Abstract
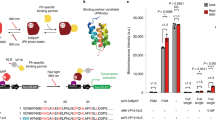
Photosensory protein domains, derived from nature, are foundational for optogenetic protein engineering. Tailoring their properties enables their full exploitation for optogenetic regulation in basic research and applied bioengineering applications. Here, we present a simple, yet powerful strategy based on random mutagenesis coupled to high-throughput screening that allowed altering the most fundamental properties of the widely used nMag/pMag photodimerization system: its light sensitivity and activation. Variants were characterized in vivo in bacteria by flow cytometry and during the entire growth curve by spectrofluorometry. We identify mutations that either increase or decrease the light sensitivity at sub-saturating light intensities, while also improving the light activation and dark-to-light fold change. Notably, light sensitivity and activation levels could be changed independently. In addition, we demonstrated that the shapes of the dose-response curves can be finely tuned. This broadens the applicability of the Magnets photosensors for optogenetic regulation strategies.
Similar content being viewed by others
Data availability
Source data are provided with this paper and are available at Zenodo [https://doi.org/10.5281/zenodo.18815874]. Previously published protein structures are available at the PDB under accession code PDB 3RH8. Source data are provided with this paper.
References
Kianianmomeni, A. Optogenetics: Methods and Protocols, Methods in Molecular Biology. vol. 1408 (Springer, 2015).
Baumschlager, A. Optogenetics: Methods and Protocols, Methods in Molecular Biology. vol. 2840 (Springer, 2025).
Baumschlager, A. Engineering light-control in biology. Front. Bioeng. Biotechnol. 10, 1–15 (2022).
Shimizu-Sato, S., Huq, E., Tepperman, J. M. & Quail, P. H. A light-switchable gene promoter system. Nat. Biotechnol. 20, 1041–1044 (2002).
Baumschlager, A. & Khammash, M. Synthetic biological approaches for optogenetics and tools for transcriptional light-control in bacteria. Adv. Biol. 5, 2000256 (2021).
Hughes, R. M. A compendium of chemical and genetic approaches to light-regulated gene transcription. Crit. Rev. Biochem. Mol. Biol. 53, 453–474 (2018).
Santos-Moreno, J. & Schaerli, Y. Using synthetic biology to engineer spatial patterns. Adv. Biosyst. 3, 1–15 (2019).
Baumschlager, A., Aoki, S. K. & Khammash, M. Dynamic blue light-inducible T7 RNA polymerases (Opto-T7RNAPs) for precise spatiotemporal gene expression control. ACS Synth. Biol. 6, 2157–2167 (2017).
Goglia, A. G. & Toettcher, J. E. A bright future: optogenetics to dissect the spatiotemporal control of cell behavior. Curr. Opin. Chem. Biol. 48, 106–113 (2019).
Johnson, H. E. & Toettcher, J. E. Illuminating developmental biology with cellular optogenetics. Curr. Opin. Biotechnol. 52, 42–48 (2018).
Johnson, H. E. & Toettcher, J. E. Signaling dynamics control cell fate in the early Drosophila embryo. Dev. Cell 48, 361–370 (2019).
Stanton, B. Z., Chory, E. J. & Crabtree, G. R. Chemically induced proximity in biology and medicine. Science 359, eaao5902 (2018).
Ullrich, A. & Schlessinger, J. Signal transduction by receptors with tyrosine kinase activity. Cell 61, 203–212 (1990).
Grusch, M. et al. Spatio-temporally precise activation of engineered receptor tyrosine kinases by light. EMBO J. 33, 1713–1726 (2014).
Inglés-Prieto, Á et al. Light-assisted small-molecule screening against protein kinases. Nat. Chem. Biol. 11, 952–954 (2015).
Zimmerman, S. P. et al. Tuning the binding affinities and reversion kinetics of a light inducible dimer allows control of transmembrane protein localization. Biochemistry 55, 5264–5271 (2016).
Sako, K. et al. Optogenetic control of nodal signaling reveals a temporal pattern of nodal signaling regulating cell fate specification during gastrulation. Cell Rep. 16, 866–877 (2016).
Chang, K. Y. et al. Light-inducible receptor tyrosine kinases that regulate neurotrophin signalling. Nat. Commun. 5, 4057 (2014).
Wend, S. et al. Optogenetic control of protein kinase activity in mammalian cells. ACS Synth. Biol. 3, 280–285 (2014).
Nihongaki, Y., Suzuki, H., Kawano, F. & Sato, M. Genetically engineered photoinducible homodimerization system with improved dimer-forming efficiency. ACS Chem. Biol. 9, 617–621 (2014).
Nihongaki, Y., Kawano, F., Nakajima, T. & Sato, M. Photoactivatable CRISPR-Cas9 for optogenetic genome editing. Nat. Biotechnol. 33, 755–760 (2015).
