Abstract
Protein complexes are fundamental to all biological processes. Public repositories have expanded to include millions of potential protein–protein interactions (PPIs) from human and diverse model organisms. Yet, large-scale structural characterization of these complexes—especially across different biological kingdoms—has lagged far behind, leaving most potential and unidentified interactions unresolved. Here, we present a comprehensive atlas of 1.1 million predicted protein–protein interaction structures generated with the AlphaFold2-based ColabFold framework. This dataset spans proteome-wide interactions from bacteria, archaea, humans, mice, plants, and human–virus pairs. Overall, we identify 181,671 high-confidence protein complex structures, especially 37,855 in the human interactome. Structural clustering revealed numerous conserved protein complex architectures shared across kingdoms, providing insights into previously uncharacterized biological functions. Supported by co-immunoprecipitation experiments, we further identify candidate viral receptors for Human mastadenovirus A and Papiine alphaherpesvirus 2. Comparative analyses integrating our complex structures with the AlphaFold monomeric structure database uncovered widespread gene fusion and fission events during evolution. Finally, we demonstrate how our dataset can enhance protein binding–surface prediction using deep learning approaches, illustrating its broad utility beyond structural modeling alone. Altogether, this atlas to our knowledge, represents one of the most extensive cross-kingdom resources and opens avenues for future discoveries in various biomedical applications.
Similar content being viewed by others
Introduction
Deep learning has transformed biological researches by enabling accurate modeling of the three-dimensional structures of proteins and their complexes. Revolutionary tools such as AlphaFold2 (AF2)1, RoseTTAFold2, and ESMFold3 have considerably enhanced our understanding of protein mechanics by predicting protein monomer structures from amino acid sequences. The extension of AlphaFold-Multimer, tailored for multimeric proteins, has significantly improved complex-structure prediction accuracy4. Additionally, combinatorial and hierarchical assembly algorithms or searching approaches with Monte Carlo tree, have shown high-confidence predictions for large protein complexes based on AF25,6. Recently, the development of a unified deep-learning framework that integrates proteins, nucleic acids, small molecules, metals, and chemical modifications, including RoseTTAFold All-Atom7 and AlphaFold38, has achieved unprecedented accuracy across the biomolecular spectrum. These technological breakthroughs are reshaping biological research, accelerating therapeutic discovery, and expanding the reach of structural modeling into nearly every aspect of molecular biology.
Proteome-scale monomer prediction has profoundly expanded our understanding of protein function, evolution, and drug design9,10,11. The AlphaFold Protein Structure Database (AFDB) now contains over 214 million structures predicted by AlphaFold2, while the ESM Metagenomic Atlas contains more than 617 million structures predicted by ESMFold3. Structural-alignment-based clustering algorithms, such as Foldseek, enable large-scale mining of these protein structure databases12,13,14. These algorithms have revealed numerous previously unidentified structures and enzymes through the clustering of a broad spectrum of protein structures14,15. Two recent studies have predicted 67,715 and more than 350,000 novel viral protein structures, revealing unprecedented architectural diversity16,17. Furthermore, AI-guided structural mining has enabled a new base-editing tool with enhanced activities and minimal off-target effects18. Collectively, these developments illustrate the growing role of large-scale computational prediction in diverse areas of biological discovery.
The assembly of both stable and transient protein complexes forms the cornerstone of virtually all biological processes. Understanding the architectures of these complexes is essential for elucidating and modifying their functional roles. Repositories of potential protein-protein interactions (PPIs) generated from high-throughput experimental techniques such as yeast two-hybrid systems or computational predictions have burgeoned, now comprising millions of entries19,20. However, while small-scale predictions and experimental studies have revealed numerous bacterial and eukaryotic heterodimers and homo-oligomers8,21,22,23, comprehensive and accurate structural characterization of complexes spanning multiple kingdoms remains limited. This gap hampers our ability to analyze interaction mechanisms, evolutionary conservation, and functional regulation across species.
Here, we develop an approach to identify potential protein-protein interacting pairs in prokaryotes using co-localization analysis and operon prediction. We compile potential protein interactions in 36 pathogenic bacteria and adjacent protein pairs in 188 prokaryotic species, identifying a total of 553,814 pairs for structural modeling, of which 108,879 are classified as high-confidence protein-protein interactions. Further, we predict 559,960 protein complexes in human, plant, mouse, and human-virus interactions. Collectively, we predict 1.1 million protein complexes, among which we identify 181,671 high-confidence protein pair structures, especially, more than 37,855 high-confidence predict protein complexes in human interactome. Building upon this extensive protein-protein interaction atlas, we conduct a series of in-depth analyses that include identifying large protein complexes, clustering these complexes across kingdoms for evolutionary analysis, coupling the structural human interactome with the human-virus interactome to assess viral interference in the context of the human protein network, and analyzing gene fusion and fission events throughout evolution. Finally, we further illustrate the applicability of the high-confidence predicted protein complex dataset for identifying protein interaction surfaces. Our results provide a unified structural complex landscape of the proteome across biological domains, revealing evolutionary patterns and mechanistic insights that could not be obtained from sequence data alone. We anticipate that our large-scale atlas of predicted protein complexes across kingdoms and host-pathogen interactions will prove valuable for further research and applications.
Results
Identification of loci-linked protein complexes in bacteria and archaea
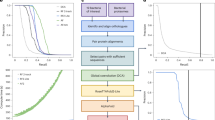
In bacteria and archaea, genes encoding proteins that participate in related biological functions are frequently co-localized on the genome, often forming operons. Such organization enables coordinated transcription and efficient regulation through a single promoter, producing polycistronic mRNA24. To identify possible physical (PPIs), we first detected operons using the Operon-mapper server25 for 36 species of widely distributed pathogenic bacteria (Supplementary Table 1) and identified 21,609 operons (gene counts >1), with an average of 3.3 genes per operon (Fig. 1a and Supplementary Fig. 1). To capture protein pairs that are not located in the same operon, we further developed a pipeline to explore co-localization of protein pairs to reflect their interdependence (Fig. 1a). Such an approach has been widely used in the discovery of CRISPR-associated proteins26,27. In this study, we systematically cataloged all pairs of proteins located within 3 kilobases (kb) of each other on the genomes of pathogenic bacteria. A comprehensive protein dataset, including information about their positions in cognate contigs, was collected. This dataset consists of 18 million contigs from bacterial genomes sourced from the National Center for Biotechnology Information (NCBI) and encompasses ~0.89 billion proteins identified in coding sequences (CDS) regions. The co-localization score was defined as the proportion of contigs where homologs of two query proteins appeared within 3 kb, normalized by the total number of contigs containing either homolog. To determine the suitable threshold, we calculated the co-localization score of proteins in a batch of biosynthetic gene clusters28. We observed a significant enrichment of protein pairs with co-localization scores exceeding 0.4. Consequently, we established a cutoff of 0.4 for subsequent analyses (Supplementary Fig. 2). Totally, we identified 164,037 potential protein pairs across 36 pathogenic bacteria using both operon analysis and co-localization mining (Fig. 1b).
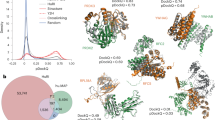
a A schematic representation of loci-linked protein pair identification through operon prediction and co-localization analysis. Operons were predicted from NCBI reference genomes, and homologs of potential protein pairs were queried against our collected NCBI Contig Protein Database. Co-localization analysis was used to identify protein pairs separated by less than 3 kb on the genome, with high co-localization score. b Distribution of protein pairs identified through the operon prediction and co-localization analysis across bacterial species. Yellow bars represent protein pairs identified exclusively through co-localization, dark purple bars indicate pairs located within operons but not identified by co-localization, and light purple bars represent overlapping pairs found by both methods. c, d Distribution of the Best Local Interaction Score (Best LIS) scores (c) and the interface predicted Template Modelling (ipTM) scores (d) for ColabFold multimer-predicted protein pairs across 36 pathogenic bacteria. e Distribution of Best LIS scores for homodimers of subunits from predicted protein pairs. c, e the dash lines denote Best LIS = 0.203 as the threshold. f Distribution of coding sequence (CDS) counts between high-confidence protein pairs. Blue represents pairs showed the CDS counts between all the predicted pairs across 36 pathogenic bacteria, while green represents those identified as the high confidence pairs. g A phylogenetic tree of representative genomes from 188 phyla, and the 36 pathogenic bacteria used in this study. Pathogens are marked with orange stars, while archaea are represented in green.
