Abstract
RNA-sequencing’s conversion of molecules to reads is inconsistent. Experiment-to-experiment variations (systemic bias) create batch effects, while gene-to-gene variations (sequence-dependent bias) invalidate inter-gene comparisons, precluding a universal scale. This confines analysis to relative fold-changes, a metric unreliable across batches. We introduce TranScale: 100 biomimetic standards with SI-traceable concentrations certified by Isotope Dilution Mass Spectrometry. Co-processed within samples, they empirically characterize systemic and sequence-dependent biases, generating a library-specific calibration curve (R² > 0.97) to convert reads into absolute quantities. This approach reveals that consistent fold-changes can mask severe absolute errors, exposing systemic biases missed by conventional QC. Across laboratories, this calibration reduced median inter-lab CV from >85% to <25% and increased biological signal-to-noise from ~0 to >7.9, outperforming the widely-used tool ComBat. By anchoring RNA-seq to the SI, our work establishes the metrological foundation for data interoperability and universal benchmarks, enabling absolute comparisons of SI-traceable quantities between any two genes.
Similar content being viewed by others
Introduction
RNA sequencing (RNA-seq) is a core methodology in biomedical research, pivotal for discovering biomarkers and elucidating disease mechanisms1. Its profound impact, however, is challenged by a significant challenge: sequencing reads are not a direct measure of molecular abundance but are instead surrogates for expression2. This is because two distinct categories of bias distort the measurement process, making the conversion of reads to absolute molecular counts impossible without proper calibration3. First, systemic biases for the same gene or transcript inherent in library preparation and sequencing protocols create severe batch effects that compromise reproducibility4, with inter-laboratory coefficients of variation (CVs) reported to reach 85.1%5. This inconsistency fundamentally hampers biomarker validation6 and the establishment of universal clinical decision thresholds7. Second, and more fundamentally, sequence-dependent biases tied to transcript features like GC-content and secondary structure render direct comparisons of gene expression levels between different genes scientifically invalid, even within a single sample. Together, these limitations confine the entire field to analyzing relative fold-changes—a metric that is itself unreliable across different batches.
The community has pursued two distinct strategies to address this. One approach relies on post-hoc computational tools like ComBat8, which, while valuable, correct statistical discrepancies without addressing the underlying measurement uncertainty, risking the conflation of biological heterogeneity with technical artifacts9. The other, more fundamental strategy has focused on physical reference materials. Early internal standards like ERCC10 were critical for assessing technical performance but, being non-biomimetic, fail to model the behavior of endogenous transcripts and thus cannot correct for sequence-specific biases11. Later, biomimetic standards like Sequins12 improved biological fidelity but, like ERCC, lacked SI-traceable value assignment, preventing them from serving as true absolute calibrators. Concurrently, landmark consortia like the MAQC/SEQC4 consortia and the Quartet project13 defined the state-of-the-art for reproducible relative quantification, reinforcing the boundaries of the existing analytical approach rather than advancing beyond them.
Thus, despite these sophisticated efforts, a universal solution for true absolute quantification across the transcriptome remained unmet. This requires a direct, metrologically traceable link between sequencing reads and molecular counts—a principle whose importance for quantitative biology is increasingly recognized14. The absence of this link is a critical barrier preventing RNA-seq from becoming a fully quantitative discipline, limiting its use where accuracy is paramount, such as defining universal diagnostic cutoffs15 or enabling systems-level stoichiometric modeling of cellular networks. While niche applications have demonstrated absolute quantification for specific RNA classes, such as AQRNA-seq for microRNAs16, their specialized designs underscore that a universal, transcriptome-wide solution has not yet been achieved.
Here, we present TranScale, a set of biomimetic RNA standards with SI-traceable certified values, and demonstrate its use within a comprehensive framework for both diagnostic validation and absolute calibration of RNA-seq workflows. Their efficacy stems from a distinct integration of three synergistic principles that directly address the aforementioned biases. By being co-processed with samples, they physically experience and thus correct for the systemic biases that cause batch effects. Their biomimetic design emulates endogenous transcripts, allowing them to accurately model and correct for the sequence-dependent biases that invalidate inter-gene comparisons. Finally, their absolute quantities, certified with SI-traceability via isotope dilution mass spectrometry (IDMS)17,18, provide the invariable anchor required to convert the entire measurement to an absolute scale. This establishes a clear metrological chain from the sequencing reads back to the mole19, contributing to the broader goal of integrating measurement science into biology14. We demonstrate that this framework not only substantially reduces inter-laboratory CV but also enables the absolute comparison of different genes across the transcriptome. This work thus provides a framework and a set of tools to address a long-standing issue in the field, facilitating the advancement of RNA-seq from a comparative towards a more quantitative discipline.
Results
A metrological framework for absolute and comparable RNA quantification
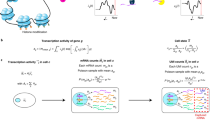
To address the major hurdle of conventional RNA sequencing, we developed a comprehensive metrological framework designed to achieve both data harmonization across samples and absolute comparability between genes (Fig. 1). The foundation of this framework is a set of 100 biomimetic RNA spike-in transcripts, engineered to mirror the complexity of the human transcriptome and designed to be non-interfering, named TranScale (Fig. 1a). Crucially, each transcript was assigned a certified copy number concentration with SI traceability using a primary reference method, isotope dilution mass spectrometry (IDMS), thus anchoring all subsequent measurements to a stable, absolute scale (Fig. 1b).
a Design of biomimetic transcripts of TranScale. The set of 100 transcripts was engineered to mimic human transcriptome complexity (e.g., multi-exon genes, alternative splicing, fusion events) and incorporated mirror sequences to prevent interference with endogenous gene detection. b Assignment of absolute copy numbers with SI traceability. The certified value for each transcript was determined using isotope dilution mass spectrometry (IDMS), a primary reference measurement procedure (see Methods for details), establishing a metrological chain traceable to the SI unit mole. c Experimental design and calibration workflow. Two biological sample sets were spiked-in and sequenced across 12 batches, varying by lab, library preparation protocol, and sequencing platform to generate substantial batch effects. The calibration workflow involves a quality screening of spike-in performance followed by the generation of a library-specific linear regression curve. Technical replicates are libraries (n = 3). d Principal component analysis (PCA) of uncalibrated data. Sample clustering is dominated by technical factors (e.g., lab, protocol) rather than biological identity. e PCA of calibrated data. After calibration, batch effects are removed, and samples cluster correctly according to their true biological groups. f Enabling inter-gene comparison. The framework converts relative expression units (e.g., Fragments Per Kilobase of transcript per Million mapped reads, FPKM) into absolute copy numbers, allowing for the direct quantitative comparison between different genes within a sample.
To rigorously test this framework, we designed a multi-laboratory study intended to generate substantial batch effects (Fig. 1c). As anticipated, principal component analysis (PCA) of the uncalibrated data from two distinct biological sample sets confirmed that technical variability effectively obscured the underlying biological differences, with samples clustering by lab and library preparation method (Fig. 1d). This result exemplifies the core challenge of data comparability in large-scale transcriptomics.
Our framework introduces a library-specific calibration strategy that leverages TranScale as SI-traceable spike-ins to address these challenges. As a proof-of-concept, we demonstrate that this calibration accomplishes the two primary goals of quantitative transcriptomics. First, it effectively removes confounding batch effects, restoring the correct clustering of samples based on their biological identity and thus enabling robust inter-sample comparability (Fig. 1e). Second, it transforms relative expression units into absolute copy numbers, permitting direct and meaningful inter-gene comparisons within any given sample (Fig. 1f). Having established the framework’s capability to address these core challenges, we proceeded with a rigorous metrological evaluation of its performance and a thorough validation of its application to endogenous human transcripts in the subsequent sections.
Design and metrological characterization of the TranScale
The foundation of our calibration strategy is a purpose-built TranScale designed for both biological relevance and metrological rigor (Fig. 2). To achieve this, we designed a library of 100 transcripts to be both biomimetic and computationally orthogonal. The biomimetic properties are detailed in Fig. 2a. Specifically, the library was designed not only with wild-type sequences but also to include clinically relevant transcript variants such as alternative splicing isoforms, single-nucleotide variants, and fusion genes, thus reflecting the complexity of biological samples. Furthermore, the 100 transcripts cover a wide length distribution (500–3800 nt) (Supplementary Table 1), and their proportional representation across different length bins was designed to be comparable to that of the human transcriptome. Similarly, the GC content of the spike-ins (40–50%) spans the range typically observed in human genes. All transcripts were confirmed to have high purity (Supplementary Table 2, Supplementary Fig. 1). Critically, we utilized inverted mirror sequences of human genes. This design ensures that the spike-ins behave similarly to endogenous transcripts during the physical steps of library preparation and sequencing, while remaining computationally distinct, thus preventing analytical ambiguity.
a Physicochemical properties of the 100 RNA spike-in transcripts of TranScale. Plots show the composition of different transcript types (pie chart, left), the distribution of transcript lengths (line plot, middle left), the proportion of transcripts in different length bins compared to human transcripts (bar chart, middle right), and the GC content compared to human transcripts (dot plot, right). b Certified values for TranScale. The heatmap shows the log2 (Mix1/Mix2) concentration ratios across different concentration tiers (left). Line plots show the certified log2 concentrations with expanded uncertainties for each of the 100 transcripts in Mix1 (top right), Mix2 (middle right), and their log2 ratio (bottom right). Technical replicates are defined as the independent measurements performed to determine the absolute copy number concentration of each gene in the TranScale (n = 12). Data are presented as reference values ± expanded uncertainties (k = 2) of TranScale. c Orthogonal validation of certified values. Scatter plots compare the copy numbers assigned by IDMS (x-axis) with those determined by RT-dPCR (y-axis) for Mix1, Mix2, and their ratio. The identity line (y = x) and Pearson’s correlation coefficient (r) are shown. d Assessment of sequencing orthogonality. The bar chart shows the percentage of total reads aligning to the TranScale sequences versus the human genome (GRCh38) alone across different library preparation (L1, L2) and sequencing platforms (ILL, DNB). Technical replicates are libraries (n = 3). Data are presented as mean values ± SD. Source data for this figure is available in the Source Data file.
