Abstract
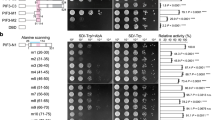
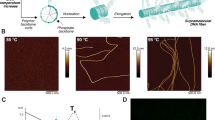
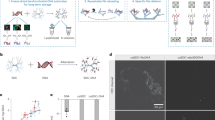
Plants tailor their architecture to warm temperatures through the central transcription factor PHYTOCHROME-INTERACTING FACTOR 4 (PIF4). Here, we dissect how structured and disordered regions of PIF4 contribute to its function in thermomorphogenesis. A long N-terminal intrinsically disordered region (IDR) enables PIF4 to form low-mobility condensates. Within this IDR, we identify an acidic transactivation domain (TAD) and an extended basic segment that carries a nuclear-localization signal and the canonical basic motif of the basic helix-loop-helix (bHLH) domain. The basic segment is both necessary and sufficient to drive PIF4 condensate formation, while the TAD tunes condensate properties. Strikingly, alanine substitutions that abolish TAD-mediated transactivation, disrupt DNA binding, or greatly reduce phase-separation propensity have no significant effect on thermomorphogenetic hypocotyl elongation. By contrast, substituting twelve basic residues within the basic segment, which disrupts both DNA binding and HLH-mediated oligomerization, abolishes thermo-induced hypocotyl growth. These findings suggest that PIF4’s oligomerization competence contributes significantly to thermomorphogenesis by enabling partner recruitment, allowing DNA-binding and transactivation functions to be supplied in trans.
Similar content being viewed by others
Data availability
The original contributions presented in the study are included in the article/Supplementary material; further inquiries can be directed to the corresponding author. Source data are provided with this paper.
References
Jagadish, S. V. K., Way, D. A. & Sharkey, T. D. Plant heat stress: concepts directing future research. Plant Cell Environ. 44, 1992–2005 (2021).
Ding, Y. & Yang, S. Surviving and thriving: how plants perceive and respond to temperature stress. Dev. Cell 57, 947–958 (2022).
Ludwig, W., Hayes, S., Trenner, J., Delker, C. & Quint, M. On the evolution of plant thermomorphogenesis. J. Exp. Bot. 72, 7345–7358 (2021).
Casal, J. J. & Balasubramanian, S. Thermomorphogenesis. Annu. Rev. Plant Biol. 70, 321–346 (2019).
Delker, C., Quint, M. & Wigge, P. A. Recent advances in understanding thermomorphogenesis signaling. Curr. Opin. Plant Biol. 68, 102231 (2022).
Park, Y.-J., Kim, J. Y., Lee, J.-H., Han, S.-H. & Park, C.-M. External and internal reshaping of plant thermomorphogenesis. Trends Plant Sci. 26, 810–821 (2021).
Koini, M. A. et al. High temperature-mediated adaptations in plant architecture require the bHLH transcription factor PIF4. Curr. Biol. 19, 408–413 (2009).
Fiorucci, A.-S. et al. PHYTOCHROME INTERACTING FACTOR 7 is important for early responses to elevated temperature in Arabidopsis seedlings. N. Phytol. 226, 50–58 (2020).
Ding, L. et al. Two B-box domain proteins, BBX18 and BBX23, interact with ELF3 and regulate thermomorphogenesis in Arabidopsis. Cell Rep. 25, 1718–1728.e4 (2018).
Chandra Malakar, B. et al. BBX24/BBX25 antagonizes the function of thermosensor ELF3 to promote PIF4-mediated thermomorphogenesis in Arabidopsis. Plant Commun. 0, 101391 (2025).
Hwang, G. et al. Overexpression of BBX18 promotes thermomorphogenesis through the PRR5-PIF4 pathway. Front. Plant Sci. 12, 782352 (2021).
Zhou, Y. et al. TCP transcription factors associate with PHYTOCHROME INTERACTING FACTOR 4 and CRYPTOCHROME 1 to regulate thermomorphogenesis in Arabidopsis thaliana. iScience 15, 600–610 (2019).
Han, X. et al. Arabidopsis transcription factor TCP5 controls plant thermomorphogenesis by positively regulating PIF4 activity. iScience 15, 611–622 (2019).
Cortijo, S. et al. Transcriptional regulation of the ambient temperature response by H2A.Z nucleosomes and HSF1 transcription factors in arabidopsis. Mol. Plant 10, 1258–1273 (2017).
