Abstract
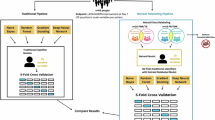
Patient recruitment remains a major bottleneck in clinical trials, calling for scalable and automated solutions. We present TrialMatchAI, an AI-powered recommendation system that automates patient-to-trial matching by processing heterogeneous clinical data, including structured records and unstructured physician notes. Built on fine-tuned, open-source large language models (LLMs) within a retrieval-augmented generation framework, TrialMatchAI ensures transparency and reproducibility and maintains a lightweight deployment footprint suitable for clinical environments. The system normalizes biomedical entities, retrieves relevant trials using a hybrid search strategy combining lexical and semantic similarity, re-ranks results, and performs criterion-level eligibility assessments using medical Chain-of-Thought reasoning. This pipeline delivers explainable outputs with traceable decision rationales. In real-world validation, 92% of oncology patients had at least one relevant trial retrieved within the top 20 recommendations. Evaluation across synthetic and real clinical datasets confirmed state-of-the-art performance, with expert assessment validating over 90% accuracy in criterion-level eligibility classification - particularly excelling in biomarker-driven matches. Designed for modularity and privacy, TrialMatchAI supports Phenopackets-standardized data, enables secure local deployment, and allows seamless replacement of LLM components as more advanced models emerge. By enhancing efficiency, interpretability and offering lightweight, open-source deployment, TrialMatchAI provides a scalable solution for AI-driven clinical trial matching in precision medicine.
Similar content being viewed by others
Data availability
The data generated in this study have been deposited in the Zenodo repository dedicated to TrialMatchAI under accession code 15045515: https://zenodo.org/records/15045515. These include the fine-tuned models (low rank adapter format), the “Ideal Candidates” synthetic patient dataset and matching results, the expert-examined patient-criterion pairs, the curated and parsed clinical trial database, the results on the TREC benchmarks, and the dictionaries used for entity normalization. Source data are provided as a Source File with this paper. The synthetic datasets used in this study for the TREC 2021 and 2022 Clinical Trials tracks are available at https://www.trec-cds.org/2021.html and http://www.trec-cds.org/2022.html. The lists of publicly accessible clinical trial identifiers used in the evaluation (ClinicalTrials.gov and CCMO) are provided in the Source Data file. The WIDE study data from the Netherlands Cancer Institute (NKI) are available under restricted access because they contain patient-level clinical information subject to privacy and institutional governance requirements. Access can be obtained by submitting a request through the Office Manager Knowledge Transfer & Contracting (KT&C) at NKI and completing the required data transfer/use agreements; requests are reviewed on a case-by-case basis. The expected response timeframe is approximately 3–6 months, and access is typically granted for one year (extensions may be requested). Source data are provided with this paper.
Code availability
The TrialMatchAI source code is available under the MIT Licence on GitHub at https://github.com/cbib/TrialMatchAI. A citable archived release is available on Zenodo80.
References
Garraway, L. A., Verweij, J. & Ballman, K. V. Precision oncology: an overview. J. Clin. Oncol. 31, 1803–1805 (2013).
Dienstmann, R., Jang, I. S., Bot, B., Friend, S. & Guinney, J. Database of genomic biomarkers for cancer drugs and clinical targetability in solid tumors. Cancer Discov. 5, 118–123 (2015).
Briel, M. et al. A systematic review of discontinued trials suggested that most reasons for recruitment failure were preventable. J. Clin. Epidemiol. 80, 8–15 (2016).
Fogel, D. B. Factors associated with clinical trials that fail and opportunities for improving the likelihood of success: a review. Contemp. Clin. Trials Commun. 11, 156–164 (2018).
Luchini, C., Lawlor, R. T., Milella, M. & Scarpa, A. Molecular tumor boards in clinical practice. Trends Cancer 6, 738–744 (2020).
Malone, E. R., Oliva, M., Sabatini, P. J. B., Stockley, T. L. & Siu, L. L. Molecular profiling for precision cancer therapies. Genome Med. 12, 8 (2020).