Next, to establish a metrological anchor for absolute quantification, we first assigned certified values with SI traceability to the individual stock solutions of each of the 100 transcripts using isotope dilution mass spectrometry (IDMS), a gold-standard reference measurement procedure (Supplementary Fig. 2, Supplementary Table 3). To ensure this traceability, the absolute concentration of each transcript was determined against primary standards from the National Institute of Metrology (NIM) (Supplementary Table 4), and the procedure demonstrated high consistency across all measurands (Supplementary Table 5). The copy number concentration of each of the 100 transcript stocks was determined by averaging 12 measurement results derived from four repeated digestions across two independent vials. This rigorous characterization process yielded high precision, with relative standard deviations (RSDs) for all transcripts ranging from 1.06% to 9.04% (all well below 10%) (Supplementary Data 1). Following the theoretical design matrix (Fig. 2b, heatmap), these certified stocks were then used to prepare two master mixes (Mix 1 and Mix 2) via precision gravimetry.
The final certified concentrations in these mixes were derived from the certified stock values and high-precision gravimetric data. The certified absolute concentrations for all 100 transcripts in both Mix 1 and Mix 2 of TranScale are presented (Fig. 2b, top and middle right plots; Supplementary Data 2). These values span the intended 10⁵-fold dynamic range, making them suitable for calibrating a wide spectrum of transcript abundances. Crucially, a comprehensive uncertainty budget was established for each transcript to define the final certified values and their associated expanded uncertainties (k = 2). This evaluation systematically combined the uncertainty contributions from the initial characterization (uchar, incorporating uncertainties from both IDMS and gravimetric preparation), between-vial homogeneity (ubb), and long-term stability (ults), as detailed in Supplementary Data 3. The final expanded relative uncertainties for the absolute copy number concentrations were confirmed to be ≤16% for Mix 1 and ≤18% for Mix 2. The resulting uncertainty for the critical Mix 1-to-Mix 2 expression ratios was confirmed to be ≤23%. Furthermore, the log₂ expression ratios between the two mixes were certified to establish the definitive metrological anchor for relative quantification. While the design targeted a nominal 256-fold range (Log₂ from −4 to +4), the certified values defined an actual 378-fold dynamic range (Log₂ from −3.84 to +4.73) (Fig. 2b, bottom right plot).
To independently confirm the accuracy of the IDMS-certified values of TranScale, we performed orthogonal validation using one-step reverse transcription digital PCR (RT-dPCR) (Supplementary Table 6, Supplementary Data 4, Supplementary Fig. 3-7), and their reverse transcription efficiencies were validated (Supplementary Table 7). For a representative subset of 20 transcripts spanning the full concentration range, the copy numbers determined by RT-dPCR showed a high degree of concordance with the IDMS-assigned values (Pearson’s r > 0.99), verifying the ground-truth accuracy of our RM (Fig. 2c, Supplementary Table 8). Furthermore, TranScale successfully met the stringent ISO criteria for homogeneity and stability (Supplementary Table 9, Supplementary Data 5–8), ensuring its reliability for widespread distribution and use.
Finally, we confirmed the sequencing orthogonality of the spike-in RM. We sequenced TranScale alone using multiple library preparations (poly(A) selection, rRNA depletion) and sequencing platforms (Illumina NovaSeq, MGI DNBSEQ-T7). In all cases, reads aligned with high efficiency (>99.1%) to the TranScale reference sequences, while a negligible fraction (<0.0015%) misaligned to the human genome (hg38) (Fig. 2d). This result confirms that TranScale is analytically “clean” and can be confidently co-processed with biological samples to enable absolute quantification without introducing cross-contamination or analytical artifacts.
Metrological diagnosis of systemic and sequence-dependent measurement errors
To directly assess the accuracy of uncalibrated RNA-seq data, we established a metrological diagnostic framework based on TranScale. A direct comparison of FPKM values against their SI-traceable reference values revealed significant transcript-specific scatter within each batch, alongside unpredictable global offsets between batches (Supplementary Fig. 8). To deconstruct these errors, we developed a dual-metric system. The first metric, Absolute ME, isolates and quantifies the transcript-specific, sequence-dependent bias for each calibrator by comparing its mean-normalized FPKM to its certified SI-traceable value. The wide dispersion of Absolute ME values observed within every batch (Fig. 3a) confirms that individual transcripts are quantified with vastly different efficiencies, providing direct evidence that uncalibrated inter-gene comparisons are fundamentally unreliable.
a Violin plots of the Measurement Error (ME) for the absolute quantification of 100 spike-in transcripts in Mix1 and Mix2 across eight experimental batches. ME is calculated as the percentage difference between the measured (mean normalized FPKM) and certified copy numbers (all determined with log2 transformed). The green line indicates the ideal value of ME = 0. b Violin plots of the ME for the ratio of measured copy numbers between Mix1 and Mix2, compared against the ratio of their certified values, for each of the 100 spike-in pairs across the eight batches. c Boxplots of the Root Mean Square Error (RMSE) for absolute quantification in Mix1 and Mix2 across the batches. Each box presented the RMSE values for 100 spike-ins between observed and reference values within a batch. d Bar plots of the Spearman Correlation Coefficient (RC) between measured and certified copy numbers for Mix1 and Mix2 across the batches. The dashed line indicates a reference value of RC = 0.89. For violin plots, the white box represents the interquartile range (IQR), the center line is the median, and the shaded area shows the probability density. For all boxplots, the center line indicates the median, the box limits represent the upper and lower quartiles (IQR), and the whiskers extend to 1.5 × IQR. Technical replicates are libraries (n = 3). Source data for this figure is available in the Source Data file.
To test the bedrock assumption of transcriptomics—the accuracy of relative quantification—we introduced a second metric, Ratio ME. This metric directly quantifies the error in the measured expression ratio between pairs of calibrators against their true SI-traceable ratio (Fig. 3b). This dual-metric diagnosis led to a crucial, paradoxical finding. The L2_DNB_P batch, despite exhibiting the most chaotic sequence-dependent biases (the largest Absolute ME dispersion), preserved relative expression ratios with high precision. Conversely, the L2_DNB_R batch, while showing high internal consistency (a narrow Absolute ME distribution), severely distorted the true quantitative ratios.
These contrasting batches provide definitive, orthogonal evidence that high relative precision can mask severe absolute measurement errors, and vice versa. This dismantles the foundational assumption that relative precision is a sufficient proxy for data quality. The magnitude of the observed Ratio ME dwarfs the certified uncertainty of TranScale (<23%), unequivocally attributing the overwhelming majority of quantification error to the RNA-seq workflow itself.
Finally, the performance hierarchy across batches identified by our dual-metric framework was independently corroborated by standard indicators like Root Mean Square Error (RMSE) and Spearman’s Correlation (RC). However, while metrics like RC assess precision, only our metrological approach measures trueness—the deviation from a known standard. This analysis also confirmed that quantification accuracy degrades for low-abundance transcripts, as seen with Mix2 (Fig. 3c, d). Collectively, these findings establish that a dual-metric, metrological evaluation is critical to fully characterize workflow performance and expose hidden vulnerabilities that conventional, relative-based QC metrics would miss.
A library-specific strategy for measurement bias correction
To correct the observed quantification errors, we implemented a library-specific calibration approach based on creating a custom calibration curve for each library. This approach first identifies a set of high-confidence internal calibrators among TranScale by filtering for spike-in transcripts with a measurement error (ME) within a predefined accuracy threshold (e.g., ±5% in log2 scale). The number of these reliable calibrators varied with batch quality, ranging from over 75 in the highest-quality batch to fewer than 40 in lower-quality batches (Fig. 4a).
a Number of high-confidence calibrators among TranScale (spike-in transcripts with a measurement error, ME, within ±5% of their certified values) were identified in each experimental batch for Mix1 and Mix2. b Representative calibration curves for a single library across different library preparation protocols and sequencing platforms. The plots show the relationship between certified input copy number and measured FPKM on a log2 scale for spike-ins from Mix1 and Mix2. The solid lines represent the linear regression fits (R² ≥ 0.97). Results from only one of the three replicate library preparations are shown. Individual points represent the calibration factors of TranScale that were retained after post-sequencing threshold filtering. Heatmaps showing the ratio of calibrated copy number (in a log2 scale) to certified reference copy number (in a log2 scale) for all 100 spike-in transcripts in Mix1 (c) and Mix2 (d) across all experimental batches after library-specific calibration. Comparison of the coefficient of variation (CV) for absolute copy number measurements before (uncalibrated) and after (calibrated) correction, for both Mix1 and Mix2. The comparisons isolate variability arising from different library preparation protocols (e), sequencing platforms (f), combinations of protocols and platforms (g), and the full cross-study dataset combining labs, protocols, and platforms (h). For violin plots, data are presented as the coefficients of variation (CV) of the observed FPKM or corrected absolute copy numbers (log2 scale) for 100 spike-ins across different sequencing batches. The white box represents the interquartile range (IQR), and the whiskers extend to 1.5 × IQR. The center line is the median, and the shaded area shows the probability density. Technical replicates are libraries (n = 3). Source data for this figure is available in the Source Data file.
Plotting the certified input copy numbers of these selected calibrators against their measured FPKM values (in a log2 scale) revealed a highly linear relationship (R² > 0.97) in every batch, irrespective of its overall accuracy (Fig. 4b). This stable, library-specific linear relationship forms the basis of our calibration method. It implies that within a given library preparation, the combined effects of systemic (batch-level) and sequence-dependent (transcript-level) biases manifest as a consistent linear transformation of the true quantities. Therefore, a library-specific linear model is sufficient to reverse this transformation and restore absolute accuracy.
