Abstract
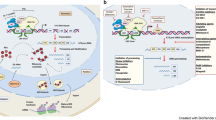
Ribosomal protein mutations are increasingly associated with cancer risk and thought to perturb ribosome function. At the same time, they reportedly activate p53, a critical anti-cancer barrier. To determine how these mutations overcome this protective block to enable tumorigenesis, we generate an in vivo model of the hotspot ribosomal protein RPS15-S138F mutation identified as a putative driver of chronic lymphocytic leukemia. Under pre-leukemic conditions, this mutation induces ribosome biogenesis defects and altered translation resulting in oxidative stress, DNA damage and induction of a p53-dependent response that promote initial cellular hypo-proliferation. However, a subset of aged mice with mutated Rps15 eventually develop B-cell leukemia (37% penetrance), which exhibits increased Myc activity with strong pro-survival and proliferation signatures. Mutant RPS15 thus induces both hypo- and hyper-proliferative signals, initially weighted towards cell cycle arrest; and that through translational rewiring, oxidative stress, DNA damage response defects and genomic instability set the stage for the acquisition of additional driving mutations, such as TP53 deletion, that can overcome this cell cycle block to trigger tumorigenesis.
Similar content being viewed by others
Data availability
The molecular data used in this study are publicly available and are included in the following patient cohorts (Fig. 1a, Supp Fig. 1a-c, Supp Table 1, Supp Data 1): Dana-Farber Cancer Institute (DFCI), German CLL Study Group (GCLLSG), French CLL Cohort, International Cancer Genome Consortium (ICGC), MD Anderson Cancer Center (MDACC), National Heart Lung and Blood Institute (NHLBI) and University of California San Diego (UCSD). Sequencing, expression, and genotyping is available at European Genome-Phenome Archive (EGA, http://www.ebi.ac.uk/ega/), which is hosted at the European Bioinformatics Institute (EBI), under accession numbers EGAS00000000092 (ICGC cohort) and in dbGaP under accession numbers: phs001473.v2.p1 (MDACC, NHLBI), phs000922.v2.p1 (GCLLSG), phs001431.v2.p1 (DFCI, UCSD), phs001091.v1.p1 (MDACC), phs000435.v3.p1 (DFCI), phs002297.v2.p1 (NHLBI), phs000879.v1.p1 (DFCI), phs002335.v1 (French), and GEO accession number GSE143673 (GCLLSG). All reported RNA-seq and Riboseq data, including raw sequencing and quantitation files, have been deposited to the GEO repository (GSE293841), and can be accessed at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE293841. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE [1] partner repository with the dataset identifier PXD071901. Source data for all figures are provided as a Source Data file with this paper. Source data are provided with this paper.
Materials availability
The Cd19-Cre/Rps15-S138F flox mouse strain will be available at the Jackson Laboratory Repository with the JAX Stock no. 037632 (http://jaxmice.jax.org/query).
Code availability
Supplementary and wrapper scripts used to conduct genomics, transcriptomics, and ribosomal sequencing analyses are made available via the following links to publicly accessible repositories: GitHub Repository: https://github.com/nruthen/RPS15_Mutant_Mouse_Multiomics. https://doi.org/10.5281/zenodo.17548003. Terra Workspace: https://app.terra.bio/#workspaces/cll-mouse/Wu_RPS15_Catherine_Analysis_Workspace.
References
Goudarzi, K. M. & Lindström, M. S. Role of ribosomal protein mutations in tumor development (Review). Int. J. Oncol. 48, 1313–1324 (2016).
Keersmaecker, K. D. et al. Exome sequencing identifies mutation in CNOT3 and ribosomal genes RPL5 and RPL10 in T-cell acute lymphoblastic leukemia. Nat. Genet. 45, 186–190 (2013).
Lawrence, M. S. et al. Discovery and saturation analysis of cancer genes across 21 tumor types. Nature. 505, 495–501 (2014).
Nieminen, T. T. et al. Germline mutation of RPS20, encoding a ribosomal protein, causes predisposition to hereditary nonpolyposis colorectal carcinoma without DNA mismatch repair deficiency. Gastroenterology. 147, 595–598.e5 (2014).
