Abstract
Vascular smooth muscle cells contribute to heritable coronary artery disease risk and undergo complex transitions to multiple disease-related phenotypes. To investigate the genetic basis of these trajectories, we develop a dense timecourse single-cell transcriptomic and epigenetic map of atherosclerosis in a murine disease model accompanied by high-plex in situ spatial data. Using temporal data and probabilistic fate modeling, we identify key transcription factors that drive cell state changes through a combination of network-based prioritization and in silico transcription factor perturbation. Parallel knockout studies of validated coronary artery disease gene Tcf21 uncover its molecular mechanisms in smooth muscle cell transition, due in part to a role regulating the transition of smooth muscle cells in the secondary heart field. Integrating the murine atlas with human coronary artery disease genetics pinpoint smooth muscle cell phenotypes that mediate disease risk, highlighting causal disease mechanisms. Together, these studies resolve atherosclerosis trajectories at single-cell resolution and identify genetic causal transcriptomic and epigenomic mechanisms of coronary artery disease risk.
Similar content being viewed by others
Introduction
Cardiovascular diseases, principally coronary artery disease (CAD) and stroke, are the worldwide leading cause of global mortality1. Therapies that modify classical environmental and metabolic factors have ameliorated a portion of CAD risk2, but more than half of the risk can be attributed to common inherited genetic variation by affecting vessel wall pathways that mediate disease pathophysiology. These genetic factors remain unidentified and untreated3,4,5,6. While extensive studies have investigated the cellular and molecular features of atherosclerosis, they have been unable to establish causality and thus hinder the translation towards vascular wall directed therapies7.
The smooth muscle cell (SMC) lineage, which appears to make the largest contribution to disease risk8,9,10, undergoes extensive complex phenotypic transitions that have not been well characterized at the cellular and molecular level, making it difficult to assign causality to specific gene programs11,12,13,14,15,16. The few SMC GWAS genes studied in detail suggest that complex transcriptional programs can direct cellular trajectories that lead to fibroblast-like (fibromyocyte, FMC) or osteochondrogenic (chondromyocyte, CMC) phenotypes, with these cellular phenotypes being proposed to mediate opposing effects on disease risk12,13,17. However, this paradigm is in conflict with genomic analyses of limited single-cell data suggesting that all FMC transition to CMC over the course of atherosclerotic lesion development9,12,18. Molecular and cellular studies of SMC transitions to human genetic data linking locus associations to individual SMC gene causality and direction of effect, are needed to promote progress in the field.
Studies reported here are aimed at the comprehensive characterization of genes and gene programs that function in SMC to mediate phenotypic transitions and disease risk. We describe a mouse transcriptomic and epigenomic atlas of atherosclerosis with single-cell RNA and chromatin accessibility (ATAC) data collected over a dense timecourse in a well-accepted mouse model, and further correlate SMC cellular molecular phenotypes with their corresponding spatial niche using high throughput in situ RNA hybridization (Xenium). We map the SMC lineage cellular trajectories with real-time course gene expression, chromatin accessibility data and advanced trajectory inference methods such as the Waddington Optimal Transport algorithm, identifying the genes and gene regulatory networks that mediate the transitions to FMC and CMC cellular phenotypes. Further, we investigate how these trajectories are affected by the CAD protective Tcf21 gene, mapping the regulatory network downstream of this gene, further identifying collaborating transcription factors (TFs) that mediate genome-wide regulation of SMC phenotype transitions and disease risk.
Results
Single-cell atherosclerosis timecourse atlas construction
We performed single-cell RNA sequencing (scRNAseq) and single-nucleus assay of transposase-accessible sequencing (scATACseq) with aortic root tissue from ApoE-/- (ApoE KO) mice to characterize the complex genetic regulatory networks that control the developmental cascade of smooth muscle cell (SMC) phenotypic transitions in the context of atherosclerotic stress13. These mice, also expressing a tamoxifen-inducible Cre recombinase driven by the SMC-specific Myh11 promoter and a Cre-activatable tdTomato reporter gene, were fed a high fat diet (HFD) across 7 timepoints for scRNAseq and 6 timepoints for scATACseq assays (Fig. 1A). Aortic root tissues were collected, digested, and subjected to droplet-based cell capture, and independent RNA and transposed DNA sequencing13. The resulting data were subjected to dimensionality reduction, clustering and visualization with Seurat, providing individual cell clusters that we and others have identified previously (Fig. 1B)11,12,13,14,15,16. For this study, we subsetted lineage traced SMC clusters, transferred scRNASeq labels onto scATAC cells and co-embedded RNA and ATAC modalities, resulting in a total of 96,195 high quality cells across both modalities (Fig. 1C, D, Supplementary Fig. 1A–K, and Supplementary Data 1, Methods)13,19.
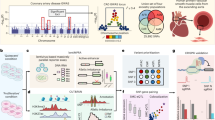
A Schematic of the single-cell RNA and ATAC sequencing data collection in the ApoE KO mouse atherosclerosis model. Created in BioRender. Li, D. (https://BioRender.com/22if1p3). Mice were tamoxifen gavaged at 8 weeks of age to induce Myh11-Cre recombination and tdTomato expression, single cells are isolated by flow cytometry and sorted on lineage traced and non-lineage traced cells. Mouse aortic root tissues were collected and scRNAseq assays performed on single cells isolated after 0, 3, 5, 7, 9, 12 or 16 weeks of high fat diet feeding. Remaining single cells from the same cellular samples underwent nuclei isolation and scATACseq assays at the 0-, 5-, 7-, 9-, 12-, and 16-week timepoints. (n = 23 scRNA libraries; n = 14 scATAC libraries, Supplementary Data 1). B UMAP embedding of all aortic root resident cells, with scRNAseq cluster assignment guided by cell-specific gene expression as we have described19. C UMAP of scRNA and scATAC clusters prior to integration. Confusion matrix showed cluster-cluster mapping between scRNA (x-axis) and scATAC (y-axis) cluster cell assignment data after CCA co-embedding. D UMAP visualization of the integrated object with scRNA and scATAC co-embedding. E–G Feature plots for specific marker genes Fbln1 (FMC-1), Fbln-2 (FMC-2), and Ibsp (CMC) in SMC. H–J Coverage plots of scATACseq open chromatin peaks for marker genes Fbln1, Fbln2, and Ibsp. K–M GSEA enrichment of cluster specific gene ontology biological process pathways for each modulated SMC cluster. N Representative slide image showing murine aorta tissue processed with Xenium Prime 5k gene expression panel. BCA – Brachiocephalic artery O Cropped image of ascending aorta with timecourse cell type labels transferred onto spatial data. P–S Image feature plot showing spatial distribution for specific marker genes Myh11 (SMC) Fbln1 (FMC-1), Fbln-2 (FMC-2), and Ibsp (CMC).
Defining molecular and spatial patterns in the SMC lineage
We expanded upon previously identified SMC cell states11,12,13,14,15,16,19 and identified six distinct SMC lineage phenotypes supported by both RNA and chromatin accessibility patterns (Fig. 1C, D, and Supplementary Fig. 1L, Supplementary Data 2). We observed high level expression of canonical SMC differentiation markers (Cnn1, Tagln, Acta2, Myh11) in three clusters (SMC-1, 2, 3). While SMC-1 primarily expressed these markers, SMC-2 demonstrated expression of markers such as Igfbp2, indicating an early phenotypic transition state20,21. A distinct SMC-3 population additionally enriched for myocardial markers Tnnt2 and Nkx2-5, identified the previously characterized population of SMC at the base of the aortic root arising from the secondary heart field(SHF)21,22. All of the phenotype clusters represented cells of the SMC lineage, as indicated by lineage tracing with the Myh11-Cre transgene (Supplementary Fig. 1H). It is important to note that recombination efficiency is quite high, ~95%, so cells expressing even low levels of Myh11 will activate the high-level expression of the tdTomato reporter, which does not really reflect endogenous Myh11 expression level that might reflect some heterogeneity in the SMC cell population.
All FMC were characterized by robust expression of fibroblast and epithelial mesenchymal transition (EMT) markers such as Vcam1, Lum, Bmp1 and Pdgfrb (Supplementary Fig. 1L). However, we identified two FMC sub-populations demonstrating discrete patterns of gene expression (Fig. 1E, F), chromatin accessibility (Fig. 1C, D, H, I), functional enrichment (Fig. 1K, L), and spatial localization (Fig. 1O–T, and Supplementary Fig. 2A–G). Taken together, these data demonstrate the integrity of each FMC molecular subtype. One group, termed FMC-1, expressed specific markers including Tcf21, Fbln1, and targets of interferon gamma signaling including H2-Aa/Ab1/Eb1, Cd74 and Il33 (Supplementary Fig. 1L). Validation with in situ RNA hybridization of cluster-specific FMC-1 markers demonstrated expression largely encompassing the media and sparsely at the fibrous cap (Fig. 1R, Supplementary Fig. 2E, and Supplementary Fig. 3A–C). Transcriptional functional enrichment with gene set enrichment analysis (GSEA) using GO biological process gene sets for FMC-1 cells identified processes such as immune activation and cytokine production (Fig. 1K, Supplementary Fig. 1L, and Supplementary Data. 3). The second cluster of cells, termed FMC-2, were identified with specific markers Thbs1 and Notch3 localized to the fibrous cap (Supplementary Fig. 3D, E) or spanning the intimal plaque (Col8a1, Fbln2, Ttc9) (Fig. 1S, Supplementary Fig. 2F, Supplementary Fig. 3F, G, H). Although FMC-2 shared a number of terms with FMC-1, expressed genes were uniquely associated with collagen fibril organization, regulation of lipid transport and wound healing (Fig. 1L, Supplementary Data. 3). Interestingly, a number of genes (Loxl1, Bmp1, Vcam1) were found to have vascular wall expression patterns encompassing those seen for both FMC-1 and FMC-2, possibly reflecting cells in transition (Supplementary Fig. 3I, J, K).
By comparison, the CMC cluster identified a distinct transition with a highly restricted chromatin accessibility pattern, and restricted expression of marker genes Col2a1 and Ibsp localized to the base of the plaque (Fig. 1G, J, Q, Supplementary Fig. 1L, Supplementary Fig. 2G, and Supplementary Fig. 3L, M) as we have shown previously13,17,19. CMC pathways were consistent with osteochondrogenic processes, as described by several groups11,12,13,14,15,16,19 (Fig. 1M, Supplementary Data. 3).
To translate these spatial findings from mice to human, we further performed label transfer onto publicly available scRNA data from human donor transplant coronary arteries and observed a similar distribution of FMC-1, FMC-2 and CMC by single-cell features and spatial transcriptomics23 (Supplementary Fig. 2H-M).
SMC fate trajectories identify cell transition gene programs
We utilized force directed layouts to visualize transcriptional relationships, in real-time, across the SMC atherosclerosis trajectories (Fig. 2A). We first observed the appearance of FMC-1 cells at 5 weeks of HFD with a subsequent increase in FMC-2. This observation is consistent with a migratory path for SMC lineage cells from the media to the fibrous cap and subsequently down into the intimal plaque24,25,26,27,28 (Fig. 1O, R, S, Fig. 2A, B, C, Supplementary Fig. 3A–H). Consistent with this possibility, FMC-2 were also enriched for osteoblast progenitor gene signatures (Fig. 1M, Supplementary Fig. 1L, and Supplementary Fig. 4A). Osteoblast progenitors have been characterized in bone development, and this module score29 determined with scRNAseq data characterizing these cells on their path toward osteochondroprogenitors. CMC abundance dramatically increased at 12 weeks of high fat diet, correlating with spatial localization at the plaque base (Supplementary Fig. 3L, M). In addition, transcriptomic patterns suggested increasing pathologic signatures including senescence, EMT, angiogenesis, apoptosis, and efferocytosis (Supplementary Fig. 4B–F, and Supplementary Data. 4) as the cells transitioned towards the CMC cell state at the plaque base, which is consistent with the acellularity in this region13,25.
A Force directed layout (FLE) representation of lineage traced SMC showing clustered cell phenotypes. B FLE embedding of lineage traced SMC split by weeks on high fat diet. C Cluster proportions of lineage traced cells split by week. D Sankey plot of Waddington optimal transport (WOT) predicted transition probability from SMC to FMC and CMC populations. Band thickness indicates relative transition probability from starting to end cell states. E FLE embedding of SMC lineage transition cell states with arrows representing WOT predicted movement of cells during phenotypic transition. F Illustrated representation of WOT predicted movement of cells during phenotypic transition. Created in BioRender. Li, D. (https://BioRender.com/22if1p3). G Transcription factor enrichment for cells fated to become each cluster-type calculated by WOT based on week 5 to week 12 transition. H CellRank2 derived predicted sustained cell states. I Optimal transport derived growth rates by cell state. J Pseudotime distribution of cells by week. K, L CellRank2 summarized expression trends across pseudotime (right) showing a representative activating gene cluster towards FMC-1 or FMC-2, TFs within this gene cluster (middle), and GO enrichment of cluster genes (left) for key genes.
To examine the translocation of FMC from media to cap to plaque, we performed additional studies to map temporal gene expression of FMC-1 and FMC-2 marker genes Fbln1 and Fbln2 across serial sections to highlight their distinct expression patterns. RNAScope performed at baseline showed that FMC-1 marker Fbln1 expression was seen in the media while expression of FMC-2 maker Fbln2 was not (Supplementary Fig. 3N). Then, at 7 weeks of high fat diet (HFD), there was enrichment of Fbln1 in the media and at lower levels at the cap, while Fbln2 was more intimally restricted with lower medial expression compared with FMC-1. (Supplementary Fig. 3O). These findings aligned with our single-cell data which showed low level FMC-1 but no FMC-2 marker expression at baseline while FMC-1 and FMC-2 expression increased after week 7 of HFD (Fig. 2A, B, C).