We used ColabFold v1.5.5 to generate 5 PDBs for each protein pair with up to 20 cycles (Supplementary Fig. 3)29. While preparing this manuscript and in step with the release of AlphaFold38, we also tried to compare the prediction accuracy of AlphaFold3 and other AlphaFold3-inspired models, including Boltz-230, Chai-131 and Protenix32, with ColabFold Multimer. We found that, on heterodimers, AlphaFold3 achieved the highest accuracy, with Boltz-2 performing comparably; for homodimers, multiple sequence alignment (MSA)–based methods showed similar accuracy (Supplementary Fig. 4). To distinguish true interactions from artifacts, we assessed each predicted complex using multiple established confidence metrics, including Predicted DockQ (pDockQ)33, pDockQ234, Local Interaction Score (LIS), Local Interaction Area (LIA)35, AlphaFold Multimer-derived interface predicted Template Modeling (ipTM) score, and linear models specifically designed for homodimer identification36. Previous studies demonstrated that35, when evaluating the top-ranked model among generated PDBs for each predicted PPI complex, the Best LIS metric provides superior Receiver Operating Characteristic (ROC) performance—indicating better discrimination between true positives and false positives—for heterodimer identification compared to ipTM, pDockQ, and pDockQ2. Furthermore, as this research recommended, we confirmed that the combination of Best LIS and Best LIA metrics yields the highest precision rate among evaluated scoring schemes (Supplementary Fig. 5a). Therefore, we adopted a threshold of Best LIS ≥ 0.203 and Best LIA ≥ 3432 as criteria to identify high-confidence heterodimeric PPIs for subsequent analyses. As a result, we identified a total of 22,216 high-confidence protein pairs across the 36 species of pathogenic bacteria (Fig. 1d). Also, we observed that increasing the Best LIS threshold improved precision but concurrently reduced in the recall rate. For instance, applying the Best LIS threshold of 0.6 resulted in a precision rate exceeding 90%, whereas the recall rate fell below 15% (Supplementary Fig. 5b). Therefore, we recommend selecting different thresholds tailored to the specific objectives and tolerance for false negatives in different studies.
In addition to heterodimer predictions, we also predicted the homo-oligomeric states of 76,429 proteins within the identified potential protein pairs. To evaluate the metrics for identifying true homodimers in ColabFold modeling, we constructed a benchmark dataset comprising 411 proteins from the PDB, curated in previous studies36 to remove redundancy and eliminate overlap with the AlphaFold2 training set. In the beginning, proteins resolved as monomers by X-ray crystallography were used as the negative dataset. Similarly, Best LIS showed best ROC performance when we analyzed the top-ranked model of each prediction (Supplementary Fig. 6a). Following previous research36, we also reclassified certain assemblies from dimers to monomers to correct for likely crystal-packing artifacts. This adjustment improved the overall classification performance, as evidenced by an increased AUC (Supplementary Fig. 6a and b). When we selected Best LIS with 0.203 and Best LIA with 3432 as the threshold (Supplementary Fig. 6c), consistent with the previous report21, a significant proportion of these proteins formed homodimers (Fig. 1e).
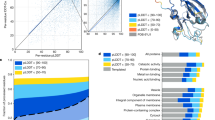
As expected, proteins encoded by adjacent CDSs exhibited the highest likelihood of forming high-confidence physical PPIs (Fig. 1f), and higher co-localization scores were positively associated with a greater prevalence of predicted high-confidence PPIs (Supplementary Fig. 7). To broaden our understanding of protein-protein interactions across the bacterial and archaeal domains, we collected representative genomes from each of the 188 prokaryotic phyla, encompassing all known archaeal or bacterial phyla, after excluding phyla related to our collected pathogens (Fig. 1g, Supplementary Fig. 8, and Supplementary Data 1).
After selecting a representative genome for each phylum, we examined the proportion of the entire proteome at the species, phylum, and domain (super-kingdom) levels that is accounted for by our selected proteins and their homologs. To this end, we performed protein clustering using MMseqs237. As expected, at the species level, whether in representative species of archaea or bacteria, the vast majority of protein clusters, apart from a few small ones, contained proteins from the target genome and collectively covered more than 90% of all proteins found in the sampled genomes of that species (Supplementary Figs. 9 and 10). When the analysis was extended to the phylum level, coverage naturally decreased but remained substantial. For example, at a cluster size threshold of ≥200 members, 84.7% of all Chlamydiota proteins were included, and more than 90% of the clusters still contained C. trachomatis proteins. Similarly, across 2,168 Cyanobacteriota genomes, applying a ≥ 1000-protein cluster threshold retained 44.4% of all proteins, and more than 90% of these clusters contained N. linckia proteins. These results suggest that numerous high-abundance, conserved proteins are shared well beyond species boundaries. At the domain level, the same trend was observed (Supplementary Fig. 11). Together, these data suggest that selecting only representative species from each phylum is sufficient to capture most evolutionarily conserved proteins, whereas additional predictions are required to resolve proteins such as strain-specific gene products that lie in the long tail of small clusters.
In total, we extracted 313,348 neighboring CDS pairs and used ColabFold to predict their corresponding protein assemblies. Notably, 47,668 predicted heterodimers were identified as high-confidence interacting pairs (Supplementary Fig. 12), underscoring the remarkable diversity of protein complexes across bacterial and archaeal kingdoms and highlighting their potential as a valuable resource for further investigation.
Beyond colocalized gene pairs, we aimed to identify interactions between proteins encoded at distant loci within prokaryotic genomes. As a proof of concept, we analyzed the Campylobacter jejuni genome and surveyed 21,676 reference genomes. Additionally, as shown in Supplementary Fig. 13, by reducing the original 200-step diffusion process to a single step, the time-efficient MSA-free Chai-1 model (MSA-free Light Chai-1 model), although lacking 3D structure generation capability, produced ipTM scores that correlated well with ColabFold Multimer predictions (r ≈ 0.68) and exhibited strong discriminatory power for PPIs (AUC = 0.783) (Supplementary Fig. 14). Therefore, all Campylobacter jejuni protein pairs that passed stringent genomic co-occurrence (score ≥ 0.5) and length filters were first evaluated using the MSA-free Light Chai-1 model, and the top-ranked candidates were subsequently modeled in 3D with ColabFold-Multimer. For example, it revealed a heterodimer between an RDD family protein and an SH3 domain–containing C40 family peptidase. Such an integrated approach could serve as a general strategy to expand the putative prokaryotic interactome (Supplementary Fig. 15).
In total, we predicted 553,814 protein pairs, including both heterodimers and homodimers, and identified 108,879 high-confidence physical protein complexes for subsequent analyses.