Novetsky A. P. et al. Frequent mutations in the RPL22 gene and its clinical and functional implications. Gynecol Oncol. 2013;128: https://doi.org/10.1016/j.ygyno.2012.10.026.
Hofman, I. J. F. et al. RPL5 on 1p22.1 is recurrently deleted in multiple myeloma and its expression is linked to bortezomib response. Leukemia. 31, 1706–1714 (2017).
Fancello, L., Kampen, K. R., Hofman, I. J. F., Verbeeck, J. & De Keersmaecker, K. The ribosomal protein gene RPL5 is a haploinsufficient tumor suppressor in multiple cancer types. Oncotarget. 8, 14462–14478 (2017).
Landau, D. A. et al. Mutations driving CLL and their evolution in progression and relapse. Nature. 526, 525–530 (2015).
Hernández-Sánchez, M. et al. CLL cells cumulate genetic aberrations prior to the first therapy even in outwardly inactive disease phase. Leukemia. 33, 518–558 (2019).
Ljungström, V. et al. Whole-exome sequencing in relapsing chronic lymphocytic leukemia: clinical impact of recurrent RPS15 mutations. Blood. 127, 1007–1016 (2016).
Burger, J. A. et al. Clonal evolution in patients with chronic lymphocytic leukaemia developing resistance to BTK inhibition. Nat. Commun. 7, 11589 (2016).
Sulima, S. O., Kampen, K. R. & De Keersmaecker, K. Cancer biogenesis in ribosomopathies. Cells. 8, 229 (2019).
Sulima, S. O., Hofman, I. J. F., De Keersmaecker, K. & Dinman, J. D. How ribosomes translate cancer. Cancer Discov. 7, 1069–1087 (2017).
Bastide, A. & David, A. The ribosome, (slow) beating heart of cancer (stem) cell. Oncogenesis. 7, 34 (2018).
Pelletier, J., Thomas, G. & Volarević, S. Ribosome biogenesis in cancer: new players and therapeutic avenues. Nat. Rev. Cancer. 18, 51–63 (2018).
Kampen, K. R. et al. The ribosomal RPL10 R98S mutation drives IRES-dependent BCL-2 translation in T-ALL. Leukemia. 33, 319–332 (2019).
Kampen, K. R., Sulima, S. O., Vereecke, S. & De Keersmaecker, K. Hallmarks of ribosomopathies. Nucleic Acids Res. 48, 1013–1028 (2020).
Sulima, S. O. et al. Ribosomal lesions promote oncogenic mutagenesis. Cancer Res. 79, 320–327 (2019).
De Keersmaecker, K., Sulima, S. O. & Dinman, J. D. Ribosomopathies and the paradox of cellular hypo- to hyperproliferation. Blood. 125, 1377–1382 (2015).
Sulima, S. O. et al. Bypass of the pre-60S ribosomal quality control as a pathway to oncogenesis. Proc. Natl. Acad. Sci. Proc. Natl. Acad. Sci. 111, 5640–5645 (2014).
Ajore, R. et al. Deletion of ribosomal protein genes is a common vulnerability in human cancer, especially in concert with TP53 mutations. EMBO Mol. Med. 9, 498–507 (2017).
Zhang, Y. & Lu, H. Signaling to p53: ribosomal proteins find their way. Cancer Cell. 16, 369–377 (2009).
Golomb, L., Volarevic, S. & Oren, M. p53 and ribosome biogenesis stress: the essentials. FEBS Lett. 588, 2571–2579 (2014).
Liu, Y., Deisenroth, C. & Zhang, Y. RP–MDM2–p53 pathway: linking ribosomal biogenesis and tumor surveillance. Trends Cancer. 2, 191–204 (2016).
Deisenroth, C. & Zhang, Y. Ribosome biogenesis surveillance: probing the ribosomal protein-Mdm2-p53 pathway. Oncogene. 29, 4253–4260 (2010).
Marechal, V., Elenbaas, B., Piette, J., Nicolas, J. C. & Levine, A. J. The ribosomal L5 protein is associated with mdm-2 and mdm-2-p53 complexes. Mol. Cell Biol. 14, 7414–7420 (1994).
