Abstract
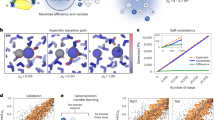
Mitochondrial outer membrane proteins (β-OMPs) serve as the first line of communication with the cellular milieu. A crucial β-OMP, the sorting and assembly machinery Sam50, is a 16-stranded transmembrane β-barrel highly conserved in all eukaryotes. Sam50 dysregulation is lethal; yet, molecular elements that regulate Sam50 remain poorly understood. Here, we identify and characterize residues that regulate Sam50 structure and function. Using single-molecule electrophysiology, in vivo function, and stability measurements, we demonstrate that the POlypeptide-TRansport Associated domain is dispensable for Sam50. Complete characterization of the folding mechanism using 165 Xaa→Ala substitutions reveals that Sam50 folds through parallel pathways, with at least two transition states. The folding nucleus is towards the N-terminus, whereas frustrated folding at the C-terminal region kinetically traps the structure. We correlate this unexpected folding of Sam50 with its SAM-assisted assembly. Additionally, our per-residue stability measurements show that destabilizing hotspots in Sam50 are linked to its gating function. Our findings suggest how the dynamic structure of Sam50 offers a functional advantage, with specific residues that regulate folding, stability, and function, also determine the protein’s sensitivity to mutations.
Similar content being viewed by others
Data availability
The source data underlying Figs. 1c, 1e, 1i, 2a, 2c, and 4a, and other SI images are provided as a Source Data file. Accession codes and hyperlinks for structures from PDB: 7BTW; 7BTY; 7VKU. Should any raw data files be needed in another format, they are available from the corresponding author. Source data are provided in this paper.
Code availability
Source files and scripts pertaining to the MD simulations are provided as a separate supplementary file, and contain the initial and final system coordinates (.PDB), lipid and force-field parameter files (.itp) and system topology files (.top), bfactor-mapped.pdb derived from per-residue fluctuations, as well as RMSD, per-residue RMSF, free energy landscape data, and distance analysis data (.txt,.xvg), are available in the Source Data file.
References
Ravi, R., Routray, D. & Mahalakshmi, R. Mitochondrial sorting and assembly machinery: chaperoning a moonlighting role?. Biochemistry 64, 312–328 (2025).
Diederichs, K. A. et al. Structural insight into mitochondrial beta-barrel outer membrane protein biogenesis. Nat. Commun. 11, 3290 (2020).
Takeda, H. et al. Mitochondrial sorting and assembly machinery operates by beta-barrel switching. Nature 590, 163–169 (2021).
Takeda, H. et al. A multipoint guidance mechanism for beta-barrel folding on the SAM complex. Nat. Struct. Mol. Biol. 30, 176–187 (2023).
Stroud, D. A. et al. Biogenesis of mitochondrial beta-barrel proteins: the POTRA domain is involved in precursor release from the SAM complex. Mol. Biol. Cell 22, 2823–2833 (2011).
Paschen, S. A. et al. Evolutionary conservation of biogenesis of beta-barrel membrane proteins. Nature 426, 862–866 (2003).
Jian, F. et al. Sam50 Regulates PINK1-parkin-mediated mitophagy by controlling PINK1 stability and mitochondrial morphology. Cell Rep. 23, 2989–3005 (2018).
Li, D., Rocha-Roa, C., Schilling, M. A., Reinisch, K. M. & Vanni, S. Lipid scrambling is a general feature of protein insertases. Proc. Natl. Acad. Sci. USA 121, e2319476121 (2024).
Nunnari, J. & Suomalainen, A. Mitochondria: in sickness and in health. Cell 148, 1145–1159 (2012).
Hohr, A. I. C. et al. Membrane protein insertion through a mitochondrial beta-barrel gate. Science 359, eaah6834 (2018).
Kutik, S. et al. Dissecting membrane insertion of mitochondrial beta-barrel proteins. Cell 132, 1011–1024 (2008).
Machin, J. M., Kalli, A. C., Ranson, N. A. & Radford, S. E. Protein-lipid charge interactions control the folding of outer membrane proteins into asymmetric membranes. Nat. Chem. 15, 1754–1764 (2023).
