Abstract
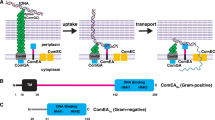
Natural transformation drives the spread of antibiotic resistance among bacteria. The DNA receptor ComEA is essential for transporting external transforming DNA into the periplasm by an unknown mechanism. Here, single-molecule optical tweezers and electron microscopy approaches show that Geobacillus stearothermophilus ComEA forms dynamic oligomers on DNA that can switch between two conformations depending on local concentration. When ComEA sparsely decorates DNA, it forms bridging oligomers that condense the DNA to generate sub-pN pulling forces. When ComEA more fully decorates DNA, it forms non-bridging oligomers that decondense DNA and cannot generate force. Mutating ComEA to favor either bridging or non-bridging conformations causes transformation deficiency in Bacillus subtilis, meaning condensation and decondensation each play mechanistic roles. Our results show that ComEA reversibly condenses DNA during natural transformation, first producing force to pull DNA into the periplasm and then abating force production to promote transport into the cytoplasm.
Similar content being viewed by others
Data availability
All data supporting the findings in this paper are provided in the main manuscript and its Supplementary files. Source data are provided within the source data file. Source data are provided with this paper.
References
Dubnau, D. & Blokesch, M. Mechanisms of DNA uptake by naturally competent bacteria. Annu. Rev. Genet. 53, 217–237 (2019).
Johnston, C., Martin, B., Fichant, G., Polard, P. & Claverys, J. Bacterial transformation: distribution, shared mechanisms and divergent control. Nat. Rev. Micro Biol. 12, 181–196 (2014).
Mell, J. C. & Redfield, R. J. Natural competence and the evolution of DNA uptake specificity. J. Bacteriol. 196, 1471–1483 (2014).
Winter, M., Buckling, A., Harms, K., Johnsen, P. J. & Vos, M. Antimicrobial resistance acquisition via natural transformation: context is everything. Curr. Opin. Microbiol. 64, 133–138 (2021).
Laxminarayan, R. et al. Antibiotic resistance—the need for global solutions. Lancet Infect. Dis. 13, 1057–1098 (2013).
Chen, I., Christie, P. J. & Dubnau, D. The ins and outs of DNA transfer in bacteria. Science 310, 1456–1460 (2005).
Damke, P. P. et al. ComFC mediates transport and handling of single-stranded DNA during natural transformation. Nat. Commun. 13, 1961 (2022).
Diallo, A. et al. Bacterial transformation: ComFA is a DNA-dependent ATPase that forms complexes with ComFC and DprA. Mol. Microbiol. 105, 741–754 (2017).
Provvedi, R. & Dubnau, D. ComEA is a DNA receptor for transformation of competent Bacillus subtilis. Mol. Microbiol. 31, 271–280 (1999).
Inamine, G. S. & Dubnau, D. ComEA, a Bacillus subtilis integral membrane protein required for genetic transformation, is needed for both DNA binding and transport. J. Bacteriol. 177, 3045–3051 (1995).
Gangel, H. et al. Concerted spatio-temporal dynamics of imported DNA and ComE DNA uptake protein during gonococcal transformation. PLoS Pathog. 10, e1004043 (2014).
Seitz, P. et al. ComEA is essential for the transfer of external DNA into the periplasm in naturally transformable Vibrio cholerae cells. PLoS Genet. 10, e1004066 (2014).
Seitz, P. & Blokesch, M. DNA transport across the outer and inner membranes of naturally transformable Vibrio cholerae is spatially but not temporally coupled. mBio 5, 01409–01414 (2014).
Takeno, M., Taguchi, H. & Akamatsu, T. Role of ComEA in DNA uptake during transformation of competent Bacillus subtilis. J. Biosci. Bioeng. 113, 689–693 (2012).
Hahn, J., DeSantis, M. & Dubnau, D. Mechanisms of transforming DNA uptake to the periplasm of Bacillus subtilis. mBio 12, 01061–21 (2021).
Ahmed, I. et al. Structure-function studies reveal ComEA contains an oligomerization domain essential for transformation in gram-positive bacteria. Nat. Commun. 13, 7724 (2022).
Odijk, T. Stiff chains and filaments under tension. Macromolecules 28, 7016–7018 (1995).
Baumann, C. G., Smith, S. B., Bloomfield, V. A. & Bustamante, C. Ionic effects on the elasticity of single DNA molecules. Proc. Natl. Acad. Sci. USA 94, 6185–6190 (1997).