Reconstruction of multi-component protein complexes in prokaryotes
To de novo chart multi-component protein assemblies from our predicted binary PPIs, we built comprehensive interaction maps across 36 pathogenic bacterial species. In these networks, proteins form discrete communities corresponding to putative complexes; for example, Klebsiella pneumoniae exhibits highly cohesive clusters (Fig. 2a). A prominent and recurring feature across species is the ribosome, which forms a robust community in most organisms (Supplementary Fig. 16). Additionally, we recovered multiple large assemblies, including the ethanolamine-utilization carboxysome, type I fimbriae, the urease complex, and the recently described propanediol-utilization metabolosome38. Across the 36 pathogens, we identified 3,803 communities comprising >2 proteins, with the per-species counts ranging from 16 to 242. The largest community contained 39 subunits (Fig. 2b). Collectively, these results indicate that community detection on predicted PPI maps can recover diverse multi-component protein assemblies at proteome scale.
a Comprehensive network view of the high-confidence interaction network for representative Klebsiella pneumoniae. The names of the advanced communities are listed on both sides of the network. b Bar graph showing the number of predicted bacterial structure-linked communities associated with various pathogens. c Genetic and structural analyses of the CopRS system in Salmonella enterica. Best LIS scores were shown above the line of the genome. d Predicted structures of homo-oligomeric assemblies of virulence factors. e–j Predicted advanced structures of multi-layer complex.
Having delineated communities largely from heterodimeric edges, we next asked whether their constituent subunits also self-associate, thereby assembling into higher-order complexes. As a case study in a virulence-linked module, we examined copper homeostasis. Bacteria can deploy copper intoxication to restrict invading pathogens, and bacteria counter with copper-sensing and efflux systems39. In Salmonella enterica, we identified a CopRS two-component system in which the sensor kinase CopS was predicted to form a homodimer, while CopS and the response regulator CopR were predicted to associate as a heterodimer. By combining these interfaces, we reconstructed a quaternary CopRS assembly using CombFold6 (Fig. 2c), providing a structural hypothesis that could inform the design of targeted inhibitors.
Building on complexes inferred from heterodimeric interactions, we next examined whether homodimerization could nucleate higher-order assemblies, especially in virulence modules. The virulence factor database encompasses 27,982 potential virulence factors across various bacterial species, including a spectrum of mechanisms such as exotoxin production, adherence, and immune modulation40. We generated a comprehensive set of 26,490 homodimeric structures and identified structural symmetries using QSproteome21. For instance, we reconstructed the hexametric assembly of the Hcp family type VI secretion system effector, which displays a hollow, ring-shaped configuration (Fig. 2d). We hypothesized that in 36 pathogenic bacteria, part of the predicted homodimers could further assemble into homo-oligomers, whose subunits might also engage in interactions with other proteins, to form advanced assemblies. To test this hypothesis, we developed a computational pipeline for the comprehensive screening (Supplementary Fig. 17 and Fig. 2e–j). For example, we discovered that in Listeria monocytogenes, proteins EutL and TIGR02536 family ethanolamine utilization protein interact with each other and can each form a trimer separately. We thereby assembled the two-layered 6-subunit complex using our pipeline. Furthermore, we reconstructed several advanced complexes, such as phage baseplate, phage tail, and ethanolamine utilization complex, revealing the potential for the assembly of higher-order complexes through the integration of homologous and heterologous dimers (Fig. 2e–j).
Collectively, these analyses show that predicted structural interactomes recover a wide range of multi-component assemblies, underscoring the architectural and mechanistic breadth of prokaryotic protein complexes.
The atlas of human and model organism protein complex structures
To extend the atlas to eukaryotic systems, we compiled candidate PPIs from multiple large-scale resources, including the HI-Union Human Reference Interactome41, which have been predicted by FoldDock23,33 and experimentally supported human protein–protein interaction candidates from the STRING database19. We filtered the candidates from the STRING database using the pLDDT score of each monomer obtained from the AFDB database, retaining only those with pLDDT scores above 70 and protein lengths between 150 and 800 amino acids. The overlap between two datasets is 3231, and totally we collected 278,167 potential candidates. Subsequently, we predicted the structures by ColabFold with up to 20 cycles to improve the accuracy. We identified 37,855 high-confidence structures (Best LIS ≥ 0.203 and Best LIA ≥ 3432), representing a 12-fold increase relative to previously published AlphaFold-based predictions of human PPIs23 (Fig. 3a-c).
a, b Distribution of ipTM scores (a) or Best LIS scores (b) for protein pairs selected from human proteins. c Hierarchical clustering heat-map of the 1000 most highly connected human high-confidence protein–protein interaction nodes. Blue pixels indicate observed interactions. d Predicted complex structures of human PRKCZ and NPM1. e Immunoprecipitation and Western blot analyses of H1299 cell lysates from purified cytoplasm (CO), nucleoplasm (NP), or nucleolus (NO). f, g Immunofluorescence staining assays assessing PRKCZ and NPM1 in H1299 cells, including co-localization quantification using Pearson’s correlation coefficient from 30 cells across three independent experiments, scale bar = 25 μm. Center line indicates the mean, and error bars indicate the standard deviation (mean ± SD). h Sankey diagram showing heterodimeric PPIs across taxonomic groups and cluster types. i Structural comparison between Brucella melitensis BolA/IbaG family iron-sulfur metabolism protein and Grx4 family monothiol glutaredoxin with human glutaredoxin-3 and human bolA-like protein 2. j Structural comparison between human and Mycobacterium tuberculosis proteasome α/β heterodimers.
Since entries from the STRING database were included without applying internal confidence score (combined score) filtering, a total of 1.47 million redundant protein–protein interaction pairs were initially considered. We further compared the STRING combined scores with our predicted LIS scores. Overall, no clear linear relationship was observed between the two measures (Pearson correlation coefficient, r = 0.155), suggesting that even interactions with low STRING scores should be considered in subsequent predictions (Supplementary Fig. 18).
Compared with proteins from prokaryotic species, human proteins are generally longer, which complicates structural prediction and reduces accuracy (Supplementary Fig. 19a). Accordingly, we observed a higher proportion of low-confidence predictions among human proteins (Supplementary Fig. 19b). To further assess whether AlphaFold-based modeling can discriminate true human protein interactions, we selected the PRKCZ–NPM1 pair for detailed analysis. As shown in Fig. 3d, although both monomers contained extensive low-confidence regions and the ipTM score was low (0.366), the Best LIS and Best LIA metrics exceeded the defined thresholds, indicating a high-confidence interaction. In endogenous assays using H1299 cells, we performed co-immunoprecipitation (Co-IP) and cellular co-localization analyses, which confirmed a significant protein–protein interaction (Fig. 3d-g).
In addition, we collected 200,558 potential PPIs across multiple model organisms, including Mus musculus and Arabidopsis thaliana, leading to the identification of a total of 19,812 high-confidence protein complex structures (Supplementary Figs. 20 and 21). Comparative structural analyses across divergent kingdoms may also reveal the evolutionary trajectories that have driven the diversification of these complexes. Remarkably, several protein complexes were found to be highly conserved across the tree of life, underscoring their fundamental biological importance (Fig. 3h). For example, Human glutaredoxin 3 (Glrx3) is an essential [2Fe-2S]-binding protein that forms [2Fe-2S]-bridged complexes with human BolA2. It plays key roles in immune cell responses, embryogenesis, cancer cell proliferation, and the regulation of cardiac hypertrophy42. We found that this complex closely resembles the BolA/IbaG family iron-sulfur metabolism protein and the Grx4 family monothiol glutaredoxin in Brucella melitensis. Notably, human Glrx3 has evolved to contain three tandem-repeat domains (Fig. 3i and j). Altogether, we totally identified 57,667 high-confidence protein-protein interaction complexes. This expansion suggests a significant potential step forward in our understanding of the structural landscape of protein complexes in humans and other model organisms.