Zhang, Y. et al. Ribosomal protein L11 negatively regulates oncoprotein MDM2 and mediates a p53-dependent ribosomal-stress checkpoint pathway. Mol. Cell Biol. 23, 8902–8912 (2003).
Dai, M.-S. et al. Ribosomal protein L23 activates p53 by inhibiting MDM2 function in response to ribosomal perturbation but not to translation inhibition. Mol. Cell Biol. 24, 7654–7668 (2004).
Chen, D. et al. Ribosomal protein S7 as a novel modulator of p53-MDM2 interaction: binding to MDM2, stabilization of p53 protein, and activation of p53 function. Oncogene. 26, 5029–5037 (2007).
Daftuar, L., Zhu, Y., Jacq, X. & Prives, C. Ribosomal proteins RPL37, RPS15 and RPS20 regulate the Mdm2-p53-MdmX network. PLoS One. 8, e68667 (2013).
Dameshek, W. Riddle: What do aplastic anemia, paroxysmal nocturnal hemoglobinuria (PNH) and “hypoplastic”: leukemia have in common? Blood. 30, 251–254 (1967).
Kang, J. et al. Ribosomal proteins and human diseases: molecular mechanisms and targeted therapy. Sig Transduct. Target Ther. 6, 1–22 (2021).
Kapralova, K. et al. Oxidative DNA damage, inflammatory signature, and altered erythrocytes properties in diamond-blackfan anemia. Int. J. Mol. Sci. 21, 9652 (2020).
Bretones, G. et al. Altered patterns of global protein synthesis and translational fidelity in RPS15-mutated chronic lymphocytic leukemia. Blood. 132, 2375–2388 (2018).
Ntoufa, S. et al. RPS15 mutations rewire RNA translation in chronic lymphocytic leukemia. Blood Adv. 5, 2788–2792 (2021).
Knisbacher B. A. et al. Molecular map of chronic lymphocytic leukemia and its impact on outcome. Nat. Genet. (2022)
Rössler, I. et al. The C-terminal tail of ribosomal protein Rps15 is engaged in cytoplasmic pre-40S maturation. RNA Biol. 19, 560–574 (2022).
Yu, L. et al. Survival of Del17p CLL depends on genomic complexity and somatic mutation. Clin. Cancer Res. 23, 735–745 (2017).
Kwok, M., Agathanggelou, A. & Stankovic, T. DNA damage response defects in hematologic malignancies: mechanistic insights and therapeutic strategies. Blood. 143, 2123–2144 (2024).
Wang, D.-M. et al. Intermediate prognosis of 6q deletion in chronic lymphocytic leukemia. Leuk. Lymphoma. 52, 230–237 (2011).
Lipreri da Silva, J. C. et al. Ezrin is highly expressed and a druggable target in chronic lymphocytic leukemia. Life Sci. 311, 121146 (2022).
Song, Y. et al. Ezrin mediates invasion and metastasis in tumorigenesis: a review. Front. Cell Dev. Biol. 8, 588801 (2020).
Li, X., Jin, F. & Li, Y. A novel autophagy-related lncRNA prognostic risk model for breast cancer. J. Cell Mol. Med. 25, 4–14 (2021).
Khan, T., Kryza, T., Lyons, N. J., He, Y. & Hooper, J. D. The CDCP1 signaling hub: a target for cancer detection and therapeutic intervention. Cancer Res. 81, 2259–2269 (2021).
Yang, G. et al. circ-BIRC6, a circular RNA, promotes hepatocellular carcinoma progression by targeting the miR-3918/Bcl2 axis. Cell Cycle. 18, 976–989 (2019).
Han, Y. & Wang, H. MiR-3918 inhibits tumorigenesis of glioma via targeting EGFR to regulate PI3K/AKT and ERK pathways. J. Mol. Neurosci. 72, 433–440 (2022).
Klintman, J. et al. Genomic and transcriptomic correlates of Richter transformation in chronic lymphocytic leukemia. Blood. 137, 2800–2816 (2021).
Moulin, C. et al. Clinical, biological, and molecular genetic features of Richter syndrome and prognostic significance: a study of the French Innovative Leukemia Organization. Am. J. Hematol. 96, E311–E314 (2021).
Nadeu F. et al. Detection of early seeding of Richter transformation in chronic lymphocytic leukemia. Nat Med. 1–10 (2022).
