Abstract
In mouse cells, constitutive heterochromatin is associated with underlying arrays of A/T-rich DNA repeat elements, called the major satellite repeats (MaSat or MSR). We examine >18,000 MSR copies in mouse ES cells and identify that heterochromatin forms only at transcriptionally competent MSR units. To directly dissect the function of MSR DNA, we insert isolated MSR units into an inert genomic region that is repeat- and gene-free. Insertion of three or more intact MSR units induces heterochromatic histone marks, recruitment of HP1 and incorporation of histone H1. Only transcriptionally competent MSR units, but not permutated MSR variants or LINE1 5’UTR elements, nucleate de novo heterochromatin. MSR-derived transcription is bi-directional and MSR-originating transcripts are attenuated by the RNAPII-associated Integrator complex. Instructively, multi-copy intact MSR units impart an unwound DNA template that facilitates RNAPII engagement. Together, this study uncovers a DNA/RNA-based logic and transcription-coupled mechanism for the nucleation of heterochromatin.
Similar content being viewed by others
Introduction
Heterochromatin has been first described nearly 100 years ago as a chromatin region that remains densely stained in interphase and mitosis1. Since then, great insight has been gained and heterochromatin has been shown to have crucial roles for the structural and functional organisation of eukaryotic chromatin, for protecting the genome from the activity of invasive DNA and RNA transposable elements and for the stabilisation of gene expression programmes2,3. Heterochromatin defines one of the two ground states of eukaryotic chromatin4. In a simplified view, heterochromatin is associated with the non-coding and repeat-rich portion of the genome, where it regulates a transcription-repressive chromatin structure. By contrast, euchromatin controls a transcription-permissive chromatin structure at the coding and gene-rich fraction of the genome. Whereas DNA-based mechanisms for transcription factor and TATA box factor binding at gene promoters and the assembly of RNA polymerase II (RNAPII) complexes are well understood5,6 and can induce transcription-permissive euchromatin7, underlying genetic (i.e. DNA-based) mechanisms for heterochromatin establishment have not been fully resolved. Heterochromatin is, however, characterised by an evolutionarily conserved biochemical mechanism, in which Suv39h KMT enzymes generate histone H3K9 methylation8 that provides a binding affinity for HP1 proteins9,10.
In mammalian genomes, around 50% of the DNA sequence is composed of repetitive elements11. Of those, 3.6% are major satellite repeats (MaSat or MSR) that are mainly organised in large arrays of a reiterated 234 bp A/T-rich MSR unit12 in pericentric heterochromatin of each mouse chromosome. These numbers imply that there are around 384,000 copies of MSR units in the mouse genome. The consensus 234 bp MSR DNA sequence does not have a bona fide promoter architecture, but is full of embedded transcription factor binding sites that were shown to allow binding of Zn-finger13,14 or homeobox15 transcription factors. A recent finding has identified non-consecutive nucleotide triplets (TTC) to provide targeting affinities in highly variable heterochromatic sequences for atypical Zn-finger factors that then recruit Suv39h enzymes16. This discovery revealed an alternative mode for a DNA-based mechanism in targeting Suv39h enzymes to heterochromatin16.
Another long-standing paradox has been that heterochromatin presents transcriptional activity, which is then attenuated and silenced17,18. Thus, it is not resolved which RNA generating mechanisms would discriminate heterochromatic from euchromatic transcriptional activity. While this has been best studied in Schizosaccharomyces pombe and revealed RNA interference (RNAi)-mediated functions3,19,20,21,22 or RNA quality control factors23,24,25, it remains unclear whether, or to what extent, RNAi or related or distinct RNA processing mechanisms would also contribute to heterochromatin establishment in somatic mammalian cells26. Dicer-deficient mouse embryonic stem cells (mESC) were shown to display chromosome segregation defects and to accumulate high levels of major satellite repeat transcripts27,28. Transcriptional activity of MSR DNA is mostly silenced in differentiated cells, but MSR transcripts are up-regulated in the fertilised mouse zygote29,30 and are required to support progression of early mouse embryogenesis prior to zygotic genome activation and the emergence of embryonic stem cells. Further, forced expression of MSR transcripts has been shown to be a tumour driver31 and aberrantly high levels of satellite RNA are found in many forms of human cancer32.
The above insights underscore the importance of MSR transcriptional regulation and suggest that MSR repeat DNA could instruct a transcription-coupled mechanism for heterochromatin formation that also discriminates RNA processing of heterochromatic from euchromatic transcripts. Using insertion of isolated MSR units into an inert genomic region, we demonstrate this function of MSR DNA and identified that only transcriptionally competent MSR units can nucleate de novo heterochromatin formation. In addition, this study also resolves a long-standing paradox between heterochromatic and euchromatic transcriptional activity and uncovers a DNA/RNA-based logic for the establishment of heterochromatin in the mouse epigenome.
Results
Intact, but not permutated MSR units, induce H3K9me3 at an inert genomic region
The reiterative nature and the complex DNA composition of pericentric satellite regions33,34 has defied the analysis of individual MSR units in contributing to heterochromatin formation. Several MSR sequences are also found at some intergenic regions11 and around 42 intergenic regions containing MSR sequences have been annotated in the mouse genome. Using our published profiles for H3K9me3 in mouse ES cells (mESC)35 and ENCODE datasets36,37, we could subgroup these intergenic MSR regions to have high H3K9me3 enrichment, intermediate H3K9me3 enrichment or no H3K9me3 enrichment (Supplementary Fig. 1a). We then analysed H3K9me3 enrichment at some of these intergenic MSR regions in mESC in more detail by using directed ChIP with SNP primers that are specific for the intergenic MSR variants or with primers from immediately adjacent unique sequences (Supplementary Fig. 1b–d). We next focused on one intergenic MSR region that contains 1.5 copies (404 bp) of largely intact MSR sequences on mouse chromosome 9 at position 35 Mbp (act-9/35 MSR variant) and on another intergenic MSR region that has 1.2 copies (300 bp) of a greatly permutated MSR sequence on mouse chromosome 3 at position 99 Mbp (per-3/99 MSR variant) (Supplementary Fig. 2a). Directed ChIP for H3K9me3 indicates enrichment at the act-9/35 MSR variant position but no significant H3K9me3 signals across or adjacent to the per-3/99 MSR variant position (Supplementary Fig. 2b). Importantly, the H3K9me3 signals at the act-9/35 MSR variant are lost in the absence of Suv39h enzymes (Supplementary Fig. 2b).
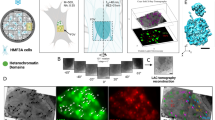
The enriched H3K9me3 signals at the act-9/35 MSR variant could suggest an intrinsic activity for this MSR sequence in guiding H3K9me3 methylation. However, the act-9/35 MSR variant is flanked by other repeat elements, such as a fragmented LTR retrotransposon and remnants of LINE repeats (Supplementary Fig. 2a), which also have been shown to be targets for H3K9 KMT enzymes35. To directly examine an intrinsic potential of MSR sequences in guiding H3K9me3, we developed a highly reductionist approach, in which we insert isolated MSR units into an inert chromosomal region that is repeat- and gene-free. We interrogated the mouse genome to identify genomic regions that have a minimal window of 20–50 kb lacking any annotated repeat or gene sequences. We then compared the epigenetic profile36,37 of these repeat- and gene-free regions and selected an intergenic region of chromosome 2 at position 116 Mbp (Chr2/116), which is largely devoid of DNA and chromatin modifications and has no RNA output (Supplementary Fig. 3a). In addition, the Chr2/116 region is in open chromatin38 and highly accessible for DNase-seq (Supplementary Fig. 3a). Neighbouring genes are 500 kb (Meis2) or 350 kb (Tmco5) 5’ or 3’ distal to the Chr2/116 region and are inactive. We therefore consider the Chr2/116 intergenic region as an inert genomic region that has a 20 kb repeat-free window (Fig. 1a).
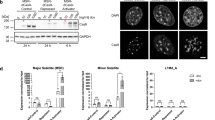
a UCSC genome browser shot of the intergenic region in mouse chromosome 2 at position 116 Mbp (Chr2/116 insertion site) that has a 20 kb repeat- and gene-free window. b Diagram of Chr2/116 homology arm DNA constructs having a control DNA sequence (human buffer, HB) (750 bp) or one or three copies of the per-3/99 (300 bp) or the act-9/35 (261 bp) MSR variant units. In the diagrams for the MSR variants, the red lines illustrate point mutations as compared to the MSR DNA consensus sequence. These constructs were inserted into the Chr2/116 integration site by hit-and-run CRISPR/Cas9 technology. Genotyping analysis for the validation of correct Chr2/116 insertions for per-3/99 and act-9/35 MSR variant copies in homozygous mESC clones. c Left panel: ChIP-qPCR for H3K9me3 with construct-specific T7 and T3 primers across the per-3/99 and act-9/35 MSR insertions. Asterisks indicate statistically significant differences compared with the HB control (mean±SD) (****p < 0.0001, one-way ANOVA, Dunnett’s test). n = 3 independent experiments. Right panel: RT-qPCR for RNA output in the 5’ and 3’ Chr2/116 homology arms of per-3/99 and act-9/35 MSR insertions. Expression is normalised to Hprt and is relative to HB control (mean±SD). (****p < 0.0001, one-way ANOVA, Dunnett’s test). n = 3 independent experiments.