Tan, W. et al. The heat response regulators HSFA1s promote Arabidopsis thermomorphogenesis via stabilizing PIF4 during the day. Sci. Adv. 9, eadh1738 (2023).
Li, B. et al. Heat shock factor A1s are required for phytochrome-interacting factor 4-mediated thermomorphogenesis in Arabidopsis. J. Integr. Plant Biol. 66, 20–35 (2024).
Ibañez, C. et al. Brassinosteroids dominate hormonal regulation of plant thermomorphogenesis via BZR1. Curr. Biol. 28, 303–310.e3 (2018).
Qiu, Y. Regulation of PIF4-mediated thermosensory growth. Plant Sci. 297, 110541 (2020).
Fan, D. et al. A multisensor high-temperature signaling framework for triggering daytime thermomorphogenesis in Arabidopsis. Nat. Commun. 16, 5197 (2025).
Raschke, A. et al. Natural variants of ELF3 affect thermomorphogenesis by transcriptionally modulating PIF4-dependent auxin response genes. BMC Plant Biol. 15, 197 (2015).
Tian, S. et al. The evening complex component ELF3 recruits H3K4me3 demethylases to repress PHYTOCHROME INTERACTING FACTOR4 and 5 in Arabidopsis. Plant Cell 37, koaf014 (2025).
Box, M. S. et al. ELF3 controls thermoresponsive growth in Arabidopsis. Curr. Biol. 25, 194–199 (2015).
Jung, J.-H. et al. Phytochromes function as thermosensors in Arabidopsis. Science 354, 886–889 (2016).
Jung, J.-H. et al. A prion-like domain in ELF3 functions as a thermosensor in Arabidopsis. Nature 585, 256–260 (2020).
Li, Z.-Y. et al. Functions of phytochrome interacting factors (PIFs) in adapting plants to biotic and abiotic stresses. Int. J. Mol. Sci. 25, 2198 (2024).
Paik, I., Kathare, P. K., Kim, J.-I. & Huq, E. Expanding roles of PIFs in signal integration from multiple processes. Mol. Plant 10, 1035–1046 (2017).
Cai, X. & Huq, E. Shining light on plant growth: recent insights into phytochrome interacting factors. J. Exp. Bot. 76, 646–663 (2025).
Sakuraba, Y. et al. Phytochrome-interacting transcription factors PIF4 and PIF5 induce leaf senescence in Arabidopsis. Nat. Commun. 5, 4636 (2014).
Leivar, P. et al. Multiple phytochrome-interacting bHLH transcription factors repress premature seedling photomorphogenesis in darkness. Curr. Biol. 18, 1815–1823 (2008).
Shin, J. et al. Phytochromes promote seedling light responses by inhibiting four negatively-acting phytochrome-interacting factors. Proc. Natl. Acad. Sci. USA 106, 7660–7665 (2009).
Lee, C.-M. & Thomashow, M. F. Photoperiodic regulation of the C-repeat binding factor (CBF) cold acclimation pathway and freezing tolerance in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 109, 15054–15059 (2012).
Li, N., Bo, C., Zhang, Y. & Wang, L. PHYTOCHROME INTERACTING FACTORS PIF4 and PIF5 promote heat stress induced leaf senescence in Arabidopsis. J. Exp. Bot. 72, 4577–4589 (2021).
Sakuraba, Y., Bülbül, S., Piao, W., Choi, G. & Paek, N.-C. Arabidopsis EARLY FLOWERING3 increases salt tolerance by suppressing salt stress response pathways. Plant J. 92, 1106–1120 (2017).
Wang, Z., Perez, V. & Hua, J. Guard cell activity of PIF4 represses disease resistance in Arabidopsis. Plant Cell Environ. 48, 1468–1478 (2025).
Zhang, X. et al. Near-infrared light and PIF4 promote plant antiviral defense by enhancing RNA interference. Plant Commun. 5, 100644 (2024).
Liu, Z. et al. FLS2-RBOHD-PIF4 module regulates plant response to drought and salt stress. Int. J. Mol. Sci. 23, 1080 (2022).
Khanna, R. et al. A novel molecular recognition motif necessary for targeting photoactivated phytochrome signaling to specific basic helix-loop-helix transcription factors. Plant Cell 16, 3033–3044 (2004).
Lambert, S. A. et al. The human transcription factors. Cell 172, 650–665 (2018).
Strader, L., Weijers, D. & Wagner, D. Plant transcription factors—being in the right place with the right company. Curr. Opin. Plant Biol. 65, 102136 (2022).