Atlas of structure-predicted human-virus protein interactions
Viruses exploit a sophisticated network of host-virus PPIs to hijack cellular processes, including endocytosis, nuclear transport, protein translation, and secretion. In response, host cells activate a complex transcriptional program mediated by PPIs, thereby triggering innate antiviral defenses, regulating viral replication, and stimulating the adaptive immune response43. High-throughput experimental and computational approaches have collectively identified a large number of PPI candidates, greatly advancing our understanding of the human-virus interactome44. However, the number of host-virus PPI complexes with experimentally resolved three-dimensional structures remains extremely limited. The comprehensive structural characterization of these PPIs, including the precise three-dimensional arrangements of the interacting proteins, will provide critical insights into the molecular mechanisms underlying viral pathogenesis and host defense strategies. Two curated databases of predicted human–virus protein–protein interactions, HVIDB45 and P-HIPSTer44, provide precise resources for predictions. Due to computational constraints (P-HIPSTer >280k interactions; HVIDB 48,643 PPIs), we retained HVIDB entries and subsampled P-HIPSTer. After removing duplicates, we totally collected 81,235 unique PPI candidates covering 3531 virus proteins and 8532 host proteins (Fig. 4a). Novelty was evaluated against the AF-Multimer training set, and 62 virus–human PPIs (at a 70% sequence identity threshold) were found to match training-set structures for both chains. Among the analyzed virus–human PPIs, 5119 (5.72%) were predicted to have Best LIS scores above 0.203 and Best LIA scores above 3432, indicating high-confidence physical interactions.
a Comprehensive interaction network depicting a wide range of protein-protein interactions between human and viral proteins. Nodes represent individual proteins, blue dots as human and red dots as virus, connected by lines showing high confidence interactions. b, c Clustered views of selected protein interactions focusing on key human proteins and viral proteins. d The visualization of diverse structural interactions between human receptors and viral proteins. Each protein complex is depicted with its respective binding partner, highlighting key interactions such as the Human immunodeficiency virus 1 envelope glycoprotein with the human T-cell surface glycoprotein CD4. Additional protein interactions from other viruses and their human targets are also shown, such as Human mastadenovirus F membrane glycoprotein E3 CR1-beta with human PD-L1. Structural quality and interaction confidence were quantified by pLDDT, ipTM, and Best LIS scores, respectively. e HEK293T cells transiently expressing HA-E3 and Flag-CD4 were subjected to co-immunoprecipitation and western blot analyses (left panel). HEK293T cells transiently expressing HA-UL44 and Flag-HLA-DRA were subjected to co-immunoprecipitation and western blot analyses (right panel). All proteins contained only the extracellular domains, with their sequences listed in Supplementary Table 2. Schematic diagram representing the dynamic protein binding competition and protein ternary complexes formation. f Schematic diagram representing the dynamic protein binding competition and protein ternary complexes formation.
In addition, we found that some human proteins could interact with distinct viral proteins, whereas certain viral proteins were capable of binding to multiple host target proteins, suggesting the potential involvement of key viral mediators and critical human target proteins (Fig. 4a). Among these interactions, the top 15 viral or human proteins are illustrated in Fig. 4b and c, where several 14-3-3 family members display broad interactions with diverse viral proteins. We specifically focused on host membrane proteins, given their interactions may mediate viral entry into cells and thus represent important targets for therapeutic intervention. CD4, a crucial receptor for Human Immunodeficiency Virus (HIV) and an important cellular biomarker, was predicted to interact with both the glycoprotein of Zaire ebolavirus and the membrane glycoprotein E3 CR1-beta of Human mastadenovirus A (Fig. 4d). These interactions exhibited similar binding sites on CD4 and we validated the interaction between the extracellular domains of CD4 and E3 from Human mastadenovirus A using co-immunoprecipitation (Co-IP) assays (Fig. 4e). Additionally, we also confirmed the interaction of human HLA-DRA and UL44 in Papiine alphaherpesvirus 2 using the same experimental approach (Fig. 4e), suggesting that our AlphaFold-based protein complex structure modeling approach could potentially identify key viral entry receptors on host cells (Supplementary Fig. 22a).
We hypothesized that viral proteins interfere with normal host protein–protein interactions by competing for binding sites or forming alternative complex assemblies. We integrated our predicted human protein complex atlas with the human–virus interactome and identified 78,191 ternary relationships with high confidence. In these relationships, viral proteins interact with human proteins that, in turn, engage additional host partners. To distinguish whether a viral protein forms a ternary complex with two host proteins or instead competes with host proteins for binding, we calculated the binding interface violations for each pairwise combination. Our analysis revealed that 57.4% human protein-protein interactions could potentially be disrupted by viral proteins, as indicated by interface violations exceeding 50% (Supplementary Fig. 22b). For example, we found that the ORF128 ankyrin repeat protein from Orf Virus can potentially interact with human SKP1 (Fig. 4f), which also binds to human FBXL20. SKP1 and FBXL20 constitute core components of the SCF (Skp1–Cul1–F-box) ubiquitin ligase complex, which regulates cell-cycle progression, DNA-damage responses, autophagy, and apoptosis46. Interestingly, the ORF128 ankyrin repeat protein targets an overlapping binding surface on FBXL20. Such competitive binding may displace FBXL20 from SKP1, thereby potentially facilitating Orf virus pathogenesis (Fig. 4f).
Collectively, we constructed a structural atlas of virus–human interactions, identifying key host factors and potential viral receptor proteins. Integration of predicted human interactomes revealed potential competitive interactions in which viral proteins may disrupt native human protein–protein interactions, underscoring the importance of the AlphaFold-derived structural atlas in elucidating viral pathogenic mechanisms. In total, we predicted 1.1 million protein complexes, covering proteome-wide interactions across bacteria, archaea, humans, mice, plants, and human–virus pairs, thereby substantially expanding structural coverage and providing a comprehensive reference dataset for diverse analyses (Supplementary Fig. 23).
Identifying protein fusion and fission events through structural alignments of protein monomers and complexes
The AlphaFold Protein Structure Database (AFDB), which contains structural predictions for over 200 million protein monomers, has garnered widespread enthusiasm for its transformative potential, not only in structural biology but across the entire field of life sciences. During evolution, proteins undergo fusion and fission events, in which open reading frames (ORFs) merge or split, thereby enabling the acquisition of new functions47. However, the discovery of protein fusion or fission events remains challenging because of the scarcity of protein complex structures and the difficulty in detecting remote sequence homology12.
We hypothesized that this evolutionary transition can be detected through comprehensive structural alignment analyses of two large datasets: the AlphaFold monomer structure database and our predicted cross-kingdom protein complex dataset, both encompassing eukaryotic and prokaryotic species. Because two subunits within a complex may share structural similarity and correspond to the same monomer position, we aligned each subunit separately to monomers in the pre-clustered AFDB50 database, which contains about 50 million protein structures. The alignment was performed using Foldseek with an overlap threshold of 30% (Fig. 5a). By comparing the predicted protein complexes from the bacterial, archaeal, human, Arabidopsis, mouse, and human–virus datasets with those in AFDB50, we identified 668,992 matches, the majority of which were found in prokaryotic species (Fig. 5b). Surprisingly, we found that thousands of AFDB50 entries also overlapped with human–viral complexes. One representative example, shown in Fig. 5c, involves thymidylate kinase from the vaccinia virus, which was predicted to bind to the human thymidylate kinase (gene: DTYMK). This complex exhibited high structural similarity to an uncharacterized protein from Bremia lactucae, a plant-pathogenic fungus, suggesting that the virus may have evolved to preserve its dimerization capability for survival. Additionally, we identified an essential protein complex in human mitochondria that resembles a hypothetical protein from Perkinsus marinus, suggesting that our comparative analysis could aid in uncovering the unknown functions of proteins (Fig. 5d). Further comparative analyses revealed that such rearrangements are widespread across kingdoms (Supplementary Fig. 24a). Conversely, in a distinct case, we observed both protein monomers and protein complexes with high sequence similarity within the same bacterial species, indicating the potential occurrence of ORF read-through interruptions caused by occasional stop codons (Supplementary Fig. 24b, c). We identified multiple instances in which protein complexes and their corresponding full-length monomers coexist within the same bacterial species, highlighting the prevalence of gene fusion and fission phenomena across prokaryotic lineages (Supplementary Fig. 24d, e). In addition to clearly identified direct protein fusions, we also discovered cases in which small protein complexes resembled partial regions of long monomers, possibly resulting from complex structural rearrangements of the proteins. Notably, a predicted protein complex from Promethearchaeota archaeon (archaea) exhibited structural similarity to a much larger protein from Macrostomum lignano (Fig. 5j).
a Workflow for identifying protein complexes involving protein fusion and fission. Two subunit monomers within the protein complex were individually searched against the AFDB50 database using Foldseek. Co-occurred monomers were further analyzed to identify the mapping positions of the two subunits on it, thereby finding monomers with structures similar to the protein complex. b Bar graph showing the numbers of identified protein fusion events. The green bars represent the number of protein hits from AFDB50 identified as involving protein fusion. c–j Predicted 3D structure of protein monomers, their resembled protein complex, and the aligned image.