Steele, A. J. et al. The sesquiterpene lactone parthenolide induces selective apoptosis of B-chronic lymphocytic leukemia cells in vitro. Leukemia. 20, 1073–1079 (2006).
Li, X. et al. Derivatisation of parthenolide to address chemoresistant chronic lymphocytic leukaemia. Medchemcomm. 10, 1379–1390 (2019).
Agathanggelou, A. et al. Targeting the Ataxia Telangiectasia mutated-null phenotype in chronic lymphocytic leukemia with pro-oxidants. Haematologica. 100, 1076–1085 (2015).
Akasaka, S. & Yamamoto, K. Hydrogen peroxide induces G:C to TA and G:C to C:G transversions in the supF gene of Escherichia coli. Mol. Gen. Genet. 243, 500–505 (1994).
Lee, D., O’Connor, T. R. & Pfeifer, G. P. Oxidative DNA damage induced by copper and hydrogen peroxide promotes CG→TT tandem mutations at methylated CpG dinucleotides in nucleotide excision repair-deficient cells. Nucleic Acids Res. 30, 3566–3573 (2002).
el-Deiry, W. S. et al. WAF1/CIP1 is induced in p53-mediated G1 arrest and apoptosis. Cancer Res. 54, 1169–1174 (1994).
Harper, J. W., Adami, G. R., Wei, N., Keyomarsi, K. & Elledge, S. J. The p21 Cdk-interacting protein Cip1 is a potent inhibitor of G1 cyclin-dependent kinases. Cell. 75, 805–816 (1993).
Deng, C., Zhang, P., Harper, J. W., Elledge, S. J. & Leder, P. Mice lacking p21CIP1/WAF1 undergo normal development, but are defective in G1 checkpoint control. Cell. 82, 675–684 (1995).
Karanjawala, Z. E., Murphy, N., Hinton, D. R., Hsieh, C. L. & Lieber, M. R. Oxygen metabolism causes chromosome breaks and is associated with the neuronal apoptosis observed in DNA double-strand break repair mutants. Curr. Biol. 12, 397–402 (2002).
Sallmyr, A., Fan, J. & Rassool, F. V. Genomic instability in myeloid malignancies: increased reactive oxygen species (ROS), DNA double strand breaks (DSBs) and error-prone repair. Cancer Lett. 270, 1–9 (2008).
Tanaka, T., Halicka, H. D., Huang, X., Traganos, F. & Darzynkiewicz, Z. Constitutive histone H2AX phosphorylation and ATM activation, the reporters of DNA damage by endogenous oxidants. Cell Cycle. 5, 1940–1945 (2006).
Letavayová, L. et al. Relative contribution of homologous recombination and non-homologous end-joining to DNA double-strand break repair after oxidative stress in Saccharomyces cerevisiae. DNA Repair. 5, 602–610 (2006).
Sharma, V. et al. Oxidative stress at low levels can induce clustered DNA lesions leading to NHEJ-mediated mutations. Oncotarget. 7, 25377–25390 (2016).
Lee, T., Yao, G., Nevins, J. & You, L. Sensing and integration of Erk and PI3K signals by Myc. PLoS Comput. Biol. 4, e1000013 (2008).
Tsai, W.-B. et al. Activation of RAS/PI3K/ERK pathway induces c-Myc stabilization to upregulate argininosuccinate synthetase, leading to arginine deiminase resistance in melanoma cells. Cancer Res. 72, 2622–2633 (2012).
Chen, J. et al. ZAP-70 constitutively regulates gene expression and protein synthesis in chronic lymphocytic leukemia. Blood. 137, 3629–3640 (2021).
Chen, J., Moore, A. & Ringshausen, I. ZAP-70 shapes the immune microenvironment in B cell malignancies. Front. Oncol. 10, 595832 (2020).
Adams, D. J. et al. Genetic determinants of micronucleus formation in vivo. Nature. 627, 130–136 (2024).
Hosea, R., Hillary, S., Naqvi, S., Wu, S. & Kasim, V. The two sides of chromosomal instability: drivers and brakes in cancer. Signal Transduct. Target. Ther. 9, 75 (2024).