Penberthy, L. T., Dahman, B. A., Petkov, V. I. & DeShazo, J. P. Effort required in eligibility screening for clinical trials. J. Oncol. Pract. 8, 365–370 (2012).
Meystre, S. M. et al. Piloting an automated clinical trial eligibility surveillance and provider alert system based on artificial intelligence and standard data models. BMC Med. Res. Methodol. 23, 88 (2023).
Ni, Y. et al. Increasing the efficiency of trial-patient matching: automated clinical trial eligibility pre-screening for pediatric oncology patients. BMC Med. Inform. Decis. Mak. 15, 28 (2015).
Hassanzadeh, H., Karimi, S. & Nguyen, A. Matching patients to clinical trials using semantically enriched document representation. J. Biomed. Inform. 105, 103406 (2020).
Weng, C. et al. An integrated model for patient care and clinical trials (IMPACT) to support clinical research visit scheduling workflow for future learning health systems. J. Biomed. Inform. 46, 642–652 (2013).
Jin, Q. et al. Matching patients to clinical trials with large language models. Nat. Commun. 15, 9074 (2024).
Gao, J., Xiao, C., Glass, L. M. & Sun, J. COMPOSE: cross-modal pseudo-Siamese network for patient trial matching. In Proceedings of the 26th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, 803–812 (ACM, 2020).
Thadani, S. R., Weng, C., Bigger, J. T., Ennever, J. F. & Wajngurt, D. Electronic screening improves efficiency in clinical trial recruitment. J. Am. Med. Inform. Assoc. 16, 869–873 (2009).
Zhang, X., Xiao, C., Glass, L. M. & Sun, J. DeepEnroll: patient-trial matching with deep embedding and entailment prediction. In Proceedings of The Web Conference 2020, 1029–1037 (Association for Computing Machinery, 2020).
Weng, C. et al. EliXR: an approach to eligibility criteria extraction and representation. J. Am. Med. Inform. Assoc. 18, i116–i124 (2011).
Yuan, C. et al. Criteria2Query: a natural language interface to clinical databases for cohort definition. J. Am. Med. Inform. Assoc. 26, 294–305 (2019).
Shi, J., Graves, K. & Hurdle, J. F. A generic rule-based system for clinical trial patient selection. Preprint at https://doi.org/10.48550/arXiv.1907.06860 (2019).
O’Regan, P. et al. Digital ECMT cancer trial matching tool: an open source research application to support oncologists in the identification of precision medicine clinical trials. JCO Clin. Cancer Inform. 7, e2200137 (2023).
Alzubaidi, L. et al. Review of deep learning: concepts, CNN architectures, challenges, applications, future directions. J. Big Data 8, 53 (2021).
Klein, H. et al. MatchMiner: an open-source platform for cancer precision medicine. Npj Precis. Oncol. 6, 69 (2022).
Rybinski, M., Kusa, W., Karimi, S. & Hanbury, A. Learning to match patients to clinical trials using large language models. J. Biomed. Inform. 159, 104734 (2024).
Vaswani, A. et al. Attention is all you need. Preprint at https://doi.org/10.48550/arXiv.1706.03762 (2023).
Lee, J. et al. BioBERT: a pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 36, 1234–1240 (2020).
Devlin, J., Chang, M.-W., Lee, K. & Toutanova, K. BERT: pre-training of deep bidirectional transformers for language understanding. Preprint at https://doi.org/10.48550/arXiv.1810.04805 (2019).
Brown, T. et al. Language models are few-shot learners. Adv. Neural Inf. Process. Syst. 33, 1877–1901 (2020).
Meng, X. et al. The application of large language models in medicine: a scoping review. iScience 27, 109713 (2024).
Kaskovich, S. et al. Automated matching of patients to clinical trials: a patient-centric natural language processing approach for pediatric leukemia. JCO Clin. Cancer Inform. 7, e2300009 (2023).