Overall, through comprehensive comparative analyses of two large protein structure datasets, we identified numerous instances of protein fusion and fission events, underscoring their pivotal roles in protein evolution.
Improving protein-binding site prediction using our predicted protein complexes dataset
The AlphaFold Database (AFDB) of protein monomer structures not only provides an extensive repository of structural information but also serves as a critical source of training data for developing advanced predictive models. For example, high-quality subsets of AFDB data have been curated to train the AlphaFold3 model8. Over 40 million high-quality monomer structures were used to train the unified structural and protein sequence embedding model SaProt48. AFDB is also applied during fine-tuning stages for protein domain segmentation annotation49. Identifying functional sites on protein surfaces is crucial for drug discovery, protein engineering, and vaccine design. Among our dataset, a substantial array of protein interaction surfaces is available. MPBind, a recently developed model for protein-binding site prediction, is built upon protein language models and equivariant graph neural networks50. We continued training MPBind, referred to as MPBind-PHC, using 9609 predicted non-redundant protein complexes that were selected according to Best LIS ≥ 0.4 and pLDDT ≥ 80.
To evaluate binding-site prediction performance, we first curated a validation dataset from predicted protein complex structures by removing redundant entries based on interface and monomer structural similarity using FoldSeek. The fine-tuned model was then tested on an independent test set, Pro_Test_315, derived from experimentally resolved structures51, as well as on a filtered version of this dataset (Filtered Pro_Test_315) constructed with the same deduplication procedure. Across all three evaluation datasets, the fine-tuned model exhibited improved AUC values, highlighting the potential of predicted protein complexes as a valuable resource for applications in biomedicine and beyond (Supplementary Fig. 25).
In sum, we demonstrated deep-learning applications built on a large-scale protein complex dataset, showing that the massive predictions generated by AlphaFold not only facilitate the discovery of individual PPIs but also provide substantial value to protein science through high-quality predicted data-driven modeling.
Discussion
In this study, we employed a large-scale, AlphaFold-based structure-prediction pipeline to build an atlas of predicted protein complexes spanning viruses, archaea, bacteria, plants, and mammals. This resource advances our understanding of key biological mechanisms, including viral pathogenesis, the evolution of protein assemblies, and complex cellular processes. The rapid progress of high-accuracy structure prediction suggests that large-scale, data-centric modeling will become a broadly applicable approach for investigating diverse biological processes. Although such modeling offers advantages when integrated with high-throughput experiments, it also presents challenges. High-throughput screening identified numerous candidate protein–protein interactions; nevertheless, only a minority received high-confidence predictions from AlphaFold-Multimer, plausibly owing to a combination of screening-derived false positives and genuine—but transient, weak, or condition-specific—interactions that elude confident modeling given incomplete MSA coverage and current algorithmic constraints. To address these issues, it will be important to improve the accuracy and sensitivity of both computational models and experimental techniques, thereby enabling more comprehensive identification and characterization of protein interactions, particularly those that are transient or underrepresented in current datasets.
Recent advances in deep learning-based structure prediction, especially all-atom models such as AlphaFold3 and RosettaFold-All-Atom7,8, extend beyond protein structure prediction to include modeling of protein-ligand and protein-nucleic acid complexes, and to account for post-translational modifications. Such interactions underpin diverse biological functions, including binding of transcription factors to DNA52, regulatory mechanisms between metabolites and enzymes53, and phosphorylation-dependent signaling54. Incorporating these interaction types into predictive frameworks enables more faithful modeling of dynamic cellular processes. We anticipate that the near future will bring an influx of expansive datasets, catalyzing a surge of biological discoveries and further enhancing the refinement and iteration of deep-learning models. This integration will not only deepen our understanding of molecular machinery but also drive innovation in drug design and genetic engineering. Taken together, our proteome-wide atlas of protein complex structures opens up new opportunities for systems biology, protein engineering, structure-guided omics, and structure-informed evolutionary analyses, ultimately advancing our understanding of the fundamental principles that govern protein interactions and link sequence, structure, and function.
Methods
Pathogenic bacteria operon prediction
We obtained the translated CDS FASTA files of the reference genomes for each of the 36 pathogenic bacterial species from the NCBI genome dataset (https://www.ncbi.nlm.nih.gov/datasets/genome/). The operons were predicted by Operon Mapper (https://biocomputo.ibt.unam.mx/operon_mapper/)25. Proteins encoded within in the same operon were pairwise combined as operonic protein pairs.
Representative genome analysis of archaeal and bacterial phyla
For each phylum of Archaea and Bacteria, a representative sequence was selected from the GTDB database (https://gtdb.ecogenomic.org/tree). The selection criteria prioritized the largest order encompassing the greatest number of classes, followed by the largest family within that order, and continuing this hierarchical approach to identify the GTDB species representative. Corresponding translated CDS FASTA files were retrieved from NCBI based on the GenBank Assembly accession numbers. When a CDS FASTA file was unavailable, alternative representative sequences were chosen. The final selection included 21 representative sequences from Archaea (spanning 21 phyla) and 167 representative sequences from Bacteria (covering 181 phyla). From the acquired FASTA files, all adjacent protein sequence pairs located on the same genome were extracted.
Phylogenetic tree analysis
Phylogenetic tree analysis was conducted using OrthoFinder 2.5.5 (https://github.com/davidemms/OrthoFinder) on the genomic proteins of 36 pathogenic bacteria, 21 representative species of Archaea, and 167 representative species of Bacteria. The resulting phylogenetic trees were visualized with iTOL v6 (https://itol.embl.de/).
Protein complex prediction using ColabFold
We employed ColabFold v1.5.5 (https://github.com/YoshitakaMo/localcolabfold) to predict protein complexes, employing databases including UniRef30 (uniref30_2302), ColabFoldDB (colabfold_envdb_202108), and the template database (pdb100_230517) for the structure inference. ColabFold operates in two main stages: Multiple Sequence Alignment (MSA) generation and protein structure prediction. For the MSA generation, we used the colabfold_search command line tool with --use-templates 1 option enabled to perform template-based searches, while keeping all other parameters at their default settings.
Subsequently, we utilized the colabfold_batch command line tool to predict protein structures, and performed the structure refinement on GPUs (--amber --use-gpu-relax) with the Alphafold2_Multimer_v3 model. When template matches were identified in the MSA, we incorporated template information into the prediction using --template option. For cases without template matching, predictions were carried out without using the --template parameter. ColabFold uses the num-recycle parameter to control the number of prediction recycles. Increasing the number of recycles can improve prediction quality but increases computation time. Without any special instructions, we used 20 recycles (--num-recycle 20) to ensure high prediction quality. For mice, Arabidopsis, and 188 representative bacterial and archaeal species, we used 3 recycles (--num-recycle 3) to significantly reduce inference time while maintaining sufficient prediction quality. With default parameter --num-models 5, Colabfold outputs five PDB-formatted files ranked from 1 to 5, for each protein complex prediction. We selected the ipTM highest-ranked PDB file as the final prediction. The corresponding confidence scores, including pLDDT, pTM, and ipTM, were recorded in the inference log files and scoring JSON files. The LIS and LIA metric scores were calculated as the guidance (https://github.com/flyark/AFM-LIS).