We applied a hit-and-run CRISPR/Cas9 technology to insert one or three copies of either a truncated act-9/35 MSR variant (261 bp) or the per-3/99 MSR variant (300 bp) (Supplementary Fig. 4) into the inert Chr2/116 region. The MSR variant copies were cloned in opposite orientations as compared to their endogenous locations (Supplementary Fig. 2a) and multiple MSR units are organised in 5’ to 3’ tandem repeat arrangements (Fig. 1b). As a control sequence (human buffer), we used a 750 bp DNA sequence from an intergenic region of human chromosome 21 (Supplementary Fig. 5a). Homology arm recombination constructs were modified with short oligonucleotides that serve as insert-specific primers and also provide binding sites for the T7 and T3 RNA polymerase (Fig. 1b). The Chr2/116 insertion site is heterozygous in the used mouse ES cells, with one allele carrying two internal deletions (Supplementary Fig. 3b). Following CRISPR/Cas9-mediated recombination of the insertion constructs, we therefore selected homozygous mouse ES cell clones (Fig. 1b).
Homozygous mouse ES cell clones carrying insertions for the human buffer (HB), one or three copies of the act-9/35 MSR variant and one or three copies of the per-3/99 MSR variant were processed for H3K9me3 ChIP-qPCR with the insert-specific T7 and T3 primers. No H3K9me3 enrichment over background levels could be detected for the HB control, the per-3/99 MSR variant (one or three copies) and for one copy of the act-9/35 MSR variant. Only for the ES cell clone with three copies of the act-9/35 MSR variant did we observe enriched H3K9me3 signals across the integration site (Fig. 1c, left panel).
We also examined whether the isolated MSR variants are transcriptionally competent and could generate MSR-originating transcripts. Only for the ES cell clone with a three-copy insertion of the act-9/35 MSR variant did we detect MSR-derived transcripts that extend into the 5’ Chr2/116 homology arm (Fig. 1c, right panel), possibly reflecting the preferred generation of MSR forward transcripts39. These data would be consistent with a transcription factor based15 and transcription-coupled model for intact MSR sequences in guiding H3K9me3. Indeed, the act-9/35 MSR variant (8 point mutations as compared to the MSR DNA consensus) maintains most of the MSR embedded transcription factor (TF) binding sites, whereas the per-3/99 MSR variant ( > 48 point mutations as compared to the MSR DNA consensus) has lost all (Supplementary Fig. 4). Intriguingly, the number and the positions of TTC triplets (around 20)16 appear unaltered in both the act-9/35 and the per-3/99 MSR variants (Supplementary Fig. 4).
Multiple MSR DNA consensus units establish a heterochromatin island and generate bi-directional RNA
We next used the 234 bp MSR DNA consensus sequence12 (Supplementary Fig. 4), which maintains all TF binding sites and should have full transcriptional competence. We designed Chr2/116 insertion constructs that have one MSR copy (MSR1), three MSR copies (MSR3) or nine MSR copies (MSR9) (Fig. 2a) and generated two independent homozygous ES cell clones for each insertion construct (Fig. 2b).
a Diagram of Chr2/116 homology arm DNA constructs having a control DNA sequence (human buffer, HB) or one, three or nine copies of the 234 bp MSR DNA consensus unit. b Genotyping analysis for the validation of correct insertions in two independent homozygous mESC clones, with clone identities indicated by the numbers. c ChIP-qPCR for H3K9me3 with construct-specific T7 and T3 primers in two independent mESC clones for MSR1, MSR3 and MSR9 insertions. Asterisks indicate statistically significant differences when compared to the HB insertion (mean±SD) (****p < 0.0001, ns = not significant, one-way ANOVA, Dunnett’s test). n = 3 independent experiments. d ChIP-qPCR for H3K9me3 with a primer walk into 4 kb 5’ and 3’ flanking regions distal to the Chr2/116 insertion site to examine extension of an H3K9me3 domain. This experiment was done with one mESC clone for MSR1, MSR3 and MSR9 insertions and with WT26 mES cells (mean±SD). n = 3 independent experiments. e RT-qPCR for RNA output in the 5’ and 3’ Chr2/116 homology arms in two independent mESC clones for HB, MSR1 and MSR3 insertions. Expression is normalised to Hprt and is relative to HB control (mean±SD). n = 3 independent experiments. On the right, qPCR cycle 40 products are validated by gel analysis. f ChIP-qPCR for enrichment of (total) RNA polymerase II. Asterisks indicate statistically significant differences compared with the HB control (mean±SD) (*p = 0.0354, **p = 0.0063, ****p < 0.0001, ns = not significant, one-way ANOVA, Dunnett’s test). n = 3 independent experiments.
H3K9me3 ChIP-qPCR with the insert-specific T7 and T3 primers indicated significant enrichment for the MSR3 and MSR9 DNA consensus repeats (Fig. 2c) that exceeded the H3K9me3 signal observed with the three-copy insertion of the act-9/35 MSR variant (Fig. 1c, left panel). No H3K9me3 enrichment above the HB control levels could be detected for the MSR1 insertions.
The ChIP-qPCR with the insert-specific T7 and T3 primers only probes for enrichment across sequences that are internal to the T7 and T3 marker sites. We used a primer walk (at 1 kb intervals) to examine H3K9me3 enrichment up to 4 kb both into the 5’ and 3’ vicinity of the Chr2/116 insertion site that expands the analysis to also include sequences flanking the homology arms (e.g. for the distal P2 (2 kb), P3 (3 kb) and P4 (4 kb) locations) (Fig. 2d, top diagram). No H3K9me3 enrichment across this 8 kb chromatin window above background wild type levels was detected for the MSR1 insertion. The MSR3 insertion had a focal H3K9me3 enrichment with elevated H3K9me3 signals within 1 kb 5’ (5’P1) and 1 kb 3’ (3’P1) to the insertion site. Intriguingly, the MSR9 insertion induced an extended H3K9me3 domain with significantly increased H3K9me3 signals spanning from the insertion site to the distal 5’P4 and 3’P4 locations (Fig. 2d). This result suggests that increasing the number of MSR units will enhance the potential of MSR DNA to nucleate a heterochromatin island and would be consistent with a model in which constitutive heterochromatin is extended and stabilised by the very high reiteration of MSR units (up to > 10,000 copies) found in the pericentric regions of mouse chromosomes.
We next analysed the transcriptional competence of the MSR1, MSR3 and MSR9 insertions by using RT-qPCR for the detection of MSR-originating transcripts that would extend into the 5’P0 and 3’P0 Chr2/116 homology arm sequences. Only for the MSR3 and MSR9 insertions, but not for the MSR1 or the HB control insertions, did we observe the generation of Chr2/116 transcripts. These MSR-originating transcripts are bi-directional, and MSR3-derived transcripts have a greater abundance as compared to MSR9-derived transcripts (Fig. 2e). In addition, we also performed ChIP-qPCR for the detection of (total) RNA polymerase II (RNAPII) that showed enrichment only across the MSR3 and MSR9 insertions, with higher signals of (total) RNAPII for the MSR3 insertions (Fig. 2f). MSR instructed transcription is specifically regulated by RNAPII, as no signals for RNAPI or RNAPIII could be detected across MSR insertions (Supplementary Fig. 6).
LINE 5’UTR repeat units do not establish heterochromatin at the inert genomic region
An important next question was to address whether H3K9me3 at the Chr2/116 insertion site could also be induced by other repeat sequences or transcriptionally competent regulatory elements. A subfamily of LINE elements (LINE L1Md_A) and ERV retrotransposons (IAP) have been shown to be targets for the SETDB1 KMT40 and are silenced by the KRAB Zn-finger (TRIM28) repressor complex41. Suv39h enzymes also target LINE L1Md_A repeat elements35. The 5’ untranslated region (5’UTR) of LINE L1Md_A is composed of a reiterating number of a 208 bp G/C-rich repeat unit that directs LINE L1Md_A transcription42. We also included the CMV promoter of the human cytomegalovirus that has been well studied as an expression element43. The CMV core promoter resides within a 228 bp long DNA sequence that has a bona fide TATA box and a defined transcriptional start site (TSS) (Supplementary Fig. 5b).
We designed Chr2/116 insertion constructs that have three copies of the LINE L1 Md_A 5’UTR repeat (L5’UTR3) or three copies of the CMV promoter (CMV3) and generated two independent homozygous ES cell clones for each insertion construct (Fig. 3a). We then compared these L5’UTR3 and CMV3 insertions with the already established MSR3 insertions. H3K9me3 ChIP-qPCR with the insert-specific T7 and T3 primers indicated intermediate enrichment for the CMV3 insertions relative to the higher H3K9me3 enrichment for the MSR3 insertions. No H3K9me3 signals above the HB control levels could be detected for the L5’UTR3 insertions (Fig. 3b, left panel). A similar profile was also observed for another heterochromatic histone mark H4K20me3 (Supplementary Fig. 7). Consistent with the different H3K9me3 levels, some enrichment for HP1α was detected across the CMV3 insertion in one of the two CMV3 ES cell clones, but was significantly augmented for the MSR3 insertions. No HP1α signal could be found across the L5’UTR3 insertions (Fig. 3b, middle panel).
a Diagram of Chr2/116 homology arm DNA constructs having a control DNA sequence (human buffer, HB), three copies of a 208 bp LINE 5’UTR repeat unit (L5’UTR3), three copies of the 228 bp CMV promoter (CMV3) or three copies of the 234 bp MSR consensus unit (MSR3). Genotyping analysis for the validation of correct insertions in two independent homozygous mESC clones, with clone identities indicated by the numbers. b ChIP-qPCR for H3K9me3 (left panel), HP1α (middle panel) and histone H1 (right panel) with construct-specific T7 and T3 primers in two independent mESC clones for HB, L5’UTR3, CMV3 or MSR3 insertions. Asterisks indicate statistically significant differences when compared to the HB control (mean±SD) (**p = 0.0042, ****p < 0.0001, ns = not significant, one-way ANOVA, Dunnett’s test). n = 3 independent experiments. c T7 promoter accessibility assay with one mESC clone for HB, L5’UTR3, CMV3 or MSR3 insertions (left panel), or for HB, MSR1, MSR3 and MSR9 insertions (right panel). Isolated nuclei were incubated with recombinant T7 RNA polymerase and RNA was detected by RT-qPCR with primers specific for the 3’ Chr2/116 homology arm. Asterisks indicate statistically significant differences when compared to the HB control (mean±SD) (****p < 0.0001, one-way ANOVA, Dunnett’s test). n = 3 independent experiments. d Left panel: T7 promoter accessibility assay with one mESC clone for HB, MSR1, MSR3 and MSR9 insertions. Asterisks indicate statistically significant differences when compared to the HB control (mean±SD) (****p < 0.0001, one-way ANOVA, Dunnett’s test). n = 3 independent experiments. Right panel: ChIP-qPCR for histone H1 with construct-specific T7 and T3 primers in one mESC clone for HB, MSR1, MSR3 and MSR9 insertions. Asterisks indicate statistically significant differences when compared to the HB control (mean±SD) (****p < 0.0001, one-way ANOVA, Tukey’s test) or for statistically significant differences between MSR3 and MSR9 insertions (**p = 0.0051, one-way ANOVA, Tukey’s test). n = 3 independent experiments.
