Abstract
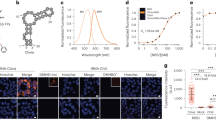
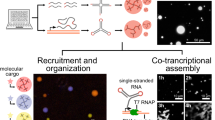
Subcellular compartments organize RNAs into phase-separated condensates, significantly influencing RNA metabolism. However, the study of how specific RNAs regulate interacting factors and their phenotypic outcomes is hindered by the lack of advanced imaging and regulation tools. To address this, we developed an orthogonal RNA aptamer system, Clivia-HT, which integrates a fluorescent imager with a ribonuclease-targeting chimera (RIBOTAC) degrader. This platform enables simultaneous RNA visualization and targeted degradation in living cells. Furthermore, we engineered light-activatable and light-inactivatable RIBOTACs to achieve temporal RNA degradation control. Applying this system, we demonstrated that Activating Transcription Factor 4 (ATF4) mRNA localizes to stress granules and regulates their dynamics. We also detected the role of Non-Coding RNA Activated by DNA Damage (NORAD) in RNA–protein condensate assembly. Our approach establishes a versatile method for spatiotemporal RNA manipulation, providing a powerful tool for probing RNA function in dynamic cellular environments.
Similar content being viewed by others
Data availability
The data supporting the findings of this study are available from the corresponding authors upon request. Source data for the figures and Supplementary Figs. are provided as a Source Data file. Source data are provided with this paper.
References
Cao, X., Zhang, Y., Ding, Y. & Wan, Y. Identification of RNA structures and their roles in RNA functions. Nat. Rev. Mol. Cell Biol. 25, 784–801 (2024).
Le, P., Ahmed, N. & Yeo, G. W. Illuminating RNA biology through imaging. Nat. Cell Biol. 24, 815–824 (2022).
Braselmann, E., Rathbun, C., Richards, E. M. & Palmer, A. E. Illuminating RNA biology: tools for imaging RNA in live mammalian cells. Cell Chem. Biol. 27, 891–903 (2020).
Trachman, R. J. & Ferré-D’Amaré, A. R. Tracking RNA with light: Selection, structure, and design of fluorescence turn-on RNA aptamers. Q. Rev. Biophys. 52, e8 (2019).
Xu, W., Chan, K. M. & Kool, E. T. Fluorescent nucleobases as tools for studying DNA and RNA. Nat. Chem. 9, 1043–1055 (2017).
Lei, X. et al. Illuminating RNA through fluorescent light-up RNA aptamers. Biosens. Bioelectron. 271, 116969 (2025).
Neubacher, S. & Hennig, S. RNA structure and cellular applications of fluorescent light-up aptamers. Angew. Chem. Int. Ed. Engl. 58, 1266–1279 (2019).
Paige, J. S., Wu, K. Y. & Jaffrey, S. R. RNA mimics of green fluorescent protein. Science 333, 642–646 (2011).
Tang, X. et al. Amplified tandem Spinach-based aptamer transcription enables low background mirna detection. Anal. Chem. 90, 10001–10008 (2018).
Filonov, G. S., Moon, J. D., Svensen, N. & Jaffrey, S. R. Broccoli: Rapid selection of an RNA mimic of green fluorescent protein by fluorescence-based selection and directed evolution. J. Am. Chem. Soc. 136, 16299–16308 (2014).
Zhang, J. et al. Tandem Spinach array for mRNA imaging in living bacterial cells. Sci. Rep. 5, 17295 (2015).
Jiang, L. et al. Large stokes shift fluorescent RNAs for dual-emission fluorescence and bioluminescence imaging in live cells. Nat. Methods. 20, 1563–1572 (2023).
Dey, S. K. & Jaffrey, S. R. RIBOTACs: Small molecules target RNA for degradation. Cell Chem. Biol. 26, 1047–1049 (2019).
Zhang, P. et al. Reprogramming of protein-targeted small-molecule medicines to RNA by ribonuclease recruitment. J. Am. Chem. Soc. 143, 13044–13055 (2021).