We further investigated the lesion cellular anatomy at 16 weeks HFD with high quality Xenium 5k geneset data for the mouse aorta. With transfer of single-cell RNA sequencing labels to the spatial clusters, we demonstrated that FMC-1 are enriched in the media and also localized in the cap after 16 weeks HFD. Subsequently high levels of FMC-2 were identified in the cap and spanning the intima to the CMCs at the base of the plaque, further validating our inference based on selected RNAScope markers (Fig. 1N-S, Supplementary Fig. 2A).
To quantitatively model these observed cell state changes, we applied the Waddington Optimal Transport (WOT) algorithm to build a probabilistic model for cell state transitions30,31. WOT is a heuristic method that models growth rates based on cell cycle and apoptosis gene expression to perform developmental trajectory inference. To visualize the WOT predicted relative transition probabilities and emphasize the actual proportion of each cell state at each time point, we have visualized the transition probability results with a Sanke plot (Fig. 2D). The exact proportion of each cell state at each time point can be observed in Fig. 2C and Supplementary Data. 5. Applying this algorithm across timepoints, we found that while SMC-1/2/3 are present at baseline (Week 0), SMC-2 are the primary cell phenotype that transitions to FMC states (Fig. 2D, E, F). WOT modeling suggested that a small FMC-1 population existed at baseline, significantly expanded by 5 weeks, and directly contributes to both FMC-2 and CMC. FMC-2 appeared to primarily become CMC which were evident by 12 weeks of HFD. However, given the higher proportion of FMC-1 present relative to FMC-2, both FMC cell states likely contributed similarly to the total number of CMC.
WOT further captured TF enrichment across cell state transitions, using weeks 5 and 12 as reference points which encapsulated the majority of the cell state transitions. This approach allowed us to highlight the crucial TFs expressed in SMC that guide these cells towards their final fates (Supplementary Data 6). We then filtered these transition TFs with significantly enriched accessible transcription factor binding motifs identified from scATACseq data. This integrative approach enabled us to identify TFs that have previously been associated with SMC phenotypic transitions, and in some cases pointed to previously unappreciated functions for these genes (Fig. 2G). For example, we found Tcf21 expression to be the most enriched TF in cells fated to become FMC-1, consistent with its early role in phenotypic transitions, while WOT-based TF enrichment also suggested a prominent role in the FMC-CMC state transition32,33. Across the enriched TFs found in our model, several patterns stand out. For example, TFs (Runx1/2, Zbtb7c, Zeb1/2, Smad3, Stat3, Hes1, Nfkb1, Cebpb) amongst this list have been readily associated in literature with AP-1, Klfs and the SMC regulatory Srf-myocardin complex. This suggests that pioneer factors and immediate-early AP-1 activation constitute a core early event that then allows for recruitment of transcription factors at specific enhancers to drive key cell state transition steps in conjunction with Srf-Myocardin. These core TFs then organize around central cellular processes such as TGFB signaling, hypoxia signaling, inflammatory response and osteochondrogenesis as summarized in Supplementary Data 734,35,36,37,38. Moreover, the top SMC-3 promoting TF was Isl1, reinforcing its SHF origins39.
The suggested trajectory paradigm is further supported by analyses with the RealTime kernel calculations in CellRank that computationally derive sustained states. This analysis recovered FMC-1, FMC-2, two CMC and the SMC-2 and SMC-3 as sustained states (Fig. 2H). Further, we visualized the WOT derived growth rates by cell state and found a greater proportion of cells with increased proliferation in the early transitioning SMC-2 cells, reduced growth rates in the FMC with FMC-2 having higher proportion of low growth rates and higher senescence score (Fig. 2I, Supplementary Fig. 4B), and lastly a higher growth rate in CMC, reflective of proliferative early-stage chondrocytes in developing bone40,41 (Fig. 2I). Given the high correlation between pseudotime and cell state stages confirmed by our real-time analysis (Pearson’s r = 0.57; spearman’s rho = 0.56, p < 2e-16) (Fig. 2J), we modeled the atherosclerosis trajectory with CellRank pseudotime kernel as an orthogonal approach to derive a probabilistic transition model.
Analyses using WOT and pseudotime suggested that both FMC-1 and FMC-2 phenotypes transition probabilities stabilize in mature lesions and are thus more likely to represent sustained phenotypes rather than a transition state. Complementary analyses with pre-existing 20 and 26 week high fat diet (ApoE and Ldlr-KO, respectively) atherosclerosis mouse scRNAseq datasets15 and brachiocephalic derived advanced lesions11 support the sustained existence of these phenotype cells into late stages of disease (Supplementary Fig. 4F–K). This is an attractive possibility, following the hypothesis that protective cells transition primarily to stable FMC and disease promoting cells undergo further transition to the CMC lineage. These findings are in contrast to previous pseudotime based analysis which have suggested that CMC are the common primary endpoint for all transitioning SMC in the plaque9,12.
The increased resolution provided by pseudotime allowed us to cluster gene expression across the different inferred trajectories, identify key TFs within trajectory gene clusters, and characterize pathways based on gene expression trends clustered over pseudotime. For example, we found that FMC-1 trajectory clustered genes demonstrate early activation of genes involved in inflammatory response and response to molecules of bacterial origin suggestive of a core set of genes involved in stress response including TFs such as Klf4, Cebpb, Runx1 and Nfkb1 (Supplementary Fig. 5A, cluster 3) followed by gene groups demonstrating epithelial cell migration and cell-substrate adhesion including TFs like Tcf21, Ar, Zeb2, and Twist1 (Fig. 2K) and more unique processes such as antigen processing and presentation and cytokine mediated signaling that includes TFs such as Ahr, Gas7 and Stat1 (Supplementary Fig. 5A, cluster 2). For FMC-2, we observed an early cluster enriched for epithelium migration and regulation of Wnt signaling (Fig. 2L, Supplementary Fig. 5B, cluster 3), while later clusters showed enrichment for cell chemotaxis, angiogenesis and collagen metabolic process (Supplementary Fig. 5B, clusters 1, 2). Similarly, for CMC, there were robust signals for biomineralization and ossification terms (Supplementary Fig. 5C).
Accessible chromatin reveals cell state specific DNA motifs
To characterize the epigenetic processes that mediate the noted transcriptional effects, we investigated chromatin accessibility in the transitioning cells with scATACseq data (Fig. 1C, D). We observed high specificity of chromatin accessibility within each label transferred tdTomato lineage traced cluster and visualized top specific peaks along pseudotime (Fig. 3A). Using GREAT for functional enrichment42, we identified specific cellular processes including smooth muscle related processes in the SMC, cell migration, inflammation and response to platelet derived growth factor in the FMC, and ossification and chondrocyte development in the CMC (Fig. 3A, Supplementary Data 8).
A Hierarchical clustering of top specific scATAC peaks arranged by cell states across 30 pseudotemporal bins. Top two bars show cell proportion by week and by cluster assignment, respectively. Heatmap displays min-max scaled mean chromatin accessibility for each SMC cell state. Representative biological processes from GREAT pathway enrichment for each cell state is displayed on the right. B Hierarchical clustering of ChromVar scores (motif accessibility deviation from background) for cluster enriched motifs across 30 pseudotemporal bins. C Representative schematic of network inference. TF binding through binding site-motif matching within accessible chromatin regions are linked to genes. The expression of TF target genes were then fitted with a bagging regression model to generate TF-target gene networks. Created in BioRender. Li, D. (https://BioRender.com/22if1p3). D Summarized PANDO predicted TF network represented with UMAP embedding. Color gradient and size represent the expression-weighted pseudotime and centrality scores, respectively. E. Differential TF activity between SMC (salmon) and modulated SMC states (purple). TF activity is calculated by multiplying the mean regulatory coefficient of each TF network with its respective average TF expression. Sign of the activity represents inferred net regulatory effect – activating (+) or repressing (-). F, G Ranked perturbation scores (product between control differentiation and KO-simulation vectors) for systematic KO simulation effect of TFs on the SMC and FMC lineages. Higher scores predict decreased likelihood of phenotypic shift from the originating cell state after TF perturbation. H, I Rank of top TFs predicted to affect the SMC to FMC or FMC to CMC transition using 1) CellOracle in silico perturbation scores, 2) Optimal Transport derived fated cell TF enrichment, or 3) aggregated ranking across both methods.
To prioritize dynamic TF binding motifs, we applied ChromVar to calculate their motif binding accessibility probability distributions. We filtered these data using a core set of overrepresented TF motifs identified by Signac and HOMER and visualized motif deviations along pseudotime to capture the dynamic shifts of TF binding activity (Fig. 3B). We used the early pseudotime bins 3-24 (PseudoEarly) and late pseudotime bins 25-30 (PseudoLate) to approximate the SMC-FMC and FMC-CMC transitions (Fig. 3A). These analyses identified temporal trends for both known and previously unrecognized regulators of smooth muscle fate. For example, Srf, Mef2, and Zeb motif deviation was observed to be higher in SMC13,43. In addition, for the early transitions, we observed a number of TFs with the greatest motif accessibility across the SMC stages, including Tead, Zbtb7c, Meox2, Rfx, and Nfi factors. Factors showing greatest motif accessibility during the late transition stages included Tcf21, Smad3, Rbpj, Stat, Nfatc, Cebpb, and Runx factors, with many of these known to be involved in endochondral bone formation. KLF factors and numerous AP-1 factors showed a bimodal pattern with accessibility early in the quiescent SMC state and later, from FMC to pre-CMC bins, likely representing their pioneer factor functions at these differentiated cell states. These analyses summarized the chromatin landscape guiding TF pathways, extending previous studies using pseudotime data derived from baseline and sustained phenotypic cell states13 (Fig. 3B).
Network prioritization identifies top SMC-FMC transition TFs
Using our co-embedded scRNA and scATAC dataset, we leveraged complementary network inference methods, Pando44 and CellOracle45, to create a custom workflow to infer transcription factor-target interaction networks (gene regulatory networks, GRNs) that direct phenotypic transition and simulate cell identity changes with in silico TF perturbations (Fig. 3C, and Supplementary Data 9,10, Methods). The dataset was divided by PseudoEarly and PseudoLate bins to infer GRNs for these analyses. We visualized a summary GRN colored by the average pseudotime-TF expression to reveal a cascade of network activation (Fig. 3D). SMC lineage phenotypes clustered separately, identifying numerous TFs that are likely to direct specific regulons that mediate cell states and transitions characteristic of the response to disease stresses (Supplementary Data 11). We computed TF activities for the PseudoEarly and PseudoLate SMC states, and a comparison of TF activity change from SMC to transitioning SMC cell state activity revealed patterns such as shifts in hypoxia inducible factor (HIF) activity highlighted by the known interaction between Hif1a and Epas146 and other factors such as Ahr that interact with HIF through the common heterodimer Arnt47 (Fig. 3E).
We then performed systematic in silico perturbations using our inferred TF-target links from SMC-FMC and FMC-CMC transitions to calculate perturbation scores measuring the potential of TFs to drive cell transition away from the originating cell state. In SMC, we identified enrichment of EMT factors such as Tcf21 and Zeb factors, and similar to the top TF activity changes, we identified enrichment for hypoxia inducible factors including Hif1a, and Epas1, reflecting their high network connectivity in early phenotypic transitions (Fig. 3F). For factors promoting change to CMC phenotypes, we identified known factors such as Klf411 and Runx1/248,49, in addition to unique factors such as Trps1, Sox6, Erg, Zbtb7c, Prrx1, Arntl2, and Snai1 that were nominated to drive the differentiation of cells from FMC to CMC (Fig. 3G).
We further created an aggregated TF transition ranking by combining normalized enrichment from WOT for core TFs promoting the FMC-1 and CMC fates and normalized SMC-FMC as well as FMC-CMC perturbation scores from CellOracle (unfiltered WOT and CellOracle heatmaps in Supplementary Fig. 6A–C). Through this combination of cell fate enrichment and network-based prioritization analyses, we found Ar, Zeb, Smad, Mecom, Prrx1, and Cebpb factors, as well as Tcf21, to be among central enriched TFs involved in the phenotypic transition from SMC to FMC (Fig. 3H, and Supplementary Fig. 6B), and Runx, Klf, Egr2, Sox9, and Zbtb7c as top drivers for FMC to CMC transition (Fig. 3I, and Supplementary Fig. 6C). Additionally, Runx and Zbtb7c factors were consistently highly ranked across both transitions.
To validate our analytical algorithms, we selected three highly ranked transcription factors predicted from our CellOracle analysis to affect the transition process, AR, EPAS1 and ZEB1and performed siRNA knockdown followed by qPCR for contractile markers including ACTA2 and TAGLN as well as FMC markers LUM and PDGFRB (Supplementary Fig. 7A–C). Knockdown of these genes provided robust evidence for increased contractile marker expression across these conditions. Furthermore, we also observed significant downregulation of LUM and PDGFRB for EPAS1 and ZEB1 knockdown while AR knockdown showed a smaller effect size but a trend towards decreased expression levels of these FMC markers. Together, this provides additional functional validation of our predictive modeling.
To provide functional validation of the integrity of the FMC-1 versus FMC-2 cellular phenotypes, we characterized cell expression signatures from primary human coronary artery smooth muscle cells (HCASMC) derived from different donors previously published in Liu, et al.50. Using top marker genes from our single-cell mouse timecourse, we performed non-negative least squares cellular deconvolution and identified cell lines 2105 and 1508 as exhibiting the top ‘FMC-1-like’ and ‘FMC-2-like’ cell states, respectively (Supplementary Fig. 7D–G). We then treated these primary cell lines with calcium phosphate media14 for seven days to simulate an osteochondrogenic CMC-like transition phenotype. Quantitative PCR identified a greater increase in CMC genes including RUNX2, SOX9, and HAPLN1 in the 1508 cell line, whereas there was minimal change in expression of these genes in the 2105 line (Supplementary Fig. 7H, I). This is in keeping with our observations that the FMC-2 population has a greater probability of transition to the CMC state. Furthermore, this important finding suggests that there are functional differences in the primary cell lines depending on their origin/disease state and they may harbor differential ability to respond to various simulated vascular stresses. Identification of these primary HCASMC lines with FMC-1 vs FMC-2 phenotype will aid future studies of molecular differences between these cell states.