In addition to repressive histone modifications and binding of HP1 proteins, another hallmark of heterochromatin is a more compact and less accessible chromatin structure3. We used ChIP-qPCR for the linker histone H1 and detected histone H1 enrichment specifically and only across the MSR3 insertions (Fig. 3b, right panel). We next probed for chromatin accessibility by adopting an approach that is using incubation of nuclei with recombinant T7 RNA polymerase44. The accessibility of chromatin targets that have an engineered binding site for T7 RNA polymerase (T7 RNAP) can be quantified by the production and amount of T7 RNAP directed transcripts. All of the Chr2/116 insertion constructs have engineered binding sites for T7 (and T3) RNA polymerases. We used this T7 RNAP assay and observed a high level production of T7 RNAP directed transcripts with the HB control insertions (Fig. 3c), consistent with the Chr2/116 region to be in highly accessible chromatin (Supplementary Fig. 3a). A comparable high-level production of T7 RNAP directed transcripts was also detected for the L5’UTR3 and CMV3 insertions. By contrast, the amount of T7 RNAP directed transcripts was significantly reduced ( > 3-fold) for the MSR3 insertions (Fig. 3c) and even further decreased ( > 7-fold) for the MSR9 insertions (Fig. 3d, left panel). In addition, MSR9 insertions also have increased signals for histone H1 incorporation as compared to MSR3 insertions (Fig. 3d, right panel). These data indicate a progressively less accessible chromatin structure with expanded numbers of MSR DNA units. These results could explain why there is reduced RNAPII recruitment (Fig. 2f) and decreased levels of MSR-derived transcripts (Fig. 2e) in the MSR9 insertions as compared to the MSR3 insertions.
MSR-originating transcripts have non-mRNA qualities
Our reductionist approach dissects factor access and transcriptional competence of regulatory DNA sequences outside the context of a canonical gene body architecture and without engaging splicing control of exon/intron boundaries or 3’ end processing of poly-A transcripts. In fact, there are no cryptic exons or bona fide poly-adenylation (poly-A) sequences within the 4 kb vicinity 5’ or 3’ to the Chr2/116 insertion site. Accordingly, the majority of transcripts that could originate from the Chr2/116 integration site and extend within the Chr2/116 region should not be exported but rather remain chromatin associated. Indeed, sub-cellular fractionation of RNA from mESC clones carrying the HB, L5’UTR3, CMV3 and MSR3 insertions indicates that any of the insert-derived Chr2/116 transcripts (if they are generated) are primarily chromatin associated (Supplementary Fig. 8).
However, there are significant differences for Chr2/116 RNA output that are dependent on whether Chr2/116 transcripts originate from L5’UTR3, CMV3 or MSR3 regulatory units. First, RT-qPCR from total RNA preparations indicates no Chr2/116 transcripts for the HB control insertions, but high level 5’ forward Chr2/116 transcripts for the L5’UTR3 and CMV3 insertions that are in the direction of the described transcriptional start sites (TSS) for the LINE 5’UTR or the CMV promoter units (Fig. 4a). By contrast, MSR3-originating Chr2/116 transcripts are detected in both the 5’ and 3’ direction and their abundance is greatly attenuated ( > 5-fold reduced) as compared to the L5’UTR3 and CMV3-derived Chr2/116 transcripts (Fig. 4a). Second, ChIP-qPCR analysis for the initiating (RNAPII-Ser5phos) or the elongating (RNAPII-Ser2phos) forms of RNAPII with primers in the 5’ vicinity (5’P0–5’P3 locations) of the Chr2/116 insertion site reveals significantly less initiating RNAPII-Ser5phos signal for the MSR3 insertions (as compared to the L5’UTR3 and CMV3 insertions) that is progressively lost and no longer detectable at the 5’P3 position (Fig. 4b, left panels). A modest signal for initiating RNAPII-Ser5phos in the 3’ vicinity of the Chr2/116 integration site is exclusively detected for the MSR3 insertion at the directly adjacent 3’P0 position, but again lost at more distal locations. Strikingly, enrichment for the elongating form of RNAPII-Ser2phos was observed only for the L5’UTR3 and CMV3 insertions, but was not found for the MSR3 insertion, neither at the immediate 5’ or 3’ vicinity of the Chr2/116 integration site (Fig. 4b, right panels). Third, using the same P0 – P3 primers in the 5’ and 3’ vicinity of the Chr2/116 integration site, we performed RT-qPCR to inspect the length of insert-derived Chr2/116 transcripts that are present in chromatin associated RNA purifications. Chromatin associated Chr2/116 transcripts progressively decline from the 5’P0 position but extend to the 5’P3 location for L5’UTR3- and CMV3-derived Chr2/116 transcripts. By contrast, MSR3-derived Chr2/116 transcripts cannot be detected at the 5’P3 or the 3’P3 locations (Fig. 4c). Fourth, bulk MSR RNA has been shown to have non-mRNA qualities, including the absence of a 5’ cap and largely lacking poly-A tails39. We used the terminator 5’ exonuclease that will degrade 5’ uncapped RNA to probe for sensitivity of insert-derived Chr2/116 transcripts to digestion with terminator 5’ exonuclease. Only for the MSR3-originating Chr2/116 transcripts (both for the 5’ and 3’ direction) did we observe degradation by the terminator 5’ exonuclease (Fig. 4d). Collectively, these data indicate that MSR-originating transcripts are bi-directional and highly attenuated, remain chromatin associated, have a short transcript length and lack a 5’ cap.
a RT-qPCR to detect insert-derived Chr2/116 transcripts in two independent mESC clones for human buffer (HB), L5’UTR3, CMV3 or MSR3 insertions. Expression is normalised to Hprt and is relative to HB control (mean±SD). n = 3 independent experiments. On the right, qPCR cycle 40 products are validated by gel analysis. b ChIP-qPCR for initiating RNA polymerase II (RNAPII-Ser5phos) (left panels) and for elongating RNA polymerase II (RNAPII-Ser2phos) in the 5’ and 3’ vicinity (P0, P1 and P3 locations) of one mESC clone for HB, L5’UTR3, CMV3 or MSR3 insertions. Asterisks indicate statistically significant differences compared with the HB control (mean±SD) (*p = 0.0117, **p = 0.0096, ***p < 0.001, ****p < 0.0001, one-way ANOVA, Dunnett’s test). n = 3 independent experiments. c RT-qPCR with flanking primers (0 kb, 1 kb and 3 kb 5’ or 3’ distal to the Chr2/116 integration site) to examine transcript length of chromatin-associated RNA in one mESC clone for HB, L5’UTR3, CMV3 or MSR3 insertions. Expression is normalised to Malat1 and is relative to HB control (mean±SD). n = 3 independent experiments. d Analysis of 5’end capping (degradation by terminator 5’exonuclease) of Chr2/116 transcripts originating form L5’UTR3, CMV3 or MSR3 Chr2/116 insertions. Asterisks indicate statistically significant differences when compared to untreated (without terminator 5’exonuclease) control (mean±SD) (****p < 0.0001, two-way ANOVA, Šídák’s test). n = 3 independent experiments.
MSR-originating transcripts are attenuated by the integrator complex
The non-mRNA qualities of MSR-originating transcripts described above could suggest an adapted transcriptional activity that would involve more than the RNAPII holoenzyme. The Integrator complex can associate with RNAPII and has been described to process small nuclear RNA45. Functions of the Integrator complex include 5’ and 3’ processing of nascent non-coding RNA, regulation of RNAPII pause-release and promoter-proximal termination of RNAPII-derived transcripts46,47,48. The Integrator is a metazoan-specific multimeric complex, which, among other subunits, has an endonuclease component (INTS11) and a PHD-finger containing module (INTS12) that can recognise histone H3 methylation46. The Integrator complex has a preference to interact with A/T-rich sequences that is further enhanced by a stem-loop configuration of the target DNA46. In addition, the structural definition of the Integrator49 revealed association not only with RNAPII, but, in a post-termination super-complex, also with the sensor of single-stranded DNA (SOSS)50.