The calibration model, derived from a subset of high-confidence calibrators, demonstrated robust generalization across the entire spike-in panel. Post-correction, the ratio of measured-to-certified copy numbers (in a log2 scale) for all 100 spike-ins converged tightly around 1.0 across every batch, including for transcripts initially measured with high error (Fig. 4c, d, Supplementary Fig. 9). The framework’s robustness was compellingly demonstrated by its successful normalization of the two most challenging batches, each representing a distinct failure mode identified in our diagnostic phase. It corrected the L2_DNB_P batch, which harbored large absolute errors masked by apparent relative precision, as well as the L2_DNB_R batch, which was defined by severe sequence-dependent biases (high Absolute ME dispersion). The ability of a single linear model to reduce both systemic and sequence-dependent biases validates the core theoretical premise of our metrological approach.
Notably, the process also identified a consistent subset of “recalcitrant” transcripts that resisted correction across all batches (Supplementary Data 9). These transcripts were designed with features known to challenge short-read quantification, including complex alternative splicing and high sequence homology arising from repetitive elements, fusion/wild-type pairs, and single nucleotide variants (SNVs). For some transcripts, such as R17 and R42, these challenges were compounded by their low abundance, where stochastic noise amplifies quantification errors. The failure to calibrate these challenging sequences indicates a breakdown in the upstream bioinformatic workflow. This highlights the framework’s utility not only for correction but also for diagnosing the inherent limitations of standard RNA-seq data processing pipelines, a finding that is further explored in the Discussion.
To quantify the impact of this harmonization on data comparability, we calculated the coefficient of variation (CV) for absolute quantity measurements across different sources of technical variability. Across various conditions—including different protocols (Fig. 4e), platforms (Fig. 4f), and combinations thereof (Fig. 4g)—calibration consistently reduced the median CV. Even in the most challenging cross-study comparison (different labs, platforms, and protocols), the median CV for absolute copy numbers was reduced from over 85% to below 15% post-calibration (Fig. 4h).
Harmonization of absolute quantification across batches
To ensure the robustness of our calibration framework, we first established an optimized input ratio for the TranScale calibrators. Our standard protocol uses 1 ng of TranScale per µg of total RNA, which resulted in TranScale-derived reads constituting approximately 1.40%-4.57% of the total library (Supplementary Data 10). Through an in silico downsampling analysis (Supplementary Fig. 10a), we confirmed that this input amount provides a safe margin above the minimum requirement for robust model fitting across different platforms and library preparation methods. Guided by this validated and stringent QC process (Supplementary Fig. 10b), we then applied the calibration framework to resolve inter-laboratory batch effects for endogenous genes. Before correction, absolute quantities (FPKM in a log2 scale) of endogenous genes in the Quartet reference materials (RM D5 and D6) exhibited high cross-batch variability (Fig. 5a, b, Supplementary Fig 11a–c), particularly across labs, platforms, and protocols where median CVs exceeded 85%. Our physical calibration framework significantly improved data concordance, reducing median CVs to consistently below 25% across all comparison strata (lab, platform, and protocol) (Fig. 5a, b).
Distribution of Coefficient of Variation (CV) for endogenous gene in Quartet samples across-lab, platform, and protocol. before and after applying our calibration method for D5 (a) and (b). c-f, Principal Component Analysis (PCA) of transcriptomic data from D5 and D6 cell lines. In all plots, samples are colored by biological group (D5 or D6) and distinguished by shape according to their technical batch. The Signal-to-Noise Ratio (SNR) for the separation of biological groups is shown below each plot. Panels show data from: c uncalibrated absolute FPKM; d FPKM corrected with SVA; e FPKM corrected with ComBat; f absolute copy numbers after our library-specific calibration workflow. PCA of transcriptomic data from a second, independent dataset comparing LCA and LCN cell lines. Samples are colored by biological group (LCA or LCN) and distinguished by shape according to technical batch. Panels show data from: (g) uncalibrated absolute FPKM; (h) absolute copy numbers after our library-specific calibration workflow. In all violin plots, the white box indicates the interquartile range (IQR), the center line is the median, and the violin shape represents the probability density of the data. Data before (observed FPKM) and after (absolute copy number measurements) correction were used to calculate the CV, respectively. Data are presented as the coefficients of variation (CV) of the observed FPKM or corrected absolute copy numbers (log2 scale) for genes detected in all libraries (N = 14,430 for D5, 14,906 for D6, 49,560 for LCA and LCN). Technical replicates are libraries (n = 3). Source data for this figure is available in the Source Data file (https://github.com/zhyu0807/TranScale/tree/main/Source_Data53).
This improvement in precision was critical for resolving biological signals. As expected, principal component analysis (PCA) of uncorrected absolute data showed samples clustering entirely by experimental batch, yielding a signal-to-noise ratio (SNR) of approximately zero (Fig. 5c). We then compared our physical calibration against conventional computational and relative normalization strategies. Widely-used computational tools offered limited improvements, with SVA failing to separate the groups (SNR = 1.2, Fig. 5d) and ComBat achieving partial separation (SNR = 5.9, Fig. 5e). In contrast, our physical calibration segregated the D5 and D6 biological groups from technical artifacts, achieving a strong SNR of 7.9, with 75.4% of the total variance explained by PC1 and PC2 (Fig. 5f). This result demonstrates that prospectively correcting the measurement data is more effective than retrospectively modeling statistical artifacts.
To further characterize our calibrator’s utility, we examined two additional strategies also enabled by TranScale. A purely ratio-based normalization, which is inherently designed to maximize relative separation, achieved a higher SNR of 9.0 (Supplementary Fig. 11d). Notably, a hybrid approach—applying a ratio-based transformation to our already calibrated absolute data—yielded the highest signal separation of all methods (SNR = 11.1, Supplementary Fig. 11e). While these ratio-based transformations excel at maximizing visual separation for discovery, our primary physical calibration is distinct in providing the absolute measurement results with SI traceability that are essential for establishing universal, quantitative benchmarks.
We validated this performance hierarchy in a clinically relevant context using paired lung cancer (LCA) and normal (LCN) cell line samples. In the uncorrected data, strong batch effects partially masked the cancer phenotype (SNR = 5.6, Fig. 5g). Applying our primary absolute calibration method eliminated this noise, resolving the tumor and normal samples and increasing the SNR to 11.8 (Fig. 5h). Confirming the trend observed in the Quartet data, the hybrid approach once again provided the most distinct visual separation, achieving an SNR of 14.7 (Supplementary Fig. 11f).
Collectively, these results demonstrate two key findings. First, our physical calibration method is more effective than the tested computational tools for removing batch effects. Second, while ratio-based transformations are powerful for optimizing relative data visualization, our framework is distinct in that it produces the foundational, absolute measurement results with SI traceability. It is this absolute quantification that is crucial for building universal diagnostic models and achieving true data interoperability in clinical genomics.
Validation of inter-gene comparability and absolute accuracy
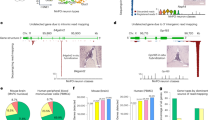
To validate the absolute accuracy and inter-gene comparability for endogenous transcripts, we upgraded the Quartet RNA reference materials from a relative ratio-based standard to one with SI-traceable absolute quantities. We defined a set of conserved, cross-batch detected genes by retaining transcripts detected in all eight batches, with an FPKM > 0.1 in all three technical replicates within each qualifying batch. High-quality batches were screened based on absolute quantitative performance evaluation and inter-batch statistical consistency. This process generated a new, dual-function reference dataset containing absolute copy numbers for 13,483 genes in RM D5 and 13,862 genes in RM D6 (Fig. 6a, Supplementary Fig. 12, Supplementary Data 12). The metrological quality of this dataset was high, with over 84% of the assigned absolute values having a relative standard uncertainty (uchar) below 20%.
a Density plots of log₂ absolute copy number distributions for quantified genes in samples D5 (blue, 11,398 genes) and D6 (green, 12,164 genes). Orthogonal validation of calibrated RNA-seq measurements. Scatter plots compare log2 absolute copy numbers from RNA-seq with those from RT-dPCR for (b) sample D5 and (c) sample D6. d Comparison of the log2 expression ratio (D5/D6) measured by RNA-seq against the reference ratio from RT-dPCR. Measurement error (ME) of the absolute quantities (with mean-normalization) with absolute datasets of D5 (e) and D6 (f) constructed by TranScale across eight sequencing batches. Data are presented for genes that were detected in every batch and covered by the established absolute quantification reference datasets (N = 10,000 for D5 and D6). Calibration performance evaluation. Plots show D5/D6 ratio MEs before (g) and after (h) library-specific calibration by TranScale. MEs are calculated by the D5/D6 ratio from the reference datasets reported previously. Data are presented for genes that were detected in every batch and covered by the external ratio-based quantification reference datasets (N = 6740 for D5 and D6). For violin plots, the white box represents the interquartile range (IQR), and the whiskers extend to 1.5 × IQR. The center line is the median, and the shaded area shows the probability density. Technical replicates are libraries (n = 3). i Comparison of quantification profiles for a set of housekeeping genes using a relative metric (log2 FPKM, left) versus calibrated absolute copy numbers (log2 copy number, right). j Direct comparison of calibrated absolute expression levels (log2 copy number) for a set of housekeeping genes (top) and tumor-associated genes (bottom). Source data for this figure is available in the Source Data file.
To rigorously validate this new absolute reference dataset, we performed extensive orthogonal and external benchmarking. First, absolute copy numbers measured by our calibrated RNA-seq showed strong concordance with quantities measured by a reference method, RT-dPCR, for 46 target genes in both D5 (Fig. 6b, Pearson’s r = 0.89) and D6 (Fig. 6c, Pearson’s r = 0.89). The expression ratios (D5/D6) derived from these absolute values were also highly concordant with those from RT-dPCR (Fig. 6d, Pearson’s r = 0.88).