Gueguen, L. et al. A prospective pragmatic evaluation of automatic trial matching tools in a molecular tumor board. Npj Precis. Oncol. 9, 28 (2025).
de Kok, J. W. T. M. et al. A guide to sharing open healthcare data under the General Data Protection Regulation. Sci. Data 10, 404 (2023).
Li, X. et al. LLM-Match: an open-sourced patient matching model based on large language models and retrieval-augmented generation. Preprint at https://doi.org/10.48550/arXiv.2503.13281 (2025).
Lin, J., Xu, H., Wang, Z., Wang, S. & Sun, J. Panacea: a foundation model for clinical trial search, summarization, design, and recruitment. Preprint at https://doi.org/10.48550/arXiv.2407.11007 (2024).
Lewis, P. et al. Retrieval-augmented generation for knowledge-intensive NLP tasks. Preprint at https://doi.org/10.48550/arXiv.2005.11401 (2021).
Jacobsen, J. O. B. et al. The GA4GH Phenopacket schema defines a computable representation of clinical data. Nat. Biotechnol. 40, 817–820 (2022).
Roberts, K., Demner-Fushman, D., Voorhees, E. M., Bedrick, S. & Hersh, W. R. Overview of the TREC 2021 Clinical Trials Track. In Proc. 30th Text REtrieval Conference (TREC, 2021).
Roberts, K., Demner-Fushman, D., Voorhees, E. M., Bedrick, S. & Hersh, W. R. Overview of the TREC 2022 Clinical Trials Track. In Proc. 31sr Text REtrieval Conference (TREC, 2022).
Samsom, K. G. et al. Study protocol: whole genome sequencing implementation in standard diagnostics for every cancer patient (WIDE). BMC Med. Genomics 13, 169 (2020).
Wei, J. et al. Chain-of-thought prompting elicits reasoning in large language models. Adv. Neural Inf. Process. Syst. 35, 24824–24837 (2022).
Ayaz, M., Pasha, M. F., Alzahrani, M. Y., Budiarto, R. & Stiawan, D. The fast health interoperability resources (FHIR) standard: systematic literature review of implementations, applications, challenges and opportunities. JMIR Med. Inform. 9, e21929 (2021).
Robertson, S. E., Walker, S., Jones, S., Hancock-Beaulieu, M. M. & Gatford, M. Okapi at TREC-3. In Proc. Third Text REtrieval Conference (TREC-3), NIST Spec. Publ. 500-225, 109–126 (1995).
Robertson, S. & Zaragoza, H. The probabilistic relevance framework: BM25 and beyond. Found. Trends Inf. Retr. 3, 333–389 (2009).
Malkov, Y. A. & Yashunin, D. A. Efficient and robust approximate nearest neighbor search using hierarchical navigable small world graphs. IEEE Trans. Pattern Anal. Mach. Intell. 42, 824–836 (2020).
Team, G. et al. Gemma 2: improving open language models at a practical size. Preprint at https://doi.org/10.48550/arXiv.2408.00118 (2024).
Abdin, M. et al. Phi-4 technical report. Preprint at https://doi.org/10.48550/arXiv.2412.08905 (2024).
Sung, M. et al. BERN2: an advanced neural biomedical named entity recognition and normalization tool. Bioinformatics 38, 4837–4839 (2022).
Sung, M., Jeon, H., Lee, J. & Kang, J. Biomedical entity representations with synonym marginalization. In Proceedings of the 58th Annual Meeting of the Association for Computational Linguistics (eds Jurafsky, D. et al.) 3641–3650 https://doi.org/10.18653/v1/2020.acl-main.335 (Association for Computational Linguistics, Online, 2020).
Lewis, P., Ott, M., Du, J. & Stoyanov, V. Pretrained language models for biomedical and clinical tasks: understanding and extending the state-of-the-art. In Proceedings of the 3rd Clinical Natural Language Processing Workshop (eds Rumshisky, A. et al.) 146–157 https://doi.org/10.18653/v1/2020.clinicalnlp-1.17 (Association for Computational Linguistics, Online, 2020).