Tcf21 loss significantly alters SMC transition probabilities
As a top TF predicted to direct SMC to FMC transition, we further examined the effect of exemplar CAD gene Tcf21 knockout on SMC trajectories with updated single-cell chemistries using the timecourse methods employed for the control dataset to elucidate Tcf21 SMC regulatory mechanisms that occur with vascular stress.
We first examined changes in SMC transition cell numbers that resulted from Tcf21-KO. In keeping with our prior work16, there was a significant decrease in the FMC populations at 12 and 16 weeks of HFD (Fig. 4A). Comparison of lineage-traced SMC proportions revealed a marked ~3-fold increase in the SMC-3 Tcf21-KO cells at 5 weeks that persisted to week 12. These Tnnt2-expressing cells have been lineage traced to the secondary heart field21,22, suggesting an early expansion of this aortic base contractile medial compartment in the context of Tcf21 loss. The possible involvement of Tcf21 in the regulation of these cells was supported by its expression in Nkx2-5 lineage traced cells as identified with scRNASeq (Supplementary Fig. 8A–E). At 12 weeks, there was a notable relative decrease in Tcf21-KO FMC-1, FMC-2 and CMC proportions that corresponded with a relative increase in SMC-2 and SMC-3 cluster proportions, suggesting that the Tcf21-KO cells were halted in disease associated transitions. At 16 weeks, there was a consistent decrease in FMC-1 without a decrease in FMC-2, suggesting potential compensation for the loss of Tcf21 expression in the SMC lineage cells over time (Fig. 4A). At both 12 and 16 weeks we also observed a decrease in CMC cell proportion in the Tcf21-KO relative to control, suggesting that Tcf21 directly promotes CMC development as suggested by our previous analyses (Figs. 2G, 3I).
A Comparison of cluster proportions in lineage traced SMC from control and Tcf21-KO mice matched by week of diet exposure. (n = 8 scRNA libraries; n = 9 scATAC libraries, Supplementary Data 1). B Sankey plot of control (top) and Tcf21-KO (bottom) WOT predicted transition probability from SMC to FMC and CMC populations. Band thickness represents relative transition probability from starting to end cell states. C Heatmap of hierarchically clustered significantly differentially expressed genes between control and Tcf21-KO cells in the PseudoLate bins. D GSEA (GO-biological process gene sets) of differentially expressed genes between control and Tcf21-KO cells in the PseudoLate bins. P-value is obtained using a running-sum statistic with gene label permutation. Size and color represent overlapping gene counts and enrichment FDR q-value, respectively and x-axis shows GSEA enrichment scores. E Over-representation analysis of observed Tcf21-KO DE genes overlap with the PANDO-predicted Tcf21 TF network, two-tailed Fisher’s exact test P < 0.0001. F Network showing predicted target TFs of Tcf21, observed Tcf21-KO effect on expression (edge, color), and network centrality of these target TFs (size). G GO-BP enrichment of Tcf21 target TF networks. H Correlation for single-cell WOT fate scores for FMC-1, FMC-2 and CMC fates with module scores for each Tcf21 target TF network. Color represents TF expression directionality with Tcf21-KO. Control correlation of TF expression only with WOT fate scores in gray. I Direct Tcf21 network summarized in context of pseudotime. Created in BioRender. Li, D. (https://BioRender.com/22if1p3).
WOT was used to investigate alterations in SMC trajectory transition probabilities in the context of Tcf21-KO (Fig. 4B). Focusing on cell state changes in the SMC lineage traced cells, there was a notable decrease in the overall transition probabilities across cell states for Tcf21-KO. The SMC-2 contribution to both FMC-1 and FMC-2 was decreased in KO diseased tissues, as was the FMC-1 contribution to the FMC-2 phenotype, and there was a low level of transition for FMC-1 and FMC-2 to CMC (Fig. 4B). Overall, there was evidence for a dramatic decrease in probability of transition to the CMC phenotype while transition to both the FMC-1 and FMC-2 phenotype was decreased but sustained. These decreased probabilities for SMC to FMC and FMC to CMC transitions in knockout mice accounted for the changes in SMC phenotype cell numbers but also highlighted the presence of compensatory processes which allowed continued phenotypic transition (Fig. 4A). As noted previously, at 16 weeks, the FMC-1 and FMC-2 states remained, suggesting that they represent a sustained phenotype rather than exclusively a transition to the CMC phenotype.
Tcf21-KO DEGs enrich for predicted network and CAD genes
Analysis of differentially expressed genes (DEGs) in the Tcf21-KO compared to control mice using DESeq2 provided insight into associated gene programs. DEGs for the FMC-CMC transition for Tcf21-KO versus control, identified 965 genes (Supplementary Data 12, 13). This list was enriched for TGFB family genes such as Ltbp1 and Tgfb2, and numerous CAD GWAS genes including Tgfb1, Zeb2, Lrp1, Palld, Col4a2, Lmod1, and Pdgfd (Fig. 4C). These DEGs were further analyzed with GSEA using GO-BP terms to gain insight into altered pathways and directionality of effect. We found Cellular response to Tgfb stimulus and actin filament bundle organization to have high positive DEG enrichment, consistent with Tcf21 promoted effects and increased representation of locomotion and wound healing possibly representing compensatory processes given the overall decreased phenotypic transition phenotype (Fig. 4D). Terms with a negative enrichment score (average downregulation with Tcf21-KO) were identified as suppressive for connective tissue development and endochondral bone morphogenesis, further suggesting that Tcf21 target genes likely have a role promoting the CMC phenotype.
SMC lineage cell state changes and related cellular trajectories can be modeled through gene-gene interactions in a GRN. To build a Tcf21 specific network we employed the regulatory network predicted from our PseudoLate control timecourse which spanned the majority of Tcf21 activation in transitioning SMC. Moreover, network targets by this method are more likely to represent direct interactions given the conditional need for epigenetically accessible Tcf21 binding sites to be linked to target genes. We compared Tcf21 target genes identified with this network analysis with Tcf21-KO DEGs and found significant overlap of DE genes and network genes, with 135 of 318 predicted network genes (42%) also showing differential expression with Tcf21-KO (Fishers exact test p < 1e-4) (Fig. 4E). Further, we applied GSEA using the predicted Tcf21 network gene set ranked by TF-gene regulatory coefficient weights and observed a significant normalized enrichment score (NES = 1.24, p = 0.031) of Tcf21-KO DEGs. These analyses showed congruence between distinct approaches and validated the utility of using predictive transcriptional modules from a comprehensive control dataset to infer perturbed pathways.
We leveraged the predictive pathway ability of inferred networks to augment the functional characterization of the Tcf21 signaling network. We selected differentially expressed TFs within the Tcf21 GRN and integrated these TF-centered GRNs to create a validated Tcf21-TF sub-network and find both repressed and activated TFs that are regulated by Tcf21 (Fig. 4F). We identified downstream TF networks with differential KO module scores and performed functional enrichment on this subset of TF networks to visualize the pathways affected. We found multiple TFs including Foxp2, Mecom, Zeb2, Meis2, Mecom and Tshz2 predicted to be involved in cell migration, angiogenesis and extracellular organization processes, while genes such as Meis1, Zeb2 and Foxp2 were also involved in contractile processes (Fig. 4G).
We computed TF fate correlations by comparing WOT transition probability with cell-level TF module scores and using TF-only expression as a control comparison (Fig. 4H). Interestingly, top modules that exhibit high correlation with FMC fates also have their central TF upregulated with Tcf21-KO, suggesting compensatory roles alongside Tcf21 given our observation of overall decreased FMC proportions. For example Zeb2, previously shown to drive phenotypic transition13, was increased in Tcf21-KO, and its later average TF-pseudotime suggested that its role is downstream of Tcf21 towards the FMC-2 fate. In contrast, TFs such as Meis1/2 have earlier average TF-pseudotime, suggesting compensatory upregulation possibly via feedback mechanisms. Conversely, top TF modules correlated with the CMC fate, including the known ossification regulator Sox9, showed decreased expression upon Tcf21-KO (Fig. 4I).
TCF21-TEAD epigenetic interactions modify CAD GWAS genes
Previous studies have shown that Tcf21 interacts with histone deacetylases to broadly alter the in vitro epigenetic landscape of human coronary artery SMC (HCASMC)51 but its effects in vivo have not been explored. We observed widespread motif accessibility deviations upon Tcf21-KO when visualizing differential ChromVar scores across pseudotime and generated differential ChromVar scores on a by cluster basis (Fig. 5A, and Supplementary Data 14, 15). We further used Jensen-Shannon divergence (JSD) scores to identify differentially deviated TF motifs across pseudotime between control and Tcf21-KO ChromVar distributions, finding significant differences in many core TF motifs including Zeb, Klf, Runx1/2, AP-1, Srf, and Ctcf, while Tcf21 was borderline significant (padj = 0.10) (Fig. 5B). TCF21 HCASMC ChIPseq was reprocessed from Zhao et al.52 and showed significant enrichment in TEAD and CEBP TF binding motifs within TCF21 peaks (Fig. 5C, and Supplementary Fig. 9A) in addition to TCF21 and AP-151. When these ChromVar scores were visualized along pseudotime in the mouse timecourse, there was enhancement of Tead1 motif accessibility with loss of Tcf21 while Cebpb shared a pattern similar to Tcf21, showing decreased accessibility with Tcf21-KO (Supplementary Fig. 9B). We performed additional ChIPseq in HCASMC for TEAD1 and confirmed significant overlap with TCF21 (Fig. 5C, Fisher’s exact test p < 1e-4). Pathway analysis of shared peaks using GREAT predicted shared biological functions related to inflammation, apoptosis, TNF, and TGFB (Fig. 5D). Peaks shared by TCF21 and CEBPB were enriched for cell adhesion, ERK signaling, and cell motility terms (Supplementary Fig. 9C, D, Fishers exact test p < 1e-4). Genome wide colocalization of TCF21 and CEBPB/TEAD1 motifs was identified by comparing the location of CEBPB/TEAD1 motifs in TCF21 ChIPseq peaks and TCF21 motifs in TEAD1 peaks (Supplementary Fig. 9D, E). Interestingly, performing enrichment analysis for GWAS SNPs at TCF21-TEAD1 co-bound loci using ChIPseq binding data and GWAS SNP localization with the GWASAnalytics package, we found that all TCF21 binding sites, including those with TEAD1 peaks, showed high level enrichment for CAD (-log p-value 10) (Fig. 5E, and Supplementary Fig. 9F), but in the absence of TEAD1 peaks showed only low level enrichment for cancer and metabolism (Supplementary Fig. 9I). Also, TEAD1 peaks including TCF21 peaks showed strong enrichment for hypertension (-log p-value 10), while TEAD1 peaks only showed low level enrichment of SNPs for diabetes and hypertension (Supplementary Figs. 9G, H). Taken together, these data suggest that the interaction of TCF21 and TEAD1 plays a significant specific role toward CAD risk. This TCF21-TEAD1 relationship corresponded with murine trajectory analysis nominating Tead1 as an FMC-2 driver (Fig. 2L) and making it an intriguing Tcf21-interacting partner for further study. These results for TCF21-TEAD1 were contrasted with similar studies for CEBPB where inclusion of TCF21 loci detracted from the CEBPB metabolism signal (Supplementary Fisg. 9J, K). Analyses for HNF1A peaks served as negative control for these studies (Supplementary Figs. 9L, M).
A Hierarchically clustered heatmap of differential ChromVar scores (Tcf21-KO vs. control) labeled with selected representative TF motifs B Jensen-Shannon divergence of observed Tcf21-KO vs. control ChromVar scores showing motifs demonstrating significant probability divergence between conditions. Control JSD is calculated by randomly splitting control dataset into two equal distributions and calculating the JSD between these distributions. C ChIPseq peak overlap between TCF21 and TEAD1. D Representative pathways identified from shared ChIPseq peaks using Genomic Regions Enrichment of Annotations Tool (GREAT). P-values obtained using binomial test with Bonferroni correction. E GWASAnalytics analysis of GWAS SNP (EMBL-EBI GWAS Catalog) enrichment of overlapping ChIPseq peaks between TCF21 and TEAD1 showing highest enrichment for CAD traits. P-values obtained using binomial test with FDR q-value. F Proximity ligation assay showing nuclear fluorescent signal enrichment in the TCF21 + TEAD1 antibody group compared with TCF21+IgG control (Scale bar 170 µm). PLA experiments were performed twice, independently. G Co-IP showing TCF21-Myc immunoprecipitation followed by western blot for TEAD1. *denotes empty spacer. Co-IP experiments were performed twice, independently. H GSEA(using GO-biological process gene sets) of differentially expressed genes between siTEAD1 and control in HCASMC. P-value is obtained using a running-sum statistic with gene label permutation. Size and color represent overlapping gene counts and enrichment FDR q-value, respectively. The x-axis shows GSEA enrichment scores. I–K Dual luciferase assay for overlapping regulatory regions SRF Intron (n = 4 for test conditions, n = 2 for control conditions – see Statistics and Reproducibility), BMP1 Intron (n = 3, per condition), and LOXL1 5’ UTR (n = 3, per condition) for TCF21 and TEAD1 demonstrating competitive repression and epigenetic fine-tuning of enhancer/promoter activity between regulatory elements. P-values are ChIPseq data shown as normalized reads (bins per million) per condition. Luciferase data are presented as mean values +/- SD and p-values obtained using one-way ANOVA with FDR multiple comparisons post-hoc test. Source data are provided in the Source Data File.
We further examined the shared genomic patterns between TCF21 and TEAD1 by partitioning their shared binding loci into TCF21 + TEAD1 or TCF21 only loci. We observed greater enhancer profiles for TCF21 + TEAD1 shared binding sites compared to TCF21 only, as indicated by overlap with H3K27ac (Supplementary Fig. 9N). TCF21 + TEAD1 shared binding sites were located farther from the TSS regions (Supplementary Fig. 9O), consistent with enhancer co-localization. Further, pathway analysis of putative genes identified in TEAD1/TCF21/H3K27 regions by GREAT revealed enriched pathway keywords including differentiation, development and endopeptidase regulation while TCF21-only+H3K27 region genes showed enrichment for immune, viral and neutrophil related keywords (Supplementary Fig. 9P, Q).
