptomics data using reference atlases. Nat. Commun. 12, 2965 (2021).
Zheng, L. et al. Pan-cancer single-cell landscape of tumor-infiltrating t cells. Science 374, abe6474 (2021).
Tooley, K. et al. Pan-cancer mapping of single cd8+ t cell profiles reveals a tcf1: Cxcr6 axis regulating cd28 co-stimulation and anti-tumor immunity. Cell Rep. Med.5, https://doi.org/10.1016/j.xcrm.2024.101640 (2024).
Chu, Y. et al. Pan-cancer t cell atlas links a cellular stress response state to immunotherapy resistance. Nat. Med. 29, 1550–1562 (2023).
Chow, A., Perica, K., Klebanoff, C. A. & Wolchok, J. D. Clinical implications of t cell exhaustion for cancer immunotherapy. Nat. Rev. Clin. Oncol. 19, 775–790 (2022).
Han, A., Glanville, J., Hansmann, L. & Davis, M. M. Linking t-cell receptor sequence to functional phenotype at the single-cell level. Nat. Biotechnol. 32, 684–692 (2014).
Pai, J. A. & Satpathy, A. T. High-throughput and single-cell t cell receptor sequencing technologies. Nat. methods 18, 881–892 (2021).
Mathewson, N. D. et al. Inhibitory cd161 receptor identified in glioma-infiltrating t cells by single-cell analysis. Cell 184, 1281–1298 (2021).
Pauken, K. E. et al. Tcr-sequencing in cancer and autoimmunity: barcodes and beyond. Trends Immunol. 43, 180–194 (2022).
Luoma, A. M. et al. Tissue-resident memory and circulating t cells are early responders to pre-surgical cancer immunotherapy. Cell 185, 2918–2935 (2022).
Zhou, Y. et al. Tumor biomarkers for diagnosis, prognosis and targeted therapy. Signal Transduct. Target. Ther. 9, 132 (2024).
Warren, J. D. et al. Septin 9 methylated dna is a sensitive and specific blood test for colorectal cancer. BMC Med. 9, 1–9 (2011).
Wu, T. D. et al. Peripheral t cell expansion predicts tumour infiltration and clinical response. Nature 579, 274–278 (2020).
Valpione, S. et al. Immune awakening revealed by peripheral t cell dynamics after one cycle of immunotherapy. Nat. cancer 1, 210–221 (2020).
Pauken, K. E. et al. Single-cell analyses identify circulating anti-tumor cd8 t cells and markers for their enrichment. J. Exp. Med. 218, https://doi.org/10.1084/jem.20200920 (2021).
Argelaguet, R., Cuomo, A. S., Stegle, O. & Marioni, J. C. Computational principles and challenges in single-cell data integration. Nat. Biotechnol. 39, 1202–1215 (2021).
Valkiers, S. et al. Recent advances in t-cell receptor repertoire analysis: bridging the gap with multimodal single-cell rna sequencing. ImmunoInformatics 5, 100009 (2022).
Chen, Y. et al. Spatiotemporal single-cell analysis decodes cellular dynamics underlying different responses to immunotherapy in colorectal cancer. Cancer Cell 42, 1268–1285 (2024).
Guo, X. et al. Contrasting cytotoxic and regulatory t cell responses underlying distinct clinical outcomes to anti-pd-1 plus lenvatinib therapy in cancer. Cancer Cell, https://doi.org/10.1016/j.ccell.2025.01.001 (2025).
Yang, K. D. et al. Multi-domain translation between single-cell imaging and sequencing data using autoencoders. Nat. Commun. 12, 31 (2021).
Amodio, M. et al. Exploring single-cell data with deep multitasking neural networks. Nat. methods 16, 1139–1145 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Schattgen, S. A. et al. Integrating t cell receptor sequences and transcriptional profiles by clonotype neighbor graph analysis (conga). Nat. Biotechnol. 40, 54–63 (2022).
Zhang, Z., Xiong, D., Wang, X., Liu, H. & Wang, T. Mapping the functional landscape of t cell receptor repertoires by single-t cell transcriptomics. Nat. methods 18, 92–99 (2021).
Drost, F. et al. Multi-modal generative modeling for joint analysis of single-cell t cell receptor and gene expression data. Nat. Commun. 15, 5577 (2024).
Zhang, X., Wang, X., Shivashankar, G. & Uhler, C. Graph-based autoencoder integrates spatial transcriptomics with chromatin images and identifies joint biomarkers for alzheimer’s disease. Nat. Commun. 13, 7480 (2022).
Kingma, D. P. et al. An introduction to variational autoencoders. Found. Trends® Mach. Learn. 12, 307–392 (2019).
Lopez, R., Gayoso, A. & Yosef, N. Enhancing scientific discoveries in molecular biology with deep generative models. Mol. Syst. Biol. 16, e9198 (2020).