Zaratiana, U., Tomeh, N., Holat, P. & Charnois, T. GLiNER: generalist model for named entity recognition using bidirectional transformer. In Proc. Conf. North Am. Chapter Assoc. Comput. Linguist. Hum. Lang. Technol. 1, 5364–5376 (2024).
Peng, H. et al. Biomedical named entity normalization via interaction-based synonym marginalization. J. Biomed. Inform. 136, 104238 (2022).
Chen, J. et al. M3-embedding: multi-linguality, multi-functionality, multi-granularity text embeddings through self-knowledge distillation. Findings of the Association for Computational Linguistics, 2318–2335 (ACL, 2024).
Gormley, C. & Tong, Z. Elasticsearch: The Definitive Guide (O’Reilly Media, Inc., 2015).
Durden, K. et al. Provider motivations and barriers to cancer clinical trial screening, referral, and operations: findings from a survey. Cancer 130, 68–76 (2024).
Davis, S. ICH GCP E6 has arrived: are we ready? Perspect. Clin. Res. 16, 171–172 (2025).
Dexheimer, J. W. et al. A time-and-motion study of clinical trial eligibility screening in a pediatric emergency department. Pediatr. Emerg. Care 35, 868–873 (2019).
Yu, C. et al. A survey on agent workflow - status and future. In 2025 8th International Conference on Artificial Intelligence and Big Data (ICAIBD) 770–781 https://doi.org/10.1109/ICAIBD64986.2025.11082076 (2025).
Xu, X. et al. A survey on knowledge distillation of large language models. Preprint at https://doi.org/10.48550/arXiv.2402.13116 (2024).
Nievas, M., Basu, A., Wang, Y. & Singh, H. Distilling large language models for matching patients to clinical trials. J. Am. Med. Inform. Assoc. 31, 1953–1963 (2024).
Schafer, J. B., Frankowski, D., Herlocker, J. & Sen, S. Collaborative Filtering Recommender Systems. In The Adaptive Web: Methods and Strategies of Web Personalization (eds Brusilovsky, P. et al.) 291–324 https://doi.org/10.1007/978-3-540-72079-9_9 (Springer, Berlin, Heidelberg, 2007).
Kundra, R. et al. OncoTree: a cancer classification system for precision oncology. JCO Clin. Cancer Inform. 5, 221–230 (2021).
Huiskens, J. et al. From registration to publication: a study on Dutch academic randomized controlled trials. Res. Synth. Methods 11, 218–226 (2020).
Gallifant, J. et al. Peer review of GPT-4 technical report and systems card. PLoS Digit. Health 3, e0000417 (2024).
Biedermann, P. et al. Standardizing registry data to the OMOP Common Data Model: experience from three pulmonary hypertension databases. BMC Med. Res. Methodol. 21, 238 (2021).
Canham, S. & Ohmann, C. A metadata schema for data objects in clinical research. Trials 17, 557 (2016).
Hamosh, A., Scott, A. F., Amberger, J., Valle, D. & McKusick, V. A. Online mendelian inheritance in man (OMIM). Hum. Mutat. 15, 57–61 (2000).
Degtyarenko, K. et al. ChEBI: a database and ontology for chemical entities of biological interest. Nucleic Acids Res. 36, D344–D350 (2008).
Meehan, T. F. et al. Logical development of the cell ontology. BMC Bioinforma. 12, 6 (2011).
Brown, G. R. et al. Gene: a gene-centered information resource at NCBI. Nucleic Acids Res. 43, D36–D42 (2015).
Bodenreider, O. The unified medical language system (UMLS): integrating biomedical terminology. Nucleic Acids Res. 32, D267–D270 (2004).
Chen, J. et al. Towards medical complex reasoning with LLMs through verifiable medical problems. Findings of the Association for Computational Linguistics, 14558–14575 (ACL, 2025).