Biran, A. et al. Activation of notch and Myc signaling via B-cell-restricted depletion of Dnmt3a generates a consistent murine model of chronic lymphocytic leukemia. Cancer Res. 81, 6117–6130 (2021).
Chang, A. et al. Recruitment of KMT2C/MLL3 to DNA damage sites mediates DNA damage responses and regulates PARP inhibitor sensitivity in cancer. Cancer Res. 81, 3358–3373 (2021).
Lv, S. et al. Loss of KMT2D induces prostate cancer ROS-mediated DNA damage by suppressing the enhancer activity and DNA binding of antioxidant transcription factor FOXO3. Epigenetics. 14, 1194–1208 (2019).
Sun, L. et al. MGA mutation as a novel biomarker for immune checkpoint therapies in non-squamous non-small cell lung cancer. Front. Pharm. 12, 625593 (2021).
Llabata, P. et al. MAX mutant small-cell lung cancers exhibit impaired activities of MGA-dependent noncanonical polycomb repressive complex. Proc. Natl. Acad. Sci. USA. 118, e2024824118 (2021).
Chan, D. K. H. et al. Biallelic FBXW7 knockout induces AKAP8-mediated DNA damage in neighbouring wildtype cells. Cell Death Discov. 9, 1–15 (2023).
Lan, H. & Sun, Y. Tumor suppressor FBXW7 and its regulation of DNA damage response and repair. Front Cell Dev. Biol. 9, 751574 (2021).
Close, V. et al. FBXW7 mutations reduce binding of NOTCH1, leading to cleaved NOTCH1 accumulation and target gene activation in CLL. Blood. 133, 830–839 (2019).
Llabata, P. et al. Multi-omics analysis identifies MGA as a negative regulator of the MYC pathway in lung adenocarcinoma. Mol. Cancer Res. 18, 574–584 (2020).
Mathsyaraja H. et al. Loss of MGA repression mediated by an atypical polycomb complex promotes tumor progression and invasiveness. eLife. 10:e64212 (2021).
Chitalia, A. et al. Descriptive analysis of genetic aberrations and cell of origin in Richter transformation. Leuk. Lymphoma. 60, 971–979 (2019).
Parry, E. M. et al. Evolutionary history of transformation from chronic lymphocytic leukemia to Richter syndrome. Nat. Med. 29, 158–169 (2023).
Léger-Silvestre, I. et al. Specific role for yeast homologs of the Diamond Blackfan Anemia-associated Rps19 protein in ribosome synthesis. J. Biol. Chem. 280, 38177–38185 (2005).
Léger-Silvestre, I. et al. The ribosomal protein Rps15p is required for nuclear exit of the 40S subunit precursors in yeast. EMBO J. 23, 2336–2347 (2004).
Ferreira-Cerca, S., Pöll, G., Gleizes, P.-E., Tschochner, H. & Milkereit, P. Roles of eukaryotic ribosomal proteins in maturation and transport of pre-18S rRNA and ribosome function. Mol. Cell. 20, 263–275 (2005).
Khairulina, J. et al. Eukaryote-specific motif of ribosomal protein S15 neighbors A site codon during elongation and termination of translation. Biochimie. 92, 820–825 (2010).
Bulygin, K., Chavatte, L., Frolova, L., Karpova, G. & Favre, A. The first position of a codon placed in the A site of the human 80S ribosome contacts nucleotide C1696 of the 18S rRNA as well as proteins S2, S3, S3a, S30, and S15. Biochemistry. 44, 2153–2162 (2005).
Pisarev, A. V. et al. Specific functional interactions of nucleotides at key −3 and +4 positions flanking the initiation codon with components of the mammalian 48S translation initiation complex. Genes Dev. 20, 624–636 (2006).
Robinson, J. T., Thorvaldsdóttir, H., Wenger, A. M., Zehir, A. & Mesirov, J. P. Variant review with the integrative genomics viewer. Cancer Res. 77, e31–e34 (2017).
Handy, D. E. et al. Glutathione peroxidase-1 regulates mitochondrial function to modulate redox-dependent cellular responses. J. Biol. Chem. 284, 11913–11921 (2009).
Zhao, Y. et al. Noncanonical regulation of alkylation damage resistance by the OTUD4 deubiquitinase. EMBO J. 34, 1687–1703 (2015).