We then investigated physical and functional interaction of these two TFs. Proximity ligation assays found that TCF21 and TEAD1 co-localized in the nucleus, suggesting direct protein-protein interaction (Fig. 5F). To detect a direct physical interaction between TCF21 and TEAD1, we performed Co-IP using a myc-tagged TCF21 transfected into HEK293 cells (Fig. 5G). We performed nuclear protein extraction followed by IP for the MYC-tag and western blot for TEAD1. These studies included IgG negative control and 5% input positive controls and provided evidence for TCF21-TEAD1 physical interaction. Further, immunohistochemistry in mice aortic root atherosclerosis sections also demonstrated intimal expression of Tead1 and Tcf21 (Supplementary Fig. 10A–C).
Given these findings, we investigated the expressed gene programs in SMC directed by TEAD1 by performing TEAD1 knockdown with siRNA in HCASMC along with bulk RNAseq. The transcriptomic changes with TEAD1 knockdown showed strong enrichment for processes such as cellular response to cAMP, extracellular matrix organization, connective tissue development, and chondrocyte development (Fig. 5H). These findings are in line with what we observed from our TEAD1 ChIPseq functional enrichment, further corroborating the hypothesis that TEAD1 plays a major role in the smooth muscle transition process to affect the development of phenotypically transitioning SMC.
Finally, to examine the functional interactions between TCF21 and TEAD1, we performed dual luciferase reporter gene transfection assays with A7r5 rat smooth muscle cells on a shared enhancer residing in an intron of the SRF gene, a master regulator of lineage contractile gene expression35, and two additional enhancers in CAD loci encoding ECM effectors of TGFB signaling, BMP1 and LOXL1. For the SRF enhancer, we showed that normal activation by SRF binding partner MYOCD was highly suppressed by TCF21 and to a greater degree by TEAD1 alone (Fig. 5I). There was an intermediate reporter activity when TCF21 and TEAD1 were both transfected, suggesting a competitive interaction between these TFs. Also, for the BMP1 and LOXL1 enhancers, both TFs showed repressor activity, but again, intermediate suppression when both were expressed in the same cells, suggesting competition (Fig. 5J, K). Taken together these data suggest that TCF21 and TEAD1 directly interact at shared regions across the genome to epigenetically regulate transcription.
Tcf21 mediates SMC CAD genetic risk via early rewiring
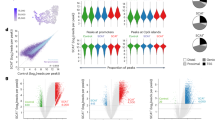
Single-cell methods have previously nominated genetic risk signals to have a unique high enrichment in SMC8. We investigated the relative disease related significance of our SMC cell states, using the scDRS algorithm5,53. At the single-cell level, we leveraged scDRS to integrate gene expression and GWAS gene z-score weights from MAGMA to generate disease relevance scores for each cell type. Because scDRS quantifies risk gene enrichment but not directionality, we also computed GWAS risk-weighted average expression by taking average individual gene expression multiplied by its scDRS gene weights and further inferred directionality using updated heritability adjusted S-PrediXcan modeling54 (Supplementary Data 16). Using this framework, we identified FMC-2 having overall statistical enrichment for excess CAD-associated risk gene expression by scDRS (Fig. 6A). Interestingly, neither the transcriptionally similar FMC-1 nor the calcification-associated CMC exhibited significant scDRS enrichment. Next, from the aggregated GWAS risk-weighted expression analysis, of genes with available predicted risk directionality, we calculated greater averaged expression of CAD risk genes in CMC relative to FMC and SMCs. Consistent with this complex biology, FMC-2 cells express genes that both promote or suppress CAD risk with the net predicted risk direction more protective, prompting further investigation of distinguishing expression patterns of these transition phenotypes.
A Scatter plot of CAD GWAS risk-weighted average expression of disease genes based on MAGMA gene weights and PrediXcan estimated directionality versus GWAS gene enrichment significance calculated by scDRS for each SMC cell state. P-value obtained from scDRS Monte Carlo test with dotted line representing FDR q-value = 0.05. B Scatter plot of individual TF and scDRS score expression correlation across pseudotime. Color represents averaged scDRS gene rank for TF gene modules and size represents overlap with GWAS genes. C Change in scDRS significance per cluster with Tcf21-KO. P-value obtained from scDRS Monte Carlo test with dotted line representing raw p-value = 0.05. D Summarized Tcf21 PseudoEarly DEG network filtered for putative GWAS genes and TFs. Color scale and node shape indicate differential expression and gene type, respectively. E GSEA(GO-biological process gene sets) of differentially expressed genes between Tcf21-KO and control SMC in the PseudoEarly bins. Size and color represent overlapping gene counts and enrichment q-value, respectively. The x-axis shows GSEA enrichment scores. F–K Gene expression feature plots and correlation of arterial tissue eQTL NES to GWAS odds risk for overlapping single nucleotide polymorphisms of selected Tcf21-KO differentially expressed genes from PseudoEarly bins with shared (F, G) or opposing (H–K) expression-disease risk directionality with Tcf21. Error bands in eQTL correlation plots represent linear regression 95% confidence interval bands.
We used the TF networks identified from the control timecourse (Methods) to ask how individual TFs and their networks associate with CAD risk. First, we synthesized full TF network modules by assimilating all unique target connections from PseudoEarly and PseudoLate predicted regulatory networks for available TFs. Second, we extracted gene-level scDRS weights to derive normalized TF-scDRS correlations. Third, we averaged gene-scDRS correlation ranks of all genes within each TF network module to create a TF network average scDRS rank. At the network level, we observed TFs with higher average scDRS rank in the SMC to FMC pseudotime such as Tcf21, Nkfb1, Runx, Zeb, Hif and Smad factors (Fig. 6B, Supplementary Data 17). Many of these core TFs also overlap with in silico TF perturbation predictions (Fig. 3F, G). Moreover, this method nominated novel TFs with network enrichment of CAD risk genes that were postulated to drive the SMC-FMC transition, including Arntl, Prrx1, Tshz2, and Mecom (Supplementary Data 8) or the FMC-CMC transition such as Trps1, Zbtb7c, Snai1, and Sox5/6/9.
To further dissect the regulatory relationships of CAD GWAS genes in phenotypic transition, we examined the scDRS enrichment in Tcf21-KO and found an increase in SMC-3 scDRS z-score, meeting nominal significance (Fig. 6C). This shift in scDRS score implicated the ability of Tcf21 to coordinate CAD GWAS genes early in the phenotypic transition timeline. In addition, this observation was in agreement with Tcf21 expression showing SMC-3 enrichment (Fig. 1C) and demonstrating a basal level of accessible Tcf21 TF binding sites in early pseudotime (Fig. 3B). We then examined the DEGs in PseudoEarly bins and observed significant overlap with a curated set of putative GWAS genes (63/640, p < 0.0001) (Fig. 6D). GSEA enrichment of DEGs not only revealed similar increased contractile processes with Tcf21-KO but also highlighted a decreased response to stress signals such as ‘cytokine stimulus’ and ‘unfolded protein response’ (Fig. 6E). Integrating these genes with human genetic signals, we focused on the differential expression of genes that shared GWAS disease risk-eQTL correlation with Tcf21. For example, increases in proliferative factors such as PDGFD and SEMA3C or decreases in inflammatory transcription factor STAT3 were associated with increased CAD risk (Fig. 6F–H). Conversely, there was also an enrichment of genes that were correlated with decreased CAD risk, such as LRP1, which has multifaceted coronary disease implications, COL4A2 which promotes basement membrane integrity or MYO9B that modulates vascular wound repair (Fig. 6I–K). Together, these relationships suggest a broad role for TCF21 in promoting risk through coordinated regulation of CAD GWAS genes in the phenotypic transition of disease SMC.
Discussion
We have conducted a comprehensive single-cell study to investigate the molecular trajectory of SMC phenotypic transitions during atherosclerosis using a combination of multi-modal single-cell sequencing at multiple timepoints, in situ hybridization and spatial transcriptomics to identify SMC phenotype niches, and disease phenotype trajectory modeling. These data provide transcriptomic, epigenomic and cellular lesion anatomical data characterizing two different FMC populations. FMC-1 arise first by 5 weeks of diet exposure, expresses inflammatory and immune markers while FMC-2 accumulation accelerates weeks later and is characterized by extracellular matrix, lipid handling, osteoblast progenitor expression profiles and a greater correlation with contractile marker expression compared to FMC-1. Trajectory modeling suggested FMC-1 contribute to FMC-2, but both phenotypes arise primarily from a modulating group of cells that maintain classical SMC contractile marker expression. FMC-1 are localized to the media and to the fibrous cap, suggesting their involvement in migration, while FMC-2 are identified primarily at the fibrous cap and intimal plaque. Both FMC contribute to CMC transition cells and are likely the sole source of these endochondral bone-like phenotype cells.
Our comprehensive multi-omic dataset has enabled the profiling of TF motif accessibility gradients across time and leveraged these data to generate regulatory networks, prioritize key driver genes and evaluate their functional pathways. Analyses using both WOT and pseudotime suggested that both FMC-1 and FMC-2 phenotypes transition probabilities stabilize in mature lesions and are thus more likely to represent sustained phenotypes rather than a transition state. Complementary analyses with pre-existing 20 and 26 week high fat diet atherosclerosis mouse scRNAseq datasets15 as well as brachiocephalic advanced lesion analysis11 support the sustained existence of these phenotype cells to late stages of disease. This is an attractive possibility, following the hypothesis that protective cells transition primarily to stable FMC and disease promoting cells undergo further transition to the CMC lineage. We further integrated trajectory analysis and in silico TF perturbation to identify critical factors which establish cell state identity through their ability to physically access genomic regulatory regions. For example, our aggregated SMC to FMC transition analysis (Fig. 3H, I) showed extensive overlap of top factors known to affect SMC phenotypic transition to FMC or CMC such as Tcf21, Ar55, Runx1/248,56, Zeb213, Smad312, and Klf411 while also nominating TFs such as Arntl, Mecom, Prrx1, Trps1, Zbtb7c, and a variety of HIF-related factors that participate in divergent functional roles for future study.
Among the prioritized TFs, Tcf21 emerged as a compelling candidate given its top rank in predicted effects on phenotypic transition as well as our previous work identifying it as a causal CAD GWAS gene and providing human genetic evidence for its CAD risk inhibition16. First generation scRNAseq studies have shown that Tcf21 loss was associated with decreased fibroblast-like SMC lineage cells that we termed fibromyocytes and histology showed decreased SMC migration from the media and decreased contribution to the fibrous cap. Therefore, our focused timecourse single-cell study in the Tcf21-KO mouse model with enhanced scRNAseq chemistry, greater transcriptomic depth, and linked chromatin accessibility data allowed further analysis of cellular trajectories and phenotypic transitions altered by Tcf21 loss. These analyses identified novel TF networks directly altered with Tcf21-KO as well as interacting epigenetic factors such as Tead1 which together with Tcf21, antagonize the differentiated SMC cell-fate and fine-tune cellular TGFB response.
We also found Tcf21 to be enriched in SMC-3 cells that emanate in part from the secondary heart field (SHF), and the Tcf21-KO mouse showed a dramatic 3-fold expansion in the Tnnt2 expressing SMC-3 cells after only 5 weeks of diet. This observation suggests that Tcf21 suppresses transition, and possibly migration, of cells from this region. SMC derived from the SHF give rise to the proximal aortic wall and to the adjacent outflow tract and exhibit well-recognized embryonic lineage specific responses to critical signaling pathways43 such as TGFB57, PDGFD58, and NFkB59. Further, increased Tcf21 expression was identified after disease initiation in the FMC-1 where its expression was noted to be inversely correlated with expression of Acta2 and other contractile markers, consistent with our previous findings that Tcf21 suppresses SMC lineage marker genes through direct transcriptional mechanisms that block MYOCD-SRF mediated transcription of lineage markers35.
We and others have used CAD GWAS findings along with gene expression or chromatin accessibility data to show that much of the risk for CAD resides in the coronary vascular SMC lineage5,8,9,53. Our high-resolution dataset extends upon previous observations and identifies FMC-2 as the cell state harboring expression of genes that mediate CAD risk. The directionality of this risk is an important consideration, since we have previously shown that TCF21 promotes the FMC phenotype transition and has a protective effect toward risk causality, and it is imperative to know which of the two FMC clusters that we have described mediates this protective effect. The FMC-1 versus FMC-2 cellular phenotypes are quite different and the mechanism for protection could be achieved through a number of different pathways in each cell type. We employed S-PrediXcan for this purpose, which uses composite eQTL data to make this determination (Fig. 6A). This algorithm is able to take GWAS results and predict the effects of each variant on expression levels of genes that are associated with the trait, and it is able to do this for every loci that is associated with the trait throughout the genome. In situ RNA staining localizes the FMC-2 population to both the fibrous cap (Notch3, Thbs1) and the intimal plaque (Fbln2, Ttc9) confirmed with label transfer using Xenium spatial transcriptomics. Further molecular analysis finds FMC-2 at the juncture of critical gene modules for senescence and apoptosis while expressing numerous CAD associated genes that we have linked to atherosclerosis, including ZEB2 and SMAD312,13. Using this scDRS score leverages the power of GWAS genes to identify likely causal cell state determining genes whose functions are critical to prime cells towards phenotypic transition.