Chemla, Y. R. et al. Mechanism of force generation of a viral DNA packaging motor. Cell 122, 683–692 (2005).
Bustamante, C., Marko, J. F., Siggia, E. D. & Smith, S. Entropic elasticity of λ-phage DNA. Science 265, 1599–1600 (1994).
Liphardt, J., Onoa, B., Smith, S. B., Tinoco Jr, I. & Bustamante, C. Reversible unfolding of single RNA molecules by mechanical force. Science 292, 733–737 (2001).
Bennink, M. L. et al. Unfolding individual nucleosomes by stretching single chromatin fibers with optical tweezers. Nat. Struct. Biol. 8, 606–610 (2001).
Foroutannejad, S. et al. The cofactor-dependent folding mechanism of Drosophila cryptochrome revealed by single-molecule pulling experiments. Nat. Commun. 14, 1057 (2023).
Hao, Y., Canavan, C., Taylor, S. S. & Maillard, R. A. Integrated method to attach DNA handles and functionally select proteins to study folding and protein-ligand interactions with optical tweezers. Sci. Rep. 7, 10843 (2017).
Mickolajczyk, K. J., Olinares, P. D. B., Chait, B. T., Liu, S. & Kapoor, T. M. The MIDAS domain of AAA mechanoenzyme Mdn1 forms catch bonds with two different substrates. eLife 11, e73534 (2022).
Smith, S. B., Cui, Y. & Bustamante, C. Overstretching B-DNA: the elastic response of individual double-stranded and single-stranded DNA molecules. Science 271, 795–799 (1996).
van Mameren, J. et al. Unraveling the structure of DNA during overstretching by using multicolor, single-molecule fluorescence imaging. Proc. Natl. Acad. Sci. USA 106, 18231–18236 (2009).
Burghard-Schrod, M., Kilb, A., Krämer, K. & Graumann, P. L. Single-molecule dynamics of DNA receptor ComEA, membrane permease ComEC, and taken-Up DNA in competent Bacillus subtilis cells. J. Bacteriol. 204, 572 (2022).
Peskin, C. S., Odell, G. M. & Oster, G. F. Cellular motions and thermal fluctuations: the Brownian ratchet. Biophys. J. 65, 316–324 (1993).
Uhl, M. & Seifert, U. Force-dependent diffusion coefficient of molecular Brownian ratchets. Phys. Rev. E 98, 022402 (2018).
Rohde, M. The Gram-positive bacterial cell wall. Microbiol. Spectr. 7, gpp3–gpp2018 (2019).
Dorman, C. J. H-NS: a universal regulator for a dynamic genome. Nat. Rev. Microbiol. 2, 391–400 (2004).
Wien, F. et al. Amyloid-like DNA bridging: a new mode of DNA shaping. Nucleic Acids Res. 53, gkaf169 (2025).
Chen, J. M. et al. Lsr2 of Mycobacterium tuberculosis is a DNA-bridging protein. Nucleic Acids Res. 36, 2123–2135 (2008).
Erkelens, A. M. et al. The B. subtilis Rok protein is an atypical H-NS-like protein irresponsive to physico-chemical cues. Nucleic Acids Res. 50, 12166–12185 (2022).
Harju, J. & Broedersz, C. P. Physical models of bacterial chromosomes. Mol. Microbiol. 123, 143–153 (2025).
Graham, T. G. et al. ParB spreading requires DNA bridging. Genes Dev. 28, 1228–1238 (2014).
Schumacher, M. A., Piro, K. M. & Xu, W. Insight into F plasmid DNA segregation revealed by structures of SopB and SopB–DNA complexes. Nucleic Acids Res. 38, 4514–4526 (2010).
Krämer, H. et al. lac repressor forms loops with linear DNA carrying two suitably spaced lac operators. EMBO J. 6, 1481 (1987).
Lobell, R. B. & Schleif, R. F. DNA looping and unlooping by AraC protein. Science 250, 528–532 (1990).
Ryu, J. et al. Bridging-induced phase separation induced by cohesin SMC protein complexes. Sci. Adv. 7, eabe5905 (2021).
Umland, T. C., Wei, S., Craigie, R. & Davies, D. R. Structural basis of DNA bridging by barrier-to-autointegration factor. Biochemistry 39, 9130–9138 (2000).