Chai, B., Huang, J., Cairns, B. R. & Laurent, B. C. Distinct roles for the RSC and Swi/Snf ATP-dependent chromatin remodelers in DNA double-strand break repair. Genes Dev. 19, 1656–1661 (2005).
Park, J.-H. et al. Mammalian SWI/SNF complexes facilitate DNA double-strand break repair by promoting gamma-H2AX induction. EMBO J. 25, 3986–3997 (2006).
Ogiwara, H. et al. Histone acetylation by CBP and p300 at double-strand break sites facilitates SWI/SNF chromatin remodeling and the recruitment of non-homologous end joining factors. Oncogene. 30, 2135–2146 (2011).
Lee, H.-S., Park, J.-H., Kim, S.-J., Kwon, S.-J. & Kwon, J. A cooperative activation loop among SWI/SNF, γ-H2AX and H3 acetylation for DNA double-strand break repair. EMBO J. 29, 1434–1445 (2010).
Peng, G. et al. BRIT1/MCPH1 links chromatin remodelling to DNA damage response. Nat. Cell Biol. 11, 865–872 (2009).
Pace, P. et al. FANCE: the link between Fanconi anaemia complex assembly and activity. EMBO J. 21, 3414–3423 (2002).
Wang, X. et al. Chk1-mediated phosphorylation of FANCE is required for the Fanconi anemia/BRCA pathway. Mol. Cell Biol. 27, 3098–3108 (2007).
Sunavala-Dossabhoy, G. & De Benedetti, A. Tousled homolog, TLK1, binds and phosphorylates Rad9; TLK1 acts as a molecular chaperone in DNA repair. DNA Repair. 8, 87–102 (2009).
TLK1B promotes repair of DSBs via its interaction with Rad9 and Asf1 Caroline Canfield, justin rains, and Arrigo De Benedetti - PMC [Internet]. [cited 7 Nov 2023]. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2803485/
Werner, S. R. et al. Enhanced cell cycle progression and down regulation of p21(Cip1/Waf1) by PRL tyrosine phosphatases. Cancer Lett. 202, 201–211 (2003).
Lunardi, A. et al. A genome-scale protein interaction profile of Drosophila p53 uncovers additional nodes of the human p53 network. Proc. Natl. Acad. Sci. USA. 107, 6322–6327 (2010).
Tompkins, V. S. et al. A novel nuclear interactor of ARF and MDM2 (NIAM) that maintains chromosomal stability. J. Biol. Chem. 282, 1322–1333 (2007).
Annibaldis, G. et al. Readthrough of stop codons under limiting ABCE1 concentration involves frameshifting and inhibits nonsense-mediated mRNA decay. Nucleic Acids Res. 48, 10259–10279 (2020).
Mangkalaphiban, K. et al. Transcriptome-wide investigation of stop codon readthrough in Saccharomyces cerevisiae. PLOS Genet. 17, e1009538 (2021).
Wangen J. R., Green R. Stop codon context influences genome-wide stimulation of termination codon readthrough by aminoglycosides. eLife. 9:e52611 (2020).
Loughran, G. et al. Evidence of efficient stop codon readthrough in four mammalian genes. Nucleic Acids Res. 42, 8928–8938 (2014).
Fromm, S. A. et al. The translating bacterial ribosome at 1.55 Å resolution generated by cryo-EM imaging services. Nat. Commun. 14, 1095 (2023).
Gromadski, K. B. & Rodnina, M. V. Kinetic determinants of high-fidelity tRNA discrimination on the ribosome. Mol. Cell. 13, 191–200 (2004).
LaRiviere, F. J., Wolfson, A. D. & Uhlenbeck, O. C. Uniform binding of aminoacyl-tRNAs to elongation factor Tu by thermodynamic compensation. Science. 294, 165–168 (2001).
Negrutskii, B. S. & Deutscher, M. P. Channeling of aminoacyl-tRNA for protein synthesis in vivo. Proc. Natl. Acad. Sci. USA. 88, 4991–4995 (1991).
Stapulionis, R. & Deutscher, M. P. A channeled tRNA cycle during mammalian protein synthesis. Proc. Natl. Acad. Sci. USA. 92, 7158–7161 (1995).