With this internally validated absolute dataset of endogenous genes in hand, we could test the central premise of our framework: whether the diagnostic paradigm developed using spike-ins holds true for the endogenous transcriptome. We applied the same dual-metric analysis to our dPCR-validated dataset, calculating the Absolute and Ratio Measurement Error (ME) distributions for thousands of endogenous genes across the eight batches (Fig. 6e–g). The results provided a powerful, transcriptome-wide validation of the two distinct and non-intuitive failure modes first identified using our calibrators. The L2_DNB_P batch, for instance, unequivocally confirmed the “subtle paradox.” Mirroring its behavior with the spike-ins, this workflow exhibited highly dispersed Absolute MEs for endogenous genes, indicating a chaotic internal measurement process. Critically, this severe flaw was again masked by a deceptively precise Ratio ME distribution (Fig. 6g). This provides definitive proof that the danger of high relative precision masking poor absolute accuracy is a real-world risk for endogenous gene quantification. Likewise, the L2_DNB_R batch confirmed the second, equally critical paradox. Just as with the calibrators, this workflow displayed a remarkably consistent internal process for endogenous genes, evidenced by its tightly compacted Absolute ME distribution (Fig. 6e, f). Yet, it again failed to preserve relative quantification, revealed by its dispersed Ratio ME distribution (Fig. 6g).
By demonstrating that the exact same paradoxical behaviors discovered in our controlled system are replicated at the transcriptome scale, we confirm that these vulnerabilities are not theoretical but are inherent risks in the measurement of endogenous genes. This establishes that a dual-metric system is not merely an improvement but a fundamental necessity for robust quality control in any RNA-seq experiment. As a final validation, we benchmarked our data against the original Quartet “ground truth” ratios13. The calibration significantly reduced the ME and tightened the distribution of measured ratios compared to the uncorrected data (Fig. 6g, h). This demonstrates that our absolute correction also improves the accuracy of relative quantification against an established external standard.
Having validated the framework, we applied it to reveal the true quantitative landscape of endogenous genes. We observed that conventional FPKM values present a compressed and distorted view of housekeeping gene expression. In contrast, our calibration transforms the data to absolute molecular counts, restoring the full dynamic range and revealing a well-defined quantitative structure (Fig. 6i). This fundamental restoration of the data landscape enables direct and meaningful comparisons between the absolute expression levels of different genes (Fig. 6j).
This new capability for absolute quantification provides novel biological insights by enabling the quantitative dissection of distinct biological relationships. First, we examined the link between a master regulator and its direct target. In the constructed absolute quantification dataset D5, our data reveal that the absolute transcript level of the MET proto-oncogene is about 150-fold lower than that of its key downstream adaptor, the GRB2, and that is highly consistent with the results verified by RT-dPCR (log₂ copy number ratio of GRB2/MET ≈ 7.3 vs. ≈ 7.7, Supplementary Fig 13). Our study enabled the expression profile quantification within any given sample, and also validated the absolute quantification of this pair of genes in dataset D6. Likewise, the copy number ratio of these two genes across both samples was consistent with the RT-dPCR results (Fig. 6b–d, Supplementary Fig 13). This observation moves beyond simple correlation to provide, using a generalized method, an estimate of the transcript abundance ratio within a critical therapeutic pathway. In a second example, our framework provides a quantitative link between genomic structure and transcriptional output for tumor samples. We observed that the absolute transcript levels of ERBB2 and GRB7, two genes known to be co-amplified on chromosome 17q12 in tumor tissue, such as breast cancer20. We verified that the basal expression level of GRB7 is low in normal sample (11.78 log2 copies per μg of total RNA for D5), but is elevated by over 400-fold (20.42 log2 copies per μg of total RNA) in the tumor sample LCA (Supplementary Fig. 13a). Furthermore, the expression levels of GRB7 and ERBB2 are approximately equal, as an excellent concordance (20.1 vs. 20.4 log2 copies per μg of total RNA) between TranScale-calibrated RNA-seq and dPCR absolute measurements (Supplementary Fig. 13a), this co-overexpression provides biological insight into the enhancement of oncogenic signaling, and quantitative evidence that a genomic co-amplification event is translated into a near 1:1 ratio of transcript output—a precise mapping from genome to transcriptome that was previously difficult to ascertain with confidence (Fig. 6i). Collectively, these results, validated by both an orthogonal method and an external reference standard, demonstrate that our application framework successfully mitigates sequence-dependent biases for endogenous transcripts. This enables robust, accurate, and direct comparisons of absolute expression levels between different genes, resolving a fundamental limitation in transcriptomics.
Demonstration of robust diagnostic classification
Finally, to illustrate the potential of TranScale in identifying biological insights and supporting clinical decision-making, we conducted a diagnostic simulation targeting the ERBB2 (HER2) oncogene across 12 independent sequencing libraries. We established a “ground truth” diagnostic cutoff (18.5 of log2 copies/µg total RNA) based on reference dPCR data, which clearly distinguished the tumor model (LCA) from normal controls (Quartet D5/D6). We observed that technical batch effects in uncalibrated relative quantification (FPKM) frequently masked the overexpression phenotype of the tumor samples, leading to inconsistent classifications. In contrast, TranScale calibration effectively neutralized these variations, recovering the true absolute abundance of ERBB2 and ensuring 100% concordance with the dPCR-defined diagnostic classification across all platforms and laboratories (Supplementary Fig. 14). These results highlight the capability of SI-anchored absolute quantification to reveal biological signals that may be obscured by technical noise in multi-center studies.
Discussion
RNA-seq’s quantitative power is constrained by equating read counts with molecular quantity1,21, a practice causing batch effects that hinder data integration22 and clinical benchmarks23,24. The field has relied on retrospective computational normalization25,26, post-hoc methods that treat statistical symptoms, not the underlying measurement problem, and risk conflating artifacts with biological signals27. We address this by establishing a metrological framework that redefines RNA-seq as an absolute quantitative method. By introducing SI-traceable, biomimetic TranScale calibrators, we create a chain of traceability from sequencing reads to the SI unit (the mole). This physical calibration reduces inter-laboratory CV from >85% to <25% and achieves a biological signal-to-noise ratio superior to ComBat (Fig. 5), demonstrating that prospective measurement correction is more effective than retrospective statistical modeling.
Our method’s efficacy stems from integrating two key principles: biomimicry in design and metrological rigor in value assignment. This integrated approach is crucial for advancing beyond the limitations of previous standards. For instance, while non-biomimetic controls like ERCC have been valuable for assessing platform performance4, prior studies have suggested their use in normalization can introduce systemic biases due to their distinct physicochemical properties28,29. Our analysis of data (Supplementary Fig. 15) from a recent, large-scale inter-laboratory study5 provides new quantitative evidence that supports this concern in the context of absolute calibration. In this “real-world” dataset, we observed that attempting to use ERCCs for absolute calibration led to a decrease in the signal-to-noise ratio (SNR) (Supplementary Fig. 15b, c), an increase in inter-batch variability (CV) (Supplementary Fig. 15d) and the derived absolute copy numbers showed poor concordance (r = 0.32) with orthogonal dPCR measurements (Supplementary Fig. 15e). Similarly, at the relative level, the copy number ratios obtained using the potential ERCC correction method also deviated significantly from those of the Quarte reference dataset M8/D613 (Supplementary Table 10). We hypothesize that this unexpected outcome is a direct consequence of ERCCs’ non-biomimetic nature, as their behavior may not be representative of endogenous transcripts during the complex sequencing workflow. This finding underscores that a standard’s ability to mimic endogenous molecules is a critical consideration for effective absolute calibration.
This principle of biomimicry is shared by other standards like Sequins12 which were designed with exon-intron structures. However, effective absolute quantification requires a second, equally important attribute: SI-traceable certified values. Sequins are provided with defined relative ratios, making them excellent tools for validating a workflow’s ability to measure fold-changes, but they are not designed to provide the absolute concentration anchor needed to build a true calibration curve from sequencing signal to molecular count30,31. TranScale is thus developed to address both requirements. It combines a biomimetic design for representative behavior—a feature empirically shown to be critical through our ERCC comparison—with the metrological foundation for true absolute scaling. While no synthetic standard can perfectly replicate all endogenous transcripts, our results indicate this dual-principled approach provides a practical and robust solution for absolute transcriptomics. Unlike synthetic controls that may lack traceability or fail to mimic endogenous mRNA behavior, TranScale combines biomimicry with SI-traceability. This unique feature allows for the correction of sequence-dependent biases that compromise traditional relative quantification (Supplementary Table 11). Furthermore, our ERBB2 analysis highlights a critical advantage of this metrological approach: the ability to define universal diagnostic cutoffs. Currently, clinical transcriptomics is often limited by the need for batch-specific controls or relative comparisons. Our data suggest that TranScale-calibrated absolute quantification allows for the establishment of fixed, transferable decision thresholds—similar to those used in clinical chemistry (e.g., blood glucose levels)—thereby bridging the gap between research-grade RNA-seq and robust clinical diagnostics.
We also distinguish the metrological utility of TranScale from that of Unique Molecular Identifiers (UMIs)32,33. While UMIs are effective at mitigating PCR duplication artifacts by collapsing identical barcodes, they track cDNA molecules and do not typically account for losses occurring upstream of UMI assignment. Such losses, arising from variations in reverse transcription efficiency, fragmentation, or adapter ligation, may contribute to discrepancies between the original RNA abundance and the amplifiable cDNA pool. In contrast, TranScale calibrators are introduced at the beginning of the workflow, thereby capturing these cumulative pre-PCR inefficiencies. Consequently, we propose that the two approaches are complementary: UMIs provide digital counting for downstream amplification, while TranScale provides a reference for correcting biases associated with the initial sample processing.
A key innovation is the framework’s diagnostic utility. By deconstructing measurement error into two orthogonal components, structural bias (quantified by Absolute ME) and ratio distortion (quantified by Ratio ME)—we uncovered a systemic challenge: that a workflow’s internal consistency and its ability to preserve quantitative ratios are fundamentally decoupled properties, meaning that excellence in one provides no guarantee of performance in the other. This was definitively demonstrated by two contrasting workflows with our TranScale: one (L2_DNB_P) showed that high relative precision can mask a chaotic internal measurement process, while another (L2_DNB_R) proved that apparent internal consistency can conceal a systemic failure to preserve quantitative ratios, a hallmark of non-linear measurement response (Fig. 3). The critical implication of this dual paradox—that reliance on either metric alone is insufficient for quality control—was then confirmed at a transcriptome-wide scale using our upgraded Quartet reference material, which revealed the same decoupled vulnerabilities for thousands of endogenous genes (Fig. 6). This challenges the foundational assumption that preserving expression ratios is sufficient for all quantitative analyses34 and suggests historical datasets may harbor undetected structural biases, compromising universal diagnostic thresholds and contributing to irreproducibility23,35.