Comparison of AlphaFold3 with protenix, Chai-1, Boltz-2 and ColabFold-multimer
Heterodimeric and homodimeric complexes deposited in PDB, since 17 May 2023, were retrieved by Dockground server55 filtered to a maximum resolution of 3.5 Å, an interface comprising of at least 12 contacting residues, and a buried surface area greater than 800 Ų. Redundancy was removed with Foldseek-multimer with default parameters, yielding 54 heterodimers and 75 homodimers that served as the definitive test set. The AlphaFold3 predictions were performed on the online server (https://alphafoldserver.com/). ColabFold-Multimer was predicted as described above. Boltz-2 was predicted as the following command line: boltz predict input_path --recycling_steps 3 --sampling_steps 200 --diffusion_samples 5 --use_msa_server. Protenix predictions with ESM embeddings were executed as protenix predict --input input.json --out_dir./output_no_msa --seeds 101 --use_esm. Protenix predictions using MSA, and Chai-1 prediction were performed with scripts available on GitHub (https://github.com/wensm77/Protein-Complex-Atlas). Prediction quality was assessed using lDDT module in OpenStructure56 to evaluate residue-level accuracy and DockQ57 to measure overall chain–chain docking performance.
Co-localization score calculation
Initially, we used each of the 36 pathogenic bacterial genomes as a query in the search against the NCBI database, which served as the target and comprised 890 million proteins (across 18 million contigs). Searches were performed using the MMseqs2 version 15.6f452, filtering for sequences with the similarity ranging from 0.3 to 0.9. The search parameter —max-seqs was set to 10,000 to ensure sufficient confidence in the subsequent co-localization score calculations. Prior to calculating this matrix, we defined co-localization as occurring between two protein sequences when both of the following conditions are met: (1) the proteins are located on the same contig, and (2) the shortest distance between the two proteins does not exceed 3 kbp.
Considering a pathogenic bacterium with n query protein sequences, we denote \({q}_{i}\) as the i-th query protein and \({T}_{i}=[{t}_{1}^{i},{t}_{2}^{i},\ldots,{t}_{{Li}}^{i}]\) as the searched target protein list after identity filtering, where \({Li}\ge 0\) denotes the number of the corresponding i-th found sequences. For query protein pair \({q}_{i},{q}_{j}\), we iterated through every possible pair of protein sequences in \({T}_{i}\) and \({T}_{j}\) and counted whether they were co-localized. In other words, \({f}_{{count}}(i,j)\) is initialized to 0 for all \(i,j\le n\). Moreover, \(\forall {t}_{u}^{i}\in {T}_{i},{t}_{v}^{j}\in {T}_{j}\), \({f}_{{count}}(i,j)\)=\(\,{f}_{{count}}(i,j)\)+1 if \({t}_{u}^{i},{t}_{v}^{j}\) are co-localized. Accordingly, calculate \({f}_{{count}}(i,j)\) for all combinations of \(i,{j}\le {n}\) except \(i=j\), since we ignore self-localization. Actually, it is unnecessary to calculate \({f}_{{count}}(i,j)\) for the \(i < j\) situation owing to \({f}_{{count}}(i,j)\)=\(\,{f}_{{count}}( \, \, j,i)\).
Based on the analysis above, the co-localization score matrix \(M=[{m}_{{ij}}]\in {{\mathbb{R}}}^{n\times n}\) is calculated as follows:
where \({contig\_cnt}({T}_{i})\) indicates the number of contigs associated with T_i. It should be noted that if at least one of \(\{{Li},{Lj}\}\) is small, then one of \(\{{m}_{{ij}},{m}_{{ji}}\}\) will be may become disproportionately high while the other is close to 0. To avoid such situation and ensure the accuracy of co-localization scores, we applied the principle of mutual interaction between co-localized protein pairs as follows: (1) \({m}_{{ij}}\ge 0.4\) or \({m}_{{ji}}\ge 0.4\); (2) \({q}_{i},{q}_{j}\) are co-localized; (3) \({Li}\ge 2000\) and \({Lj}\ge 2000\). Protein pairs satisfying these conditions were considered associated co-localized protein pairs.
Cell culture and plasmids transfection
H1299 (CRL-5803) and 293 T (CRL-3216) were obtained from ATCC (Manassas, VA, USA) and cultured in DMEM medium (Gibco, Rockville, MD, USA) supplemented with 10% fetal bovine serum (FBS; Hyclone, Logan, UT, USA), 100 units/mL penicillin (GIBCO), and 100 μg/mL streptomycin (GIBCO). Cells were maintained in a humidified 37 °C incubator under a 5% CO₂ atmosphere. All cell lines used in this study were routinely tested negative for mycoplasma contamination and were maintained at low passage numbers to maintain their identity, and were authenticated by morphology check and growth curve analysis.
For plasmids transfection, 293 T cells at 60% confluence were transfected using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). The plasmids expressing Flag- or HA-tagged proteins (Flag-E3A, Flag-UL44, HA-CD4, and HA-HLA) were synthesized and constructed by GenScript Biotech (Sequences showed in Supplementary Table 2).
Immunofluorescence staining
Cells grown on coverslips were fixed with 4% polyformaldehyde in PBS, permeabilized with 0.1% Triton X-100 in PBS, blocked with 4% bovine serum albumin in PBS, hybridized to an appropriate primary antibody (PKCζ: 1:100, 340815, Zenbio; NPM1:1:200, MA5-12508, ThermoFisher), followed by incubation with a second antibody (Goat anti-Mouse Alexa Fluor 488, A-11029 or Goat anti- Rabbit Alexa Fluor 514, A-31558, ThermoFisher). The cells were counterstained with ProLong® Gold Antifade Reagent with DAPI (82961, CST) prior to visualization and photographed using a Leica TCS SP5II confocal laser scanning microscope. LAS X (version3.3.0) was used to analyze fluorescent images. To evaluate the PKCζ or NPM1 co-localization with DAPI, the free software Image J. Fiji, coupled with the Coloc 2 plugin and Pearson’s correlation coefficient were used to calculate double fluorescence correlation coefficients, and co-localized fluorescence quantifications were presented by scatter plots.
Co-immunoprecipitation (Co-IP) and western blot
Cells were lysed in IP buffer (1 mM Tris pH 7.5, 5 mM NaCl, 0.25% Nonidet P-40, 0.1% deoxycholate, and protease inhibitors), and equal amounts of total protein were incubated with primary antibodies, or normal indicated IgG, overnight at 4 °C, and then 30 µl of protein A/G beads were added for an additional 2 h of incubation. For exogenous Co-IP, anti-HA beads were added to equal amounts of total protein and incubated overnight. Beads were centrifuged (500 × g for 30 s) and washed three times using wash buffer (20 mM Tris-HCl, 250 mM NaCl, 0.2 mM EGTA, and 0.1% Nonidet P-40). The beads were heated at 100 °C for 10 min before western blot analyses. Anti-HA magnetic beads (88836) were obtained from Thermo Fisher Scientific (Waltham, MA, USA).
For western blot, cell lysates were loaded, separated by SDS-PAGE, transferred to PVDF membranes (Millipore), and hybridized to an appropriate primary antibody and horseradish peroxidase (HRP)-conjugated secondary antibody for subsequent detection by enhanced chemiluminescence (Bio-Rad). Western blot images were analyzed using Image Lab Software 5.0. Antibody for GAPDH (AB0037,1:5000) was purchased from Abways (Shanghai, China). Antibodies for SP1 (SC-59,1:1000) were purchased from Santa Cruz (USA). Antibody for PKCζ (PRKCZ) (340815, 1:1000) was purchased from Zenbio (Chengdu, China). Antibody for KPNA2 (ET1705-61, 1:1000) was purchased from Huabio (Hangzhou, China). Antibody for NPM1 (MA5-125081:1000) was purchased from ThermoFisher Scientific (Wilmington, DE, USA). Antibodies for HA (3724, 1:1000), rabbit lgG (2729, 1:1000), and Flag (117935, 1:1000) were purchased from Cell Signaling Technology (Danvers, MA, USA).