Maurya, S. R. & Mahalakshmi, R. N-helix and Cysteines inter-regulate human mitochondrial VDAC-2 function and biochemistry. J. Biol. Chem. 290, 30240–30252 (2015).
Srivastava, S. R., Zadafiya, P. & Mahalakshmi, R. Hydrophobic mismatch modulates stability and plasticity of human mitochondrial VDAC2. Biophys. J. 115, 2386–2394 (2018).
Surrey, T., Schmid, A. & Jahnig, F. Folding and membrane insertion of the trimeric beta-barrel protein OmpF. Biochemistry 35, 2283–2288 (1996).
Kruger, V. et al. Identification of new channels by systematic analysis of the mitochondrial outer membrane. J. Cell Biol. 216, 3485–3495 (2017).
Anfinsen, C. B. Principles that govern the folding of protein chains. Science 181, 223–230 (1973).
Pocanschi, C. L., Patel, G. J., Marsh, D. & Kleinschmidt, J. H. Curvature elasticity and refolding of OmpA in large unilamellar vesicles. Biophys. J. 91, L75–L77 (2006).
George, A. et al. Engineering a hyperstable Yersinia pestis outer membrane protein Ail using thermodynamic design. J. Am. Chem. Soc. 144, 1545–1555 (2022).
Moon, C. P., Zaccai, N. R., Fleming, P. J., Gessmann, D. & Fleming, K. G. Membrane protein thermodynamic stability may serve as the energy sink for sorting in the periplasm. Proc. Natl. Acad. Sci. USA 110, 4285–4290 (2013).
Schiffrin, B., Brockwell, D. J. & Radford, S. E. Outer membrane protein folding from an energy landscape perspective. BMC Biol. 15, 123 (2017).
Marx, D. C. & Fleming, K. G. Membrane proteins enter the fold. Curr. Opin. Struct. Biol. 69, 124–130 (2021).
Diederichs, K. A., Buchanan, S. K. & Botos, I. Building Better Barrels - beta-barrel Biogenesis and Insertion in Bacteria and Mitochondria. J. Mol. Biol. 433, 166894 (2021).
Huysmans, G. H., Baldwin, S. A., Brockwell, D. J. & Radford, S. E. The transition state for folding of an outer membrane protein. Proc. Natl. Acad. Sci. USA 107, 4099–4104 (2010).
Paslawski, W. et al. Cooperative folding of a polytopic alpha-helical membrane protein involves a compact N-terminal nucleus and nonnative loops. Proc. Natl. Acad. Sci. USA 112, 7978–7983 (2015).
Kozjak, V. et al. An essential role of Sam50 in the protein sorting and assembly machinery of the mitochondrial outer membrane. J. Biol. Chem. 278, 48520–48523 (2003).
Wang, Q. et al. Structural insight into the SAM-mediated assembly of the mitochondrial TOM core complex. Science 373, 1377–1381 (2021).
Chamachi, N. et al. Chaperones Skp and SurA dynamically expand unfolded OmpX and synergistically disassemble oligomeric aggregates. Proc. Natl. Acad. Sci. USA 119, e2118919119 (2022).
Calabrese, A. N. et al. Inter-domain dynamics in the chaperone SurA and multi-site binding to its outer membrane protein clients. Nat. Commun. 11, 2155 (2020).
Jefferson, R. E., Blois, T. M. & Bowie, J. U. Membrane proteins can have high kinetic stability. J. Am. Chem. Soc. 135, 15183–15190 (2013).
Moon, C. P., Kwon, S. & Fleming, K. G. Overcoming hysteresis to attain reversible equilibrium folding for outer membrane phospholipase A in phospholipid bilayers. J. Mol. Biol. 413, 484–494 (2011).
Moon, C. P. & Fleming, K. G. Using tryptophan fluorescence to measure the stability of membrane proteins folded in liposomes. Methods Enzymol. 492, 189–211 (2011).
Chan, N. C. & Lithgow, T. The peripheral membrane subunits of the SAM complex function codependently in mitochondrial outer membrane biogenesis. Mol. Biol. Cell 19, 126–136 (2008).
Metzger, M. B., Scales, J. L., Dunklebarger, M. F., Loncarek, J. & Weissman, A. M. A protein quality control pathway at the mitochondrial outer membrane. Elife 9, e51065 (2020).