Cohen Kalafut, N., Huang, X. & Wang, D. Joint variational autoencoders for multimodal imputation and embedding. Nat. Mach. Intell. 5, 631–642 (2023).
Lin, X., Tian, T., Wei, Z. & Hakonarson, H. Clustering of single-cell multi-omics data with a multimodal deep learning method. Nat. Commun. 13, 7705 (2022).
Lagattuta, K. A. et al. The t cell receptor sequence influences the likelihood of t cell memory formation. Cell Rep. 44 (2025).
Hudson, D., Fernandes, R. A., Basham, M., Ogg, G. & Koohy, H. Can we predict t cell specificity with digital biology and machine learning? Nat. Rev. Immunol. 23, 511–521 (2023).
Cabaniols, J.-P., Fazilleau, N., Casrouge, A., Kourilsky, P. & Kanellopoulos, J. M. Most α/β t cell receptor diversity is due to terminal deoxynucleotidyl transferase. J. Exp. Med. 194, 1385–1390 (2001).
Arstila, T. P. et al. A direct estimate of the human αβ t cell receptor diversity. Science 286, 958–961 (1999).
McInnes, L., Healy, J. & Melville, J. Umap: Uniform manifold approximation and projection for dimension reduction. Journal of Open Source Software, 3, 861 (2018).
Moore, D. S., McCabe, G. P., Alwan, L. C. & Craig, B. A. The practice of statistics for business and economics (Springer, 2016).
Fang, Z., Liu, X. & Peltz, G. Gseapy: a comprehensive package for performing gene set enrichment analysis in python. Bioinformatics 39, btac757 (2023).
Hu, M. et al. Evaluation of large language models for discovery of gene set function. Nat. Methods 22, 82–91 (2025).
Lee, Y. K., Mukasa, R., Hatton, R. D. & Weaver, C. T. Developmental plasticity of th17 and treg cells. Curr. Opin. Immunol. 21, 274–280 (2009).
Daniel, B. et al. Divergent clonal differentiation trajectories of t cell exhaustion. Nat. Immunol. 23, 1614–1627 (2022).
Schattgen, S. et al. Diverse modes of T cell receptor sequence convergence define unique functional and cellular phenotypes. bioRxiv (2025).
Lagattuta, K. A. et al. Repertoire analyses reveal t cell antigen receptor sequence features that influence t cell fate. Nat. Immunol. 23, 446–457 (2022).
Chen, X. et al. Variational lossy autoencoder. In International Conference of Learning Representations (2017).
Imambi, S., Prakash, K. B. & Kanagachidambaresan, G. Pytorch. Programming with TensorFlow: solution for edge computing applications 87–104 (2021).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotech. https://doi.org/10.1038/s41587-023-01767-y (2023).
Acknowledgements
We thank Adrienne M. Luoma and Jonathan D. Schoenfeld for helpful discussions, and Elvira Forte for scientific input and manuscript editing. CH and MA were supported by a fellowship from the Eric and Wendy Schmidt Center at the Broad Institute. KWW was partially supported by the NIH (R01 CA238039, R01CA251599, P01 CA236749, and P01 CA163222). KWW is a member of the Parker Institute for Cancer Immunotherapy (PICI). This work was supported by a grant from the National Institutes of Health (DK043351, DK135492, and AI110495) to RJX. CU was partially supported by NCCIH/NIH (1DP2AT012345), ONR (N00014-22-1-2116 and N00014-24-1-2687), DOE (DE-SC0023187), the MIT-IBM Watson AI Lab, MIT J-Clinic for Machine Learning and Health, the Eric and Wendy Schmidt Center at the Broad Institute, and a Simons Investigator Award.
Author information
Authors and Affiliations
Contributions
C.H., M.A., O.A., and C.U. designed the research. C.H. and M.A. developed and implemented the algorithms and performed model and data analysis. C.H., M.A., O.A., K.W.W., R.J.X., and C.U. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
KWW serves on the scientific advisory boards of DEM BioPharma, Solu Therapeutics, D2M Biotherapeutics, DoriNano, Inc., and Nextechinvest. He is a co-founder of Immunitas Therapeutics and receives sponsored research funding from Fate Therapeutics. He holds equity in TScan Therapeutics. RJX is Board Director at MoonLake Immunotherapeutics, and Scientific Advisory Board member at Nestle, Magnet Biomedicine, and Arena Bioworks, Co-founder of Convergence Bio; these organizations had no role in this study. CU serves on the Scientific Advisory Board of Immunai and Relation Therapeutics and has received sponsored research support from AstraZeneca and Janssen Pharmaceuticals. These activities are not related to the research reported in this study. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Benny Chain and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
He, C., Amodio, M., Ashenberg, O. et al. Multimodal framework for the joint analysis of single-cell RNA and T cell receptor sequencing data predicts T cell response to cancer immunotherapy. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70505-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70505-0