Homo oligomer-contained large protein complex assembly
To construct a large homo-oligomeric protein complex, we adopted a strategy that combines QSproteome with AlphaFold2-bigbang21. Specifically, we first used QSproteome to truncate the flexible regions of homodimers using the default parameters, and then calculated the symmetry-related structure based on this truncation. After determining the optimal symmetry, we used AnAnaS to generate the untruncated symmetric complex structure. This untruncated symmetric structure, along with the MSA and sequence information of the homologous dimers, was provided as input to AlphaFold2-bigbang to predict the full-length symmetric complex structure. This approach allowed us to overcome the negative impact of flexible region truncation on AlphaFold2’s prediction accuracy.
In the high-quality protein structure prediction database, for a heterodimer composed of two distinct monomers, a and b, we first identified homodimers composed solely of a and b, respectively. Then, using the aforementioned method, we calculated the full-length optimal symmetric structures for these two homodimers and determined their optimal symmetries. If the two homodimers share the same optimal symmetry, we used CombFold (https://github.com/dina-lab3D/CombFold) as instructed to assemble them into a complete homo-oligomer containing a large protein complex.
Finetuning MPBind
We fine-tuned MPBind to accommodate our specific task requirements. We curated a subset comprising 71,667 high-quality complexes by filtering based on Best LIS ≥ 0.4 and pLDDT ≥ 80. To ensure diversity and non-redundancy in the training data, we implemented a deduplication strategy based on structural similarity. Utilizing FoldSeek, we eliminated complexes where the TM-score between any two protein monomers was ≥ 0.5. Following this process, the final non-redundant training set consisted of 9,609 dimeric complexes. Based on this dataset, we conducted continued training of the MPBind model (https://github.com/jianlin-cheng/MPBind) for 45 epochs on an NVIDIA RFX3090 GPU. Subsequently, we conducted deduplication on partial of remaining samples based on interface similarity and monomer similarity. In the interface deduplication phase, we employed a strategy that integrates structural alignment with interface site information to identify and eliminate proteins that, despite differing overall structures, exhibit highly similar key functional interfaces. We first utilized FoldSeek to compute structural alignment information between all protein single chains in the candidate samples (command: foldseek easy-search query.pdb targetdb out.m8 tmp --format-output “query,target,evalue,bits,qstart,qend,tstart,tend” -s 9.5). For any pair of proteins (denoted as A and B), we examined the extent of overlap between their structural alignment regions and the pre-defined interface residues. Interface residues in these complexes were defined as surface residues with a relative solvent accessibility greater than 5% that lost more than 1 Ų of absolute solvent accessibility upon protein-protein complex formation. If the overlap between the alignment region of protein A and the entire interface region of protein B exceeds 50%, or if the overlap between the alignment region of protein B and the entire interface region of protein A exceeds 50%, then the pair of proteins is deemed to exhibit interface similarity. Any complex containing protein chains determined to have interface similarity will be removed from the validation set candidate pool. Subsequently, in the monomer deduplication phase, we aimed to thoroughly exclude any single chains in the validation set complexes that display structural similarity with those in the training set. We employed FoldSeek to perform structural clustering of all monomers in the dataset, using the command: foldseek easy-cluster pdb res tmp -c 0.9. Each monomer was mapped to a unique structural cluster representative. Any validation-candidate dimer sharing a representative (from either chain) with the training set was discarded, yielding a validation set whose dimers are structurally independent of all training chains. We ultimately obtained a validation set comprising 205 monomers. Similarly, we performed deduplication on the independent dataset Pro_Test_315, resulting in the construction of a test set containing 28 monomers derived from PDB structures. The predicted residues were selected as the threshold with 0.5 as before.
Genomic co-occurrence rate calculation
Reference genomes were downloaded from NCBI. The homologs were searched by MMseqs2 with similarity between 0.3 and 0.9. Then the genomic co-occurrence was calculated as the number of genomes in which both genes are present, divided by the total number of genomes in which either gene is present.
Protein complex clustering and Gene fusion hit discovery
Protein complexes were predicted by Foldseek Multimer as the command: foldseek easy-multimercluster INPUT_PDB_DIR OUTPUT_CLUSTER_DIR TMP_WORK_DIR --multimer-tm-threshold 0.65 --chain-tm-threshold 0.5 --cov-mode 2 -e 0.01 --exhaustive-search. AFDB50 Foldseek database file was downloaded from https://foldseek.steineggerlab.workers.dev/. Each monomer extracted from the protein complexes was searched against AFDB50 with Foldseek (command: foldseek easy-search query.pdb targetdb out.m8 tmp --format-output “query,target,evalue,bits,qstart,qend,tstart,tend” -s 9.5). For each hit, we considered the aligned query and target segments as intervals, measured the length of their intersection (overlap) and union (combined coverage), and calculated an overlap/union ratio; only hits with a ratio <0.30 were accepted to avoid mapping to the same region.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The 1.1 million predicted protein structures generated in this study have been deposited in the ModelScope database (https://www.modelscope.cn/collections/protein_complex_atlas-2ae5e7d4f4a343). The processed, curated high-confidence PPI structures are available at a companion website (https://www.biopredictnavigator.cn). Accession codes for analysed genomes of representative prokaryotes are available in Supplementary Data 1. Source data are provided with this paper.
Code availability
The code for this manuscript is provided in GitHub repository: https://github.com/wensm77/Protein-Complex-Atlas, and on Zenodo: https://doi.org/10.5281/zenodo.18630539.
References
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876 (2021).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Evans, R. et al. Protein complex prediction with AlphaFold-Multimer. bioRxiv, 2021.10.04.463034 (2021).
Bryant, P. et al. Predicting the structure of large protein complexes using AlphaFold and Monte Carlo tree search. Nat. Commun. 13, 6028 (2022).
Shor, B. & Schneidman-Duhovny, D. CombFold: predicting structures of large protein assemblies using a combinatorial assembly algorithm and AlphaFold2. Nat. Methods 21, 477–487 (2024).
Krishna, R. et al. Generalized biomolecular modeling and design with RoseTTAFold All-Atom. Science 384, eadl2528 (2024).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Akdel, M. et al. A structural biology community assessment of AlphaFold2 applications. Nat. Struct. Mol. Biol. 29, 1056–1067 (2022).
Bouatta, N. & AlQuraishi, M. Structural biology at the scale of proteomes. Nat. Struct. Mol. Biol. 30, 129–130 (2023).
Hammack, A. T. & Blaby-Haas, C. E. Machine learning sheds light on microbial dark proteins. Nat. Rev. Microbiol. 22, 63–63 (2024).
van Kempen, M. et al. Fast and accurate protein structure search with Foldseek. Nat. Biotechnol. 42, 243–246 (2024).
Barrio-Hernandez, I. et al. Clustering predicted structures at the scale of the known protein universe. Nature 622, 637–645 (2023).
Durairaj, J. et al. Uncovering new families and folds in the natural protein universe. Nature 622, 646–653 (2023).
Bordin, N. et al. AlphaFold2 reveals commonalities and novelties in protein structure space for 21 model organisms. Commun. Biol. 6, 160 (2023).
Nomburg, J. et al. Birth of protein folds and functions in the virome. Nature 633, 710–717 (2024).
Kim, R. S., Levy Karin, E., Mirdita, M., Chikhi, R. & Steinegger, M. BFVD—a large repository of predicted viral protein structures. Nucleic Acids Res. 53, D340–D347 (2024).