Schindler, S. E. et al. Photo-activatable Cre recombinase regulates gene expression in vivo. Sci. Rep. 5, 13627 (2015).
Tucker, C. L., Vrana, J. D. & Kennedy, M. J. Tools for controlling protein interactions using light. Curr. Protoc. Cell Biol. 64, 17.16.1–17.16.20 (2014).
Müller, K. & Weber, W. Optogenetic tools for mammalian systems. Mol. Biosyst. 9, 596–608 (2013).
Müller, K., Engesser, R., Timmer, J., Zurbriggen, M. D. & Weber, W. Orthogonal Optogenetic Triple-Gene Control in Mammalian Cells. ACS Synth. Biol. 3, 796–801 (2014).
Kawano, F., Suzuki, H., Furuya, A. & Sato, M. Engineered pairs of distinct photoswitches for optogenetic control of cellular proteins. Nat. Commun. 6, 6256 (2015).
Zoltowski, B. D. et al. Conformational switching in the fungal light sensor vivid. Science 316, 1054–1057 (2007).
Nihongaki, Y. et al. CRISPR-Cas9-based photoactivatable transcription systems to induce neuronal differentiation. Nat. Methods 14, 963–966 (2017).
Nihongaki, Y., Otabe, T., Ueda, Y. & Sato, M. A split CRISPR–Cpf1 platform for inducible genome editing and gene activation. Nat. Chem. Biol. 15, 882–888 (2019).
Kawano, F., Okazaki, R., Yazawa, M. & Sato, M. A photoactivatable Cre-loxP recombination system for optogenetic genome engineering. Nat. Chem. Biol. 12, 1059–1064 (2016).
Sheets, M. B., Wong, W. W. & Dunlop, M. J. Light-inducible recombinases for bacterial optogenetics. ACS Synth. Biol. 9, 227–235 (2020).
Benedetti, L. et al. Light-activated protein interaction with high spatial subcellular confinement. Proc. Natl. Acad. Sci. USA 115, E2238–E2245 (2018).
Chen, F. & Wegner, S. V. Blue light switchable bacterial adhesion as a key step toward the design of biofilms. ACS Synth. Biol. 6, 2170–2174 (2017).
Wang, X., Chen, X. & Yang, Y. Spatiotemporal control of gene expression by a light-switchable transgene system. Nat. Methods 9, 266–269 (2012).
Raghavan, G., Hidaka, K., Sugiyama, H. & Endo, M. Direct observation and analysis of the dynamics of the photoresponsive transcription factor GAL4. Angew. Chem. - Int. Ed. 58, 7626–7630 (2019).
Guinn, M. T. & Balázsi, G. Noise-reducing optogenetic negative-feedback gene circuits in human cells. Nucleic Acids Res. 47, 7703–7714 (2019).
Yao, S. et al. RecV recombinase system for in vivo targeted optogenomic modifications of single cells or cell populations. Nat. Methods 17, 422–429 (2020).
Li, X. et al. A single-component light sensor system allows highly tunable and direct activation of gene expression in bacterial cells. Nucleic Acids Res. 48, e33 (2020).
Romano, E. et al. Engineering AraC to make it responsive to light instead of arabinose. Nat. Chem. Biol. 17, 817–827 (2021).
Crosson, S. & Moffat, K. Structure of a flavin-binding plant photoreceptor domain: Insights into light-mediated signal transduction. Proc. Natl. Acad. Sci. USA 98, 2995–3000 (2001).
Vaidya, A. T., Chen, C.-H. H., Dunlap, J. C., Loros, J. J. & Crane, B. R. Structure of a light-activated LOV protein dimer that regulates transcription. Sci. Signal 4, 1–8 (2011).
Zoltowski, B. D., Vaccaro, B. & Crane, B. R. Mechanism-based tuning of a LOV domain photoreceptor. Nat. Chem. Biol. 5, 827–834 (2009).
Crosson, S., Rajagopal, S. & Moffat, K. The LOV domain family: Photoresponsive signaling modules coupled to diverse output domains. Biochemistry 42, 2–10 (2003).
Conrad, K. S., Manahan, C. C. & Crane, B. R. Photochemistry of flavoprotein light sensors. Nat. Chem. Biol. 10, 801–809 (2014).
Benedetti, L. et al. Optimized vivid-derived magnets photodimerizers for subcellular optogenetics in mammalian cells. Elife 9, 1–49 (2020).
Hockberger, P. E. et al. Activation of flavin-containing oxidases underlies light-induced production of H2O2 in mammalian cells. Proc. Natl. Acad. Sci. USA 96, 6255–6260 (1999).
Reshetnikov, V. V., Smolskaya, S. V., Feoktistova, S. G. & Verkhusha, V. V. Optogenetic approaches in biotechnology and biomaterials. Trends Biotechnol. 40, 858–874 (2022).