To address the potential concern about the complexity of our method, it is essential to distinguish between the one-time metrological characterization of the TranScale material and its routine laboratory application. The complex IDMS and statistical workflows are confined to the foundational production of the standard by a specialized institution. For the end-user, the process is straightforward, involving the spike-in of a pre-calibrated mixture followed by standard sequencing, as illustrated in our operational guide (Supplementary Fig. 10b, upper panel). The data analysis then simplifies to applying a linear calibration. This user-friendly application is further distinguished from the more rigorous, upfront diagnostic validation of a laboratory’s entire workflow. For that purpose, our dual-metric framework (using two distinct mixes) is provided to certify a measurement system’s reliability (Supplementary Fig. 10b, lower panel). Once a workflow is validated, the routine use of a single TranScale mix is sufficient for robust batch correction and absolute quantification. This tiered approach provides a practical and scalable path towards absolute and comparable transcriptomic data.
Correcting these biases enables mapping an absolute molecular landscape. This method facilitates direct, quantitative inter-gene comparison on an absolute scale. This allows analyzing quantitative relationships with high clarity, providing promising biological insights, which signals a transition toward constructing quantitative, systems-level models of cellular networks.
This metrological method provides a prospective solution for harmonizing large-scale projects like TCGA36 and the Human Cell Atlas37. Incorporating an SI-traceable calibrator from the outset anchors datasets to a common, absolute scale, providing a more robust foundation for meta-analysis than post-hoc methods38. To demonstrate this, we upgraded the Quartet RNA RMs13 by assigning SI-traceable values to over 13,000 transcripts, creating a dual-function reference for absolute and ratio-metric analysis. The TranScale RNA RM is publicly available39.
Transitioning to an absolute measurement system illuminates the nature of measurement bias. Our calibration’s premise is that most biases in an RNA-seq workflow can be modeled as a linear transformation (y ≈ ax + b + ε), a premise validated by the successful harmonization of thousands of endogenous genes across variable batches (Figs. 5, 6). The model is effective because the slope a captures multiplicative biases (e.g., efficiency variations), while the intercept b accounts for additive background noise. Deviations from this model are not limitations but a diagnostic feature, pinpointing “recalcitrant” sequences (Fig. 4c, d) designed with features known to confound quantification, such as repetitive elements or high homology from fusion/WT pairs and SNVs. For these sequences, ambiguous read alignments are resolved probabilistically, making a transcript’s quantification dependent on its homologs. This upstream process severs the link between abundance and read count, creating a non-linear signal that no subsequent calibration can correct. Identifying where these foundational assumptions of read-count quantification fail provides an empirical basis for improving future alignment and quantification algorithms40,41.
The principles of this work should be extended to low-input, single-cell42, and spatial transcriptomics43, where absolute quantification is critical for defining cell states44,45. In summary, by anchoring RNA-seq to the SI system, we provide the tools to transition the field from relative comparisons to absolute measurements, akin to the standardized assays of clinical chemistry46. This paradigm shift enables universal diagnostic cutoffs and robust data integration, accelerating the translation of transcriptomic discoveries into precision medicine47.
Methods
Design and generation of TranScale RNA transcripts
The TranScale panel was meticulously developed through a three-stage process to create a high-fidelity biomimetic standard for RNA sequencing.
Biomimetic design and sequence selection
Our design process targeted a panel of approximately 100 transcripts that collectively mirror the human transcriptome. The selection criteria were multifaceted:
-
(i)
Biomimetic sequence strategy: To preserve structural characteristics while eliminating sequence homology, each transcript was designed by inverting a selected human coding sequence (CDS). This strategy maintains native sequence complexity, such as a GC content similar to the hg38 genome, while ensuring that TranScale reads do not misalign during bioinformatic analysis.
-
(ii)
Functional diversity: The selected sequences represented a range of biological features, including wild-type genes, clinically relevant fusion genes, and alternative splicing isoforms.
-
(iii)
Physicochemical range: The initial design aimed for a broad length distribution from ~500 nt to over 4000 nt to cover the majority of the human mRNA length spectrum.
Production, optimization, and quality control
This stage focused on translating the computational design into physical RNA molecules of the highest metrological quality.
-
(i)
Template generation: The full DNA sequence for each calibrator—comprising a 5’ BamHI site, a T7 promoter, the inverted CDS, a 3’ 28 bp poly(A) tail, a T7 terminator, and a final EcoRI site—was synthesized and cloned into a pMV vector (BGI Genomics). Following sequence verification (Supplementary Table 1), recombinant vectors were transformed into E. coli DH5α. Template DNA for in vitro transcription (IVT) was obtained by BamHI/EcoRI digestion and purified with the Universal DNA Purification Kit (Tiangen).
-
(ii)
In vitro transcription and pragmatic optimization: IVT was performed using the T7 High Yield RNA Synthesis Kit (Yeasen). A 20 µL reaction containing 1 µg of template DNA, 10 mM of each NTP, 1× reaction buffer, and 2 µL T7 RNA Polymerase Mix was incubated at 37 °C for 4 hours, followed by DNase I treatment. At this critical stage, we observed that some of the longer target transcripts (>3800 nt) yielded impure products due to premature termination. Prioritizing analytical validity, we systematically shortened these sequences to a maximal length that permitted consistent, high-purity synthesis.
-
(iii)
Purification and verification: The resulting full-length RNA was purified using the MEGAclear Kit (Thermo Fisher), and its concentration was estimated with a Nanodrop. The integrity and purity of each of the 100 final transcripts were rigorously verified as a single, sharp peak on an Agilent 2100 Bioanalyzer (RNA 6000 Nano kit).
Final panel composition and sufficiency
The final TranScale panel consists of 100 well-defined, high-purity RNA transcripts. Despite the necessary length adjustments for quality control, the panel spans a significant range of lengths (500–3800 nt). Importantly, the utility of the panel is not contingent on the detection of all 100 calibrators. Our operational analyses show that robust calibration models (R² > 0.95) can be reliably constructed using as few as 20-30 well-distributed calibrators, confirming the sufficiency of the final panel for its intended application (Supplementary Fig. 10).
Isotope Dilution Mass Spectrometry (IDMS) for TranScale
The absolute concentration (copy number) of each of the 100 purified RNA transcripts of TranScale RM was certified using Isotope Dilution Mass Spectrometry (IDMS), a primary reference measurement procedure recognized by the Joint Committee for Traceability in Laboratory Medicine (JCTLM). The certification workflow was as follows:
NMP standards preparation
As standards, the certified RMs of adenosine 5′-monophosphate (AMP), guanosine 5′-monophosphate disodium salt (GMP), cytidine 5′-monophosphate (CMP), and uridine 5′-monophosphate disodium salt (UMP) were obtained from the National Institute of Metrology, China (NIM) (NMPs). The corresponding isotope-labeled (13 C, 15 N) nucleotide monophosphates (LNMPs) (Silantes, Germany) were used as internal controls.
Quantification of NMPs standards
For quantification of transcripts, NMPs in the digested samples were separated completely using an SB-AQ C18 column (Agilent, USA), 0.1% formic acid (v/v) was used as mobile phase in a flow rate of 0.2 mL/min maintained at 30 °C. Signals of the well-separated NMPs were detected by SCIEX QTRAP® 6500 + LC-MS/MS in positive ion and multiple reaction monitoring (MRM) mode. Other instrumental and mass spectrometer data acquisition parameters, including Q1, Q3, DP, EP, CE and CXP, were optimized for the NMPs and LNMPs detection (Supplementary Table 3). A known amount of a corresponding stable isotope-labeled internal standard (LNMPs) for each ribonucleoside (NMPs). Mixtures with mass ratios of NMPs to LNMPs of 0.4, 0.8, 1.2, 1.6, and 2.0 were performed, respectively. NMP concentrations (μg/g) in transcripts were determined according to the standard curves of mass ratio and IDMS peak area ratio of NMPs to LNMPs (Supplementary Fig. 2). The raw mass spectrometry data were processed for absolute quantification using Analyst software (version 1.6, AB Sciex).
Hydrolysis of RNA transcripts
An aliquot of the purified RNA transcript was subjected to complete enzymatic hydrolysis to break it down into its constituent ribonucleosides. According to the protocol of our previous studies, briefly, each of the generated transcripts was diluted to a concentration of 1 ng/μL, and then 50 μL RNA sample was mixed with Phosphodiesterase I from Crotalus adamanteus venom (SVP, Sigma-Aldrich, USA) at the final concentration of 0.002 U/μL, 5 μL of LNMPs mixture was added. All reactions were formulated by the gravimetric method. After 25 min of incubation at 25 °C, the reaction was terminated by incubation at 80 °C for 15 min48.
Quantification of target transcripts
The molar concentration of each of the four NMPs was determined with high precision and accuracy by measuring the abundance ratio of the natural analyte to its corresponding LNMPs. According to the quantification method of NMPs, each RNA sample was measured twice independently, and each digestion was analyzed in triplicate. All the above measurements were independently conducted by two operators. The final mass fraction of each NMP in the digested RNA sample was expressed as Eq. (1):
where WRNA is the mass fraction of the RNA sample in micrograms per gram, Wx is the mass fraction of the selected NMP in the RNA sample, MRNA is the molecular mass of the RNA molecule, MNMP is the molecular mass of the selected NMP, and N is the number of the selected NMP in the RNA sample. The copy number of RNA (n, in copies per microgram) was expressed as Eq. (2) according to the Avogadro constant and RNA molecular weight48. Where NA is Avogadro’s constant. A density of 1.00 g/mL was taken into account in the calculation. The copy numbers (copies/μL) of transcripts were calculated.
Consistency check of IDMS
Concentration of a specific RNA transcript was calculated independently from the concentration of each of the four nucleosides, based on the known sequence of that transcript (e.g., [RNA Transcript] = [Adenosine] / number of ‘A’s in sequence). The final certified value for each transcript was assigned as the average of the four independent calculations, provided they showed high consistency (e.g., relative standard deviation <1%). This cross-validation ensures the certified value is robust against potential biases in hydrolysis or measurement for any single nucleoside.