Huang, J. et al. Discovery of deaminase functions by structure-based protein clustering. Cell 187, 4426–4428 (2024).
Szklarczyk, D. et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Oughtred, R. et al. The BioGRID database: A comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci. 30, 187–200 (2021).
Schweke, H. et al. An atlas of protein homo-oligomerization across domains of life. Cell 187, 999–1010 e15 (2024).
Humphreys, I. R. et al. Computed structures of core eukaryotic protein complexes. Science 374, eabm4805 (2021).
Burke, D. F. et al. Towards a structurally resolved human protein interaction network. Nat. Struct. Mol. Biol. 30, 216–225 (2023).
Shine, M. et al. Co-transcriptional gene regulation in eukaryotes and prokaryotes. Nat. Rev. Mol. Cell Biol. 25, 534–554 (2024).
Taboada, B., Estrada, K., Ciria, R. & Merino, E. Operon-mapper: a web server for precise operon identification in bacterial and archaeal genomes. Bioinformatics 34, 4118–4120 (2018).
Altae-Tran, H. et al. Uncovering the functional diversity of rare CRISPR-Cas systems with deep terascale clustering. Science 382, eadi1910 (2023).
Makarova, K. S. et al. Evolutionary classification of CRISPR-Cas systems: a burst of class 2 and derived variants. Nat. Rev. Microbiol. 18, 67–83 (2020).
Terlouw, B. R. et al. MIBiG 3.0: a community-driven effort to annotate experimentally validated biosynthetic gene clusters. Nucleic Acids Res. 51, D603–D610 (2023).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Passaro, S. et al. Boltz-2: towards accurate and efficient binding affinity prediction. bioRxiv, 2025.06.14.659707 (2025).
Discovery, C. et al. Chai-1: Decoding the molecular interactions of life. bioRxiv, 2024.10.10.615955 (2024).
Team, B. A. A. S. et al. Protenix—advancing structure prediction through a comprehensive AlphaFold3 reproduction. bioRxiv, 2025.01.08.631967 (2025).
Bryant, P., Pozzati, G. & Elofsson, A. Improved prediction of protein-protein interactions using AlphaFold2. Nat. Commun. 13, 1265 (2022).
Zhu, W., Shenoy, A., Kundrotas, P. & Elofsson, A. Evaluation of AlphaFold-multimer prediction on multi-chain protein complexes. Bioinformatics 39, btad424 (2023).
Kim, A.-R. et al. Enhanced protein-protein interaction discovery via AlphaFold-multimer. bioRxiv, 2024.02.19.580970 (2024).
Schweke, H. et al. An atlas of protein homo-oligomerization across domains of life. Cell 187, 999–1010.e15 (2024).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Yang, M. et al. Biogenesis of a bacterial metabolosome for propanediol utilization. Nat. Commun. 13, 2920 (2022).
Chandrangsu, P., Rensing, C. & Helmann, J. D. Metal homeostasis and resistance in bacteria. Nat. Rev. Microbiol. 15, 338–350 (2017).
Liu, B., Zheng, D., Zhou, S., Chen, L. & Yang, J. VFDB 2022: a general classification scheme for bacterial virulence factors. Nucleic Acids Res. 50, D912–D917 (2022).
Luck, K. et al. A reference map of the human binary protein interactome. Nature 580, 402–408 (2020).
Li, H., Mapolelo, D. T., Randeniya, S., Johnson, M. K. & Outten, C. E. Human glutaredoxin 3 forms [2Fe-2S]-bridged complexes with human BolA2. Biochemistry 51, 1687–1696 (2012).
Liang, G. & Bushman, F. D. The human virome: assembly, composition and host interactions. Nat. Rev. Microbiol. 19, 514–527 (2021).
Lasso, G. et al. A structure-informed atlas of human-virus interactions. Cell 178, 1526–1541 e16 (2019).
Yang, X. et al. HVIDB: a comprehensive database for human-virus protein-protein interactions. Brief. Bioinform. 22, 832–844 (2021).
Xiao, J. et al. FBXL20-mediated Vps34 ubiquitination as a p53 controlled checkpoint in regulating autophagy and receptor degradation. Genes Dev. 29, 184–196 (2015).
Arnold, B. J., Huang, I. T. & Hanage, W. P. Horizontal gene transfer and adaptive evolution in bacteria. Nat. Rev. Microbiol. 20, 206–218 (2022).
Su, J. et al. SaProt: protein language modeling with structure-aware vocabulary. bioRxiv, 2023.10.01.560349 (2023).
Lau, A. M., Kandathil, S. M. & Jones, D. T. Merizo: a rapid and accurate protein domain segmentation method using invariant point attention. Nat. Commun. 14, 8445 (2023).
Wang, Y., Boadu, F. & Cheng, J. MPBind: multitask protein binding site prediction by protein language models and equivariant graph neural networks. bioRxiv, 2025.04.12.648527 (2025).
Fang, Y. et al. DeepProSite: structure-aware protein binding site prediction using ESMFold and pretrained language model. Bioinformatics 39, btad718 (2023).
Spitz, F. & Furlong, E. E. M. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Martínez-Reyes, I. & Chandel, N. S. Mitochondrial TCA cycle metabolites control physiology and disease. Nat. Commun. 11, 102 (2020).
Lee, J. M., Hammarén, H. M., Savitski, M. M. & Baek, S. H. Control of protein stability by post-translational modifications. Nat. Commun. 14, 201 (2023).
Douguet, D., Chen, H.-C., Tovchigrechko, A. & Vakser, I. A. Dockground resource for studying protein–protein interfaces. Bioinformatics 22, 2612–2618 (2006).
Biasini, M. et al. OpenStructure: an integrated software framework for computational structural biology. Acta Crystallogr. D. Biol. Crystallogr. 69, 701–709 (2013).
Mirabello, C. & Wallner, B. DockQ v2: improved automatic quality measure for protein multimers, nucleic acids, and small molecules. Bioinformatics 40, (2024).
Acknowledgements
This work was supported by National Key R&D Program of China (No.2023YFF1205400 to D.M.), National Natural Science Foundation of China under grant (No. 32571689 and No. 32301230 to D.M., No. 82573859 to Y. Y. and No. 72174172 to D. E.), Zhejiang Laboratory PI start program, Fundamental and Interdisciplinary Disciplines Breakthrough Plan of the Ministry of Education of China (JYB2025XDXM502 to J.Z.), the Noncommunicable Chronic Diseases-National Science and Technology Major Project (No. 2024ZD0525100 to J.Z.) and the Scientific and Technological Innovation Team for Qinghai-Tibetan Plateau Research in Southwest Minzu University (Grant No.2024CXTD20). Authors thank Yuanzhao Pan (Beijing National Day School) for valuable support and insightful discussions. Xitong Li (Jiangnan University), Weizhen Ou (Jiangnan University), Jijun Fan (Jiangnan University), Wenbo Deng (China University of Mining and Technology), and Shuhao Niu (Jiangnan University) provided suggestions for language revisions.
Author information
Authors and Affiliations
Contributions
D.M. conceived this project. X.Q., C.Y., J.L., S.W., Yuanyuan L., K.D., Yongfu H., J.F., W.M., L.L., Z.L., Y.S., H.Z., Yayun H., R.Z., P.J., Yafei L., B.L., H.W., Yuxuan C., Z.M., P.Y., X.X., J.W., Y.Z., Q.Z., W.Z., K.Y., S. L., H.X., D.E. performed the computational analysis. J.F. and Y.Y. performed wet-lab experiments. D.M., Y.Y., Ying C., and C.S. supervised the project. D.M., R.Z., Z.X., J.Z., D.E., H.X., and W.Z. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Qi, X., Ye, C., Liang, J. et al. Atlas of predicted protein complex structures across kingdoms. Nat Commun 17, 4397 (2026). https://doi.org/10.1038/s41467-026-70884-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70884-4