Abstract
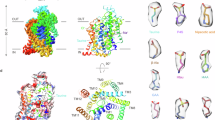
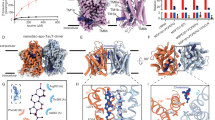
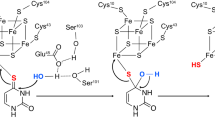
Taurine is a sulfur-containing amino acid that plays several crucial roles in the body. Its uptake is mediated by the taurine transporter (TauT). Genetic mutations and dysregulation of TauT have been linked to various neurological disorders, cardiomyopathy, childhood progressive retinal degeneration, and cancer, making TauT a promising target for therapeutic intervention in these diseases. However, the structure and mechanism of TauT remain poorly understood. In this study, we present the structures of the human taurine transporter (hTauT) under four conditions: the substrate-free state, the taurine-bound state, the β-alanine-bound state, and the cyclic inhibitor piperidine-4-sulfonate (P4S)-bound state. These structures reveal that taurine binds at the central substrate-binding site of hTauT. Notably, β-alanine and the cyclic P4S inhibitors also mimic taurine, occupying the same substrate-binding site. In the substrate-free and P4S-bound forms, hTauT also adopt an inward-open conformation, where transmembrane helix TM1a bends toward the membrane, facilitating the opening of the intracellular gate for ion and substrate release. These structural insights enhance our understanding of the mechanisms underlying substrate and ion recognition and transport in hTauT, paving the way for the future development of taurine transporter substrate analogues or selective inhibitors.
Similar content being viewed by others
Data availability
Cryo-EM maps of hTauT generated in this study have been deposited in the Electron Microscopy Data Bank under the accession codes: EMD-62738 (the inward-open conformation of the substrate-free state), EMD-62739 (the inward-occluded conformation of the substrate-free state), EMD-62740 (the taurine-bound state), EMD-62741 (the β-alanine-bound state), EMD-62742 (the inward-open conformation of the P4S-bound state), and EMD-62743 (the inward-occluded conformation of the P4S-bound state), respectively. Atomic models of hTauT generated in this study have been deposited in the Protein Data Bank under the accession codes: 9L19 (the inward-open conformation of the substrate-free state), 9L1A (the inward-occluded conformation of the substrate-free state), 9L1B (the taurine-bound state), 9L1C (the β-alanine-bound state), 9L1D (the inward-open conformation of the P4S-bound state), and 9L1E (the inward-occluded conformation of the P4S-bound state), respectively. The entries used in this study are available in the Protein Data Bank under accession code: 9EO4, 7Y7W, 4US3, 7Y7V, 9CP5, 9J8B, 3TT3, 8HFF, 8WFI, 8ZPB, and 8WFL. Source data are provided with this paper.
References
Schaffer, S. & Kim, H. W. Effects and mechanisms of taurine as a therapeutic agent. Biomol. Ther. 26, 225–241 (2018).
Hayes, K. C., Carey, R. E. & Schmidt, S. Y. Retinal degeneration associated with taurine deficiency in the cat. Science 188, 949–951 (1975).
Qaradakhi, T. et al. The anti-inflammatory effect of taurine on cardiovascular disease. Nutrients 12, 2847 (2020).
Wharton, B. A., Morley, R., Isaacs, E. B., Cole, T. J. & Lucas, A. Low plasma taurine and later neurodevelopment. Arch. Dis. Child. Fetal Neonatal Ed. 89, F497–498 (2004).
Marcinkiewicz, J. & Kontny, E. Taurine and inflammatory diseases. Amino Acids 46, 7–20 (2014).
Baliou, S. et al. Significance of taurine transporter (TauT) in homeostasis and its layers of regulation (Review). Mol. Med. Rep. 22, 2163–2173 (2020).
Warskulat, U. et al. Phenotype of the taurine transporter knockout mouse. Methods Enzymol. 428, 439–458 (2007).
Ito, T. et al. Cardiac and skeletal muscle abnormality in taurine transporter-knockout mice. J. Biomed. Sci. 17, S20 (2010).
Ansar, M. et al. Taurine treatment of retinal degeneration and cardiomyopathy in a consanguineous family with SLC6A6 taurine transporter deficiency. Hum. Mol. Genet. 29, 618–623 (2020).
Xia, Y. F. et al. miR-3156-3p is downregulated in HPV-positive cervical cancer and performs as a tumor-suppressive miRNA. Virol. J. 14, 20 (2017).
Yasunaga, M. & Matsumura, Y. Role of SLC6A6 in promoting the survival and multidrug resistance of colorectal cancer. Sci. Rep. 4, 4852 (2014).
Cao, T. et al. Cancer SLC6A6-mediated taurine uptake transactivates immune checkpoint genes and induces exhaustion in CD8+ T cells. Cell 187, 2288–2304.e2227 (2024).