We performed ChIP-qPCR to detect INTS11 and INTS12 enrichment across the different Chr2/116 insertions. Only for the MSR3 insertion, but not for the L5’UTR3 and CMV3 insertions, did we identify INTS11 and INTS12 signals that were significantly above the HB control levels (Fig. 5a). We next decreased INTS11 by siRNA and analysed relative abundance of insert-specific Chr2/116 transcripts 48 hrs after Ints11 knock-down. MSR3-derived Chr2/116 transcripts (both in the 5’ and 3’ direction) were significantly elevated ( > 4-fold), whereas levels of L5’UTR3- and CMV3-derived Chr2/116 transcripts were unresponsive to Ints11 knock-down (Fig. 5b). These data expose the Integrator as a novel regulator for the processing of MSR transcripts. Intriguingly, the enrichment of MSR3-derived Chr2/116 transcripts following the Ints11 siRNA occurs without a reduction of H3K9me3 and only a modest decrease of HP1α across the Chr2/116 insertion site (Supplementary Fig. 9).
a ChIP-qPCR for INTS11 and INTS12 at the Chr2/116 integration site in two independent mESC clones for HB, L5’UTR3, CMV3 or MSR3 insertions. Asterisks indicate statistically significant differences when compared to the HB control (mean±SD) (***p = 0.0002, ****p < 0.0001, ns = not significant, one-way ANOVA, Dunnett’s test). n = 3 independent experiments. b RT-qPCR transcript analysis at the Chr2/116 integration site in one mESC clone for HB, L5’UTR3, CMV3 or MSR3 insertions 48 hrs after knock-down of Ints11. Asterisks indicate statistically significant differences when compared to the negative control (unspecific siRNA knock-down) (mean±SD) (****p < 0.0001, ns = not significant, two-way ANOVA, Šídák’s test). n = 3 independent experiments. c Heat maps of RNA-seq (total RNA, 150 bp paired-end) for MSR transcripts (left panel), and ChIP-seq for H3K9me3 (middle panel) and HP1α (right panel) in mouse ES cells. Data are shown at 0 h and 24 h following depletion of INTS11-degron. MSR sequences were classified into two groups based on sequence identity (95–100% conservation to MSR DNA consensus and below 95% conservation to MSR DNA consensus). Within each group, the signals were ordered according to their level of enrichment. Western blot analysis for the depletion of INTS11 is shown on top. Blot was probed with α-HA antibody and GAPDH was used as loading control. MSR transcripts and ChIP-seq reads are aligned and binned according to their sequence identity (decreasing from 100% to 95% and then to 75%) with the MSR DNA consensus sequence. Two biological replicates are shown for each analysis.
We next explored the scope of MSR transcriptional regulation by the Integrator complex. For this, we first established a bioinformatic pipeline that allows for the de novo assembly of MSR repeat regions from long-read DNA sequencing of mouse ES cell genomic DNA. With this de novo assembly, we could build 89 MSR containing DNA contigs that together provide sequence information for 18,018 MSR variants (Supplementary Fig. 10). We then used an engineered mouse ES cell line that expresses a degron-tagged INTS11 endonuclease (INTS11-3HA-ecDHFR) from the endogenous Ints11 locus. HiSeq RNA sequencing (150 bp paired-end) from total RNA was performed 0 or 24 h after depletion of the INTS11-degron. The HiSeq RNA sequencing data were filtered for MSR transcripts and the MSR RNA reads were aligned to our assembly of 18,018 MSR DNA variants. This alignment was binned according to sequence identity with the MSR DNA consensus sequence and is represented as a heat map starting with high sequence identity (100–95%) of MSR transcripts to the MSR DNA consensus and then gradually decreasing to MSR variants with only 95−75% identity. The data show that MSR transcript levels were considerably increased for the segment of intact MSR units (100–95% identity with the MSR DNA consensus) after INTS11-degron depletion, whereas for the great majority of permutated MSR variants ( < 95% identity with the MSR DNA consensus) only very few or no MSR transcripts were found (Fig. 5c, left panel). These data are coherent with the enrichment of INTS11 at intact MSR units that we can detect by ChIP-seq in wt mESC (Supplementary Fig. 11). As described above for the MSR3 Chr2/116 insertions, elevated MSR transcript levels following INTS11-degron depletion occur without alterations for H3K9me3 (Fig. 5c, middle panel). Interestingly, HP1α is specifically decreased for the segment of intact MSR units but not over permutated MSR variants (Fig.5c, right panel). We conclude from these analyses that around 10-15% of MSR units in the mouse epigenome preserve transcriptional competence. The data further indicate that RNA output at these transcriptionally competent MSR units is attenuated by the Integrator complex, which also appears to modulate HP1α association.
Multi-copy MSR units expose an unwound DNA template
The MSR DNA repeat unit is full of embedded TF binding sites (Supplementary Fig. 4), but has no canonical promoter architecture. Still, the A/T-richness and proposed non-B form configuration of MSR DNA51 could mimic a promoter-like activity, either by exposing single-stranded DNA or by enabling RNA:DNA hybrid or R-loop formation, both of which have been suggested to imitate RNAPII engagement and unwinding of the DNA template52. Intriguingly, R-loops not only induce anti-sense transcription52 but also contribute to RNAPII pause site termination in protein-coding genes53. We performed ChIP-qPCR with recombinant RPA-eGFP (Supplementary Fig. 12) to detect single-stranded DNA and also used ChIP-qPCR with recombinant HBD-eGFP54 to probe for RNA:DNA hybrids across the different Chr2/116 insertions. While no enrichment was observed across the human buffer (HB) control, significant and comparable levels of RPA-eGFP signals were found for the L5’UTR3, CMV3 and MSR3 insertions, suggesting a transcriptionally unwound DNA template (Fig. 6a). In addition, only for the MSR3 insertion did we identify robust accumulation of RNA:DNA hybrids (Fig. 6b).
a ChIP-qPCR with recombinant RPA-eGFP to detect exposed single-stranded DNA across human buffer (HB), L5’UTR3, CMV3 and MSR3 Chr2/116 insertions. Asterisks indicate statistically significant differences for RPA-eGFP signals compared with the HB control (mean±SD) (*p ≤ 0.0.0302, **p = 0.0022, one-way ANOVA, Dunnett’s test). n = 3 independent experiments. b ChIP-qPCR with recombinant HBD-eGFP to detect RNA:DNA hybrids across HB, L5’UTR3, CMV3 and MSR3 Chr2/116 insertions. Asterisks indicate statistically significant differences for HBD-eGFP signals compared with the HB control (mean±SD) (****p < 0.0001, one-way ANOVA, Dunnett’s test). n = 3 independent experiments. c Diagram of Chr2/116 homology arm DNA constructs having a control DNA sequence (human buffer, HB) or one, two, three or nine copies of the 234 bp MSR DNA consensus unit. d ChIP-qPCR for RPA-eGFP with construct-specific T7 and T3 primers across HB, MSR1, MSR2, MSR3 and MSR9 Chr2/116 insertions. Asterisks indicate statistically significant differences for RPA-eGFP signals compared with the HB control (mean±SD) (**p = 0.0029, ****p < 0.0001, one-way ANOVA, Dunnett’s test). n = 3 independent experiments. e ChIP-qPCR for HBD-eGFP with construct-specific T7 and T3 primers across HB, MSR1, MSR2, MSR3 and MSR9 Chr2/116 insertions. Asterisks indicate statistically significant differences for HBD-eGFP signals compared with the HB control (mean±SD) (**p = 0.0015, ****p < 0.0001, one-way ANOVA, Dunnett’s test). n = 3 independent experiments. f RT-qPCR to detect insert-derived 5’ and 3’ Chr2/116 transcripts in one mESC clone for HB, MSR1, MSR2, MSR3 and MSR9 insertions following a 24 hrs incubation with etoposide. Expression is normalised to Hprt and is relative to HB control (mean±SD). Asterisks indicate statistically significant differences when compared to DMSO (without etoposide) control (mean±SD) (****p < 0.0001, two-way ANOVA, Šídák’s test). n = 3 independent experiments.
It remained an open question why single copy Chr2/116 insertions of an intact MSR unit are inactive to nucleate heterochromatin formation. To address this, we also generated two copy insertions of the MSR consensus sequence (MSR2) at the Chr2/116 integration site. With this, we can compare MSR1, MSR2, MSR3 and MSR9 insertions (Fig. 6c) and probe for their potential to generate transcripts and instruct a heterochromatic signature. As with MSR1 insertions, two copies of the MSR consensus sequences are not transcriptionally competent and did not induce H3K9me3 (Supplementary Fig. 13). We also did not detect enriched RPA-eGFP signals at MSR1 or MSR2 insertions, which, however, are evident for the MSR3 and, at even higher levels, for the MSR9 insertions (Fig. 6d). Similarly, ChIP-qPCR with recombinant HBD-eGFP identified the presence of RNA:DNA hybrids for the MSR3 and MSR9 insertions, but not for MSR1 or MSR2 insertions (Fig. 6e). Recently, endogenous MSR sequences were shown to be under topological control, since MSR transcription can be stimulated by topoisomerase inhibition that would allow more non-B form DNA55. We thus examined whether the Chr2/116 MSR insertions also respond to topoisomerase dysfunction by using the topoisomerase II (TOP2) poison etoposide. Only for MSR3 and MSR9 insertions, but not for MSR1 and MSR2 insertions did we detect significantly elevated 5’ and 3’ Chr2/116 transcripts following exposure to etoposide (Fig. 6f). Together, these data validate that only multi-copy insertions of the MSR consensus unit can induce an altered topology of chromatin-embedded A/T-rich MSR repeat DNA that is reflected by an unwound DNA template and bi-directional transcriptional activity responsive to topoisomerase inhibition.
Discussion
We show that isolated MSR units can nucleate heterochromatin at an inert genomic region (Chr2/116) in the mouse epigenome that is repeat- and gene-free. In addition, we identify the Integrator as a novel regulator to control MSR RNA output. Collectively, the data allow for a model connecting an altered topology and promoter-mimic activity of the MSR DNA template with bi-directional and attenuated transcription and in which MSR RNA remains chromatin associated and contributes to a more compact nucleosome structure that also incorporates histone H1 (Fig. 7). This model is focused on constitutive heterochromatin in mouse ES cells and, for reasons of clarity, does not detail the important functions of DNA methylation56, RNA interference, histone variants deposition, nucleosome remodeling and additional heterochromatin principles2,3,22. It also does not explain facultative heterochromatin or other forms of a repressed or silenced chromatin state at gene promoters57.