Blanc-Mathieu, R., Dumas, R., Turchi, L., Lucas, J. & Parcy, F. Plant-TFClass: a structural classification for plant transcription factors. Trends Plant Sci. 29, 40–51 (2024).
Soto, L. F. et al. Compendium of human transcription factor effector domains. Mol. Cell 82, 514–526 (2022).
Morffy, N. et al. Identification of plant transcriptional activation domains. Nature 632, 166–173 (2024).
Jolma, A. et al. DNA-binding specificities of human transcription factors. Cell 152, 327–339 (2013).
Már, M., Nitsenko, K. & Heidarsson, P. O. Multifunctional intrinsically disordered regions in transcription factors. Chemistry 29, e202203369 (2023).
Miao, J. & Chong, S. Roles of intrinsically disordered protein regions in transcriptional regulation and genome organization. Curr. Opin. Genet. Dev. 90, 102285 (2025).
Boija, A. et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell 175, 1842–1855.e16 (2018).
Funnell, A. P. W. & Crossley, M. Homo- and heterodimerization in transcriptional regulation. Adv. Exp. Med. Biol. 747, 105–121 (2012).
Frank, F., Liu, X. & Ortlund, E. A. Glucocorticoid receptor condensates link DNA-dependent receptor dimerization and transcriptional transactivation. Proc. Natl. Acad. Sci. USA 118, e2024685118 (2021).
Mignon, K. et al. DNA-binding and dimerization of the SOG1 NAC domain are functionally linked with its ability to undergo liquid-liquid phase separation. Nucleic Acids Res. 54, gkaf1522 (2026).
Toledo-Ortiz, G., Huq, E. & Quail, P. H. The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell 15, 1749–1770 (2003).
Vayssières, A., Huettel, B., Franzen, R., Richter, R. & Chai, J. PIF4 enhances DNA binding of CDF2 to co-regulate target gene expression and promote Arabidopsis hypocotyl cell elongation. Nat. Plants 8, 1082–1093 (2022).
Bu, Q., Castillon, A., Chen, F., Zhu, L. & Huq, E. Dimerization and blue light regulation of PIF1 interacting bHLH proteins in Arabidopsis. Plant Mol. Biol. 77, 501–511 (2011).
Sun, J., Qi, L., Li, Y., Chu, J. & Li, C. PIF4-mediated activation of YUCCA8 expression integrates temperature into the auxin pathway in regulating Arabidopsis hypocotyl growth. PLoS Genet. 8, e1002594 (2012).
Zhu, J.-Y., Oh, E., Wang, T. & Wang, Z.-Y. TOC1–PIF4 interaction mediates the circadian gating of thermoresponsive growth in Arabidopsis. Nat. Commun. 7, 13692 (2016).
Bajracharya, A. et al. PHYTOCHROME-INTERACTING FACTOR 4/HEMERA-mediated thermosensory growth requires the Mediator subunit MED14. Plant Physiol. 190, 2706–2721 (2022).
Shapulatov, U. et al. The Mediator complex subunit MED25 interacts with HDA9 and PIF4 to regulate thermomorphogenesis. Plant Physiol. 192, 582–600 (2023).
Yoo, C. Y. et al. Direct photoresponsive inhibition of a p53-like transcription activation domain in PIF3 by Arabidopsis phytochrome B. Nat. Commun. 12, 1–16 (2021).
Huai, J. et al. JASMONATE ZIM-domain protein 3 regulates photomorphogenesis and thermomorphogenesis through inhibiting PIF4 in Arabidopsis. Plant Physiol. 195, 2274–2288 (2024).
Luo, F. et al. A Phytochrome B-PIF4-MYC2/MYC4 module inhibits secondary cell wall thickening in response to shaded light. Plant Commun. 3, 100416 (2022).
Peng, M. et al. Activation and memory of the heat shock response is mediated by prion-like domains of sensory HSFs in Arabidopsis. Mol. Plant 18, 457–467 (2025).
Xie, Y. et al. Phytochrome B inhibits the activity of phytochrome-interacting factor 7 involving phase separation. Cell Rep. 42, 113562 (2023).
Erdős, G., Pajkos, M. & Dosztányi, Z. IUPred3: prediction of protein disorder enhanced with unambiguous experimental annotation and visualization of evolutionary conservation. Nucleic Acids Res. 49, W297–W303 (2021).
Chong, S. et al. Imaging dynamic and selective low-complexity domain interactions that control gene transcription. Science 361, eaar2555 (2018).