Stary, D. & Bajda, M. Structural studies of the taurine transporter: a potential biological target from the GABA transporter subfamily in cancer therapy. Int. J. Mol. Sci. 25, 7339 (2024).
Han, X. Targeting taurine transporter (TauT) for cancer immunotherapy of p53 mutation mediated cancers - molecular basis and preclinical implication. Adv. Exp. Med. Biol. 1155, 543–553 (2019).
Pramod, A. B., Foster, J., Carvelli, L. & Henry, L. K. SLC6 transporters: structure, function, regulation, disease association and therapeutics. Mol. Asp. Med. 34, 197–219 (2013).
Terrill, J. R., Pinniger, G. J., Graves, J. A., Grounds, M. D. & Arthur, P. G. Increasing taurine intake and taurine synthesis improves skeletal muscle function in the mdx mouse model for Duchenne muscular dystrophy. J. Physiol. 594, 3095–3110 (2016).
Jangra, A. et al. Emergence of taurine as a therapeutic agent for neurological disorders. Neural Regen. Res. 19, 62–68 (2024).
Jakaria, M. et al. Taurine and its analogs in neurological disorders: Focus on therapeutic potential and molecular mechanisms. Redox Biol. 24, 101223 (2019).
Miyamoto, T. A. et al. Taurine-mediated cardioprotection is greater when administered upon reperfusion than prior to ischemia. Adv. Exp. Med. Biol. 643, 27–36 (2009).
Tu, S. et al. Effect of taurine on the proliferation and apoptosis of human hepatocellular carcinoma HepG2 cells. Exp. Ther. Med. 10, 193–200 (2015).
Froger, N. et al. Taurine: The comeback of a neutraceutical in the prevention of retinal degenerations. Prog. Retinal Eye Res. 41, 44–63 (2014).
Larsson, O. M., Griffiths, R., Allen, I. C. & Schousboe, A. Mutual inhibition kinetic analysis of gamma-aminobutyric acid, taurine, and beta-alanine high-affinity transport into neurons and astrocytes: evidence for similarity between the taurine and beta-alanine carriers in both cell types. J. Neurochem. 47, 426–432 (1986).
Richter, M., Moroniak, S. J. & Michel, H. Identification of competitive inhibitors of the human taurine transporter TauT in a human kidney cell line. Pharmacol. Rep. 71, 121–129 (2019).
Yahara, T., Tachikawa, M., Akanuma, S. -i, Kubo, Y. & Hosoya, K. -i Amino acid residues involved in the substrate specificity of TauT/SLC6A6 for taurine and γ-aminobutyric acid. Biol. Pharm. Bull. 37, 817–825 (2014).
Hattori, M., Hibbs, R. E. & Gouaux, E. A fluorescence-detection size-exclusion chromatography-based thermostability assay for membrane protein precrystallization screening. Structure 20, 1293–1299 (2012).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2585 (2014).
Yamashita, A., Singh, S. K., Kawate, T., Jin, Y. & Gouaux, E. Crystal structure of a bacterial homologue of Na+/Cl-dependent neurotransmitter transporters. Nature 437, 215–223 (2005).
Motiwala, Z. et al. Structural basis of GABA reuptake inhibition. Nature 606, 820–826 (2022).
Han, X., Budreau, A. M. & Chesney, R. W. Ser-322 is a critical site for PKC regulation of the MDCK cell taurine transporter (pNCT). J. Am. Soc. Nephrol. 10, 1874–1879 (1999).
Cheng, M. H. & Bahar, I. Monoamine transporters: structure, intrinsic dynamics and allosteric regulation. Nat. Struct. Mol. Biol. 26, 545–556 (2019).
Nielsen, J. C. et al. Structure of the human dopamine transporter in complex with cocaine. Nature 632, 678–685 (2024).
Nayak, S. R. et al. Cryo-EM structure of GABA transporter 1 reveals substrate recognition and transport mechanism. Nat. Struct. Mol. Biol. 30, 1023–1032 (2023).
Ji, W. et al. Substrate binding and inhibition mechanism of norepinephrine transporter. Nature 633, 473–479 (2024).
Li, Y. et al. Dopamine reuptake and inhibitory mechanisms in human dopamine transporter. Nature 632, 686–694 (2024).
Zhu, A. et al. Molecular basis for substrate recognition and transport of human GABA transporter GAT1. Nat. Struct. Mol. Biol. 30, 1012–1022 (2023).
Malinauskaite, L. et al. A mechanism for intracellular release of Na+ by neurotransmitter/sodium symporters. Nat. Struct. Mol. Biol. 21, 1006–1012 (2014).
Anderson, C. M., Howard, A., Walters, J. R., Ganapathy, V. & Thwaites, D. T. Taurine uptake across the human intestinal brush-border membrane is via two transporters: H+-coupled PAT1 (SLC36A1) and Na+- and Cl(-)-dependent TauT (SLC6A6). J. Physiol. 587, 731–744 (2009).