In this model, multi-copy units of the A/T-rich MSR DNA consensus sequence are intrinsically prone to form an altered topology that exposes single-stranded DNA and/or favours R-loop formation (indicated by the red and blue extensions). This partially unwound DNA template can serve as a promoter-mimic and facilitates RNA polymerase II (RNAPII) engagement, which is further guided by transcription factor (TF) binding to MSR DNA. MSR-derived transcription is bi-directional and attenuated by the RNAPII-associated Integrator (INT) complex. MSR transcripts (wavy dashed lines) have non-mRNA qualities and largely remain chromatin associated. MSR transcripts assist in the recruitment of HP1 and Suv39h enzymes and can stabilise an RNA-nucleosome scaffold. Heterochromatin establishment is matured by robust Suv39h-mediated H3K9me3 (Me), HP1 binding and histone H1 (H1) incorporation, which together form a more compact nucleosome structure and silence transcriptional activity. Abbreviations are: MSR (major satellite repeat), TF (transcription factor), Pol II (RNA polymerase II), INT (Integrator complex), Suv39h (H3K9 KMT), Me (H3K9me3), HP1 (heterochromatin protein 1), RNA (chromatin-associated MSR transcripts), H1 (histone H1). See text for detailed explanation.
The Integrator, like RNAPII, is not a core component of heterochromatin. While the Integrator is present at endogenous (intact) MSR units, it is also detected at other repeat classes (e.g. ERV retrotransposons)58, genes and many regulatory sites in the mouse epigenome59. Further, INTS11 depletion does not change a H3K9me3 signature but selectively reduces HP1α at transcriptionally competent MSR units. While it has been shown that H3K9me3 can persist even after the Suv39h enzymes are removed60, it is currently not resolved whether the decrease of HP1α is triggered by altered protein-protein contacts or by increased RNA output, which could modulate chromatin association of HP1α61.
From our contig analysis of 18,018 MSR sequences, we estimate that less than 15% of MSR units maintain high sequence identity to the MSR consensus sequence and preserve transcriptional competence, and this quality is sufficient to instruct de novo heterochromatin formation when transcriptionally competent MSR units are isolated and inserted at an inert genomic location. We thus consider transcriptionally competent MSR units as heterochromatin nucleating MSR copies. By contrast, highly permutated MSR variants are transcriptionally inert and cannot instruct de novo heterochromatin formation. There is also a third subpopulation of MSR sequences, which are moderately permutated and are positive for H3K9me3 and HP1α but lack an alignment for MSR transcripts (Fig. 5c). These intermediate MSR variants are often interspersed with heterochromatin nucleating MSR units in the complex arrangements of pericentric MSR arrays (Supplementary Fig. 11). Consistent with the proposed models for a self-enforcing heterochromatin state3,22,62,63, heterochromatin nucleating MSR units would initiate heterochromatin establishment, which is then extended into adjacent regions including moderately permutated MSR variants.
Classic work in Drosophila has shown that expansions of transgene repeats can cause gene silencing via the induction of repressive chromatin64. It has been proposed that topological pairing of transgene copies could underlie this repeat-dependent gene silencing and contribute to heterochromatin formation64. An intriguing result from our reductionist approach is that single or two copy Chr2/116 insertions of an MSR unit are not sufficient to induce H3K9me3. For the three copy Chr2/116 MSR insertions, the DNA units are positioned in tandem (head-to-tail) arrangements. Tandem repeat arrangements favour non-B form DNA51 that is further enhanced by A/T-richness of the DNA sequence. However, A/T-richness or the presence of TTC triplets, per se, do not appear to be a sufficient quality, as act-9/35 and per-3/99 MSR variants have comparable A/T-richness and a similar number and positioning of TTC triplets. The per-3/99 MSR variant has lost all TF binding sites and cannot promote H3K9me3, strongly supporting the necessary function of transcription factor binding for MSR-dependent heterochromatin formation15. To what extent TF binding or HP1 association would recruit heterochromatin components independently of MSR transcript production is currently not resolved. The three copy tandem insertions of an MSR unit also provide for a longer DNA template (potentially spanning four to five nucleosomes) that could present more non-B form DNA and/or would have an extended ability to form exposed single-stranded DNA and RNA:DNA hybrids. Consistent with a proposed role of RNA:DNA hybrids or R-loops to mimic a promoter-like activity52, we would like to interpret the data to suggest that three copies, but not one or two copies, of an MSR unit can expose sufficient topological alterations, facilitate RNAPII engagement further promoted by TF binding and allow for bi-directional transcription that is then sensed by the Integrator complex and silencing machinery. Notably, R-loops also induce repressive H3K9 methylation at RNAPII pause site termination regions53.
The non-mRNA qualities of MSR-derived transcripts and their attenuation by the Integrator complex provide unanticipated mechanistic insight into the regulation of heterochromatin in mammalian cells. They also suggest that the Integrator complex and heterochromatin could function as RNA quality machines that can sense compromised (uncoordinated, less processive and/or bi-directional) transcriptional activity and discriminate non-coding vs. coding transcription46,47. The RNA quality control mechanisms for heterochromatin formation described in this study appear to be particularly relevant for repeat-rich genomes (such as e.g. mammalian cells) and are regulated by other RNA quality factors in the repeat-poor genomes of unicellular organisms (such as e.g. S. pombe)23,25. Intriguingly, the cleavage and polyadenylation specificity factors (CPSF) that cooperate in promoting heterochromatin assembly in S. pombe24 are orthologous to the INTS9/INTS11 subunits of Integrator46.
Chromatin-associated MSR RNA has been shown to stabilise a RNA-nucleosome scaffold39. Suv39h enzymes can bind MSR RNA39,65 and were also reported to interact with histone H166. MSR3-derived transcripts are co-regulated by the Integrator complex and it is possible, although currently not examined, that components of the Integrator complex could bridge and facilitate an RNA-Suv39h-histone H1 nucleosome interface. Whether this nucleosome interface would be further strengthened by m6A RNA methylation54,67, if present on the MSR3-derived Chr2/116 transcripts, or whether there is also reduced histone turnover68 remains to be investigated. These are interesting questions that require future studies with isolated MSR or other satellite repeat elements. In sum, our study connects the DNA, RNA and chromatin principles for the de novo establishment of heterochromatin and resolves a long-standing paradox between heterochromatic and euchromatic transcriptional activity.
Methods
Tissue culture and cell lines
Wild type (WT26) mouse embryonic stem cells (mESC) (129/Sv x C57Bl/6 J, male) and Suv39h double-null (DN57) mESC (129/Sv x C57Bl/6 J, male) were generated as previously described69. WT26 and DN57 mESC were cultured on gelatin-coated dishes in High Glucose DMEM (Sigma) supplemented with 15% serum replacement (Thermo Fisher), 2 mM L-glutamine (Sigma), 1X non-essential amino acids (Sigma), 1 mM sodium pyruvate (Sigma), 0.1 mM 2-mercaptoethanol (Gibco), and 1 ml supernatant (per 500 ml medium) from LIF-producing COS-7 cells (in-house production). Cells were cultured at 37 °C in 5% CO2 and were routinely verified to be mycoplasma free.
INTS11-degron mESC were generated by CRISPR/Cas9-mediated knock-in of a fusion construct connecting the full-length INTS11 (600 amino acids) with the E. coli dihydrofolate reductase (ecDHFR) degron. Homozygous replacement of the endogenous Ints11 locus (chromosome 4) by the INTS11-3HA-ecDHFR degron was verified by genotyping59. INTS11-degron mESC were cultured on gelatin-coated dishes in High Glucose DMEM (Sigma) supplemented with 15% FBS (Gibco), 2 mM L-glutamine (Sigma), 1X non-essential amino acids (Gibco), 1 mM sodium pyruvate (Gibco), 100 U/mL penicillin-streptomycin, 0.1 mM 2-mercaptoethanol (Gibco), 1000 U/ml mouse recombinant LIF (Stemcell, 78056) and 10 μM Trimethoprim (Cayman Chemical, 16473). Trimethoprim maintains the stability of the ecDHFR degron.
Identification of repeat-free intergenic regions in the mouse genome
Repeat- and gene-free regions in the mouse mm10 genome70 were identified using a custom Python script. The repeat masker gtf file of the mouse mm10 genome was indexed and all genomic regions that have a 20–50 kb repeat-free window were extracted. Next, the genecode GRCm39 Ensembl GTF file was indexed and iterated through each repeat-free region. Only genomic regions that did not contain genes within and 50 kb distal (5’ and 3’) to a repeat-free region were extracted. The custom Python script can be found at: https://github.com/gerikson/Find_repeatFree_region.
Chr2/116 homology arm DNA constructs and guide RNA-Cas9 plasmids
Donor plasmids containing the various DNA elements flanked by 1 kb 5’ and 3’ homology arms of the Chr2/116 integration site were synthesized by GenScript and cloned into the pUC19 backbone plasmid. All constructs were verified by sequencing and the Chr2/116 homology arm sequences (Supplementary Table 2) and the different insert sequences (Supplementary Table 3) are listed.