Trojanowski, J. et al. Transcription activation is enhanced by multivalent interactions independent of phase separation. Mol. Cell 82, 1878–1893.e10 (2022).
Dalton, J. C., Bätz, U., Liu, J., Curie, G. L. & Quail, P. H. A modified reverse one-hybrid screen identifies transcriptional activation domains in PHYTOCHROME-INTERACTING FACTOR 3. Front. Plant Sci. 7, 881 (2016).
Hope, I. A. & Struhl, K. Functional dissection of a eukaryotic transcriptional activator protein, GCN4 of yeast. Cell 46, 885–894 (1986).
Hope, I. A., Mahadevan, S. & Struhl, K. Structural and functional characterization of the short acidic transcriptional activation region of yeast GCN4 protein. Nature 333, 635–640 (1988).
Staller, M. V. et al. Directed mutational scanning reveals a balance between acidic and hydrophobic residues in strong human activation domains. Cell Syst. 13, 334–345.e5 (2022).
Staller, M. V. et al. A high-throughput mutational scan of an intrinsically disordered acidic transcriptional activation domain. Cell Syst. 6, 444–455.e6 (2018).
Kotha, S. R. & Staller, M. Clusters of acidic and hydrophobic residues can predict acidic transcriptional activation domains from protein sequence. Genetics 225, iyad131 (2023).
Hong, Y. et al. Hydrophobicity of arginine leads to reentrant liquid-liquid phase separation behaviors of arginine-rich proteins. Nat. Commun. 13, 7326 (2022).
Vernon, R. M. et al. Pi-Pi contacts are an overlooked protein feature relevant to phase separation. Elife 7, e31486 (2018).
Voronova, A. & Baltimore, D. Mutations that disrupt DNA binding and dimer formation in the E47 helix-loop-helix protein map to distinct domains. Proc. Natl. Acad. Sci. USA 87, 4722–4726 (1990).
Ferré-D’Amaré, A. R., Pognonec, P., Roeder, R. G. & Burley, S. K. Structure and function of the b/HLH/Z domain of USF. EMBO J. 13, 180–189 (1994).
Al-Sady, B., Kikis, E. A., Monte, E. & Quail, P. H. Mechanistic duality of transcription factor function in phytochrome signaling. Proc. Natl. Acad. Sci. USA 105, 2232–2237 (2008).
Udupa, A., Kotha, S. R. & Staller, M. V. Commonly asked questions about transcriptional activation domains. Curr. Opin. Struct. Biol. 84, 102732 (2024).
Erijman, A. et al. A high-throughput screen for transcription activation domains reveals their sequence features and permits prediction by deep learning. Mol. Cell 78, 890–902.e6 (2020).
Sanborn, A. L. et al. Simple biochemical features underlie transcriptional activation domain diversity and dynamic, fuzzy binding to Mediator. Elife 10, e68068 (2021).
Arnold, C. D. et al. A high-throughput method to identify trans-activation domains within transcription factor sequences. EMBO J. 37, e98896 (2018).
Hummel, N. F. C. et al. The trans-regulatory landscape of gene networks in plants. Cell Syst. 14, 501–511.e4 (2023).
Smith, N. C. & Matthews, J. M. DNA-binding, multivalent interactions and phase separation in transcriptional activation. Aust. J. Chem. 76, 351–360 (2023).
Davis, R. L., Cheng, P., Lassar, A. & Weintraub, H. The MyoD DNA binding domain contains a recognition code for muscle-specific gene activation. Cell 60, 733–746 (1990).
Hao, Y., Oh, E., Choi, G., Liang, Z. & Wang, Z.-Y. Interactions between HLH and bHLH factors modulate light-regulated plant development. Mol. Plant 5, 688–697 (2012).
Roig-Villanova, I. et al. Interaction of shade avoidance and auxin responses: a role for two novel atypical bHLH proteins. EMBO J. 26, 4756–4767 (2007).
Porcher, C., Liao, E. C., Fujiwara, Y., Zon, L. I. & Orkin, S. H. Specification of hematopoietic and vascular development by the bHLH transcription factor SCL without direct DNA binding. Development 126, 4603–4615 (1999).
Amoutzias, G. D., Robertson, D. L., Van de Peer, Y. & Oliver, S. G. Choose your partners: dimerization in eukaryotic transcription factors. Trends Biochem. Sci. 33, 220–229 (2008).
Ortega, E. et al. Transcription factor dimerization activates the p300 acetyltransferase. Nature 562, 538–544 (2018).