Skoko, D. et al. Barrier-to-autointegration factor (BAF) condenses DNA by looping. Proc. Natl. Acad. Sci. USA 106, 16610–16615 (2009).
Bell, N. A. & Molloy, J. E. Single-molecule force spectroscopy reveals binding and bridging dynamics of PARP1 and PARP2 at DNA double-strand breaks. Proc. Natl. Acad. Sci. USA 120, e2214209120 (2023).
Gien, H. et al. HIV-1 nucleocapsid protein binds double-stranded DNA in multiple modes to regulate compaction and capsid uncoating. Viruses 14, 235 (2022).
Van den Broek, B. et al. Visualizing the formation and collapse of DNA toroids. Biophys. J. 98, 1902–1910 (2010).
Taylor, J. A. et al. Specific and non-specific interactions of ParB with DNA: implications for chromosome segregation. Nucleic Acids Res. 43, 719–731 (2015).
Tišma, M. et al. Dynamic ParB–DNA interactions initiate and maintain a partition condensate for bacterial chromosome segregation. Nucleic Acids Res. 51, 11856–11875 (2023).
Keenholtz, R. A. et al. Oligomerization and ATP stimulate condensin-mediated DNA compaction. Sci. Rep. 7, 1–13 (2017).
Ganji, M. et al. Real-time imaging of DNA loop extrusion by condensin. Science 360, 102–105 (2018).
Strick, T. R., Kawaguchi, T. & Hirano, T. Real-time detection of single-molecule DNA compaction by condensin I. Curr. Biol. 14, 874–880 (2004).
Wu, M. et al. Human topoisomerase iiα promotes chromatin condensation via a phase transition. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/2024.10.15.618281v1.full (2024).
Joyeux, M. Role of salt valency in the switch of H-NS proteins between DNA-bridging and DNA-stiffening modes. Biophys. J. 114, 2317–2325 (2018).
Amit, R., Oppenheim, A. B. & Stavans, J. Increased bending rigidity of single DNA molecules by H-NS, a temperature and osmolarity sensor. Biophys. J. 84, 2467–2473 (2003).
Liu, Y., Chen, H., Kenney, L. J. & Yan, J. A divalent switch drives H-NS/DNA-binding conformations between stiffening and bridging modes. Genes Dev. 24, 339–344 (2010).
Smith, S. B., Cui, Y. & Bustamante, C. in Methods in Enzymology 134–162 (Elsevier, 2003).
Gieseler, J. et al. Optical tweezers—from calibration to applications: a tutorial. Adv. Opt. Photonics 13, 74–241 (2021).
Mickolajczyk, K. J. et al. Long-range intramolecular allostery and regulation in the dynein-like AAA protein Mdn1. Proc. Natl. Acad. Sci. USA 117, 18459–18469 (2020).
Mickolajczyk, K. J. et al. Force-dependent stimulation of RNA unwinding by SARS-CoV-2 nsp13 helicase. Biophys. J. 120, 1020–1030 (2021).
Gao, X. et al. Functional characterization of core components of the Bacillus subtilis cyclic-di-GMP signaling pathway. J. Bacteriol. 195, 4782–4792 (2013).
Albano, M., Hahn, J. & Dubnau, D. Expression of competence genes in Bacillus subtilis. J. Bacteriol. 169, 3110–3117 (1987).
Acknowledgements
We thank members of the Mickolajczyk, Neiditch, and Dubnau Labs for useful discussions. We thank Rajesh Patel for his technical knowledge and the RWJMS Core EM facility that he manages. J.S. was supported by Rutgers IRACDA K12 GM093854 from the NIH. Support for this work was provided by the National Institutes of Health through grants R01GM057720 (M.B.N. and D.D.) and R35GM157075 (K.J.M.). Additional support was provided by a Busch Biomedical Grant from the Office for Research at Rutgers (K.J.M.).
Author information
Authors and Affiliations
Contributions
M.B.N., D.D., and K.J.M. conceived of the project. J.S., G.A., and K.J.M. carried out optical tweezer data acquisition and analysis. J.M., H.C., and A.R. carried out fluorescence polarization and electron microscopy experiments. I.A. prepared recombinant protein samples with guidance from M.B. J.H. carried out B. subtilis experiments with guidance from D.D. K.J.M. wrote the paper together with M.B.N. and D.D., and with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Santiago, J.I., Ahmed, I., Hahn, J. et al. Reversible DNA condensation drives natural transformation. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70559-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70559-0