Tong, Y. et al. Programming inactive RNA-binding small molecules into bioactive degraders. Nature 618, 169–179 (2023).
Tong, Y. et al. Decreasing the intrinsically disordered protein α-synuclein levels by targeting its structured mRNA with a ribonuclease-targeting chimera. Proc. Natl Acad. Sci. USA 121, e2306682120 (2024).
Han, H.-H. et al. The design of small-molecule prodrugs and activatable phototherapeutics for cancer therapy. Chem. Soc. Rev. 52, 879–920 (2023).
Long, K. et al. Near-infrared light-triggered prodrug photolysis by one-step energy transfer. Nat. Commun. 14, 8112 (2023).
Liu, J. et al. Light-induced control of protein destruction by opto-PROTAC. Sci. Adv. 6, eaay5154 (2020).
Xue, G., Wang, K., Zhou, D., Zhong, H. & Pan, Z. Light-induced protein degradation with photocaged PROTACs. J. Am. Chem. Soc. 141, 18370–18374 (2019).
Naro, Y., Darrah, K. & Deiters, A. Optical control of small molecule-induced protein degradation. J. Am. Chem. Soc. 142, 2193–2197 (2020).
Klán, P. et al. Photoremovable protecting groups in chemistry and biology: Reaction mechanisms and efficacy. Chem. Rev. 113, 119–191 (2013).
Zhou, W., Jin, J., Chen, H., Zhang, W. & Luan, X. Fueling the revolution: RIBOTACs manipulating RNA decay. Trends Cancer 11, 1141–1144 (2025).
Ramaswamy Krishnan, S., Roy, A. & Michael Gromiha, M. R-sim: A database of binding affinities for RNA-small molecule interactions. J. Mol. Biol. 435, 167914 (2023).
Velagapudi, S. P. et al. Approved anti-cancer drugs target oncogenic non-coding RNAs. Cell Chem. Biol. 25, 1086–1094.e1087 (2018).
Sztuba-Solinska, J. et al. Identification of biologically active, HIV TAR RNA-binding small molecules using small molecule microarrays. J. Am. Chem. Soc. 136, 8402–8410 (2014).
Warner, K. D., Hajdin, C. E. & Weeks, K. M. Principles for targeting RNA with drug-like small molecules. Nat. Rev. Drug Discov. 17, 547–558 (2018).
Joly, J. P. et al. Artificial nucleobase-amino acid conjugates: A new class of TAR RNA binding agents. Chemistry 20, 2071–2079 (2014).
Han, Y. et al. Structure of human RNase L reveals the basis for regulated RNA decay in the IFN response. Science 343, 1244–1248 (2014).
Costales, M. G. et al. Small-molecule targeted recruitment of a nuclease to cleave an oncogenic RNA in a mouse model of metastatic cancer. Proc. Natl Acad. Sci. USA 117, 2406–2411 (2020).
Meyer, S. M. et al. DNA-encoded library screening to inform design of a ribonuclease targeting chimera (RiboTAC). J. Am. Chem. Soc. 144, 21096–21102 (2022).
Liu, J. L. & Gall, J. G. U bodies are cytoplasmic structures that contain uridine-rich small nuclear ribonucleoproteins and associate with P bodies. Proc. Natl Acad. Sci. USA 104, 11655–11659 (2007).
Braselmann, E. et al. A multicolor riboswitch-based platform for imaging of RNA in live mammalian cells. Nat. Chem. Biol. 14, 964–971 (2018).
Lee, A. K. et al. Translational repression of G3BP in cancer and germ cells suppresses stress granules and enhances stress tolerance. Mol. Cell 79, 645–659.e649 (2020).
Anderson, P. & Kedersha, N. RNA granules. J. Cell Biol. 172, 803–808 (2006).
Mateju, D. et al. Single-molecule imaging reveals translation of mRNAs localized to stress granules. Cell 183, 1801–1812.e1813 (2020).