For the guide RNA-Cas9 plasmids, four sets of guide RNA (gRNA) targeting +/− 50 bp of the Chr2/116 integration site were identified by the CRISPR gRNA design tool CHOPCHOP71 and cloned into the pSpCas9(BB)−2A-Puro (px459) backbone plasmid (Addgene, 62988). The gRNA sequence, 5’-ACAGAGCACTAAGTCTGACT-3’, showed the highest targeting efficiency (as assessed by Surveyor assay) and this guide RNA-Cas9 plasmid (px1480) was chosen for targeting all Chr2/116 insertions.
Hit-and-run CRISPR/Cas9 insertions at the Chr2/116 integration site and generation of homozygous mES cell clones
Wild-type mESC were grown in six-well plates (3.5 × 105 cells per well) and transfected (Lipofectamine 3000, Thermo Fisher Scientific) with a 3:1 molar ratio (total of 5 μg DNA) of the donor plasmid and the guide RNA-Cas9 px1480 plasmid. 48 h post transfection, cells were selected with 1.25 μg/ml puromycin (Sigma) for 2 days. Puromycin was washed out and surviving cells were allowed to recover reaching near confluency. A cell suspension was prepared and single cells were FACS sorted into 96-well plates. Growing single cell colonies were expanded and cultured for an additional 10-14 days in mESC medium without selection. PCR-genotyping with flanking primers outside the Chr2/116 homology arms (Supplementary Table 4) was performed. To address inherent clonal variability and randomness of CRISPR-mediated repeat insertion at the Chr2/116 locus, a minimum of 2 independent clones verified to contain their respective homozygous insertions were selected for further experimentation. PCR amplicons derived with the flanking primers were sequence verified to ensure that the Chr2/116 insertions were error-free.
Chromatin immunoprecipitation (ChIP)
Directed ChIP was performed as described35. For single cross-linking ChIP, 2.5 × 107 cells were resuspended in 10 ml PBS and incubated with 1% formaldehyde (Sigma) at RT for 15 min. The reaction was quenched by adding 1:20 volume of 2.5 M glycine (Sigma) for 5 min and chromatin was fragmented by sonication. 4 μg of anti-H3K9me3 (Abcam, ab8898) antibody was used for single cross-linking ChIP.
For double cross-linking ChIP, 2.5 × 107 cells were resuspended in PBS and incubated with 2 mM DSG (ChemScene, 79642-50-5) at RT for 45 min. Cells were washed twice with PBS and processed by 1% formaldehyde fixation (15 min at RT) and glycine quenching. Chromatin was fragmented at 4 °C to an average size of 1 kb (ChIP across the Chr2/116 integration site) or 250 bp (ChIP for the Chr2/116 primer walk) using a COVARIS S220 focused-ultrasonicator. For each ChIP reaction, 10 μg of sonicated chromatin was incubated with antibody on a rotating wheel at 4 °C O/N and captured with magnetic Protein G Dynabeads (Thermo Fisher, 10004D). Beads were washed and eluted in EB buffer (1% SDS and 0.1 M NaHCO3) and incubated in the presence of 50 μg RNase A (Thermo Fisher Scientific, EN0531) and 60 μg proteinase K (Thermo Fisher Scientific, EO0491) at 37 °C for 1 h, followed by additional incubation at 65 °C O/N in a Thermomixer (Eppendorf). Decross-linked DNA was purified using a PCR purification kit (Macherey-Nagel, 740609.250). Antibodies for double cross-linking ChIP were: 3 μg of anti-HP1α (Cell Signaling, 2616), 4 μg of anti-Histone H1 (Santa Cruz, sc-8030), 3 μg of anti-Rpb1 NTD (Cell Signaling, 14958), 1 μg of anti-phospho-Rpb1 CTD (Ser5) (Cell Signaling, 13523), 1 μg of anti-phospho-Rpb1 CTD (Ser2) (Cell Signaling, 13499), 5 μg of anti-INTS11 (Abcam, ab75276) and 5 μg of anti-INTS12 (Sigma, HPA035772).
qPCR for Chr2/116 insertions (and primer walk)
ChIP enrichment across the Chr2/116 insertions was analysed by qPCR with 20 ng of input DNA and insert-specific T7 and T3 primers (250 nM each) on a QuantStudio 6 Flex Real-Time PCR System (Applied Biosystems) using PrimeSTAR Max DNA polymerase (Takara Bio, R045) and EvaGreen Dye (Biotium, 31000).
For the ChIP-qPCR primer walk 0, 1, 2, 3 and 4 kb into the 5’ and 3’ homology arm and flanking regions of the Chr2/116 integration sites, ChIP enrichment was analysed by qPCR with 20 ng of input DNA and 250 nM of forward and reverse Chr2/116 primers (2/116: 5’P0, 5’P1, 5’P2, 5’P3, 5’P4; 2/116: 3’P0, 3’P1, 3’P2, 3’P3, 3’P4) using 2X SYBR Select Master Mix (Thermo Fisher Scientific, 4472920). Relative ChIP enrichment was calculated as percentage of input DNA. qPCR Chr2/116 forward and reverse primers are listed in Supplementary Table 4.
RNA extraction and RT-qPCR
Total RNA was extracted from 1 × 106 cells using TRIreagent (Sigma). RNA pellets were resuspended in nuclease-free water (Qiagen) and RNA concentration was measured on a Nanodrop 1000 (Thermo Fisher Scientific). The quality of the RNA was examined by Bioanalyzer RNA 6000 Nano Kit (Agilent). To remove contaminating DNA, 10 μg of the RNA samples were incubated with 4 U of TURBO DNase (Thermo Fisher Scientific, AM2238) at 37 °C for 1 h. RNA was purified with a RNA Clean & Concentrator kit (Zymo Research, R1016). The DNase digestion and RNA purification was performed twice. cDNA was generated by incubating 1 μg of total RNA with 200 U of SuperScript II Reverse Transcriptase (Thermo Fisher Scientific, 18064014), 0.5 mM of dNTP mix (Thermo Fisher Scientific, R0191), and 200 ng of random hexamers (Thermo Fisher Scientific, N8080127) at RT for 10 min, followed by an additional incubation at 42 °C for 50 min. The reaction was stopped by heating at 70 °C for 15 min. The resulting cDNA was stored at −20 °C. 16 ng cDNA was incubated with 2X SYBR Select Master Mix (Thermo Fisher Scientific, 4472920) and 250 nM of forward and reverse primers in a total volume of 10 μl. RT-qPCR was performed using a QuantStudio 6 Flex Real-Time PCR System (Applied Biosystems). Cycle threshold (Ct) values and Hprt expression were used to calculate normalized expression (ΔΔCt). RT-qPCR primers for detection of 5’P0 and 3’P0 Chr2/116 transcripts and for Chr2/116 primer walking (2/116: 5’P0, 5’P1, 5’P3 3’P0, 3’P1, 3’P3) are listed in Supplementary Table 4.
T7 RNA polymerase assay for chromatin accessibility
T7 RNA polymerase (T7 RNAP) assay for examining chromatin accessibility was adapted from a previous protocol44. 5 × 107 cells were PBS washed and resuspended for 5 min on ice in cold HP-20 Buffer (20 mM Tris pH 8.0, 20 mM KCL, 8 mM MgCl2, 1 mM CaCl2, 2 mM DTT, 2 mM NaCl, 0.3 M Sucrose, and 0.2% NP-40) to break cell membranes. Crude nuclei were then washed in cold Buffer T (20 mM Tris pH 8.0, 20 mM KCL, 8 mM MgCl2, 1 mM CaCl2, 2 mM DTT, 2 mM NaCl, and 0.3 M Sucrose) and centrifuged at 20,000 × g for 10 min at 4 °C. The supernatant was discarded and the number of cell nuclei was counted using Trypan Blue (Sigma) staining on a Countess 3 Cell Counter (Thermo Fisher Scientific). 1 × 107 nuclei were incubated with 25 mM rNTPs (NEB, N0466S) and 160 U T7 RNAP (NEB, M0658S) at 37 °C for 1 h. The reaction was stopped and RNA was extracted using TRIreagent (Sigma). T7 RNAP derived transcripts were analyzed by RT-qPCR with Chr2/116 5’P0 and 3’P0 primer pairs.
RNA 5’ cap analysis
The presence of an RNA 5’ cap (m7GpppN) on Chr2/116 insert-derived transcripts was examined with a Terminator 5’-Phosphate Exonuclease kit (Epicenter, TER51020). 2 μg of total RNA was incubated with or without 1 U of Terminator Exonuclease at 30 °C for 1 hrs. Reaction was stopped by purifying RNA with a RNA Clean & Concentrator kit (Zymo Research, R1016) and then cDNA was generated with SuperScript II Reverse Transcriptase (Thermo Fisher Scientific, 18064014) as described above. RT-qPCR for the detection of 5’P0 and 3’P0 Chr2/116 transcripts was done as described above.
Detection of single-stranded DNA with recombinant RPA-eGFP
For ChIP-qPCR, 10 μg of double cross-linked and sonicated chromatin was incubated with 10 μg of recombinant RPA-eGFP or eGFP at 4 °C O/N on a rotating wheel and then captured with GFP-Trap magnetic agarose beads (ChromoTek, gtma-20) at 4 °C for 1 h. Beads washes, chromatin decross-linking and DNA purification were performed as described above. ChIP enrichment across the Chr2/116 insertion site was analysed by qPCR using PrimeSTAR Max DNA polymerase (Takara Bio, R045) and EvaGreen Dye (Biotium, 31000) with insert-specific T7 and T3 primers.
Detection of RNA:DNA hybrids with recombinant HBD-eGFP
Expression and purification of eGFP-tagged mouse recombinant hybrid binding domain (HBD-eGFP) was done as described54 and ChIP-qPCR with recombinant HBD-eGFP was performed as explained for RPA-eGFP.