Preparation of TranScale RNA RM
Based on the certified copy numbers of individual transcript stocks determined by IDMS, the two master mixes, named Mix1 and Mix2, each containing all the components of 100 transcripts, were prepared via precision gravimetry. The two mixtures were presented at defined copy number ratios, described by several sub-mixtures, and each of them contains at least 10 spike-ins with the balanced transcript size distribution and GC content. The two mixtures were prepared by diluting all 100 in-vitro transcribed RNAs with RNA storage solution gravimetrically (Thermo Fisher, USA). During the mixing process, the mass of each component was recorded using a balance for accurate proportioning. Mix1 as well as Mix2 were formulated across a concentration range of 5 orders of magnitude, spanning approximately 105 and 106-fold concentration range of transcripts, respectively.
Validation of TranScale RNA RM
RT-dPCR assays
To further verify the concentrations of transcripts of each gradient of TranScale and the ratio between Mix1 and Mix2, the One-step reverse transcription digital PCR (RT-dPCR) assays for probes were established on a Sniper dPCR platform (DQ24, Sniper, China). The 22 μL reaction mixture included 11 μL of 2×dPCR probe master mix, 5 μL of transcripts, primers, and probe with the optimized concentrations, and RNase-free water. The RNA template was gravimetrically diluted before it was added to the reaction. Specifically, RT-dPCR assays for two transcripts were established in each concentration gradient of the consortium (20 assays in total), all primers and probes were listed in Supplementary Data 4.
The RT-dPCR thermocycling conditions were as follows: 45°C for 15 min of reverse transcription, 95 °C for 5 min of enzyme activation, 40 cycles of 95 °C for 15 s of denaturation and 30 s of annealing/extension at the optimized temperature, following 60 °C for 1 min of fluorescence signal acquisition. For each optimized RT-dPCR assay, annealing temperatures of 54, 56, 58, and 60 °C were tested, and following an orthogonal optimization with final probe concentrations of 125, 250, and 500 nM and final primer concentrations of 500 and 900 nM was performed. Furthermore, the specificity of the established RT-dPCR method was validated with a mixture of 99 transcripts that did not contain the target template.
Specificity of RT-dPCR assays
When performing gene-specific PCR amplification using a multiplexed system containing 100 reference genes as template, rigorous confirmation of PCR specificity must be achieved. To ensure specificity of RT-dPCR assays, we used a mixture containing 99 transcripts except the target spike-in as the PCR template; the observed negative droplets that was same as the no template control will explain no cross-reaction with the RT-dPCR assays.
Dynamic range of RT-dPCR assays
The dynamic ranges of RT-dPCR assays were investigated by a serial dilution. Each of the 20 RNA transcripts was diluted in gradients according to the gravimetric method and subsequently subjected to RT-dPCR amplification reactions, with 3-6 replicates per gradient. A good linearized correlation within 5 orders of magnitude of RNA concentrations between gravimetric dilution and RT-dPCR determination was expected.
Reverse transcription efficiency
The in vitro transcribed RNA mass fraction obtained by IDMS was converted into copy number concentration, which was used as the absolute copy number concentration of the RNA molecule. The copy number concentration of the cDNA molecule was reversed as determined by the established RT-dPCR. The efficiency of reverse transcription of the RNA template into cDNA was determined and expressed as the ratio of the RNA copy number concentration of RT-dPCR and IDMS.
Assessment of homogeneity
With reference to JJF 1343-202249 Characterization, Homogeneity and Stability Assessment of RMs (equivalent to ISO 33405), homogeneity testing was performed through random sampling of sub-packaged samples by RT-dPCR. Eleven vials of TranScale RMs Mix1 and Mix2 were randomly selected, and the established RT-dPCR assays were performed. Each vial was measured with three replicates. F test was used to assess the homogeneity. Mbetween and Mwithin were calculated according to Eqs. (3) and (4). If F < Fɑ, it indicates that the RM is well homogenized. Where a is the total number of RM units. For the ith unit, n replicate measurements are performed, xij represents the jth observed value of unit i. Mbetween and Mwithin are between-unit and within-unit mean squares.
Assessment of stability
Stability of TranScale RM was evaluated by 20 established RT-dPCR assays. For short-term stability, the RMs were stored at 4 °C and 20 °C for 0, 3,7 and 14 days. Students’ t-test was performed on the assay results to assess the short-term stability. For long-term stability, the RMs were placed at −80 °C and measured at 1, 2, 3, 6, and 12 months. The long-term stability was evaluated according to Eq. (5) and (6). A linear regression model is employed, where β₁ is the slope coefficient, xᵢ is the observation time, yᵢ corresponds to the observed value at time xᵢ, x̄ signifies the mean value of all observation time points, s is the standard deviation of the residuals at each data point along the fitted regression line.
Certified values and measurement uncertainty
IDMS was used for value assignment of all 100 mother solutions of RNA transcripts; the certified value for each transcript is established by taking the arithmetic mean of 12 independent measurements. These values are derived through gravimetric determination, where absolute quantities in the RM are calculated based on precisely weighed masses of high-purity RNA transcripts.
The uncertainty of our TranScale RM (Uc-Mix1 and Uc-Mix1) can be classified into characterization uncertainties (uchar, combining uncertainties from both IDMS and gravimetric dilution), sample inhomogeneities (uhom) and instabilities (us), and then combined to form the combined uncertainties (uc) by using Eq. (7). The expanded uncertainties (UMix1 and UMix1) are calculated from the uC multiplied by the coverage factor (k = 2) at a confidence level of 95%. Mix1/Mix2 represented the ratio of copy number concentration of reference gene in Mix1 and Mix2, and its uncertainty is the arithmetic square root of the sum of the squared uncertainties of Mix1 and Mix2, as Eq. (8).
Sequencing orthogonality validation for RNA-seq
Considering the potential for spike-in reads to cross-align to the reference human genome, we initially sequenced a neat mixture of 100 spike-ins of the RM without a natural RNA sample, at equimolar concentrations to ensure sufficient coverage for validation of spike-in design with experimental reads. For the neat RM sample, four batches of RNA-seq were generated in two laboratories (L1 and L2) using PolyA and RiboZero library construction protocols, and two platforms Illumina NovaSeq 6000 and DNBSEQ-T7 with 150 bp paired-end reads. 1 µg of RNA was used in each library preparation. The reference genome and gene model annotation files were prepared, the FASTA file of genome index was combined with hg38 (https://ftp.ensembl.org/pub/release-94/fasta/homo_sapiens/dna/Homo_sapiens.GRCh38.dna.toplevel.fa.gz) and all the 100 artificial DNA template sequences of RNA spike-ins, as well as the combined GTF file comprising both of the annotated human genes (http://ftp.ensembl.org/pub/release94/gtf/homo_sapiens/Homo_sapiens.GRCh38.93.gtf.gz) and the artificial genes. The clean reads were aligned to a combined genome index and hg38, respectively. Hisat2 v2.0.5 was used as the mapping tool, and the respective mapping rate was calculated.
TranScale as spike-ins for RNA-seq
Strand-specific library
RNA integrity was assessed using the Bioanalyzer 2100 system, RNA library for strand-specific RNA-seq was prepared. Chinese Quartet RMs D5 and D6 (from NCRM39) were used as target samples in this study. We precisely spiked 1 ng of the RM into 1 μg total RNA sample; this was executed gravimetrically. We used TranScale Mix1 mixed to D5 and Mix2 mixed to D6. mRNA was purified from total RNA using polyT capture or rRNA deletion and then fragmented into 300 ~ 350 bp fragments. The first strand cDNA was reverse-transcribed using fragmented RNA and dNTPs (dATP, dTTP, dCTP and dGTP), and the second strand cDNA was synthesized using DNA polymerase I and dNTPs (dATP, dUTP, dCTP and dGTP). Remaining overhangs of double-strand cDNA were converted into blunt ends via exonuclease/polymerase activities. After adenylation of the 3’ ends of DNA fragments, sequencing adaptors were ligated to the cDNA, and the library fragments were purified. The template without U was enriched by PCR, and the PCR product was purified to obtain the final library with a concentration of no less than 1.5 nM. In this cross-batch transcriptomic sequencing study, PCR cycle numbers were first optimized to ensure linear-range amplification. Ultimately, 13 cycles were selected for the subsequent eight experimental batches (Supplementary Fig. 16). After library construction, the concentration of the library was measured by the qubit fluorometer. The accurate concentration of the cDNA library was again examined using qPCR.
Data quality control
Raw reads of FASTQ format were first processed by the fastp software. In this step, clean data were obtained by removing reads with adapters, reads with more than 3 N, and reads with more than 20% nucleotides with Qphred≤5. At the same time, Q20, Q30, and GC content of the clean data were calculated. Then, map the clean reads to the SILVA database to remove the rRNA. All the downstream analyses were based on the clean data without rRNA.
Reads mapping to the reference genome
The same reference genome and gene model annotation files were used with the neat TranScale RM. An index of the reference genome that contains TranScale reference sequences and the human reference genome GRCh38 was built using Hisat2 v2.0.5, and paired-end clean reads were aligned to the reference genome using Hisat2 v2.0.5. For the mixed sample of spike-ins and target D5 or D6 samples, the combined genome index was used for read mapping. Counts obtained for different dosages of total input spike-ins through simulated data comparison.
Quantification of gene expression
FeatureCounts v1.5.0-p3 was used to count the number of reads mapped to each gene. And then FPKM (number of Fragments Per Kilobase of transcript sequence per Millions base pairs sequenced) of each gene was calculated based on the length of the gene and reads count mapped to this gene.
Performance metrics for absolute quantification of RNA-seq
ME. ME introduced in this study as a TranScale RM-based metric, serves as the most intuitive indicator for assessing RNA-seq absolute quantification performance. It directly quantifies deviations between observed values and certified reference values. ME of all three replicates of 100 transcripts among TranScale in each sample were respectively calculated according to Eq. (9), which represents the percentage difference between FPKM with mean normalization (Normalized FPKM) and its reference copy numbers (Certified). All data is in a log2 scale. Additionally, a Ratio ME was calculated from the ratio of the measurements for each transcript pair in Mix1 and Mix2 against the ratio of their certified values. This metric is designed to be insensitive to global systematic shifts, thus isolating sequence-dependent biases.