The scDRS analysis did not identify CAD risk in the CMC phenotype cells. This is surprising, given that mouse knockout disease models of orthologs of human CAD associated genes SMAD3 and PDGFD12,17, and other genes not yet linked to CAD by GWAS, KLF4 and AHR11,14, have demonstrated a significant correlation between CMC number, plaque burden, and vascular calcification. This is true for both disease promoting and disease protective gene functions, as determined by effect allele identification and genetics of gene expression data. It is possible that there is a dearth of informative allelic variation in CAD loci that determine the CMC phenotype. We did observe that aggregate expression of CAD disease risk genes in the CMC cell state showed a higher relative average expression of CAD GWAS risk genes, but this did not provide a statistically significant result. This observation suggests that a bias against regulatory variation at the CMC determining gene loci resulting in a low number of informative genes may explain the lack of an scDRS finding. Also, an important consideration is that FMC-2 are the most similar to CMC in terms of gene expression phenotype, and the risk identified in these cells may not be protective, but may in fact promote disease risk through directing differentiation to the CMC phenotype. eQTL based directionality inference is limited by the number of genes that have statistically significant eQTL links and this can bias results. Finally, the observation that Tcf21 promotes transition to the CMC phenotype is difficult to reconcile with these considerations but may simply reflect that the protective effect of increased FMC number outweighs the disease promoting effect of increased CMC.
We have undertaken these studies in order to better understand cellular and molecular mechanisms of CAD risk by identifying and characterizing individual genes and gene programs that modulate SMC phenotype transitions. While such studies do not prove causality, they inform on possible mechanisms of disease risk that reside in the vessel wall. Although not explored in the work discussed here, there are a number of approaches related to mapping causality in human data that warrant further investigation, and can serve to strengthen the genetic data derived in our single-cell studies of mouse models. For instance, more detailed study of rare coding variation, when present, can provide human disease phenotype and direction of effect for CAD genes that are identified with GWAS studies. A good example is the CAD GWAS MFGE8 (lactadherin) gene which was identified in the FinnGen biobank to have an association between an inframe insertion rs534125149 and protection against coronary atherosclerosis60. Along this line of reasoning, a recently described approach from the Pritchard lab employs loss of function burden tests along with relevant Perturb-seq data to bridge the gap between genetic association and biological mechanism61. By combining these two forms of data, their approach builds causal graphs in which the directional associations of genes with a trait can be explained by their regulatory effects on gene programs. It is important to note of course that rare coding variation in the protective gene TCF21 would be expected to increase the risk for CAD, and indeed likely cardiovascular developmental defects that may be inconsistent with fetal survival. Also, TCF21 has important developmental roles in the kidney and lung, and mice lacking Tcf21 die at birth due to failure of lung function. This is one possible reason that we have not found coding region mutations in the TCF21 gene that are associated with protein function. Such mutations are likely removed from the genome through purifying selection.
Another approach to enriching the pool of CAD causal genes in SMC transitions is the possibility of using human somatic mutation to implicate genes of relevance to vascular disease. The NIH program Somatic Mosaicism Across Human Tissues (SMaHT) Network is focused on cataloging naturally occurring DNA mutations (somatic mosaicism) in healthy subjects to understand aging and disease. This network is not currently studying arterial tissues, and the tasks of collecting and subjecting tissue from multiple humans and then pursuing a very deep sequencing effort is daunting, but possibly well worth pursuing.
Computational algorithms used in these studies were identified to be those most aligned and broadly tested with the genomic and genetic data types provided in our timecourse study. In particular, novel trajectory methods that become available may continue to offer useful and more nuanced details regarding the complex cell state changes that the SMC undergo during the disease process, and we make our data available for such analyses. While the WOT approach was specifically designed for timecourse scRNAseq data, the CellRank262 algorithm is now considered to potentially yield more accurate and robust results by integrating additional biological information, such as metabolic labeling, and can use optimal transport as one component of its broader multivariate analysis. We did in fact utilize orthogonal methods from the original CellRank63, such as its pseudotime kernel, to generate a probabilistic model, which appeared to follow similar patterns as our WOT approach to generate interpretable summary models.
We have created these high order genomic data sets to map the epigenetic and transcriptomic mechanisms that mediate the SMC lineage transitions and their contribution to disease risk. This study focused on genes that are expressed by transition SMC and linked to phenotypic cell state changes through TFs and other high content signaling molecular pathways that mediate disease trajectories. We have identified a number of such genes that reside in CAD associated loci and are candidates for causal relationships with SMC transitions (Supplementary Data 11). These genes were not validated as causal with genome editing or animal model studies, as such efforts are beyond the scope of the present work, but many show colocalization of expression quantitative trait and CAD associated variation suggesting causality. These example candidate genes allow a number of observations relating SMC transition and CAD gene association. Although appreciated previously, it is clear from our analysis that numerous TGFB pathway genes represent a significant component of disease risk in relation to SMC phenotypic transitions. Other represented gene programs include chondrogenesis, hypoxia response, vascular development, and epithelial mesenchymal transition, in addition to proliferation and migration. Importantly, while most processes have expression across multiple SMC lineage phenotypes, they are all enriched in FMC-2 cells.
Our study uses a rigorous workflow that identifies reproducible cell cluster phenotypes. However, this rigor is implemented at the risk of losing rare cell states including those that respond to metabolic or lipid stress and most notably, phenotypes that represent progenitor cells such as those that undergo clonal expansion in the disease setting. Future studies will be directed at identification of these putative precursor cells of interest utilizing novel lineage tracing methods (e.g., in vivo barcoding, somatic mutation detection, or novel lineage tracers using putative driver genes) that can better capture these rare populations while using our dataset as a source for validation. This way, descriptive characterization of novel cell phenotypes coupled with experiments using such lineage tracing will allow robust characterization of how these precursor cells contribute to the plaque and fibrous cap, and how the deletion of specific markers alters the course of disease.
We are compelled to note that our extensive characterization of SMC phenotypic transitions does not provide evidence that this cell lineage can adopt the macrophage phenotype. Initial speculation that SMC could transition to a macrophage phenotype was understandable given extensive data that lineage traced SMC express a number of macrophage genes such as Lgals3, but single-cell studies have indicated that these are a rare events16. Whether SMC transition to foam cells is currently understudied. Lipid loading studies with SMC have shown that they can take up lipid in vitro64, and we and others have shown that they can take up lipid in vivo16,65, but the probability and to what extent vascular SMC can transition to the foam cell phenotype remains unknown. SMC foam cells are reported to have a different phenotype than macrophage foam cells, due to a relative deficiency of lysosomal acid lipase in SMC, and retain lipid droplets in their lysosomes rather than in the cytoplasm as seen in macrophages66. This would predict SMC foam cells have an altered gene expression pattern and phenotype different from macrophage foam cells. The literature is replete with discussions regarding the impact of SMC-derived foam cells to plaque, and indeed the role of such cells toward disease risk could be significant and the avenues for manipulation important for therapeutic considerations substantial, but this cell must be characterized with modern genetic and genomic methods. A recent comprehensive single-cell study of carotid plaque has identified cells with gene expression features of both SMC and macrophage cellular phenotype, and further study of this cluster could address the current need in the field67. Without question, identification of the molecular pathways by which SMC lineage foam cells take up lipid and contribute to destabilizing the plaque could have significant importance for targeted therapy development. Moreover, additional multi-omic approaches incorporating DNA methylation, proteomic, metabolic and lipidomic data will allow us to better interrogate cellular physiology in the context of genomic analyses.
Future studies are needed to confirm the functions of nominated genes implicated in the cell state changes characterized through these studies. Only through study of the larger causal gene regulatory network will we be able to understand which aspects of the complex cellular phenotypic changes, migratory behaviors and cell-cell interactions that are responsible for the risk that is modulated by the SMC lineage. Specifically, studies are needed to characterize in greater detail the FMC-2 phenotype genes that drive CAD risk, what is the nature of CMC disease risk, and how can disease protective genes like TCF21 promote CMC formation without increasing risk. The required expansive causal CAD gene characterization can only come from large-scale in vitro CRISPR screening with highly relevant disease cellular models, and similar screens conducted in disease model mice, which will provide in vivo disease transcriptomic phenotype information regarding the function of SMC genes that both regulate SMC phenotype and CAD risk.
Methods
Mouse strains, induction of lineage marker, and sample collection
Our research complies with all relevant ethical regulations, with our animal study protocols (Protocol ID -10020, 10054) approved by the Institutional Animal Care and Use Committee at Stanford University.
Control (final genotype - TgMyh11-CreERT2, Tcf21+/+, ROSAtdT/+, ApoE−/−) and Tcf21 (final genotype - TgMyh11-CreERT2, Tcf21ΔSMC/ΔSMC, ROSAtdT/+, ApoE−/−) knockout mice with floxed tdTomato fluorescent reporter to allow for SMC-specific lineage tracing were bred onto a C57BL/6 ApoE-/- background as previously described16. For all subsequent lineage tracing experiments, two doses of tamoxifen 48 h apart via gavage was carried out when the mice reached 7.5 weeks of age. In the Tcf21-flox group, tamoxifen gavage also induces the Tcf21 knockout in addition to tdTomato lineage tracing. Following tamoxifen treatment, all mice were placed on a high-fat diet. At the designated timepoints, animals are euthanized after CO2 administration followed by cervical dislocation, consistent with the recommendations of the Panel on Euthanasia of the American Veterinary Medical Association (AVMA).The study involved two primary high fat diet experimental arms: one for single-cell RNA sequencing (scRNA) and the other for Assay for Transposase-Accessible Chromatin sequencing (ATAC).
Animals were housed in an AAALAC-accredited, specific-pathogen–free barrier facility in individually ventilated cages under a 12-h light/12-h dark cycle (lights on 0700–1900). Ambient temperature was maintained at 20-26 °C with relative humidity at 30–70% per institutional standards. Sterile bedding was changed regularly, and mice had ad libitum access to standard rodent chow or high fat diet (21% anhydrous milk fat and 0.15% cholesterol (Dyets no. 101511; Dyets) and water. Environmental enrichment (e.g., nesting material and shelters) was provided.
For scRNA experiments, samples were collected from Control mice at multiple time points named by number of weeks on high fat diet: 0, 3, 5, 7, 9, 12, and 16 weeks. Samples from the Tcf21 knockout mice were collected at 5, 12, and 16 weeks. For ATAC experiments, samples were collected from the Control mice on the high-fat diet at 0, 5, 7, 9, 12, and 16 weeks and for Tcf21 knockout mice at 5, 12, and 16 weeks. 3-4 male mice are pooled for each single-cell collection as the Myh11CreERT2 transgene is inserted in the Y chromosome, limiting lineage tracing studies to the male sex.
Aortic digestion for 10x genomics microfluidics
Samples from mouse aorta were dissociated into single cells for RNA and ATAC sequencing using the 10x Genomics Chromium platform. Euthanized mice are perfused with phosphate buffered saline and dissected to obtain the aortic root up to the level of the brachiocephalic artery. The tissue is washed with PBS and incubated in an enzymatic dissociation cocktail containing Liberase (5401127001; Sigma-Aldrich) and Elastase (LS002279; Worthington) in Hank’s Balanced Salt Solution with calcium (HBSS+ Ca2+) for 30 min. Mechanical dissociation is performed for 5 min followed by visualization under the microscope to ensure single-cell suspension. This suspension is strained through a 35 µm nylon mesh snap cap into falcon test tubes (352235 BD) followed by FACS sorting (Sony SH800) for tdTomato positive and negative cells in parallel. For single-cell ATAC, nuclei was isolated per 10X recommended protocol and captured on the 10X scATAC platform. Each individually sorted cell suspension was loaded onto 10x GEM G orH chips with remainder per 10x protocols (Chromium Single Cell 3’ RNA V3.1, Chromium Single Cell ATAC V2). For both scRNA and scATAC runs, samples at some intermediate time points were processed as pooled samples with tdTomato and non-tdTomato loaded at 1:1 ratio after FACS and listed in Supplementary Data 1. Libraries were sequenced on the Illumina NovaSeq6000 platform with targeted depth of 40-50,000 reads per cell for RNA and 75,000 reads/cell for ATAC.
10x RNA and ATAC data preprocessing
RNA fastq files were processed using Cellranger v7.0.1 (10x Genomics) to obtain transcript count matrices and aligned to mouse transcriptome mm10-2020-A-2.0.0 (10x Genomics) with custom addition of lineage tracing tdTomato transcript. Samples at each timepoint were aggregated and analyzed with R package with Seurat (v4.3)68. Low-quality cells and mitochondrial-rich cells were filtered with parameters mitochondrial percentage <6%, ribosomal percentage <25%, and nFeature_RNA > 1250 and <8000. Gene expression count matrices underwent log-transformation and library-specific scaling. Importantly, no additional batch correction for visualization was required in the pre-processing steps given the uniform processing of mice samples. Principal component analysis was used for dimensionality reduction followed by clustering using the Louvain algorithm.
For the full timecourse dataset, aligned scRNA files from Control and Tcf21-KO data were merged and processed with identical QC parameters as above. Clustering was then performed on this merged dataset in order to ensure comparable differential gene analysis and pseudotime comparisons. The processed dataset is then split into Control and Tcf21-KO objects for independent analysis. We further applied logistic regression to determine the optimal tdTomato expression cutoff and subset on SMC-derived lineage cells exclusive to tdTomato fluorescence-activated cell sorting (FACS).
For force directed layouts, we followed methods as described in Schiebinger et al. 50 dimensional diffusion components are calculated with SCANPY (1.9.3) with default parameters. For each cell, its 20 nearest neighbors was used to produce a nearest neighbor map. Then we applied leiden clustering at a resolution of 0.36 and generated cluster connectivities via PAGA and visualized the force-directed layout on the k-NN graph using ForceAtlas2.
scATAC Raw fastq files were uniformly processed using Cellranger-atac-2.1.0 (10x Genomics) and aligned to cellranger-arc-mm10-2020-A-2.0.0 reference genome (10x Genomics). The data were processed to remove low-quality cells and reads with low mapping quality with parameters peak_region_fragments >2500, peak_region_fragments <100000, pct_reads_in_peaks >30, nucleosome_signal <= 4, TSS.enrichment >2. The remaining reads were then used to construct a sparse binary matrix, representing chromatin accessibility states across individual cells and genomic regions. A unified set of peaks from all samples (Control and Tcf21-KO) were merged using overlapping and adjacent peaks using CellRanger-ATAC aggr protocol. This unified peak set was re-quantified by term frequency-inverse document frequency (TF-IDF) using Signac69 (1.10) RunTFIDF.
