Abstract
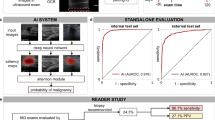
AI models show potential to improve breast cancer screening, however detailed subgroup evaluations to uncover the strengths and weaknesses of models are lacking. This study presents a granular evaluation of a commercial AI model for cancer detection on digital breast tomosynthesis (DBT) on a retrospective cohort of 167,860 screening exams in female patients. Performance in distinguishing screen detected cancers (1,368 exams) from negative exams (166,387 exams) is stratified across demographic, imaging, and pathologic subgroups to identify disparities. The overall AUROC is 0.91 and sensitivity is 0.73 with robust performance across demographics. In-situ cancers (AUROC: 0.85, sensitivity: 0.55), calcifications (AUROC: 0.80, sensitivity: 0.66), and dense breast tissue (AUROC: 0.88, sensitivity: 0.63) are associated with lower performance, while masses (AUROC: 0.93, sensitivity: 0.85) and architectural distortions (AUROC: 0.90, sensitivity: 0.83) are associated with higher performance. These results highlight the need for detailed evaluations and vigilance in adopting new clinical tools.
Similar content being viewed by others
Data availability
The clinical data used in this study, the Emory Breast Imaging Dataset (EMBED), is not available due to medical institutional data policies. 20% of the dataset is available under restricted access through the AWS Open Data program for non-commercial research use. Access can be obtained by submitting a request through an online form. All use of EMBED Open Data is subject to the data use agreement (Available: https://github.com/Emory-HITI/EMBED_Open_Data/blob/main/EMBED_license.md). Lunit INSIGHT DBT model outputs are considered proprietary and cannot be publicly released. Source data for tables and figures cannot be released publicly due to these restrictions on sharing EMBED data and Lunit INSIGHT DBT model outputs.
Code availability
The code used to perform the data processing and label assignment for this study is hosted on Github (Available: Emory-HITI/Lunit-Model-Evaluation)37.
References
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2020. CA: A Cancer J. Clinicians 70, 7–30 (2020).
Gøtzsche and K. Jørgensen, P. Screening for breast cancer with mammography. Cochrane Database Syst. Rev. 2013, 6 (2013).
Marmot, M. et al. The benefits and harms of breast cancer screening: An independent review. Br. J. Cancer 108, 2205–2240 (2013).
Broeders, M. et al. The impact of mammographic screening on breast cancer mortality in europe: a review of observational studies. J. Med. Screen. 19, 14–25 (2012).
Autier, P., Jørgensen, K. J., Smans, M. and Støvring, H. Effect of screening mammography on the risk of breast cancer deaths and of all-cause deaths: A systematic review with meta-analysis of cohort studies. J. Clin. Epidemiol. 172 (2024).
ACR, 2013 ACR BI-RADS Atlas: Breast Imaging Reporting and Data System. American College of Radiology. (2014).
Lehman, C. D. et al. National performance benchmarks for modern screening digital mammography: update from the Breast Cancer Surveillance Consortium. Radiology 283, 49–58 (2017).
Cunningham, M. P. The breast cancer detection demonstration project 25 years later. CA: A Cancer J. Clin. 47, 131–133 (1997).
Houssami, N. & Skaane, P. Overview of the evidence on digital breast tomosynthesis in breast cancer detection. Breast 22, 101–108 (2013).
Dhamija, E., Gulati, M., Deo, S. V. S., Gogia, A. & Hari, S. Digital breast tomosynthesis: an overview. Indian J. Surg. Oncol. 12, 315–329 (2021).
Richman, I. B. et al. Adoption of digital breast tomosynthesis in clinical practice. JAMA Intern. Med. 179, 1292–1295 (2019).
Astley, S. et al. A comparison of image interpretation times in full field digital mammography and digital breast tomosynthesis. Med. Imaging 8673, 189–196 (2013).
Zuley, M. L. et al. Time to diagnosis and performance levels during repeat interpretations of digital breast tomosynthesis: preliminary observations. Acad. Radiol. 17, 450–455 (2010).
Chen, Y. et al. Measuring reader fatigue in the interpretation of screening digital breast tomosynthesis (DBT). Br. J. Radiol. 96, 20220629 (2023).
Yoon, J. H. et al. Standalone AI for breast cancer detection at screening digital mammography and digital breast tomosynthesis: a systematic review and meta-analysis. Radiology 307, e222639 (2023).
Pinto, M. C. et al. Impact of artificial intelligence decision support using deep learning on breast cancer screening interpretation with single-view wide-angle digital breast tomosynthesis. Radiology 300, 529–536 (2021).
Romero-Martín, S. et al. Stand-alone use of artificial intelligence for digital mammography and digital breast tomosynthesis screening: a retrospective evaluation. Radiology 302, 535–542 (2022).
Shoshan, Y. et al. Artificial intelligence for reducing workload in breast cancer screening with digital breast tomosynthesis. Radiology 303, 69–77 (2022).
Conant, E. F. et al. Improving accuracy and efficiency with concurrent use of artificial intelligence for digital breast tomosynthesis. Radiology: Artif. Intell. 1, e180096 (2019).