RMSE. RMSE50 was calculated using observed values of spike-ins and the corresponding certified values, representing the distances of absolute expression profiles. All three replicates of 100 transcripts among TranScale in each sample were calculated after mean centering analysis of FPKM and certified copy numbers. All data was transformed using log2 scaling. The calculation first determined squared residuals for all 100 spike-ins, averaged these squared differences, then took the square root to express error in original concentration units as Eq. (10), where n is the number of measured genes, yi is the i-th observed value, and xi is the i-th reference value.
RC. RC51 was calculated based on the Spearman correlation coefficient between the observed absolute expressions of observed spike-in values and the corresponding certified values of Mix1 and Mix2 of the RMs. It is referred to as the ‘correlation with reference values’ metric, representing the numerical consistency of the absolute expression profiles. To improve reliability, all three replicates of each sample group were calculated after mean centering analysis of FPKM and certified copy numbers of TranScale. All data was transformed using log2 scaling.
SNR13. The absolute expressions-based SNR is defined as the average distance between the two sample groups of D5 and D6 (or LCA and LCN) from the average distance among technical replicates of the same groups. Based on PCA, distances of two samples in the space defined by the first two PCs were used to represent distances between the two samples. SNR was calculated as Eq. (11) described previously. Where m is the number of donors, and n is the number of replicates in each donor. Wp represents the p-th principal component of variances. PCp,i,x, PCp,j,x and PCp,j,y represent the p-th component values of replicate i and replicate j in donor x or donor y, respectively.
Justification for the QC-gated calibration parameters
The parameters for our QC-gated calibration framework were selected to strike a critical balance between stringency (ensuring only high-fidelity data informs the model) and robustness (retaining a sufficient number of calibrators to build a statistically sound model).
Measurement Error (ME) threshold (±5%)
The choice of a ± 5% ME threshold is central to the framework’s function. A more lenient threshold (e.g., ±10% or ±20%) would risk including calibrators that exhibit significant non-linear behavior or are subject to strong sequence-specific biases. This would “pollute” the linear regression, leading to an inaccurate slope and intercept that do not reflect the true systemic bias of the workflow. Conversely, an overly stringent threshold (e.g., ±1% or ±2%) would be overly punitive, potentially discarding a large number of valid calibrators due to inherent stochastic noise in the sequencing process. This could lead to an insufficient number of data points for a robust regression, or even the false rejection of an otherwise acceptable library. The ±5% ME threshold was therefore established as an optimal setpoint that effectively isolates the dominant linear response trend of the workflow while tolerating a reasonable degree of measurement noise.
FPKM threshold (>0.1)
This threshold was empirically determined based on the performance of the lowest-abundance spike-in calibrators. We observed that transcripts with FPKM values below this level exhibited poor reproducibility across technical replicates and, critically, their corrected values after calibration showed significant deviation from their certified SI-traceable values. Therefore, the FPKM > 0.1 threshold was established to exclude these unreliable data points, ensuring that the calibration model is constructed only from robust and accurately quantified calibrators.
Dynamic Range threshold (>2¹⁰-fold)
This criterion ensures that the calibration model is not built on a narrow concentration range. A wide dynamic range is essential for the model to have sufficient leverage to accurately determine the slope, ensuring its validity when applied to endogenous transcripts across the full expression spectrum.
Coefficient of determination (R²) threshold (>0.95)
An R² > 0.95 is a widely accepted standard for demonstrating a strong linear relationship. Setting this as the final gate serves as the ultimate confirmation that the foundational assumption of linearity holds true for the specific library being processed, providing high confidence in the subsequent calibration.
Minimum number of calibrators for robust model fitting
A critical, implicit QC gate in our framework is the number of calibrators that successfully pass the ME, FPKM, and dynamic range criteria. While a linear model can be generated from a few points, its reliability and predictive power for the entire transcriptome depend on a statistically robust foundation. Based on our multi-batch analysis, even lower-quality libraries that were ultimately salvageable for calibration consistently yielded a substantial number of calibrators (typically >30, as shown in Fig. 4a). Therefore, we establish a practical guideline: a library should yield a minimum of 20–30 high-confidence calibrators to proceed with high-confidence absolute quantification.
This recommendation is based on the following rationale: (1) Statistical Stability: A sufficient number of calibrators ensures that the regression parameters (slope and intercept) are stable and accurately reflect the library’s systemic bias, rather than being skewed by random noise or a few outliers. (2) Diagnostic Power: The number of passing calibrators serves as a powerful meta-indicator of overall library quality. A failure to meet this minimum threshold strongly suggests underlying issues (e.g., significant non-linear biases, poor library preparation) that make the data unsuitable for absolute quantification, even if a model with a high R² can be formally generated. Libraries falling below this guideline should be flagged, and their use should be limited to relative analyses.
Construction and metrological characterization of endogenous gene reference datasets
To establish robust reference datasets for endogenous gene expression (designated D5 and D6), we processed data from eight independent measurement batches through a stringent filtering and characterization workflow. Beginning with the complete Ensembl annotation (n = 58,735), genes were retained only if detected in at least six of eight batches, present in all three technical replicates per batch, and exhibited an FPKM > 0.1 across all replicates. For this filtered set, FPKM values were converted to absolute copy numbers using the library-specific linear models derived from the co-processed TranScale calibrators. The final certified reference value for each gene was assigned as the arithmetic mean of its corrected absolute copy numbers from all qualifying batches. For the purpose of this study, the measurement uncertainty was evaluated based on the dominant experimental component. This characterization uncertainty (u_char), a Type A evaluation, was calculated as the relative standard deviation (RSD) of the final calibrated copy numbers for each gene across all contributing measurement batches. This approach is based on the principle that for a complex, multi-stage workflow like RNA-seq, the experimentally observed reproducibility (u_char) is the largest and most practically relevant contributor to the combined uncertainty, significantly outweighing the Type B uncertainties propagated from the calibration standards. Therefore, u_char provides a direct and robust measure of the quality and consistency of the final reference values. Accordingly, the final reference gene sets were filtered based on this primary uncertainty metric, retaining only those with a relative u_char of ≤ 20%. For the D5/D6 ratio dataset, values and their combined uncertainties were calculated by propagating the respective u_char values using the standard formula for division according to Eq. (12).
Validation of reference datasets using RT-dPCR
The absolute gene expressions in the reference datasets were further validated by dPCR; primers of target genes in D5 and D6 were listed in Supplementary Data 12. First, initial cDNA synthesis was performed by incubating 2 μL RNA with 4 μL of 5× PrimeScript IV cDNA Synthesis Mix (Takara #6215 A) containing PrimeScript IV RTase, RNase Inhibitor, Oligo dT Primer, and dNTPs supplemented with 1 μL random hexamers. Nuclease-free water was added to achieve a 20-μL reaction volume. This reaction mixture was incubated at 30 °C for 10 min and then for 15 min at 42 °C and finally for 5 min at 95 °C for termination. Second, dPCR reactions were employed by the Bio-Rad QX200 Droplet Digital PCR System with 20-μL mixtures containing: 10 μL EvaGreen Supermix, 2 μL primer pair, 2 μL cDNA template, and 6 μL RNase-free ddH₂O. Following droplet generation with 70 μL oil, 40-μL droplets were transferred to a 96-well plate. Amplification conditions comprised: 5 min at 95 °C; 40 cycles of 95 °C for 30 s and 60 °C for 1 min; followed by signal stabilization (4 °C for 5 min and 90 °C for 5 min). Signals were acquired using the Droplet Reader, with all reactions performed in duplicate. Absolute quantities were verified by comparing measured absolute copy numbers of selected genes in D5 and D6 against absolute copy numbers in the reference dataset, and also assessing D5/D6 copy number ratios versus established reference ratios for Ratio consistency.
Diagnostic robustness simulation
To simulate a clinical diagnostic scenario, we utilized ERBB2 expression data from 12 sequencing libraries. A diagnostic cutoff was empirically defined as the midpoint separating the dPCR-quantified absolute copy numbers of the tumor model (LCA) and normal controls (Quartet D5 and D6). For the uncalibrated analysis, raw FPKM values were log2-transformed. For the calibrated analysis, TranScale-derived absolute copy numbers were used. We calculated the consistency of diagnostic classification (Tumor vs. Normal) across all batches relative to the dPCR-defined ground truth.
Statistics & Reproducibility
Study design and Sample size
No statistical method was used to predetermine sample size. The sample sizes were 12 technical replicates for initial measurement of each transcript of TranScale, and 3 independent technical replicates for values validation to ensure sufficient precision for measuring absolute copy number concentrations and to rigorously evaluate batch-effect correction as per metrological standards. We sequenced 3 replicates of each of the four RNA samples (D5, D6, LCA, and LCN) using 2 commercially available short-read sequencing protocols: PolyA and RiboZero, generating 12 data batches. Each batch included 6 libraries, resulting in a total of 72 libraries. These sample sizes are sufficient to provide within-batch technical replication, cross-protocol comparisons, and cross-batch/laboratory reproducibility assessment. Details are illustrated explicitly in Fig. 1.
Data exclusion
All data from planned experiments have been included. All attempts at replication were successful.
Randomization
The experiments were not randomized, as the study did not involve group comparisons requiring random allocation. Samples were allocated by a pre-specified, balanced design. Each batch contained 6 libraries; there were 3 technical replicates for 2 paired samples (samples D5 and D6, or samples LCA and LCN).
Blinding
The investigators were not blinded to allocation during experiments and outcome assessment, as the study relied on objective bioinformatic pipelines and predefined calibration standards.
Data analysis
All attempts at replication were successful. Data processing and statistical analyses were performed using R (version 4.5.0) and associated packages, including ggplot2 (version 3.5.2) for visualization. Graphs were generated using GraphPad Prism (version 10.1.2), Origin (version 2021), and Adobe Illustrator (version 2025). Specific statistical tests used for homogeneity and stability assessments are detailed in the relevant results and methods sections above.