To preprocess scRNA and scATAC data for integration, gene activities are first calculated from chromatin accessibility data using GeneActivity() function from Signac with default parameters and log normalization. Datasets were integrated using canonical correlation analysis (CCA) with the Seurat RunCCA() function upon 2000 features using the SelectIntegrationFeatures() in Seurat. We then use the integrated dataset to perform pseudotime ordering using Slingshot70 (2.6) and split the dataset into 30 pseudotime bins for uniform downstream analyses.
scATAC functional and motif analyses
To generate identify overrepresented TF binding motifs, we performed hypergeometric enrichment on the top 20,000 variable peaks with GC-matched controls using the FindMotifs function in Signac. We then further merged this motif set with HOMER’s (4.10) de novo motif enrichment method on a per-cluster basis taking the top overrepresented motifs along with all similar motifs above a similarity score of 0.7. This allowed identification of a comprehensive set of enriched transcription factor motifs to aid summarization of ATAC data. For mouse ATAC data, the background peak set utilizing the entire merged peak set generated by CellRanger-ATAC (version 2.0.1) Aggr.
We then calculate a cluster specificity score by dividing the detection percentage of accessible chromatin for each cluster by the detection percentage of all other clusters with a minimum 10% cutoff for within cluster accessibility. We filtered peaks with a cluster specificity of >1.5 for FMC-1, FMC-2, and CMC and specificity >1.25 for SMCs. From these peaks, we selected the top 5,000 peaks with the highest specificity score in the FMC-1, FMC-2, and CMC clusters, while for the aggregated SMC cluster (SMC-1, SMC-2, SMC-3), we use the top 2500 peaks. ChromVAR71 was used to calculate the transcription factor specific enrichment scores of accessible motifs across each pseudotime bin through the Signac wrapper RunChromVAR.
Trajectory analysis with Waddington optimal transport and CellRank2
To model and infer cellular trajectories in our data, we employed the Waddington optimal transport algorithm30. This approach infers cell growth rates using single-cell gene expression to generate a transition probability distribution, enabling the identification of cellular transitions and differentiation paths.
We applied the default command line implementation of Waddington WOT as described in Schiebinger et al. with cell scores derived from updated KEGG cell cycle and Apoptosis gene signatures for estimation of cell growth and death. Growth rate tables are extracted and added to single-cell metadata for additional analyses. We generated a ‘fate correlation by comparing WOT predicted cell transition probability for the FMC-1, FMC-2 and CMC fate with each Tcf21 downstream TF network module score in the combined Tcf21-KO and control dataset. As negative control, we also correlated respective TF expression with fate probability to identify consistently more robust correlation with TF module scores compared to its central TF. Fate tables in Supplementary Data 6 are arranged in order of fraction expressed ratio.
Orthogonal trajectory analyses was performed using CellRank2. We applied the RealTimeKernel using default settings to identify sustained cell states and calculate driver gene trends plotted across our imported Slingshot derived pseudotime.
Data integration, gene regulatory network generation and in silico transcription factor perturbation
We utilize the Pando44 (1.0.1) to generate gene regulatory networks and CellOracle45 (0.18.0) for in silico TF perturbation experiments. By combining Pando’s network construction capabilities with CellOracle’s in silico perturbation analysis, this approach enables a comprehensive exploration of gene regulatory dynamics in single-cell data. We divided the integrated dataset into PseudoEarly (pseudotime bins 3-24) and PseudoLate (pseudotime bins 25-30) for further analyses.
To generate GRNs from our scRNA and scATAC integrated dataset, we identified transcription factors and their respective binding sites in regions of accessible chromatin through motif scanning, and these transcription factor module candidate regions linked to genes by proximity through the PANDO framework. The bagging regression model was selected to infer the relationships between TF expression, binding-site accessibility and target gene expression to generate cell state-specific networks.
We utilized the extended transcription factor motif database from the original Pando manuscript (including JASPAR202072, CIS-BP73 as well as TFs without known motifs assigned by sequence homology) and further included all JASPAR2020 database mouse reference motifs for downstream motif scanning. Gene regulatory network inference was then performed using default PANDO parameters for selection of candidate regulatory regions from scATAC data, transcription factor motif scanning, selection of region-TF pairs, while the final regression model, we substitute the bagging ridge model for the default generalized linear model to match that of CellOracle. The output coefficient table was extracted and filtered for adjusted p-value < 0.05, R2 > 0.1, minimum number of variables (region-TF pairs) per model >10, and minimum genes per module >5 to generate the final regulatory graph that is provided as input for downstream analysis including:
-
1.
Weighted pseudotime x TF module UMAP, is calculated by the get_network_graph() function in Pando to generate a subgraph with significant module TFs as features to generate a UMAP embedding which takes into account the ingoing connection of each node (TF) as well as coexpression of TFs. Additionally, the product of average pseudotime per cell by TF expression is colorized onto each TF module to provide a visual representation of the TF module relationships in the context of pseudotime.
-
2.
TF activity score – calculated by the averaging the sum of TF expression x target gene coefficient for each TF module in the early (pseudotime bin 3-24) or late (pseudotime bin 25-30) SMC gene regulatory network.
For in silico perturbation, the merged control timecourse integrated dataset is downsampled to 40,000 cells to allow for optimal computation speed along with the top 3000 most variable genes calculated in Seurat and extended TF database from Pando converted into CellOracle compatible format to generate a baseline ‘Oracle’ file. The Pando derived TF-gene regulatory network table is converted into ‘links’ network file format as input for CellOracle’s link_data parameter for in silico systemic TF perturbation modeling with negative PS sums visualized as described in the CellOracle tutorial.
Xenium slide processing and analysis
Fresh frozen tissue sections were prepared following the 10x Genomics Xenium In Situ for Fresh Frozen Tissues Protocol (CG000579 Rev F). Briefly, tissues arranged in order of aortic root, ascending aorta, brachiocephalic artery with right subclavian branch, per-diaphragmatic descending, and abdominal descending aorta from a control (TgMyh11-CreERT2, Tcf21+/+, ROSAtdT/+, ApoE−/−) mouse were embedded in OCT compound immediately after dissection. Embedded tissue blocks were frozen on dry ice and immediately stored at -80 °C prior to sectioning. Sections of 10μm thickness were cut using a Leica CM1860 cryostat. Sections were mounted within the sample area (10.45 × 22.45 mm) of Xenium slides (PN-3000941) without overlap and stored at -80 °C until ready for use. All subsequent Xenium steps were processed at the Stanford Functional Genomics Core. Tissue morphology and quality was assessed using Hematoxylin and Eosin (H&E) and DAPI staining prior to downstream Xenium In Situ.
Gene expression assays
Processed Xenium datasets were imported into Seurat V5 for normalization and integration. Feature-barcode matrices were normalized using the SCTransform workflow with spatial coordinates retained for downstream deconvolution. To perform single-cell reference label transfer to the Xenium data, canonical correlation analysis (CCA) was implemented following strategies from the Seurat tutorial “Analysis, visualization, and integration of spatial datasets with Seurat” (https://satijalab.org/seurat/articles/spatial_vignette.html). RCTD (Robust Cell Type Decomposition) was applied to further deconvolve spot-level spatial data based on input cell reference.
Mouse to human scRNA label transfer and Spatial Slideseq analysis
Data is obtained and re-processed from the CZI Arterial Atlas courtesy of Zhao et al.23. We processed single-cell RNA data based on parameters published in Zhao et al. and subset the SMC clusters. We then transferred labels from our control smooth muscle control dataset and transferred labels using CCA with 1:30 dim and assigned SMC cell states to the human scRNA data. Slide-Seq object was aligned to hg38 by curioseeker pipeline and again underwent processing as described in Zhao et al.23. Robust Cell Type Decomposition was then applied to integrate this label transferred reference with spot-level data from the spatial transcriptomic sequencing using Seurat V574. Visualization was performed using Seurat built-in functions including SpatialDimPlot to visualize the label transferred cell states.
Processing of external datasets, Pan et al., Alencar et al., Cheng et al.
Data from Pan et al. (GSE155513)15, Alencar et al. (GSE150644)11, Cheng et al. (PRJNA794806)12 were downloaded and reprocessed using standard 10x CellRanger pipeline as noted above to facilitate label transfer analysis with our timecourse dataset. All subsetting parameters and clustering parameters decisions were made based on available published methods, following prior settings as closely as possible. For the Pan et al. dataset, we performed further data pruning, subsetting based on fluorescent marker expression as previously described in Sharma et al.19. We then applied cutoffs of nFeature >1900, percent.mt <7.5 to match our control timecourse study. For clustering, we identified top 1500 variable genes and generated UMAP with 20 principal components (PCs) as previously described. For Alencar et al. we filtered on nFeature_RNA > 200 and <5000, percent.mt <10. 19 PCswere selected for clustering. The clustering resolution was not specified. This does not affect our downstream analysis, as the goal for this analysis is to understand the distribution of label-transfer clusters from our control timecourse study. For Cheng et al. we requested original processed script from the author to recreate the R-Smad clustering. After clustering of these datasets, we then performed CCA label transfer followed by generation of confusion matrices to compare and contrast the relationship between clusters and identify existing cell states.
ChIPseq analyses
Fastq files were mapped to hg38/GRCh37 genome with Bowtie2 (1.2.3). ChIP-seq peaks were then called with MACS2 (2.2.7.1) using default parameters. From this output, ‘robust’ peaks were selected by specifying a minimum fold-enrichment of 5.
GWAS trait SNP enrichment analyses
The intersection of GWAS loci and transcription factor binding (TEAD1 + TCF21) was defined as the SNPs located within any overlapping region with ChIP-seq peaks. Direct overlap of SNPs assimilated from GWAS Catalog + MVP CAD GWAS was performed with GWASanalytics script (https://github.com/zhaoshuoxp/GWASanalytics). The binomial overlap performed by this script has been described previously75.
scDRS (Single cell disease relevance score)
The scDRS53 algorithm calculates a disease relevance score for each cell by comparing its gene expression pattern with a reference disease gene signature obtained from the MVP coronary artery disease GWAS data. We utilized summary statistics obtained from the coronary artery disease GWAS meta-analysis from the Million Veterans Program (MVP) which also incorporates transancestry genetic data. MVP CAD GWAS was munged using (MAGMA) to obtain a weighted CAD-associated gene list that is applied towards the mouse timecourse data. Remainder scDRS analysis was performed as described in Zhang et al.53.
Bulk RNASeq deconvolution
Bulk RNAseq fastq files were downloaded from GSE113348 and processed through a uniform pipeline with cutadapt (5.0), STAR (2.7.10b) and featureCounts (2.0.6) for trimming, alignment and generation of count matrices, respectively. Data was then converted to TPM with Kallisto (0.51.1) for comparison between cell lines. Following the prior publication, we removed nine cell lines from the original dataset due to poor data quality for a total of 52 cell lines used. We then applied non-negative least squares deconvolution using the python implementation of nnls from scipy to estimate the contributions of SMC-derived phenotypic clusters based on our tdTomato SMC subset reference. The top 100 marker genes per cluster (ranked by pct.1/pct.2 with at least 25% expression in the reference cluster) were used as features in the cell type signature matrix.
Curated CAD GWAS gene list
A curated list of nominated GWAS genes based on prior CAD GWAS meta-analysis was tabulated from Erdmann 2018, Koyama 2019, Matsunaga 2020, Tcheandjieu 2022, Aragam 20225,6,76,77,78.
Heritability adjusted PrediXcan, LocusZoom and eQTL correlation visualization
We estimated gene expression risk directionality inferred from the updated heritability adjusted S-PrediXcan modeling54. LocusZoomR79 (0.3.8) was used to visualize gene locus plots. eQTL colocalization was performed by extracting positional coordinates from MVP CAD GWAS and retrieving SNPs from dbGaP. SNP with lowest p-value near nominated GWAS gene was selected and all SNPs meeting GWAS significance 5e-8 and within 50 kb up and downstream of lead SNP were selected. We then identify which SNPs had corresponding eQTLs using the V10 GTEx release. The beta for each SNP was correlated with the corresponding NES of each eQTL and plotted as a scatter graph with color representing the R2 (calculated with LDlinkR) with lead SNP.
In situ RNA hybridization (RNAScope)
Slides were processed according to the manufacturer’s instructions, using reagents from ACD Bio (ACD 322360-USM). Slides were washed in PBS, then immersed in 1 × Target Retrieval reagent at 100 °C for 5 min. After washing twice in deionized water, slides were immersed in 100% ethanol, air-dried, and sections were encircled with a liquid-blocking pen. Sections were incubated with Protease III reagent at 40 °C for 30 min, then washed twice with deionized water. Sections were incubated with probes against Fbln1 (ACD 502881), Fbln2 (ACD 447931), Ibsp (ACD 415501), C3 (ACD 417841), Tcf21 (ACD 508661), Thbs1 (ACD 457891), Notch3(ACD 425171), Col8a1 (ACD 518071), Ttc9 (ACD 1113921-C1), Loxl1 (ACD 492531), Bmp1 (ACD 311151), Vcam1 (ACD 438641), Col2a1 (ACD 407221) or a negative control probe (ACD 310043) for 3 h at 40 °C. Multiplex fluorescence and colorimetric assays were performed per the manufacturer’s instructions.
HCASMC and A7r5 cell culture
Primary HCASMCs and HCASMC-hTERT80 were maintained in SMC basal medium (SmBM) supplemented with SmGM-2 SingleQuots kit (Lonza CC-3182) including human epidermal growth factor, insulin, human basic fibroblast growth factor and 5% FBS, according to the manufacturer’s instructions.