Challen, R. et al. Artificial intelligence, bias and clinical safety. BMJ Qual. Saf. 28, 231–237 (2019).
Ghassemi, M., Oakden-Rayner, L. & L. Beam, A. The false hope of current approaches to explainable artificial intelligence in health care. Lancet Dig. Health 3, e745–e750 (2021).
Kelly, C. J., Karthikesalingam, A., Suleyman, M., Corrado, G. & King, D. Key challenges for delivering clinical impact with artificial intelligence. BMC Med. 17, 195 (2019).
Rajpurkar, P., Chen, E., Banerjee, O. & Topol, E. J. AI in health and medicine. Nat. Med. 28, 31–38 (2022).
Purkayastha, S., Trivedi, H. & Gichoya, J. W. Failures hiding in success for artificial intelligence in radiology. J. Am. Coll. Radiol. 18, 517–519 (2021).
Lång, K. et al. Artificial intelligence-supported screen reading versus standard double reading in the Mammography Screening with Artificial Intelligence trial (MASAI): A clinical safety analysis of a randomised, controlled, non-inferiority, single-blinded, screening accuracy study. Lancet Oncol. 24, 936–944 (2023).
Hernström, V. et al. Screening performance and characteristics of breast cancer detected in the Mammography Screening with Artificial Intelligence trial (MASAI): A randomised, controlled, parallel-group, non-inferiority, single-blinded, screening accuracy study. Lancet. Dig. Health, S2589–7500 (2025).
Dembrower, K., Crippa, A., Colón, E., Eklund, M. & Strand, F. Artificial intelligence for breast cancer detection in screening mammography in Sweden: A prospective, population-based, paired-reader, non-inferiority study. Lancet Dig. Health 5, e703–e711 (2023).
Jeong, J. J. et al. The EMory BrEast imaging Dataset (EMBED): a racially diverse, granular dataset of 3.4 million screening and diagnostic mammographic images. Radiology: Artif. Intell. 5, e220047 (2023).
U.S. Food and Drug Administration, Center for Devices and Radiological Health. Lunit INSIGHT DBT v1.1 K242652 approval letter. (2024).
Park, E. K. et al. Impact of AI for digital breast tomosynthesis on breast cancer detection and interpretation time. Radiol. Artif. Intell. 6, e230318 (2024).
Spangler, M. L. et al. Detection and classification of calcifications on digital breast tomosynthesis and 2D digital mammography: a comparison. Am. J. Roentgenol. 196, 320–324 (2011).
Park, G. E., Kang, B. J., Kim, S. H. & Lee, J. Retrospective review of missed cancer detection and its mammography findings with artificial-intelligence-based, computer-aided diagnosis. Diagnostics 12, 387 (2022).
Hovda, T. et al. True and missed interval cancer in organized mammographic screening: a retrospective review study of diagnostic and prior screening mammograms. Acad. Radiol. 29, S180–S191 (2022).
Nanaa, M. et al. Accuracy of an artificial intelligence system for interval breast cancer detection at screening mammography. Radiology 312, e232303 (2024).
Freeman, K. et al. Use of artificial intelligence for image analysis in breast cancer screening programmes: Systematic review of test accuracy. BMJ 374, n1872 (2021).
Byng, D. et al. AI-based prevention of interval cancers in a national mammography screening program. Eur. J. Radiol. 152, 110321 (2022).
Brown-Mulry, B. & Isaac, R. Emory-HITI/Lunit-Model-Evaluation: V1.0.0 Release. Zenodo. (2025).
Acknowledgements
This study was funded by Lunit, Inc. and, in part, the National Institutes of Health (NIH) Agreement No. 1OT2OD032581. The views and conclusions contained in this document are those of the authors and should not be interpreted as representing the official policies, either expressed or implied, of the NIH. We also acknowledge support from the Emory University AI Image Extraction Core Facility (RRID:SCR_026693). J.W.G. declares support from NHLBI Award Number R01HL167811.
Author information
Authors and Affiliations
Contributions
The study was conceptualized by H.T., S.H.L., and A.S. Imaging data preparation was conducted by B.B-M. and A.M. Model inference was conducted by K.M., A.S., and S.H.L. Data engineering, analyses, and visualization were conducted by B.B-M. and R.S.I. under the supervision of H.T. and J.W.G. with input from T.D. and F.L. Statistical analysis was performed by R.S.I. with input from M.W. and B.B-M. C.A.F-R. and B.P. were responsible for clinical interpretation. The first manuscript draft was prepared by B.B-M., R.S.I., and H.T. All authors contributed to the editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
This study was funded by Lunit Inc., however all scientific evaluation and analysis was performed solely by personnel at Emory University. The authors declare no other competing interests.
Peer review
Peer review information
Nature Communications thanks Manisha Bahl, and Aldana Rosso for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Brown-Mulry, B., Isaac, R.S., Lee, S.H. et al. Subgroup performance of a commercial digital breast tomosynthesis model for breast cancer detection. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70637-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70637-3