Wei, Y. et al. Transport mechanism and pharmacology of the human GlyT1. Cell 187, 1719–1732.e1714 (2024).
Song, A. & Wu, X. Mechanistic insights of substrate transport and inhibitor binding revealed by high-resolution structures of human norepinephrine transporter. Cell Res. 34, 810–813 (2024).
Hu, T. et al. Transport and inhibition mechanisms of the human noradrenaline transporter. Nature 632, 930–937 (2024).
Holopainen, I. & Kontro, P. High-affinity uptake of taurine and beta-alanine in primary cultures of rat astrocytes. Neurochem. Res. 11, 207–215 (1986).
Yadav, R., Han, G. W. & Gati, C. Molecular basis of human GABA transporter 3 inhibition. Nat. Commun. 16, 3830 (2025).
Li, N. et al. Modulation of the human GlyT1 by clinical drugs and cholesterol. Nat. Commun. 16, 2412 (2025).
Krishnamurthy, H. & Gouaux, E. X-ray structures of LeuT in substrate-free outward-open and apo inward-open states. Nature 481, 469–474 (2012).
Yang, D. & Gouaux, E. Illumination of serotonin transporter mechanism and role of the allosteric site. Sci. Adv. 7, eabl3857 (2021).
Grouleff, J., Søndergaard, S., Koldsø, H. & Schiøtt, B. Properties of an inward-facing state of LeuT: conformational stability and substrate release. Biophys. J. 108, 1390–1399 (2015).
Bhatt, M. et al. Unveiling the crucial role of betaine: modulation of GABA homeostasis via SLC6A1 transporter (GAT1). Cell. Mol. Life Sci. 81, 269 (2024).
Scholze, P. et al. Transporter-mediated release: a superfusion study on human embryonic kidney cells stably expressing the human serotonin transporter. J. Pharm. Exp. Ther. 293, 870–878 (2000).
Shahsavar, A. et al. Structural insights into the inhibition of glycine reuptake. Nature 591, 677–681 (2021).
Guo, W., Wang, M. & Chen, L. A co-expression vector for baculovirus-mediated protein expression in mammalian cells. Biochem. Biophys. Res. Commun. 594, 69–73 (2022).
Song, K. et al. Structural basis for human TRPC5 channel inhibition by two distinct inhibitors. eLife 10, e63429 (2021).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Wang, N. et al. Structural basis of human monocarboxylate transporter 1 inhibition by anti-cancer drug candidates. Cell 184, 370–383.e313 (2021).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Meng, E. C. et al. UCSF ChimeraX: tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. Sect. D. Biol. Crystallogr. 66, 486–501 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. Sect. D. Struct. Biol. 74, 531–544 (2018).
DeLano, W. L. Pymol: An open-source molecular graphics tool. CCP4 Newsl. Protein Crystallogr. 40, 82–92 (2002).
Acknowledgements
We thank all of Wu Lab’s members for their kind help. The authors appreciate the AKTA purification platform provided by labs of Pingping Li and Bing Cui, and the centrifuge experimental platform provided by the Biological Analysis Center at Institute of Materia Medica of Chinese Academy of Medical Sciences. Our work was supported by the Electron Microscopy Laboratory, and Cryo-EM Platform of Peking University, and we would be grateful to Xuemei Li, Zhenxi Guo, Changdong Qin, Xiaojuan Hui, and Guopeng Wang for their help in making EM samples and taking/analyzing EM images. We also thank Shuimu BioSciences for Cryo-EM facility access and technical support during image acquisition. The work is supported by grants from the Non-profit Central Research Institute Fund of Chinese Academy of Medical Sciences (no. 2022-RC350-01 to J.-X.W.), the CAMS Innovation Fund for Medical Sciences (CIFMS) (2023-I2M-2-006 to J.-X.W. and 2021-I2M-3-001 to R.W.), and the National Natural Science Foundation of China (32522046 and 32371266 to J.-X.W.).
Author information
Authors and Affiliations
Contributions
J.-X.W. initiated the project. Y.Q., Y. Zhang, D.W., and X.C. screened the expression constructs. Y.Q. and Y. Zhang purified protein, prepared the cryo-EM sample, and screened the cryo-EM sample. Y.Q., Y. Zhang, and W. J. collected the cryo-EM data. Y.Q. and J.-X.W. processed the cryo-EM data. Y.Q., Y. Zhou, and J.-X.W. built and refined the atomic model. Y.Q., Y. Zhang, D.W., and J.L. performed the uptake assay. L.L. and R.W. contributed to chemicals. All authors contributed to the manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Aravind Penmatsa and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Qi, Y., Zhang, Y., Wang, D. et al. Structural mechanism of substrate binding and inhibition of human taurine transporter. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70772-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70772-x