Zhang, Y., Wang, L., Wang, F., Chu, X. & Jiang, J. H. G-quadruplex mRNAs silencing with inducible ribonuclease targeting chimera for precision tumor therapy. J. Am. Chem. Soc. 146, 15815–15824 (2024).
Zhang, Y. et al. Stimulus-activated ribonuclease targeting chimeras for tumor microenvironment activated cancer therapy. Nat. Commun. 16, 1288 (2025).
Zou, Z. et al. Photoactivatable fluorescent probes for spatiotemporal-controlled biosensing and imaging. Trends Anal. Chem. 125, 115811 (2020).
Chen, L.-L. & Kim, V. N. Small and long non-coding RNAs: Past, present, and future. Cell 187, 6451–6485 (2024).
Yin, Y. et al. U1 snRNP regulates chromatin retention of noncoding RNAs. Nature 580, 147–150 (2020).
Tichon, A. et al. A conserved abundant cytoplasmic long noncoding RNA modulates repression by Pumilio proteins in human cells. Nat. Commun. 7, 12209 (2016).
Elguindy, M. M. & Mendell, J. T. NORAD-induced Pumilio phase separation is required for genome stability. Nature 595, 303–308 (2021).
Lee, S. et al. Noncoding RNA NORAD regulates genomic stability by sequestering pumilio proteins. Cell 164, 69–80 (2016).
You, X., He, R., Liu, S.-Y. & Dai, Z. Advances in fluorescence imaging of RNAs in living cells based on functional nucleic acid probes. Biomed. Anal. 1, 1–27 (2024).
Mondal, S. et al. Multivalent interactions between molecular components involved in fast endophilin mediated endocytosis drive protein phase separation. Nat. Commun. 13, 5017 (2022).
Povedano, J. M. et al. Engineering forward genetics into cultured cancer cells for chemical target identification. Cell Chem. Biol. 26, 1315–1321.e1313 (2019).
Zhou, Z. et al. VCAM-1 secreted from cancer-associated fibroblasts enhances the growth and invasion of lung cancer cells through AKT and MAPK signaling. Cancer Lett. 473, 62–73 (2020).
Duan, Y. et al. Protein modifications throughout the lung cancer proteome unravel the cancer-specific regulation of glycolysis. Cell Rep. 37, 110137 (2021).
Elguindy, M. M. et al. Pumilio, but not RBMX, binding is required for regulation of genomic stability by noncoding RNA NORAD. eLife 8, e48625 (2019).
Mito, M., Kawaguchi, T., Hirose, T. & Nakagawa, S. Simultaneous multicolor detection of RNA and proteins using super-resolution microscopy. Methods 98, 158–165 (2016).
Arney, J. W. & Weeks, K. M. RNA-ligand interactions quantified by surface plasmon resonance with reference subtraction. Biochemistry 61, 1625–1632 (2022).
Acknowledgements
This work was supported by grants from the National Key R&D Program of China (2025YFC3409700 to T.D.), the Chinese Academy of Medical Sciences Innovation Fund for Medical Sciences (2025-I2M-XHXX-087 to X.L.), the National Key Research and Development Program of China (2023YFC3404102 to X.L.), the National Natural Science Foundation of China (82373784 to X.L.), and the Beijing-Tianjin-Hebei Basic Research Cooperation Special Project (J230018 to X.L.).
Author information
Authors and Affiliations
Contributions
J.W., K.M., and X.C. performed the experimental studies and conducted manuscript writing. H.W., H.S., Y.G., M.W., W.Z., Y.W., J.R., and M.L. carried out data analysis and manuscript validation. T.D., P.Z., and X.L. supervised the work.
Corresponding authors
Ethics declarations
Competing interests
X.L., T.D., K.M., J.W., H.Y., X.C., C.A., H.S., and K.L. have filed a patent application (CN202610149905.8) related to this work. The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jingxin Wang and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, J., Ma, K., Cao, X. et al. An orthogonal RNA aptamer for targeted RNA regulation in phase-separated subcellular compartments. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70638-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70638-2