Ethics Statement
This research complies with all relevant ethical regulations. The use of RNA materials derived from established clinical cell lines in this study was reviewed and approved by the Ethics Committee of the National Cancer Center/ Cancer Hospital, Chinese Academy of Medical Sciences (Approval no. 24/427-4707).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequence data in this paper have been deposited in the Genome Sequence Archive (GSA) under the accession code GSA-Human: HRA01324452. Source data are provided with this paper.
Code availability
Source code for gene expression analysis and the TranScale calibration pipeline have been deposited on GitHub, available at https://github.com/zhyu0807/TranScale/tree/main53, and have been archived in Zenodo under: https://doi.org/10.5281/zenodo.1844619054.
References
Stark, R., Grzelak, M. & Hadfield, J. RNA sequencing: the teenage years. Nat. Rev. Genet. 20, 631–656 (2019).
Zheng, Y. et al. Multi-omics data integration using ratio-based quantitative profiling with Quartet reference materials. Nat. Biotechnol. 42, 1133–1149 (2024).
Mortazavi, A., Williams, B. A., Mccue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods (2008).
SEQC Consortium A comprehensive assessment of RNA-seq accuracy, reproducibility and information content by the Sequencing Quality Control Consortium. Nat. Biotechnol. 32, 903–914 (2014).
Wang, D. et al. A real-world multi-center RNA-seq benchmarking study using the Quartet and MAQC reference materials. Nat. Commun. 15, 6167 (2024).
Begley, C. G. & Ellis, L. M. Raise standards for preclinical cancer research. Nature 483, 531–533 (2012).
Byron, S. A. et al. Translating RNA sequencing into clinical diagnostics: opportunities and challenges. Nat. Rev. Genet. 17, 257–271 (2016).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Nygaard, V., Rødland, E. A. & Hovig, E. Methods that remove batch effects while retaining group differences may lead to exaggerated confidence in downstream analyses. Biostatistics 17, 29–39 (2016).
External RNA Controls Consortium The External RNA Controls Consortium: a progress report. Nat. Methods 2, 731–734 (2005).
Risso, D., Ngai, J., Speed, T. P. & Dudoit, S. Normalization of RNA-seq data using factor analysis of control genes or samples. Nat. Biotechnol. 32, 896–902 (2014).
Hardwick, S. A. et al. Spliced synthetic genes as internal controls in RNA sequencing experiments. Nat. Methods 13, 792–798 (2016).
Yu, Y. et al. Quartet RNA reference materials improve the quality of transcriptomic data through ratio-based profiling. Nat. Biotechnol. 42, 1118–1132 (2023).
Better research through metrology. Nat. Methods 15, 395 https://doi.org/10.1038/s41592-018-0035-x (2018).
Milavec, M. et al. Metrological framework to support accurate, reliable, and reproducible nucleic acid measurements. Anal. Bioanal. Chem. 414, 791–806 (2022).
Hu, J. F. et al. Quantitative mapping of the cellular small RNA landscape with AQRNA-seq. Nat. Biotechnol. 39, 978–988 (2021).
Fu, L. et al. Simultaneous quantification of methylated cytidine and adenosine in cellular and tissue RNA by nano-flow liquid chromatography–tandem mass spectrometry coupled with the stable isotope-dilution method. Anal. Chem. 87, 7653–7659 (2015).
Fassett, J. D. & Paulsen, P. J. Isotope dilution mass spectrometry for accurate elemental analysis. Anal. Chem. 61, 643A–649A (1989).
De Bièvre, P., Dybkaer, R., Fajgelj, A. & Hibbert, D. B. Metrological traceability of measurement results in chemistry. Pure Appl. Chem. 79, 1755–1761 (2007).
Sircoulomb, F. et al. Genome profiling of ERBB2-amplified breast cancers. BMC Cancer 10, 539–539 (2010).
Shi, L. et al. Microarray scanner calibration curves: characteristics and implications. BMC Bioinforma. 6, S11 (2005).
Luecken, M. D. & Theis, F. J. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol. Syst. Biol. 15, e8746 (2019).
McNutt, M. Reproducibility. Science 343, 229–229 (2014).
Baker, M. Reproducibility crisis. Nature 533, 353–366 (2016).
Zhang, Y., Parmigiani, G. & Johnson, W. E. ComBat-seq: batch effect adjustment for RNA-seq count data. NAR Genom. Bioinform. 2, lqaa078 (2020).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 12 (2020).
Hicks, S. C., Townes, F. W., Teng, M. & Irizarry, R. A. Missing data and technical variability in single-cell RNA-sequencing experiments. Biostatistics 19, 562–578 (2018).
Cole, M. B. et al. Performance assessment and selection of normalization procedures for single-cell RNA-seq. Cell syst. 8, 315–328. e318 (2019).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Svensson, V., Vento-Tormo, R. & Teichmann, S. A. Exponential scaling of single-cell RNA-seq in the past decade. Nat. Protoc. 13, 599–604 (2018).
Lun, A. T., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75 (2016).
Fu, Y., Wu, P.-H., Beane, T., Zamore, P. D. & Weng, Z. Elimination of PCR duplicates in RNA-seq and small RNA-seq using unique molecular identifiers. BMC Genomics 19, 531 (2018).
Kivioja, T. et al. Counting absolute numbers of molecules using unique molecular identifiers. Nat. Methods 9, 72–74 (2012).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Begley, C. G. & Ioannidis, J. P. Reproducibility in science: improving the standard for basic and preclinical research. Circ. Res. 116, 116–126 (2015).
Hoadley, K. A. et al. Cell-of-origin patterns dominate the molecular classification of 10,000 tumors from 33 types of cancer. Cell 173, 291–304. e296 (2018).
Travaglini, K. J. et al. A molecular cell atlas of the human lung from single-cell RNA sequencing. Nature 587, 619–625 (2020).
Caldas, J. & Vinga, S. Global meta-analysis of transcriptomics studies. PLoS One 9, e89318 (2014).
National Center for Research Platform. NCRM [Internet]. [cited 2025-08-19]. Available from: https://www.ncrm.org.cn/.
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Robert, C. & Watson, M. Errors in RNA-Seq quantification affect genes of relevance to human disease. Genome Biol. 16, 177 (2015).
Lähnemann, D. et al. Eleven grand challenges in single-cell data science. Genome Biol. 21, 31 (2020).
Moses, L. & Pachter, L. Museum of spatial transcriptomics. Nat. Methods 19, 534–546 (2022).
Townes, F. W., Hicks, S. C., Aryee, M. J. & Irizarry, R. A. Feature selection and dimension reduction for single-cell RNA-Seq based on a multinomial model. Genome Biol. 20, 295 (2019).
Eraslan, G. et al. Single-nucleus cross-tissue molecular reference maps toward understanding disease gene function. Science 376, eabl4290 (2022).
Vesper, H. W., Myers, G. L. & Miller, W. G. Current practices and challenges in the standardization and harmonization of clinical laboratory tests. Am. J. Clin. Nutr. 104, 907S–912S (2016).
Saadatpour, A., Lai, S., Guo, G. & Yuan, G. C. Single-cell analysis in cancer genomics. Trends Mol. Med. 31, 576–586 (2015).
Niu, C. et al. Accurate quantification of SARS-CoV-2 RNA by isotope dilution mass spectrometry and providing a correction of reverse transcription efficiency in droplet digital PCR. Anal. Bioanal. Chem. 414, 6771–6777 (2022).
State Administration for Market Regulation. General and statistical principles for characterization of reference materials. JJF 1343-2022. (Standardization Administration of China, 2022).
Chai, T. & Draxler, R. R. Root mean square error (RMSE) or mean absolute error (MAE)?–Arguments against avoiding RMSE in the literature. Geosci. Model. Dev. 7, 1247–1250 (2014).
Ali Abd Al-Hameed, K. Spearman’s correlation coefficient in statistical analysis. Int. J. Nonlinear Anal. Appl. 13, 3249–3255 (2022).
Zhang, Y. et al. A metrological foundation for absolute transcriptomics using International System of Units-anchored calibrators, Genome Sequence Archive (GSA), https://ngdc.cncb.ac.cn/gsa-human/browse/HRA013244 (2026).
Zhang, Y. et al. A metrological foundation for absolute transcriptomics using International System of Units-anchored calibrators, https://github.com/zhyu0807/TranScale/tree/main (2026).
Zhang, Y. et al. A metrological foundation for absolute transcriptomics using International System of Units-anchored calibrators. Zenodo, https://doi.org/10.5281/zenodo.18446190 (2026).
Acknowledgements
This work was supported in part by the National Key Research and Development Program of China (grant 2023YFF0613300 to L.D.), the Basic Research Fund Project of the National Institute of Metrology, China (grant AKYZD2202 to L.D.), and the National High-level Metrology Talent Cultivation Program (grant JLTD2601 to L.D.).
Author information
Authors and Affiliations
Contributions
L.D., L.S. and X.F. conceived and supervised the study. Y.Z., B.Y., Y.Y., X.W., C.N., Y.Z.Z., Y.L., J.L., C.Z., J.Y., J.T., Z.L., Z.T., Y.G., Y.T.Z., Y.Q.L., T.X. and R.Z. performed data analysis and/or interpretation. Y.Q.L. and T.X. provided paired cell line samples. Y.L. and J.L. cultured the cell lines and prepared RNA reference materials. Y.Z. and C.Z. characterized the reference materials TranScale. B.Y. and Y.L. performed RNA library preparation. X.W., Y.Y., Y.Z. and J.T. performed RT-dPCR validation. L.D. and Y.Z. managed the TranScale and reference datasets. Y.Z. generated most figures. Y.Z. and L.D. wrote the initial draft. L.D., L.S. and X.F. critically revised the manuscript. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Wei Xu and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, Y., Yang, B., Yu, Y. et al. A metrological foundation for absolute transcriptomics using International System of Units-anchored calibrators. Nat Commun 17, 2747 (2026). https://doi.org/10.1038/s41467-026-70582-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70582-1