Rat aortic smooth muscle cells (A7r5) were purchased from ATCC and cultured in Dulbecco’s Modified Eagle Medium (DMEM) high glucose (Fisher Scientific, #MT10013CV) with 10% FBS at 37 °C and 5% CO2. A7r5 at passage 6-9 were used for experiments.
Calcification assay
Primary cell lines (1508 and 2105) were split into 6 well plates at 60k cells per well. Cells were allowed to equilibrate over 12 h prior to media change with SmBM with supplements as noted above. After 24 h, once cells are noted to be confluent, we applied a calcification media cocktail consisting of SmBM basal media supplemented with 0.4% FBS, (Lonza CC-3182 without additives). Cells were incubated in calcification media for 3 days followed by media change with calcification media and allowed to grow to 7 days prior to RNA collection as previously described14.
siRNA experiments
Vascular SMCs were transfected with siRNAs targeting TEAD1, AR, EPAS1, and ZEB1(Dharmacon ON-TARGETplus SMARTpool siRNA L-012603-00-0005, L-003400-00-0005, L-004814-00-0005, L-006564-01-0010; Horizon Discovery). Silencer Select negative control siRNA (ThermoFisher 4390844) was used and has been previously tested in our laboratory to ensure no cellular physiology changes. siRNA pool transfection was subsequently performed using Lipofectamine RNAiMax Transfection Reagent (ThermoFisher 13778150) according to manufacturer’s instructions and incubated for 12 h post-transfection in serum free media followed by an additional 12 h of recovery in SmBM supplemented media prior to RNA collection.
RNA extraction and quantitative PCR (qPCR) experiments
RNA extraction was performed using RNAEasy Mini kits (Qiagen 74106). 500 ng of RNA per sample is then aliquoted for reverse transcription with High-Capacity RNA-to-cDNA Kit (Life Technologies 4388950). Quantitative PCR reactions were conducted with Taqman Universal Master Mix (44440048) and qPCR FAM probes for genes of interest (ThermoFisher) on a QuantStudio 6 Pro Real-Time PCR System. Relative transcript abundance was determined by the comparative Ct (Δ ΔCt) method, using the housekeeping gene UBC.
CEBPB, TEAD, H3K27ac ChIPseq
Approximately 1,000,000 HCASMCs were cross linked in 1% formaldehyde for 10 min and washed with PBS and replaced with hypotonic buffer (20 mM Hepes (pH 7.9), 10 mM KCl, 1 mM EDTA (pH 8) and 10% glycerol) and incubated on ice for 6 min. Cells were then sonicated using a Branson 250 Sonifier (using power setting 5, constant duty for 10 rounds of 30-second pulses) with confirmation of chromatin fragments at 250-400 base pairs. Lysates were then incubated overnight with 5 µg of anti-CEBPB (Santa Cruz sc-150), TEAD(Cell Signaling 12292S), or H3K27ac(Abcam 4729). Protein-DNA complexes were captured with Protein G agarose beads (Sigma 8104) and eluted in 1% SDS TE buffer at 65 °C. After reverse cross-linking followed by RNase A and proteinase K digestion, chromatin was purified using a QIAquick PCR purification kit (Qiagen 28706). ChIP DNA sequencing libraries were prepared using the Kappa HyperPrep (Roche 07962347001) and sequenced on a NovaSeq6000 with 150-base pair paired-end reads.
Proximity ligation assay
PLA was performed using manufacturer provided protocols for Sigma DuoLink In Situ Red Start Kit Mouse/Rabbit (DUO92101) with antibodies to TCF21 (Sigma AV33421, anti-rb), TEAD1 (sc-376113, anti-ms) and anti-mouse IgG (Vector I-2000-1).
Co-Immunoprecipitation (Co-IP)
Co-IP was performed using a myc-Tagged Tcf21 construct cloned into the PwPI vector. HEK293 cells were transfected with Tcf21-myc followed by protein extraction and nuclear isolation following manufacturer’s instructions (Active Motif Nuclear Complex Co-IP Kit 54001). 5% input control was kept as a positive control and samples were incubated with anti-rb TEAD1 (Cell Signaling 12292S) or anti-rb IgG (Abcam ab171870) as negative control. Co-IP was performed following instructions from the Active Motif Co-IP Kit, briefly, samples were incubated with 1:100 of TEAD1 antibody according to manufacturer recommendations and 5ug of anti-rb IgG (which is in excess compared to amount of TEAD1 antibody used). After overnight incubation, samples are washed and protein collected for western blotting. An anti-mouse HRP secondary antibody targeting the light chain was used for primary antibody detection as TEAD1 size is approximately 47kDA which would overlay with heavy chain antibody fragments.
Immunohistochemistry (IHC)
IHC was performed for TEAD1 and TCF21 in mouse (TgMyh11-CreERT2, ROSAtdT/+, ApoE−/−) aortic root sections fed 16 weeks of high fat diet to characterize their protein localization. Briefly, antigen retrieval was performed on sections after dissolving OCT in distilled water, and fixing in 4% paraformaldehyde for 5 min. After two PBS washes, antigen retrieval buffer (Biocare, DV2004) diluted in deionized water was preheated to 97–100 °C in an Oster steamer. Sections were pre-treated with RNAscope hydrogen peroxide for 5 min twice, then transferred into antigen retrieval buffer for 6 min, ensuring the buffer remained at least 97 °C. Slides were immediately moved to distilled water and washed twice by gentle dipping, air-dried, and a hydrophobic barrier was drawn around tissue with a hydrophobic pen. Section are incubated with Rodent Block M (Biocare, RBM961) for 30 min and washed in TBS. Primary anti-TEAD1 (Abcam 133533) and anti-TCF21 (Sigma, HPA013189) was diluted 1:100 in DaVinci Green Diluent (Biocare, PD900) and incubated overnight at 4 °C. The next day, sections were washed in TBS, incubated with Rabbit-on-Rodent HRP Polymer for 30 min at room temperature, and washed in TBS. Vina Green Chromogen kit was prepared per manufacturer instructions, applied at 50 µL per section for 4 min, then slides were rinsed in deionized water twice, air-dried, and mounted with a xylene-based medium under coverslip for microscopy.
Dual luciferase assays
Enhancer/Promoter elements (SRF 2nd intron; chr15:73633709-73634109, BMP1 6th intron; chr8:22178981-22179700, Loxl1 5’ UTR; chr15:73926050-73926450) demonstrating overlapping between Tcf21 and Tead ChIP-seq sites were cloned into pWPI and transfected into A7r5 cells. A7r5 cells were seeded into 24 well plate (1.5 × 104 cells/well) in DMEM containing 10% FBS and incubated at 37 °C and 5% CO2 overnight. Cells were transfected with varying combinations of luciferase reporter plasmids (pLuc-MCS (empty), pLuc-enhancer, cDNAs (pWPI (empty), pWPI-TCF21, pWPI-TEAD1 and pWPI-MYOCD), and Renilla luciferase plasmid using Lipofectamine 3000 (Invitrogen, #L3000015). Six h after transfection, the media was changed to fresh complete media. Relative luciferase activity (firefly/Renilla luciferase ratio) was measured by SpectraMax L luminometer (Molecular Devices) 24 h after transfection. All experiments were conducted in at least triplicate and normalized to the reporter plasmid after subtracting empty luciferase construct luminescence.
Statistics & reproducibility
To identify differentially expressed genes in the scRNA-Seq data, we split our data by PseudoEarly (pseudotime bins 3-24) and PseudoLate (pseudotime bins 25-30) and employed the FindMarkers function with the DESeq2 wrapper and filter for genes with absolute value Log2FC > 0.15. For differential scATAC peak analysis, we applied the likelihood ratio test ‘LR’ from FindMarkers and used nCount_peaks as the latent.variable and min.pct as 0.001 on a per cell type cluster basis. For differential ChromVar analysis, we calculated row means and calculated the average difference in Z-score between conditions using the wilcox test from FindMarkers.
For Jensen-shannon divergence (JSD) calculation we compared the ChromVar motif deviation distribution for each motif across pseudotime by calculating the Kullback-Leibler (KL) divergence from each distribution to the midpoint distribution where:
Where p and q represent the two distributions being compared and m is the midpoint distribution calculated as the average of two normalized distributions
Then the JSD was computed as the square root of the average of the two KL divergences.
We first randomly split the control dataset equally and calculated the JSD divergence to derive a control distribution for each motif. We then calculated JSD divergence between Tcf21-KO and control.
To determine statistical significance, we constructed a null distribution using the control JSD measurements by performing 10,000 random sampling with replacement from the control values. One-sided empirical p-values were calculated for each TF using the formula p = (number of permuted values >observed value + 1) / (total number of permutations + 1).
Fisher’s exact test was used to assess the significance of overlaps between genomic regions. Adjusted P-values are corrected with FDR method of Benjamini-Hochberg and <0.05 were considered statistically significant. All data are presented as mean and error bars represent standard deviation (SD).
For dual luciferase experiments, the BMP1 and LOXL1 enhancers were tested in biological triplicates. For SRF 2nd intron, the test conditions were performed in quadruplicate, while control and MYOCD-only conditions were performed in duplicate. For SRF, the control and MYOCD-only conditions have been previously validated by Nagao et al.35. Each dual luciferase experiment was repeated three times independently with representative results shown.
No statistical method was used to predetermine sample sizes for in vitro and single-cell experiments. No data were excluded from the analyses. The experiments were not randomized. The Investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The single-cell processed RNA and ATAC data generated in this study have been deposited in CellxGene under accession code 7a3044e4-6b16-4693-9504-212d9a573f80 (https://cellxgene.cziscience.com/collections/7a3044e4-6b16-4693-9504-212d9a573f80). The raw data is deposited to National Center for Biotechnology Information Gene Expression Omnibus (GEO) under accession code GSE321762. The Xenium mouse aorta spatial transcriptomic data, all human coronary artery smooth muscle ChIPseq data (CEBPB, H3K27ac, TEAD1), and Bulk RNASeq data (TEAD1) generated in this study are deposited to GEO under the following accession codes. Xenium: GSE316666, ChIPseq: GSE316714, RNASeq: GSE316713). For previously published data, TCF21-pooled ChIPseq and HNF1A ChIPseq, scRNA data from Pan et al., Alencar et al., and Cheng et al., and Bulk RNASeq primary HCASMC data from Liu et al. are downloaded from GEO: GSE141752, GSE59395, GSE155513, GSE150644, PRJNA794806, GSE113348, respectively. Human spatial data from Zhao et al. downloaded from CellxGene: 8f17ac63-aaba-44b5-9b78-60f121da4c2f (https://cellxgene.cziscience.com/collections/8f17ac63-aaba-44b5-9b78-60f121da4c2f).GWAS Catalog data were downloaded from (https://www.ebi.ac.uk/gwas/) and Million Veteran Program (MVP) were downloaded from dbGap with accession number phs001672.v3.p1 (https://dbgap.ncbi.nlm.nih.gov/beta/study/phs001672.v13.p1/#study). Source data are provided in the Source Data File. Source data are provided with this paper.
References
Roth, G. A. et al. Global burden of cardiovascular diseases and risk factors, 1990-2019: update from the GBD 2019 study. J. Am. Coll. Cardiol. 76, 2982–3021 (2020).
Khera, A. V. & Kathiresan, S. Genetics of coronary artery disease: discovery, biology and clinical translation. Nat. Rev. Genet 18, 331–344 (2017).
Zdravkovic, S. et al. Heritability of death from coronary heart disease: a 36-year follow-up of 20,966 Swedish twins. J. Intern Med 252, 247–254 (2002).
Marenberg, M. E., Risch, N., Berkman, L. F., Floderus, B. & de Faire, U. Genetic susceptibility to death from coronary heart disease in a study of twins. N. Engl. J. Med. 330, 1041–1046 (1994).
Tcheandjieu, C. et al. Large-scale genome-wide association study of coronary artery disease in genetically diverse populations. Nat. Med. 28, 1679–1692 (2022).
Aragam, K. G. et al. Discovery and systematic characterization of risk variants and genes for coronary artery disease in over a million participants. Nat. Genet 54, 1803–1815 (2022).
Quertermous, T. et al. Genome-wide genetic associations prioritize evaluation of causal mechanisms of atherosclerotic disease risk. Arterioscler Thromb. Vasc. Biol. 44, 323–327 (2024).
Turner, A. W. et al. Single-nucleus chromatin accessibility profiling highlights regulatory mechanisms of coronary artery disease risk. Nat. Genet 54, 804–816 (2022).
Ord, T. et al. Dissecting the polygenic basis of atherosclerosis via disease-associated cell state signatures. Am. J. Hum. Genet 110, 722–740 (2023).
Zhang, K. et al. A single-cell atlas of chromatin accessibility in the human genome. Cell 184, 5985–6001.e5919 (2021).
Alencar, G. F. et al. The stem cell pluripotency genes Klf4 and Oct4 regulate complex SMC phenotypic changes critical in late-stage atherosclerotic lesion pathogenesis. Circulation 142, 2045–2059 (2020).
Cheng, P. et al. Smad3 regulates smooth muscle cell fate and mediates adverse remodelling and calcification of the atherosclerotic plaque. Nat. Cardiovasc. Res. 4, 322–333 (2022).
Cheng, P. et al. ZEB2 shapes the epigenetic landscape of atherosclerosis. Circulation https://doi.org/10.1161/CIRCULATIONAHA.121.057789 (2022).
Kim, J. B. et al. Environment-sensing aryl hydrocarbon receptor inhibits the chondrogenic fate of modulated smooth muscle cells in atherosclerotic lesions. Circulation 142, 575–590 (2020).
Pan, H. et al. Single-cell genomics reveals a novel cell state during smooth muscle cell phenotypic switching and potential therapeutic targets for atherosclerosis in mouse and human. Circulation https://doi.org/10.1161/CIRCULATIONAHA.120.048378 (2020).
Wirka, R. et al. Single cell analysis of smooth muscle cell phenotypic modulation in vivo reveals a critical role for coronary disease gene TCF21 in mice and humans. Nat. Med. 25, 1280–1289 (2019).