Hinde, E. et al. Quantifying the dynamics of the oligomeric transcription factor STAT3 by pair correlation of molecular brightness. Nat. Commun. 7, 1–14 (2016).
Sayou, C. et al. A SAM oligomerization domain shapes the genomic binding landscape of the LEAFY transcription factor. Nat. Commun. 7, 11222 (2016).
Shimizu, S. et al. Multimerization of the GATA4 transcription factor regulates transcriptional activity and cardiomyocyte hypertrophic response. Int. J. Biol. Sci. 18, 1079–1095 (2022).
Minucci, S. et al. Oligomerization of RAR and AML1 transcription factors as a novel mechanism of oncogenic activation. Mol. Cell 5, 811–820 (2000).
Stenger, J. E. et al. p53 oligomerization and DNA looping are linked with transcriptional activation. EMBO J. 13, 6011–6020 (1994).
Qiu, Y., Li, M., Kim, R. J.-A., Moore, C. M. & Chen, M. Daytime temperature is sensed by phytochrome B in Arabidopsis through a transcriptional activator HEMERA. Nat. Commun. 10, 140 (2019).
Jin, H., Lin, J. & Zhu, Z. PIF4 and HOOKLESS1 impinge on common transcriptome and isoform regulation in thermomorphogenesis. Plant Commun. 1, 100034 (2020).
Huai, J. et al. SEUSS and PIF4 coordinately regulate light and temperature signaling pathways to control plant growth. Mol. Plant 11, 928–942 (2018).
Zhou, N. et al. Histone methylation readers MRG1/2 interact with PIF4 to promote thermomorphogenesis in Arabidopsis. Cell Rep. 43, 113726 (2024).
Kim, J., Lee, H., Lee, H. G. & Seo, P. J. Get closer and make hotspots: liquid–liquid phase separation in plants. EMBO Rep. 22, e51656 (2021).
Ryu, K., Park, G. & Cho, W.-K. Emerging insights into transcriptional condensates. Exp. Mol. Med. 56, 820–826 (2024).
Powers, S. K. et al. Nucleo-cytoplasmic partitioning of ARF proteins controls auxin responses in Arabidopsis thaliana. Mol. Cell 76, 177–190.e5 (2019).
Poudyal, M. et al. Intermolecular interactions underlie protein/peptide phase separation irrespective of sequence and structure at crowded milieu. Nat. Commun. 14, 6199 (2023).
Hannon, C. E. & Eisen, M. B. Intrinsic protein disorder is insufficient to drive subnuclear clustering in embryonic transcription factors. Elife 12, RP88221 (2024).
Kim, H., Lee, N., Kim, Y. & Choi, G. The phytochrome-interacting factor genes PIF1 and PIF4 are functionally diversified due to divergence of promoters and proteins. Plant Cell 36, 2778–2797 (2024).
Acknowledgements
We thank all the Qiu lab members for their critical comments and suggestions regarding the manuscript. This work was supported by the National Science Foundation (NSF) grants IOS-2200200 and IOS-2239963 to Y.Q., and the National Institutes of Health (NIH) grant R01GM087388 to M.C. The work related to the confocal microscope was supported by an Institutional Development Award (IDeA) from the National Institute of General Medical Sciences (NIGMS) of the NIH under award number P20GM130460 to the Imaging Research Core of the Glycoscience Center of Research Excellence (GlyCORE) at the University of Mississippi. We would like to thank Drs. Gregg Roman, Ruofan Cao, and John Adams in the Imaging Research Core of GlyCORE at the University of Mississippi for their assistance with using the Leica SP8 Inverted Confocal Microscope.
Author information
Authors and Affiliations
Contributions
Y.Q. initiated the project in M.C.’s laboratory; Y.Q., H.X., and A.B. conceived the original research plan; Y.Q., S.R.B., and J.X. supervised the experiments; H.X., A.B., R.O., E.E.B, A.S., A.W., J.X., and Y.Q. performed the experiments; H.X., A.B., R.O., J.X., and Y.Q. analyzed the data; M.C. provided instructions, materials, and facilities on the completion of yeast-related data in Fig. 2b and Supplementary Fig. 1; Y.Q., H.X., and A.B. wrote the article with the contributions from all authors; all authors approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ziqiang Zhu, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xiong, H., Bajracharya, A., Odari, R. et al. Oligomerization-competent PIF4 drives thermomorphogenesis through functional redundancy in transactivation and DNA binding. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70748-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70748-x