Etoposide treatment
Etoposide treatment was performed as previously described55. 3 × 105 cells were plated in 10 cm dishes and cultured for 24 h before being incubated with 1 μM etoposide (Sigma, E1383) for 24 h. RT-qPCR for the detection of 5’P0 and 3’P0 Chr2/116 transcripts was done as described above.
siRNA knock-down for Ints11
Ints11 knock-down was guided by the ON-TARGETplus Mouse Ints11 SMARTpool siRNA (Dharmacon, 71957). 2 × 105 cells in 6 well plates were transfected using DharmaFECT (Dharmacon, T200102) with a mix of 4 siRNA targeted against Ints11 at a final concentration of 25 nM and incubated at 37 °C for 24 h. Without wash, cells were re-transfected for another 24 h. The transfection medium was replaced with normal ESC medium and cells were harvested for total RNA extraction.
Western blot for the detection of INTS11 after depletion
For INTS11-degron depletion, 1 × 106 cells were washed twice with PBS to remove Trimethoprim and then cultivated in fresh mESC medium that contained DMSO (1 μl/ml) instead of Trimethoprim for 24 hrs. Cells were washed twice with PBS, harvested and lysed in 200 μl of 2X Laemmli buffer (BioRad, 1610747). Samples were then boiled at 95 °C for 10 min. 20 μl of protein samples per loading well were separated by 4–20% SDS-PAGE electrophoresis (BioRad mini-PROTEAN, 4561094). Blots were imaged using a ChemiDoc system (BioRad, XRS + ). Primary antibodies were against HA-tag (Cell Signaling, 3724, 1:1000) and GAPDH (Santa Cruz, sc-32233, 1:2000).
RNA-sequencing in INTS11-degron mESC
For RNA-sequencing, total RNA was extracted with TRIzol reagent (Thermo Fisher Scientific) and ethanol precipitation 24 h after removal of Trimethoprim. To eliminate contaminating DNA, 2 U of TURBO DNase (Invitrogen, AM1907) was added to 10 μg of RNA at 37 °C for 30 min and RNA was purified. 1 μg of DNase-treated input RNA was used to generate RNA-seq libraries (150 bp paired-end) with the TruSeq Stranded Total RNA library preparation kit (Illumina, 20020596) and incorporation of the ERCC RNA Spike-in Mix (Invitrogen). Libraries were pooled and sequenced on an Illumina NextSeq 500 platform. The average sequencing depth was 50 million reads per sample.
ChIP-sequencing for H3K9me3 and HP1α in INTS11-degron mESC
ChIP sequencing was performed by the Deep Sequencing unit of the MPI-IE using the RELACS nuclei barcoding protocol72. 5 × 106 INTS11-degron mESC were fixed by single cross-linking (1% formaldehyde) for H3K9me3 and by double cross-linking (2 mM DSG and 1% formaldehyde) for HP1α, snap-frozen in liquid nitrogen and stored at −150 °C. Chromatin was digested inside nuclei and A-tailed using the NEBNext Ultra II DNA library preparation kit (NEB, E7645L). Hairpin adapter barcodes (IDT) were then ligated, samples were pooled and lysed by sonication. 100 μl of chromatin (containing around 200.000 lysed nuclei per pooled sample) was immunoprecipitated with 4 μg of anti-H3K9me3 (Abcam, ab8898) or 3 μg of anti-HP1α (Cell Signaling, 2616). Libraries with insert sizes between 200 to 1000 bp were generated and sequenced with a coverage of > 250 million reads on a NovaSeq 6000 platform (Illumina).
HiSeq RNA sequencing read alignment to the de novo assembly of MSR variants
Illumina 150 bp paired-end HiSeq RNA sequencing reads were trimmed using cutadapt v.2.573 and then aligned to the de novo assembly of MSR variants (GE_MSR_contigassembly4.0.fa) (Supplementary Fig. 10a) with bwa mem v.0.7.1774. The resulting BAM files were sorted using samtools v.1.9.075 and filtered only reads with mapq>1 were considered for downstream analysis. Bigwig files were generated through deeptools v.3.3.076 with parameters “--samFlagExclude 384 --bs 5 --extendReads 5 --normalizeUsing RPGC --effectiveGenomeSize 2652783500”. Heat maps were made by sorting MSR alignments into intact ( > 95% conservation to MSR consensus) and permutated (less than 95% conservation to MSR consensus) MSR sequence bins. Signals from the bigwig files were visualised using deeptools computeMatrix reference-point with parameters “--skipZeros --binSize 10 -a 200 -b 200 --referencePoint center”, followed by the use of plotHeatmap76.
Statistical analysis
Statistical analysis was done with GraphPad Prism V10. One-way ANOVA statistical testing was used for all ChIP-qPCR experiments and for RT-qPCR analysis of Chr2/116 5’P0 and 3’P0 transcripts and T7 RNAP chromatin accessibility assay. Two-way ANOVA statistical testing was used for RT-qPCR experiments, including RNA 5’ cap analysis, Ints11 siRNA knock-down, and etoposide treatment. *p < 0.05 was considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Fastq files for RNA-seq libraries (150 bp paired-end reads) in INTS11-degron ES cells have been deposited in the GEO database under accession number GSE299895. The de novo MSR contig assembly (GE_MSR_contigassembly4.0.fa) can be accessed upon request through the Bioinformatics unit of the MPI-IE. The fastq files for H3K9me3 and HP1α ChIP-seq in INTS11-degron ES cells have been deposited in the GEO database under accession numbers GSE300664 and GSE300945.The fastq files for INTS11 ChIP-seq in WT26 ES cells have been deposited in the GEO database under accession number GSE293226. All oligonucleotides used in this study and relevant sequence information are provided in Supplementaty Tables 1-5. Source data are provided as a Source Data file with this paper. The data generated in this study can be accessed at the following databases: GSE299895GSE300664GSE300945GSE293226 Source data are provided with this paper.
Code availability
The custom Python script is listed in the Methods and is publicly available. https://github.com/gerikson/Find_repeatFree_region.
References
Heitz, E. Das Heterochromatin der Moose. Jahrücher F.ür. Wissenschaftliche Botanik 69, 762–818 (1928).
Janssen, A., Colmenares, S. U. & Karpen, G. H. Heterochromatin: guardian of the genome. Annu. Rev. Cell Dev. Biol. 34, 265–288 (2018).
Allshire, R. C. & Madhani, H. D. Ten principles of heterochromatin formation and function. Nat. Rev. Mol. Cell Biol. 19, 229–244 (2018).
Allis, C. D. & Jenuwein, T. The molecular hallmarks of epigenetic control. Nat. Rev. Genet. 17, 487–500 (2016).
Orphanides, G., Lagrange, T. & Reinberg, D. The general transcription factors of RNA polymerase II. Genes Dev. 10, 2657–2683 (1996).
Kadonaga, J. T. Regulation of RNA polymerase II transcription by sequence-specific DNA binding factors. Cell 116, 247–257 (2004).
Li, B., Carey, M. & Workman, J. L. The role of chromatin during transcription. Cell 128, 707–719 (2007).
Rea, S. et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406, 593–599 (2000).
Lachner, M., O’Carroll, D., Rea, S., Mechtler, K. & Jenuwein, T. Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410, 116–120 (2001).
Bannister, A. J. et al. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410, 120–124 (2001).
Mouse Genome Sequencing, C. et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Hörz, W. & Altenburger, W. Nucleotide sequence of mouse satellite DNA. Nucleic Acids Res 9, 683–696 (1981).
Vassen, L., Fiolka, K. & Moroy, T. Gfi1b alters histone methylation at target gene promoters and sites of gamma-satellite containing heterochromatin. EMBO J. 25, 2409–2419 (2006).
Yamashita, K., Sato, A., Asashima, M., Wang, P. C. & Nishinakamura, R. Mouse homolog of SALL1, a causative gene for Townes-Brocks syndrome, binds to A/T-rich sequences in pericentric heterochromatin via its C-terminal zinc finger domains. Genes Cells 12, 171–182 (2007).
Bulut-Karslioglu, A. et al. A transcription factor-based mechanism for mouse heterochromatin formation. Nat. Struct. Mol. Biol. 19, 1023–1030 (2012).
Ma, R. et al. Targeting pericentric non-consecutive motifs for heterochromatin initiation. Nature 631, 678–685 (2024).
Huisinga, K. L., Brower-Toland, B. & Elgin, S. C. The contradictory definitions of heterochromatin: transcription and silencing. Chromosoma 115, 110–122 (2006).
Grewal, S. I. & Elgin, S. C. Transcription and RNA interference in the formation of heterochromatin. Nature 447, 399–406 (2007).
Volpe, T. A. et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–1837 (2002).
Hall, I. M. et al. Establishment and maintenance of a heterochromatin domain. Science 297, 2232–2237 (2002).
Verdel, A. et al. RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303, 672–676 (2004).
Grewal, S. I. S. The molecular basis of heterochromatin assembly and epigenetic inheritance. Mol. Cell 83, 1767–1785 (2023).
Lee, N. N. et al. Mtr4-like protein coordinates nuclear RNA processing for heterochromatin assembly and for telomere maintenance. Cell 155, 1061–1074 (2013).
Vo, T. V. et al. CPF recruitment to non-canonical transcription termination sites triggers heterochromatin assembly and gene silencing. Cell Rep. 28, 267–281.e265 (2019).
Khanduja, J. S. et al. RNA quality control factors nucleate Clr4/SUV39H and trigger constitutive heterochromatin assembly. Cell 187, 3262–3283 (2024).
Saksouk, N. et al. Redundant mechanisms to form silent chromatin at pericentromeric regions rely on BEND3 and DNA methylation. Mol. Cell 56, 580–594 (2014).
Kanellopoulou, C. et al. Dicer-deficient mouse embryonic stem cells are defective in differentiation and centromeric silencing. Genes Dev. 19, 489–501 (2005).