Smith, L. et al. Overview of BioCreative II gene mention recognition. Genome Biol. 9, S2 (2008).
Krallinger, M. et al. The CHEMDNER corpus of chemicals and drugs and its annotation principles. J. Cheminform. 7, S2 (2015).
Caufield, J. H. et al. A reference set of curated biomedical data and metadata from clinical case reports. Sci. Data 5, 180258 (2018).
Kim, J.-D., Ohta, T., Tsuruoka, Y., Tateisi, Y. & Collier, N. Introduction to the bio-entity recognition task at JNLPBA. In Proceedings of the International Joint Workshop on Natural Language Processing in Biomedicine and its Applications - JNLPBA ’04 70 https://doi.org/10.3115/1567594.1567610 (Association for Computational Linguistics, Geneva, Switzerland, 2004).
Li, J. et al. BioCreative V CDR task corpus: a resource for chemical disease relation extraction. Database J. Biol. Databases Curation 2016, baw068 (2016).
Wei, C.-H. et al. tmVar 2.0: integrating genomic variant information from literature with dbSNP and ClinVar for precision medicine. Bioinforma. Oxf. Engl. 34, 80–87 (2018).
Doğan, R. I., Leaman, R. & Lu, Z. NCBI disease corpus: a resource for disease name recognition and concept normalization. J. Biomed. Inform. 47, 1–10 (2014).
Dettmers, T., Pagnoni, A., Holtzman, A. & Zettlemoyer, L. QLoRA: efficient finetuning of quantized LLMs. Adv. Neural Inf. Process. Syst. 36 (2023).
Romanov, A. & Shivade, C. Lessons from natural language inference in the clinical domain. Preprint at https://doi.org/10.48550/arXiv.1808.06752 (2018).
Parthasarathy, V. B., Zafar, A., Khan, A. & Shahid, A. The ultimate guide to fine-tuning LLMs from basics to breakthroughs: an exhaustive review of technologies, research, best practices, applied research challenges and opportunities. Preprint at https://doi.org/10.48550/arXiv.2408.13296 (2024).
Abdallah, M., Georges, M. & Nikolski, M. TrialMatchAI: an end-to-end AI-powered clinical trial recommendation system to streamline patient-to-trial matching. cbib/TrialMatchAI. https://doi.org/10.5281/zenodo.18329084 (2026).
Acknowledgements
This work was supported by the European Union under the EOSC4Cancer project, funded by the European Research Executive Agency (REA) under the European Union’s Horizon Europe program (grant agreement ID:101058427; M.A., M.G., M.A.R., S.C., S.N., M.N., R.D., R.F., G.M., M.B., L.M., E.H., J.G., A.G., and S.K.). The grant supported research costs and personnel, including M.A. This work also benefited from access to the computing resources of the “CALI 3” cluster. This cluster is operated and hosted by the University of Limoges. It is part of the HPC network in the Nouvelle-Aquitaine Region in France, funded by the French government and the Region.
Author information
Authors and Affiliations
Contributions
M.A. developed the TrialMatchAI system and performed the main experiments. M.A. and M.G. conducted the benchmarking analyses. M.A.R. and S.C. designed the inclusion/exclusion criteria splitting methodology. M.A., S.N., and M.N. performed data analysis and interpretation. M.A., M.N., R.D., R.F., and G.M. contributed to the analysis and interpretation of the real patient’s cohort data, with M.B. and L.M. supporting curation and clinical context. J.G. assessed the clinical plausibility of the generated eligibility rationales. A.G. and S.K. contributed to system design choices and implementation discussions. E.H. contributed to scientific discussion and project oversight. M.N. supervised and directed the project. M.A. and M.N. wrote the manuscript. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Kirk Roberts, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Abdallah, M., Nakken, S., Georges, M. et al. TrialMatchAI: an end-to-end AI-powered clinical trial recommendation system to streamline patient-to-trial matching. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70509-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70509-w