Kim, H. J. et al. Molecular mechanisms of coronary artery disease risk at the PDGFD locus. Nat. Commun. 14, 847 (2023).
Shao, X. et al. Integrated single-cell RNA-seq analysis reveals the vital cell types and dynamic development signature of atherosclerosis. Front Physiol. 14, 1118239 (2023).
Sharma, D. et al. Comprehensive integration of multiple single-cell transcriptomic data sets defines distinct cell populations and their phenotypic changes in murine atherosclerosis. Arterioscler Thromb. Vasc. Biol. 44, 391–408 (2024).
Lin, C. J. et al. Distinct patterns of smooth muscle phenotypic modulation in thoracic and abdominal aortic aneurysms. J. Cardiovasc. Dev. Dis. 11, 349 (2024).
Pedroza, A. J. et al. Embryologic origin influences smooth muscle cell phenotypic modulation signatures in murine marfan syndrome aortic aneurysm. Arterioscler Thromb. Vasc. Biol. 42, 1154–1168 (2022).
Shukla, S. et al. Single-cell transcriptomics identifies selective lineage-specific regulation of genes in aortic smooth muscle cells in mice. Arterioscler Thromb Vasc. Biol. https://doi.org/10.1161/ATVBAHA.124.321482 (2025).
Zhao, Q. et al. A cell and transcriptome atlas of human arterial vasculature. Cell Genom. https://doi.org/10.1016/j.xgen.2025.101034 (2025).
Acharya, A. et al. The bHLH transcription factor Tcf21 is required for lineage-specific EMT of cardiac fibroblast progenitors. Development 139, 2139–2149 (2012).
Misra, A. et al. Integrin beta3 regulates clonality and fate of smooth muscle-derived atherosclerotic plaque cells. Nat. Commun. 9, 2073 (2018).
Worssam, M. D. et al. Cellular mechanisms of oligoclonal vascular smooth muscle cell expansion in cardiovascular disease. Cardiovasc Res. 119, 1279–1294 (2023).
Jacobsen, K. et al. Diverse cellular architecture of atherosclerotic plaque derives from clonal expansion of a few medial SMCs. JCI Insight https://doi.org/10.1172/jci.insight.95890 (2017).
Chappell, J. et al. Extensive proliferation of a subset of differentiated, yet plastic, medial vascular smooth muscle cells contributes to neointimal formation in mouse injury and atherosclerosis models. Circ. Res. 119, 1313–1323 (2016).
Haseeb, A. et al. SOX9 keeps growth plates and articular cartilage healthy by inhibiting chondrocyte dedifferentiation/osteoblastic redifferentiation. Proc. Natl. Acad. Sci. USA https://doi.org/10.1073/pnas.2019152118 (2021).
Schiebinger, G. et al. Optimal-transport analysis of single-cell gene expression identifies developmental trajectories in reprogramming. Cell 176, 928–943.e922 (2019).
Zhang, S., Afanassiev, A., Greenstreet, L., Matsumoto, T. & Schiebinger, G. Optimal transport analysis reveals trajectories in steady-state systems. PLoS Comput Biol. 17, e1009466 (2021).
Witzenbichler, B. et al. Regulation of smooth muscle cell migration and integrin expression by the Gax transcription factor. J. Clin. Invest. 104, 1469–1480 (1999).
Jeon, B. N. et al. KR-POK interacts with p53 and represses its ability to activate transcription of p21WAF1/CDKN1A. Cancer Res. 72, 1137–1148 (2012).
Tanaka, T. et al. Runx2 represses myocardin-mediated differentiation and facilitates osteogenic conversion of vascular smooth muscle cells. Mol. Cell Biol. 28, 1147–1160 (2008).
Nagao, M. et al. Coronary disease associated gene TCF21 inhibits smooth muscle cell differentiation by blocking the myocardin-serum response factor pathway. Circ. Res. 126, 517–529 (2019).
Bonnet, S. et al. The nuclear factor of activated T cells in pulmonary arterial hypertension can be therapeutically targeted. Proc. Natl. Acad. Sci. USA 104, 11418–11423 (2007).
Li, M. et al. Sildenafil inhibits calcineurin/NFATc2-mediated cyclin A expression in pulmonary artery smooth muscle cells. Life Sci. 89, 644–649 (2011).
Canalis, E., Schilling, L., Eller, T. & Yu, J. Role of nuclear factor of activated T cells in chondrogenesis osteogenesis and osteochondroma formation. J. Endocrinol. Invest. 45, 1507–1520 (2022).
Engleka, K. A. et al. Islet1 derivatives in the heart are of both neural crest and second heart field origin. Circ. Res. 110, 922–926 (2012).
Liu, C. F., Samsa, W. E., Zhou, G. & Lefebvre, V. Transcriptional control of chondrocyte specification and differentiation. Semin Cell Dev. Biol. 62, 34–49 (2017).
Mackie, E. J., Ahmed, Y. A., Tatarczuch, L., Chen, K. S. & Mirams, M. Endochondral ossification: how cartilage is converted into bone in the developing skeleton. Int J. Biochem Cell Biol. 40, 46–62 (2008).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Bennett, M. R., Sinha, S. & Owens, G. K. Vascular smooth muscle cells in atherosclerosis. Circ. Res. 118, 692–702 (2016).
Fleck, J. S. et al. Inferring and perturbing cell fate regulomes in human brain organoids. Nature 621, 365–372 (2022).
Kamimoto, K. et al. Dissecting cell identity via network inference and in silico gene perturbation. Nature 614, 742–751 (2023).
Lee, S. Y. et al. Differential but complementary roles of HIF-1alpha and HIF-2alpha in the regulation of bone homeostasis. Commun. Biol. 7, 892 (2024).
Salminen, A. et al. Mutual antagonism between aryl hydrocarbon receptor and hypoxia-inducible factor-1alpha (AhR/HIF-1alpha) signaling: Impact on the aging process. Cell Signal 99, 110445 (2022).
Lambert, J. et al. Network-based prioritization and validation of regulators of vascular smooth muscle cell proliferation in disease. Nat. Cardiovasc Res. 3, 714–733 (2024).
Lin, M. E. et al. Runx2 deletion in smooth muscle cells inhibits vascular osteochondrogenesis and calcification but not atherosclerotic lesion formation. Cardiovasc Res. 112, 606–616 (2016).
Liu, B. et al. Genetic regulatory mechanisms of smooth muscle cells map to coronary artery disease risk loci. Am. J. Hum. Genet 103, 377–388 (2018).
Zhao, Q. et al. TCF21 and AP-1 interact through epigenetic modifications to regulate coronary artery disease gene expression. Genome Med. 11, 23 (2019).
Zhao, Q. et al. Molecular mechanisms of coronary disease revealed using quantitative trait loci for TCF21 binding, chromatin accessibility, and chromosomal looping. Genome Biol. 21, 135 (2020).
Zhang, M. J. et al. Polygenic enrichment distinguishes disease associations of individual cells in single-cell RNA-seq data. Nat. Genet 54, 1572–1580 (2022).
Liang, Y., Nyasimi, F. & Im, H. K. Pervasive polygenicity of complex traits inflates false positive rates in transcriptome-wide association studies. bioRxiv https://doi.org/10.1101/2023.10.17.562831 (2024).
Huang, C. K. et al. Androgen receptor promotes abdominal aortic aneurysm development via modulating inflammatory interleukin-1alpha and transforming growth factor-beta1 expression. Hypertension 66, 881–891 (2015).
Sun, Y. et al. Smooth muscle cell-specific runx2 deficiency inhibits vascular calcification. Circ. Res. 111, 543–552 (2012).
Topouzis, S. & Majesky, M. W. Smooth muscle lineage diversity in the chick embryo. two types of aortic smooth muscle cell differ in growth and receptor-mediated transcriptional responses to transforming growth factor-beta. Dev. Biol. 178, 430–445 (1996).
Madura, J. A. et al. Regional differences in platelet-derived growth factor production by the canine aorta. J. Vasc. Res. 33, 53–61 (1996).
Trigueros-Motos, L. et al. Embryological-origin-dependent differences in homeobox expression in adult aorta: role in regional phenotypic variability and regulation of NF-kappaB activity. Arterioscler Thromb. Vasc. Biol. 33, 1248–1256 (2013).
Ruotsalainen, S. E. et al. Inframe insertion and splice site variants in MFGE8 associate with protection against coronary atherosclerosis. Commun. Biol. 5, 802 (2022).
Ota, M. et al. Causal modelling of gene effects from regulators to programs to traits. Nature 650, 399–408 (2025).
Weiler, P., Lange, M., Klein, M., Pe’er, D. & Theis, F. CellRank 2: unified fate mapping in multiview single-cell data. Nat. Methods 21, 1196–1205 (2024).
Lange, M. et al. CellRank for directed single-cell fate mapping. Nat. Methods 19, 159–170 (2022).
Rong, J. X., Shapiro, M., Trogan, E. & Fisher, E. A. Transdifferentiation of mouse aortic smooth muscle cells to a macrophage-like state after cholesterol loading. Proc. Natl. Acad. Sci. USA 100, 13531–13536 (2003).
Wang, Y. et al. Smooth muscle cells contribute the majority of foam cells in ApoE (Apolipoprotein E)-deficient mouse atherosclerosis. Arterioscler Thromb. Vasc. Biol. 39, 876–887 (2019).
Dubland, J. A. et al. Low LAL (Lysosomal Acid Lipase) expression by smooth muscle cells relative to macrophages as a mechanism for arterial foam cell formation. Arterioscler Thromb. Vasc. Biol. 41, e354–e368 (2021).
Bashore, A. C. et al. High-dimensional single-cell multimodal landscape of human carotid atherosclerosis. Arterioscler Thromb. Vasc. Biol. 44, 930–945 (2024).
Hao, K. et al. Integrative prioritization of causal genes for coronary artery disease. Circ. Genom. Precis Med. 15, e003365 (2022).
Stuart, T., Srivastava, A., Madad, S., Lareau, C. A. & Satija, R. Single-cell chromatin state analysis with Signac. Nat. Methods 18, 1333–1341 (2021).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Fornes, O. et al. JASPAR 2020: update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 48, D87–D92 (2020).
Weirauch, M. T. et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443 (2014).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Kim, J. B. et al. TCF21 and the environmental sensor aryl-hydrocarbon receptor cooperate to activate a pro-inflammatory gene expression program in coronary artery smooth muscle cells. PLoS Genet. 13, e1006750 (2017).
Erdmann, J., Kessler, T., Munoz Venegas, L. & Schunkert, H. A decade of genome-wide association studies for coronary artery disease: the challenges ahead. Cardiovasc Res. 114, 1241–1257 (2018).
Koyama, S. et al. Population-specific and trans-ancestry genome-wide analyses identify distinct and shared genetic risk loci for coronary artery disease. Nat. Genet. 52, 1169–1177 (2020).
Matsunaga, H. et al. Transethnic meta-analysis of genome-wide association studies identifies three new loci and characterizes population-specific differences for coronary artery disease. Circ. Genom. Precis Med. 13, e002670 (2020).
Lewis, M. J. & Wang, S. locuszoomr: an R package for visualizing publication-ready regional gene locus plots. Bioinform Adv. 5, vbaf006 (2025).
Wong, D. et al. FHL5 controls vascular disease-associated gene programs in smooth muscle cells. Circ. Res. 132, 1144–1161 (2023).
Acknowledgements
Support was provided to DL through the NIH grants F32HL165819, K08HL177173, and the Sarnoff Scholar Career Development Award. This work was supported by National Institutes of Health grants R01HL171045 (T.Q.), R01HL134817 (TQ), R01HL139478 (T.Q.), R01HL156846 (T.Q.), R01HL158525 (T.Q.), UM1HG011972 (T.Q.), U01HG011762 (T.Q.), R01HL171275 (R.W.), K08HL152308 (R.W.), R01HL171045 (A.K.), U01HG012069 (A.K.), K08HL153798 (P.C.), R01HL179083 (P.C.), R01HL181441(P.C.), K08HL167699 (C.W.), K08HL177251 (B.P.). This work was supported by American Heart Association Grants 23POST1018991 (W.G.), 24POST1187860 (J.M.), 24SCEFIA1248386 (P.C.), 20CDA35310303 (P.C.), the William G. Irwin Foundation (T.Q.), the Marfan Foundation Everest Award (P.C.) as well as a Human Cell Atlas grant (ZF2019-002437) from the Chan Zuckerberg Foundation (T.Q.). “Supplementary Figs.” created in BioRender. Li, D. (https://BioRender.com/22if1p3) is licensed under CC BY 4.0.
Author information
Authors and Affiliations
Contributions
T.Q., R.W., and A.K. conceived and supervised the research plan. R. W., D. L., P.C., S. K., A.Y., J.M., W.G., W.J., S.D., R.C., B.P., M.R., C.W., performed single-cell captures and single-cell analyses, D. L., and T. N. performed experiments with cultured cells, and helped with genomic analyses. M.W. collected samples for spatial transcriptomics, and D.L. and Q.Z. analyzed data. D.L., R.K., R.W., P.C., W.J., maintained mouse colonies and performed RNAScope experiments, D.L., S.K., R.W., P.C., A.Y., and Q.Z. performed analyses. D.L. and T. Q. wrote the manuscript, R.W. and S.K. contributed to writing and proofreading.
Corresponding author
Ethics declarations
Competing interests
T.Q. is on the scientific advisory board of Amgen. A.K. is a scientific co-founder Immunera; on the scientific advisory board of SerImmune, TensorBio; is a consultant with Bristol Myers Squibb, Arcardia Science, Inari, Precede Biosciences; and has a financial stake in DeepGenomics, Immunai, SerImmune, Freenome, Immunera and TensorBio. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Muredach Reilly and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, D.Y., Kundu, S., Cheng, P. et al. Vascular smooth muscle cell state trajectories mediate molecular mechanisms of coronary disease risk. Nat Commun 17, 4059 (2026). https://doi.org/10.1038/s41467-026-70530-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70530-z