Salis, H. M., Mirsky, E. A. & Voigt, C. A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
Cahn, J. K. B., Baumschlager, A., Brinkmann-Chen, S. & Arnold, F. H. Mutations in adenine-binding pockets enhance catalytic properties of NAD(P)H-dependent enzymes. Protein Eng. Des. Selection 29, 31–38 (2015).
Ward, J. H. Hierarchical grouping to optimize an objective function. J. Am. Stat. Assoc. 58, 236–244 (1963).
Kille, S. et al. Reducing codon redundancy and screening effort of combinatorial protein libraries created by saturation mutagenesis. ACS Synth. Biol. 2, 83–92 (2013).
Tang, L. et al. Construction of ‘small-intelligent’ focused mutagenesis libraries using well-designed combinatorial degenerate primers. Biotechniques 52, 149–158 (2012).
Rousseeuw, P. J. Silhouettes: a graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 20, 53–65 (1987).
Hunt, S. M., Elvin, M., Crosthwaite, S. K. & Heintzen, C. The PAS/LOV protein VIVID controls temperature compensation of circadian clock phase and development in Neurospora crassa. Genes Dev. 21, 1964–1974 (2007).
Yee, E. F., Chandrasekaran, S., Lin, C. & Crane, B. R. Physical Methods for Studying Flavoprotein Photoreceptors. Methods in Enzymology vol. 620 (Elsevier, 2019).
Zhou, H., Dong, Z., Verkhivker, G., Zoltowski, B. D. & Tao, P. Allosteric mechanism of the circadian protein vivid resolved through Markov state model and machine learning analysis. PLoS Comput. Biol. 15, 1–28 (2019).
Dasgupta, A. et al. Biological significance of photoreceptor photocycle length: vivid photocycle governs the dynamic VIVID-white collar complex pool mediating photo-adaptation and response to changes in light intensity. PLoS Genet. 11, 1–23 (2015).
Dionisi, S., Piera, K., Baumschlager, A. & Khammash, M. Implementation of a novel optogenetic tool in mammalian cells based on a split T7 RNA polymerase. ACS Synth. Biol. 11, 2650–2661 (2022).
Chung, C. T., Niemela, S. L. & Miller, R. H. One-step preparation of competent Escherichia coli: Transformation and storage of bacterial cells in the same solution (recombinant DNA). Pnas 86, 2172–2175 (1989).
Sambrook, J. Molecular Cloning: A Laboratory Manual. (Cold Spring Harbor Laboratory Press, 2001).
Cline, J. PCR fidelity of PFU DNA polymerase and other thermostable DNA polymerases. Nucleic Acids Res. 24, 3546–3551 (1996).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Maechler, M., Rousseeuw, P., Struyf, A., Hubert, M. & Hornik, K. cluster: cluster analysis basics and extensions. R package version 2.0.5. https://cran.r-project.org/web/packages/cluster/index.html (2016).
Kassambara, A. & Mundt, F. factoextra: extract and visualize the results of multivariate data analyses. R package version 1.0.7. https://cran.r-project.org/web/packages/factoextra/index.html (2020).
Galili, T. Dendextend: an R package for visualizing, adjusting and comparing trees of hierarchical clustering. Bioinformatics 31, 3718–3720 (2015).
Wickham, H. Ggplot2: Elegant Graphics for Data Analysis. (Springer, 2016).
Acknowledgements
We thank Dr. Tsvetan Kardashliev for helpful discussions and Dr. Luzius Pestalozzi for the testing and the supply of the polymerase and buffer used for error-prone PCR. We further thank Dr. Stephanie Aoki for helpful discussions. We thank the Single Cell and Lab Automation Facility of the DBSSE, ETH Zurich, in particular Dr. Gregor Schmidt, Dr. Aleksandra Gumienny, and Dr. Mariangela Di Tacchio for their excellent support throughout the project. This article is dedicated to the memory of Josep (Pepe) Casadesús. We acknowledge funding from FET-Open research and innovation actions grant under the European Union’s Horizon 2020 research and innovation program (CyGenTiG; grant agreement 801041) to M.K. D.C. was a recipient of an EMBO Short-Term Fellowship (Grant number 8903).
Funding
Open access funding provided by Swiss Federal Institute of Technology Zurich.
Author information
Authors and Affiliations
Contributions
A.B. conceived, planned, and coordinated the project and wrote the manuscript with contributions from all authors. A.B. and Y.W. generated the libraries and performed the FACS. A.B., Y.W., and D.C. designed and performed bacterial experiments and analyzed the corresponding data. D.C. performed the PC analysis and hierarchical clustering. S.D. performed experiments in mammalian cells and analyzed the corresponding data. M.K. supervised the project and provided funding.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Baumschlager, A., Weber, Y., Cánovas, D. et al. Enhancing the performance of Magnets photosensors. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70695-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70695-7