Godinic-Mikulcic, V. et al. Archaeal aminoacyl-tRNA synthetases interact with the ribosome to recycle tRNAs. Nucleic Acids Res. 42, 5191–5201 (2014).
Hausmann, S., Ramirez, A., Schneider, S., Schwer, B. & Shuman, S. Biochemical and genetic analysis of RNA cap guanine-N2 methyltransferases from Giardia lamblia and Schizosaccharomyces pombe. Nucleic Acids Res. 35, 1411–1420 (2007).
Nguyen T.-T., Stahl G., Déquard-Chablat M., Contamine V., Denmat SH-L. The eukaryotic ribosomal protein S15/uS19 is involved in fungal development and its C-terminal tail contributes to stop codon recognition [Internet]. bioRxiv; 2020 [cited 11 Dec 2022]. pp. 2020.02.09.940346. Available from: https://www.biorxiv.org/content/10.1101/2020.02.09.940346v1
Zeeshan, H. M. A., Lee, G. H., Kim, H.-R. & Chae, H.-J. Endoplasmic reticulum stress and associated ROS. Int. J. Mol. Sci. 17, 327 (2016).
Yin, S. et al. A murine model of chronic lymphocytic leukemia based on B cell-restricted expression of Sf3b1 mutation and Atm deletion. Cancer Cell. 35, 283–296.e5 (2019).
Lazarian, G. et al. A hotspot mutation in transcription factor IKZF3 drives B cell neoplasia via transcriptional dysregulation. Cancer Cell. 39, 380–393.e8 (2021).
Morgado-Palacin, L. et al. Partial loss of Rpl11 in adult mice recapitulates diamond-blackfan anemia and promotes lymphomagenesis. Cell Rep. 13, 712–722 (2015).
Vitale, I., Galluzzi, L., Castedo, M. & Kroemer, G. Mitotic catastrophe: a mechanism for avoiding genomic instability. Nat. Rev. Mol. Cell Biol. 12, 385–392 (2011).
Gujar, V., Li, H., Paull, T. T., Neumann, C. A. & Weyemi, U. Unraveling the nexus: genomic instability and metabolism in cancer. Cell Rep. 44, 115540 (2025).
Barlow, J. L. et al. A p53-dependent mechanism underlies macrocytic anemia in a mouse model of human 5q- syndrome. Nat. Med. 16, 59–66 (2010).
Montoro, M. J. et al. Influence of TP53 gene mutations and their allelic status in myelodysplastic syndromes with isolated 5q deletion. Blood. 144, 1722–1731 (2024).
Schneider, R. K. et al. Rps14 haploinsufficiency causes a block in erythroid differentiation mediated by S100A8 and S100A9. Nat. Med. 22, 288–297 (2016).
Yu, L., Yu, T.-T. & Young, K. H. Cross-talk between Myc and p53 in B-cell lymphomas. Chronic Dis. Transl. Med. 5, 139–154 (2019).
Campo, E. et al. The 2008 WHO classification of lymphoid neoplasms and beyond: evolving concepts and practical applications. Blood. 117, 5019–5032 (2011).
Kowarz, E., Löscher, D. & Marschalek, R. Optimized Sleeping Beauty transposons rapidly generate stable transgenic cell lines. Biotechnol. J. 10, 647–653 (2015).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods. 11, 783–784 (2014).
Doench, J. G. et al. Rational design of highly active sgRNAs for CRISPR-Cas9–mediated gene inactivation. Nat. Biotechnol. 32, 1262–1267 (2014).
Quijada-Álamo, M. et al. Dissecting the role of TP53 alterations in del(11q) chronic lymphocytic leukemia. Clin. Transl. Med. 11, e304 (2021).
McGlincy, N. J. & Ingolia, N. T. Transcriptome-wide measurement of translation by ribosome profiling. Methods. 126, 112–129 (2017).
Chothani, S. et al. deltaTE: detection of translationally regulated genes by integrative analysis of Ribo-seq and RNA-seq data. Curr. Protoc. Mol. Biol. 129, e108 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 29, 15–21 (2013).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. 102, 15545–15550 (2005).