Gutbrod, M. J. et al. Dicer promotes genome stability via the bromodomain transcriptional co-activator BRD4. Nat. Commun. 13, 1001 (2022).
Probst, A. V. et al. A strand-specific burst in transcription of pericentric satellites is required for chromocenter formation and early mouse development. Dev. Cell 19, 625–638 (2010).
Burton, A. et al. Heterochromatin establishment during early mammalian development is regulated by pericentromeric RNA and characterized by non-repressive H3K9me3. Nat. Cell Biol. 22, 767–778 (2020).
Zhu, Q. et al. Heterochromatin-encoded satellite RNAs induce breast cancer. Mol. Cell 70, 842–853.e847 (2018).
Ting, D. T. et al. Aberrant overexpression of satellite repeats in pancreatic and other epithelial cancers. Science 331, 593–596 (2011).
Thakur, J., Packiaraj, J., Henikoff, S. Sequence, chromatin and evolution of satellite DNA. Int. J. Mol. Sci. 22, (2021).
Packiaraj, J. & Thakur, J. DNA satellite and chromatin organization at mouse centromeres and pericentromeres. Genome Biol. 25, 52 (2024).
Bulut-Karslioglu, A. et al. Suv39h-dependent H3K9me3 marks intact retrotransposons and silences LINE elements in mouse embryonic stem cells. Mol. Cell 55, 277–290 (2014).
He, Y. et al. Spatiotemporal DNA methylome dynamics of the developing mouse fetus. Nature 583, 752–759 (2020).
Sethi, A. et al. Supervised enhancer prediction with epigenetic pattern recognition and targeted validation. Nat. Methods 17, 807–814 (2020).
Bonev, B. et al. Multiscale 3D genome rewiring during mouse neural development. Cell 171, 557–572.e524 (2017).
Velazquez Camacho, O., et al. Major satellite repeat RNA stabilize heterochromatin retention of Suv39h enzymes by RNA-nucleosome association and RNA:DNA hybrid formation. Elife 6, (2017).
Matsui, T. et al. Proviral silencing in embryonic stem cells requires the histone methyltransferase ESET. Nature 464, 927–931 (2010).
Rowe, H. M. et al. KAP1 controls endogenous retroviruses in embryonic stem cells. Nature 463, 237–240 (2010).
Zhou, M. & Smith, A. D. Subtype classification and functional annotation of L1Md retrotransposon promoters. Mob. DNA 10, 14 (2019).
Isomura, H. et al. A cis element between the TATA Box and the transcription start site of the major immediate-early promoter of human cytomegalovirus determines efficiency of viral replication. J. Virol. 82, 849–858 (2008).
Jenuwein, T., Forrester, W. C., Qiu, R. G. & Grosschedl, R. The immunoglobulin mu enhancer core establishes local factor access in nuclear chromatin independent of transcriptional stimulation. Genes Dev. 7, 2016–2032 (1993).
Baillat, D. et al. Integrator, a multiprotein mediator of small nuclear RNA processing, associates with the C-terminal repeat of RNA polymerase II. Cell 123, 265–276 (2005).
Kirstein, N., Gomes Dos Santos, H., Blumenthal, E. & Shiekhattar, R. The Integrator complex at the crossroad of coding and noncoding RNA. Curr. Opin. Cell Biol. 70, 37–43 (2021).
Lykke-Andersen, S. et al. Integrator is a genome-wide attenuator of non-productive transcription. Mol. Cell 81, 514–529.e516 (2021).
Stein, C. B. et al. Integrator endonuclease drives promoter-proximal termination at all RNA polymerase II-transcribed loci. Mol. Cell 82, 4232–4245.e4211 (2022).
Fianu, I. et al. Structural basis of Integrator-dependent RNA polymerase II termination. Nature 629, 219–227 (2024).
Xu, C. et al. R-loop-dependent promoter-proximal termination ensures genome stability. Nature 621, 610–619 (2023).
Kasinathan, S. & Henikoff, S. Non-B-Form DNA Is Enriched at Centromeres. Mol. Biol. Evol. 35, 949–962 (2018).
Tan-Wong, S. M., Dhir, S. & Proudfoot, N. J. R-loops promote antisense transcription across the mammalian genome. Mol. Cell 76, 600–616.e606 (2019).
Skourti-Stathaki, K., Kamieniarz-Gdula, K. & Proudfoot, N. J. R-loops induce repressive chromatin marks over mammalian gene terminators. Nature 516, 436–439 (2014).
Duda, K. J. et al. m6A RNA methylation of major satellite repeat transcripts facilitates chromatin association and RNA:DNA hybrid formation in mouse heterochromatin. Nucleic Acids Res. 49, 5568–5587 (2021).
Fuhrmann, T. et al. The isoflavone genistein selectively stimulates major satellite repeat transcription in mouse heterochromatin. Epigenet. Chromatin 18, 58 (2025).
Pantier, R. et al. MeCP2 binds to methylated DNA independently of phase separation and heterochromatin organisation. Nat. Commun. 15, 3880 (2024).
Tatarakis, A. et al. Requirements for establishment and epigenetic stability of mammalian heterochromatin. Mol. Cell 85, 3388–3406.e3312 (2025).
Torre, D. et al. Nuclear RNA catabolism controls endogenous retroviruses, gene expression asymmetry, and dedifferentiation. Mol. Cell 83, 4255–4271.e4259 (2023).
Edupuganti, R. R. et al. Integrator promotes the association of TFIID and RNA polymerase II to maintain pluripotency during development. Mol. Cell 85, 2937–2955.e2910 (2025).
Zhang, J. et al. Distinct H3K9me3 heterochromatin maintenance dynamics govern different gene programmes and repeats in pluripotent cells. Nat. Cell Biol. 26, 2115–2128 (2024).
Wadsworth, G. M. et al. RNA-driven phase transitions in biomolecular condensates. Mol. Cell 84, 3692–3705 (2024).
Martienssen, R. & Moazed, D. RNAi and heterochromatin assembly. Cold Spring Harb. Perspect. Biol. 7, a019323 (2015).
Reinberg, D. & Vales, L. D. Chromatin domains rich in inheritance. Science 361, 33–34 (2018).
Dorer, D. R. Henikoff S. Expansions of transgene repeats cause heterochromatin formation and gene silencing in Drosophila. Cell 77, 993–1002 (1994).
Johnson, W. L., et al. RNA-dependent stabilization of SUV39H1 at constitutive heterochromatin. Elife 6, (2017).
Healton, S. E. et al. H1 linker histones silence repetitive elements by promoting both histone H3K9 methylation and chromatin compaction. Proc. Natl. Acad. Sci. USA 117, 14251–14258 (2020).
Xu, W. et al. METTL3 regulates heterochromatin in mouse embryonic stem cells. Nature 591, 317–321 (2021).
Aygun, O., Mehta, S. & Grewal, S. I. HDAC-mediated suppression of histone turnover promotes epigenetic stability of heterochromatin. Nat. Struct. Mol. Biol. 20, 547–554 (2013).
Lehnertz, B. et al. Suv39h-mediated histone H3 lysine 9 methylation directs DNA methylation to major satellite repeats at pericentric heterochromatin. Curr. Biol. 13, 1192–1200 (2003).
Nassar, L. R. et al. The UCSC Genome Browser database: 2023 update. Nucleic Acids Res. 51, D1188–D1195 (2023).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Arrigoni, L. et al. RELACS nuclei barcoding enables high-throughput ChIP-seq. Commun. Biol. 1, 214 (2018).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Li H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv 1303.3997 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Acknowledgements
We thank Devon Ryan for the initial interrogation of repeat-free regions in the mouse genome and other members of the Bioinformatics team at the MPI-IE for the analysis of genome–wide data sets. We also thank members of the Deep-Sequencing facility of the MPI-IE for the generation and processing of RNA-seq and long-read DNA-seq libraries. Preliminary long-read DNA sequencing was provided as a customised service from the Christine van Broeckhoven lab (University of Antwerp). We are indebted to Danny Reinberg for insightful discussions on transcriptional regulation and to Giacomo Cavalli for advice on the 3D organization of the mouse epigenome. Research in the laboratory of R.S. is supported by funding from the University of Miami Miller School of Medicine, Sylvester Comprehensive Cancer Center, and grants R01GM078455 from the National Institute of Health. Research in the laboratory of T.J. is supported by the Max Planck Society and by additional funds from the German Research Foundation (DFG) within the CRC992 consortium ‘MEDEP’. This manuscript is dedicated to the memory of David Allis, a pioneer for the discovery of histone modifying enzymes and a friend to many of us in the field of chromatin and epigenetic research.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
T.J. conceived the study with initial work from D.P. and then continuous inputs from R.W.C. and Y-H.L. Y-H.L. performed most experiments and analysed the data. N.S. generated and verified all mESC clones with CRISPR/Cas9-mediated insertions. G.E. performed bioinformatics analyses. R.R.E. generated the INTS11-degron mESC. L.J. generated and purified HBD-eGFP and RPA-eGFP recombinant proteins and performed the EMSA assay. R.R.E. and R.S. provided guidance and expertise for the Integrator complex. T.J., Y-H.L., R.W.C. and G.E. wrote and edited the manuscript with feedback from R.R.E. and R.S. T.J. and R.S. acquired funding. T.J. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interest.
Peer review
Peer review information
Nature Communications thanks Yamini Dalal, who co-reviewed with Anne Gilbert and Sweta Sikder, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lo, YH., Shukeir, N., Erikson, G. et al. Transcriptional competence defines the heterochromatin nucleating potential of isolated MSR units. Nat Commun 17, 2653 (2026). https://doi.org/10.1038/s41467-026-70991-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70991-2