Gentle, I., Gabriel, K., Beech, P., Waller, R. & Lithgow, T. The Omp85 family of proteins is essential for outer membrane biogenesis in mitochondria and bacteria. J. Cell Biol. 164, 19–24 (2004).
Pfanner, N., Warscheid, B. & Wiedemann, N. Mitochondrial proteins: from biogenesis to functional networks. Nat. Rev. Mol. Cell Biol. 20, 267–284 (2019).
Ott, C. et al. Sam50 functions in mitochondrial intermembrane space bridging and biogenesis of respiratory complexes. Mol. Cell Biol. 32, 1173–1188 (2012).
Monteiro-Cardoso, V. F. et al. ORP5/8 and MIB/MICOS link ER-mitochondria and intra-mitochondrial contacts for non-vesicular transport of phosphatidylserine. Cell Rep. 40, 111364 (2022).
Doyle, M. T. et al. Cryo-EM structures reveal multiple stages of bacterial outer membrane protein folding. Cell 185, 1143–1156 (2022).
Noinaj, N. et al. Structural insight into the biogenesis of beta-barrel membrane proteins. Nature 501, 385–390 (2013).
Lee, J. et al. Characterization of a stalled complex on the beta-barrel assembly machine. Proc. Natl. Acad. Sci. USA 113, 8717–8722 (2016).
Tomasek, D. et al. Structure of a nascent membrane protein as it folds on the BAM complex. Nature 583, 473–478 (2020).
Dubey, A. A., Singh, M. I. & Jain, V. Rapid and robust PCR-based all-recombinant cloning methodology. PLoS ONE 11, e0152106 (2016).
Srivastava, S. R. & Mahalakshmi, R. Evolutionary selection of a 19-stranded mitochondrial beta-barrel scaffold bears structural and functional significance. J. Biol. Chem. 295, 14653–14665 (2020).
Iyer, B. R. et al. Molecular switch between structural compaction and thermodynamic Stability by the Xxx-Pro Interface in Transmembrane beta-Barrels. Biochemistry 59, 303–314 (2020).
Matouschek, A. & Fersht, A. R. Application of physical organic chemistry to engineered mutants of proteins: Hammond postulate behavior in the transition state of protein folding. Proc. Natl. Acad. Sci. USA 90, 7814–7818 (1993).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Acknowledgements
We thank Prof. Dr. Doron Rapaport (University of Tuebingen, Germany) for the S. cerevisiae SAM50 gene. We thank Prof. Dr. Nils Wiedemann (University of Freiburg, Germany) and Prof. Dr. Bernard Guiard (Center de Génétique Moléculaire, CNRS, France) for S. cerevisiae YPH499 sam50::ade2 (sam50Δ shuffle strain) and Prof. Dr. Wiedemann for the pFL39-SAM50 E. coli-yeast shuttle vector. P.B. Tiwari generated pET3a-SAM50Δ120 from the FL variant. R.R. and J.K. thank the University Grants Commission, and AR thanks the Prime Minister’s Research Fellowship, Govt. of India, for a research fellowship. R.M. is a Wellcome Trust-DBT India Alliance Senior Fellow. This work was supported primarily by the Wellcome Trust-Department of Biotechnology India Alliance grant IA/S/20/2/505182, and in part by the Science and Engineering Research Board grant SPR/2021/000018 and Department of Biotechnology grant BT/PR28858/BRB/10/1718/2018, to R.M.
Author information
Authors and Affiliations
Contributions
R.M.: research design, funding, study supervision, interpretation of results, and manuscript writing. R.R.: library preparation (DNA and protein), electrophysiology measurements, yeast experiments, equilibrium titrations, kinetics measurements, MD simulations, data analysis, and interpretation of results. S.G. and J.K.: library preparation (DNA and protein), equilibrium titrations, data analysis. A.R.: electrophysiology measurements. MD simulations, and associated data analysis. All authors have approved the manuscript contents.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ravi, R., Gupta, S., Kumari, J. et al. Linking multipoint folding and stability with functional regulation in the mitochondrial transmembrane β-barrel Sam50. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70904-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70904-3