Iyer M. K. et al. The landscape of long noncoding RNAs in the human transcriptome. Nat. Genet. 47: 199–208 (2015).
Li H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM [Internet]. arXiv; 2013 [cited 19 Dec 2025]. http://arxiv.org/abs/1303.3997
Fantini, D., Vidimar, V., Yu, Y., Condello, S. & Meeks, J. J. MutSignatures: an R package for extraction and analysis of cancer mutational signatures. Sci. Rep. 10, 18217 (2020).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Kim S. et al. Strelka2: fast and accurate calling of germline and somatic variants. Nat. Methods. 15:591–594 (2018).
Ching, T., Huang, S. & Garmire, L. X. Power analysis and sample size estimation for RNA-Seq differential expression. RNA. 20, 1684–1696 (2014).
Benjamin D. et al. Calling Somatic SNVs and Indels with Mutect2 [Internet]. bioRxiv; 2019 [cited 19 Dec 2025]. 861054. Available from: https://www.biorxiv.org/content/10.1101/861054v1
Acknowledgements
We are grateful to Nicoletta Cieri, Nira Krasnow, Chip Stewart and Martin Aryee for helpful discussions. We acknowledge Sam Pollock, Lan Nguyen, Fanny Dao, and Candace Patterson for expert project management. We thank the Dana-Farber Flow Cytometry Core, the Broad Institute Walk-Up Sequencing Core, and the Dana-Farber Cancer Institute animal research facility technical team for their technical support. This study was supported by a grant from the National Institutes of Health (NIH)/National Cancer Institute (NCI) (P01 CA206978 and P01-CA081534). CJW is the Lavine Family Chair of Preventative Therapies at Dana-Farber Cancer Institute. C.J.W. acknowledges support from the NIH/NCI (R01 CA216273, U10 CA180861) and from the CLL Global Foundation, and is a member of the Parker Institute for Cancer Immunotherapy at Dana-Farber Cancer Institute, whose work is supported, in part, by the Parker Institute for Cancer Immunotherapy. C.G. is a Scholar through the American Society of Hematology MMSAP Program and the F31 Diversity Individual Predoctoral Fellowship program through the NCI (5F31CA239443-03). M.K. is supported by a Cancer Research UK Clinician Scientist Fellowship (RCCFELCSF-May21\100002). S.L. is supported by the NCI Research Specialist Award (R50CA251956).
Author information
Authors and Affiliations
Contributions
C.G. and M.K. designed the murine and cell line models, performed experiments, and analyzed data. C.G., N.R., M.K., P.W., T.O., D.F., S.C., and S.L. participated in library preparation and analysis of RNA-seq, Ribo-seq, proteomics, and WGS data. C.G., M.K., P.W., C.C., B.S., S.S., A.B., A.N., A.L., E.W., N.D., E.T.H., G.B.H., M.J.L., E.W., B.W., L.P., M.H.S., S.Y., A.A. and S.L. performed experiments involving the murine models. F.L., F.G., T.S., J.P., M.Z. and R.C. processed and analyzed immunostaining. G.L., B.A.K., Z.L., and F.C. provided patient samples and clinical data. B.A.K., E.S.L., T.O., and D.F. performed analyses of patient data. A.A., L.W., S.T., K.J.L., D.N., F.C., G.G., A.R., S.C., E.T.H., R.C., S.S., and C.J.W. helped to design and guide the research. D.N. supervised statistical analyses. C.G., N.R., M.K. and C.J.W. wrote the manuscript. All authors discussed and interpreted results.
Corresponding author
Ethics declarations
Competing interests
C.J.W. holds equity in BioNtech, Inc., receives research funding from Pharmacyclics, and is a scientific advisory board member of Repertoire, Adventris, Aethon Therapeutics and Nature’s Toolbox, Inc. G.G. receives research funds from IBM and Pharmacyclics and is an inventor of several bioinformatics-related patents, including those related to MuTect and ABSOLUTE. All other authors do not have any relevant conflict of interest.
Peer review
Peer review information
Nature Communications thanks Rosa Lapalombella and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gutierrez, C., Kwok, M., Ruthen, N. et al. Mutant ribosomal protein RPS15 drives B cell malignancy through oxidative stress and genomic instability. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70655-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70655-1